Abstract
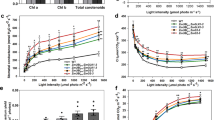
Rubisco catalyses a rate-limiting step in photosynthesis and has long been a target for improvement due to its slow turnover rate. An alternative to modifying catalytic properties of Rubisco is to increase its abundance within C4 plant chloroplasts, which might increase activity and confer a higher carbon assimilation rate. Here, we overexpress the Rubisco large (LS) and small (SS) subunits with the Rubisco assembly chaperone RUBISCO ASSEMBLY FACTOR 1 (RAF1). While overexpression of LS and/or SS had no discernable impact on Rubisco content, addition of RAF1 overexpression resulted in a >30% increase in Rubisco content. Gas exchange showed a 15% increase in CO2 assimilation (ASAT) in UBI-LSSS-RAF1 transgenic plants, which correlated with increased fresh weight and in vitro Vcmax calculations. The divergence of Rubisco content and assimilation could be accounted for by the Rubisco activation state, which decreased up to 23%, suggesting that Rubisco activase may be limiting Vcmax, and impinging on the realization of photosynthetic potential from increased Rubisco content.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data generated and analysed during this study are available from the corresponding author on reasonable request. Raw data would include photosynthesis and enzyme activity analyses.
References
Sharwood, R. E. Engineering chloroplasts to improve Rubisco catalysis: prospects for translating improvements into food and fiber crops. New Phytol. 213, 494–510 (2017).
Hauser, T., Popilka, L., Hartl, F. U. & Hayer-Hartl, M. Role of auxiliary proteins in Rubisco biogenesis and function. Nat. Plants 1, 15065 (2015).
Furbank, R. T. et al. Genetic manipulation of key photosynthetic enzymes in the C4 plant Flaveria bidentis. Func. Plant Biol. 24, 477–485 (1997).
Ghannoum, O. et al. Faster Rubisco is the key to superior nitrogen-use efficiency in NADP-malic enzyme relative to NAD-malic enzyme C4 grasses. Plant Physiol. 137, 638–650 (2005).
Sharwood, R. E., Ghannoum, O. & Whitney, S. M. Prospects for improving CO2 fixation in C3-crops through understanding C4-Rubisco biogenesis and catalytic diversity. Curr. Opin. Plant. Biol. 31, 135–142 (2016).
Feller, U., Anders, I. & Mae, T. Rubiscolytics: fate of Rubisco after its enzymatic function in a cell is terminated. J. Exp. Bot. 59, 1615–1624 (2008).
Farquhar, G. On the nature of carbon isotope discrimination in C4 species. Func. Plant Biol. 10, 205–226 (1983).
Furbank, R., Jenkins, C. & Hatch, M. C4 photosynthesis—quantum requirement, C4 acid overcycling and Q-cycle involvement. Aust. J. Plant. Physiol. 17, 1–7 (1990).
Bracher, A., Whitney, S. M., Hartl, F. U. & Hayer-Hartl, M. Biogenesis and metabolic maintenance of Rubisco. Annu. Rev. Plant. Biol. 68, 29–60 (2017).
Suzuki, Y. et al. Increased Rubisco content in transgenic rice transformed with the ‘sense’ rbcS gene. Plant Cell Physiol. 48, 626–637 (2007).
Wostrikoff, K., Clark, A., Sato, S., Clemente, T. & Stern, D. Ectopic expression of rubisco subunits in maize mesophyll cells does not overcome barriers to cell type-specific accumulation. Plant Physiol. 160, 419–432 (2012).
Barkan, A. Nuclear mutants of maize with defects in chloroplast polysome assembly have altered chloroplast RNA metabolism. Plant Cell 5, 389–402 (1993).
Brutnell, T. P., Sawers, R. J., Mant, A. & Langdale, J. A. BUNDLE SHEATH DEFECTIVE2, a novel protein required for post- translational regulation of the rbcL gene of maize. Plant Cell 11, 849–864 (1999).
Feiz, L. et al. Ribulose-1,5-Bis-Phosphate Carboxylase/Oxygenase Accumulation Factor1 is required for holoenzyme assembly in maize. Plant Cell 24, 3435–3446 (2012).
Feiz, L. et al. A protein with an inactive pterin-4a-carbinolamine dehydratase domain is required for Rubisco biogenesis in plants. Plant J. 80, 862–869 (2014).
Aigner, H. et al. Plant RuBisCo assembly in E. coli with five chloroplast chaperones including BSD2. Science 358, 1272–1278 (2017).
Kolesinski, P. et al. Insights into eukaryotic Rubisco assembly - crystal structures of RbcX chaperones from Arabidopsis thaliana. Biochim. Biophys. Acta 1830, 2899–2906 (2013).
Hauser, T. et al. Structure and mechanism of the Rubisco-assembly chaperone Raf1. Nat. Struct. Mol. Biol. 22, 720–728 (2015).
Whitney, S. M., Birch, R., Kelso, C., Beck, J. L. & Kapralov, M. V. Improving recombinant Rubisco biogenesis, plant photosynthesis and growth by coexpressing its ancillary RAF1 chaperone. Proc. Natl Acad. Sci. USA 112, 3564–3569 (2015).
Wheatley, N. M., Sundberg, C. D., Gidaniyan, S. D., Cascio, D. & Yeates, T. O. Structure and Identification of apterin dehydratase-like protein as a Ribulose-bisphosphate Carboxylase/Oxygenase (RuBisCO) assembly factor in the alpha-carboxysome. J. Biol. Chem. 289, 7973–7981 (2014).
Bhat, J. Y., Thieulin-Pardo, G., Hartl, F. U. & Hayer-Hartl, M. Rubisco activases: AAA+ chaperones adapted to enzyme repair. Front. Mol. Biosci. 4, 20 (2017).
Mueller-Cajar, O. The diverse AAA+ machines that repair inhibited rubisco active sites. Front. Mol. Biosci. 4, 31 (2017).
Sharwood, R. E., Sonawane, B. V., Ghannoum, O. & Whitney, S. M. Improved analysis of C4 and C3 photosynthesis via refined in vitro assays of their carbon fixation biochemistry. J. Exp. Bot. 67, 3137–3148 (2016).
von Caemmerer, S., Ghannoum, O., Pengelly, J. J. L. & Cousins, A. B. Carbon isotope discrimination as a tool to explore C4 photosynthesis. J. Exp. Bot. 65, 3459–3470 (2014).
Rodermel, S., Haley, J., Jiang, C. Z., Tsai, C. H. & Bogorad, L. A mechanism for intergenomic integration:abundance of ribulose bisphosphate carboxylase small-subunit protein influences the translation of the large-subunit mRNA. Proc. Natl Acad. Sci. USA 93, 3881–3885 (1996).
Johnson, X. et al. MRL1, a conserved pentatricopeptide repeat protein, is required for stabilization of rbcL mRNA in Chlamydomonas and Arabidopsis. Plant Cell 22, 234–248 (2010).
Kanevski, I. & Maliga, P. Relocation of the plastid rbcL gene to the nucleus yields functional ribulose-1,5-bisphosphate carboxylase in tobacco chloroplasts. Proc. Natl Acad. Sci. USA 91, 1969–1973 (1994).
Kubien, D. S., von Caemmerer, S., Furbank, R. T. & Sage, R. F. C4 photosynthesis at low temperature. A study using transgenic plants with reduced amounts of Rubisco. Plant Physiol. 132, 1577–1585 (2003).
Salesse-Smith, C., Sharwood, R. E., Sakamoto, W. & Stern, D. B. The Rubisco chaperone BSD2 may regulate chloroplast coverage in maize bundle sheath cells. Plant Physiol. 175, 1624–1633 (2017).
Friso, G., Majeran, W., Huang, M., Sun, Q. & van Wijk, K. J. Reconstruction of metabolic pathways, protein expression and homeostasis machineries across maize bundle sheath and mesophyll chloroplasts; large scale quantitative proteomics using the first maize genome assembly. Plant Physiol. 152, 1219–1250 (2010).
von Caemmerer, S. & Furbank, R. T. Strategies for improving C4 photosynthesis. Curr. Opin. Plant. Biol. 31, 125–134 (2016).
Carmo-Silva, E., Scales, J. C., Madgwick, P. J. & Parry, M. A. Optimizing Rubisco and its regulation for greater resource use efficiency. Plant Cell Environ. 38, 1817–1832 (2015).
Suzuki, Y., Miyamoto, T., Yoshizawa, R., Mae, T. & Makino, A. Rubisco content and photosynthesis of leaves at different positions in transgenic rice with an overexpression of RBCS. Plant Cell Environ. 32, 417–427 (2009).
Ishikawa, C., Hatanaka, T., Misoo, S., Miyake, C. & Fukayama, H. Functional incorporation of sorghum small subunit increases the catalytic turnover rate of Rubisco in transgenic rice. Plant Physiol. 156, 1603–1611 (2011).
Morita, K., Hatanaka, T., Misoo, S. & Fukayama, H. Unusual small subunit that is not expressed in photosynthetic cells alters the catalytic properties of Rubisco in rice. Plant Physiol. 164, 69–79 (2014).
Crafts-Brandner, S. J. & Salvucci, M. E. Sensitivity of photosynthesis in a C4 plant, maize, to heat stress. Plant Physiol. 129, 1773–1780 (2002).
Millard, P. The accumulation and storage of nitrogen by herbaceous plants. Plant Cell Environ. 11, 1–8 (1988).
Millard, P. & Grelet, G. -a Nitrogen storage and remobilization by trees: ecophysiological relevance in a changing world. Tree Physiol. 30, 1083–1095 (2010).
Sweetlove, L. J., Nielsen, J. & Fernie, A. R. Engineering central metabolism – a grand challenge for plant biologists. Plant J. 90, 749–763 (2017).
Farooq, M., Aziz, T., Wahid, A., Lee, D. & Siddique, K. H. M. Chilling tolerance in maize: agronomic and physiological approaches. Crop Pasture Sci. 60, 501–516 (2009).
Long, S. P. C4 photosynthesis at low temperatures. Plant Cell Environ. 6, 345–363 (1983).
Wang, D., Portis, A. R. Jr., Moose, S. P. & Long, S. P. Cool C4 photosynthesis: pyruvate Pi dikinase expression and activity corresponds to the exceptional cold tolerance of carbon assimilation in Miscanthus x giganteus. Plant Physiol. 148, 557–567 (2008).
Naidu, S. L., Moose, S. P., AK, A. L.-S., Raines, C. A. & Long, S. P. Cold tolerance of C4 photosynthesis in Miscanthus x giganteus: adaptation in amounts and sequence of C4 photosynthetic enzymes. Plant Physiol. 132, 1688–1697 (2003).
Cousins, A. B. et al. The role of phosphoenolpyruvate carboxylase during C4 photosynthetic isotope exchange and stomatal conductance. Plant Physiol. 145, 1006–1017 (2007).
Brown R. H. in C 4 Plant Biology 1st edn (eds Sage, R. & Monson, R.) 473–509 (Academic Press, Cambridge, 1999).
Ort, D. R. et al. Redesigning photosynthesis to sustainably meet global food and bioenergy demand. Proc. NatlAcad. Sci. USA 112, 8529–8536 (2015).
Niyogi, K. K. Editorial overview: Physiology and metabolism: Light responses from photoreceptors to photosynthesis and photoprotection. Curr. Opin. Plant Biol. 37, 4–7 (2017).
Sarlikioti, V., De Visser, P. H., Buck-Sorlin, G. & Marcelis, L. How plant architecture affects light absorption and photosynthesis in tomato: towards an ideotype for plant architecture using a functional–structural plant model. Ann. Bot. 108, 1065–1073 (2011).
Glowacka, K. et al. Photosystem II Subunit S overexpression increases the efficiency of water use in a field-grown crop. Nat. Commun. 9, 868 (2018).
Feng, L. et al. Overexpression of SBPase enhances photosynthesis against high temperature stress in transgenic rice plants. Plant Cell Rep. 26, 1635–1646 (2007).
Sattarzadeh, A. et al. Transgenic maize lines with cell-type specific expression of fluorescent proteins in plastids. Plant. Biotechnol. J. 8, 112–125 (2010).
Markelz, N. H., Costich, D. E. & Brutnell, T. P. Photomorphogenic responses in maize seedling development. Plant Physiol. 133, 1578–1591 (2003).
Barkan, A. Approaches to investigating nuclear genes that function in chloroplast biogenesis in land plants. Meths. Enzymol. 297, 38–57 (1998).
Lilley, R. M. & Walker, D. A. An improved spectrophotometric assay for ribulosebisphosphate carboxylase. Biochim. Biophys. Acta 358, 226–229 (1974).
Sharwood, R. E., von Caemmerer, S., Maliga, P. & Whitney, S. M. The catalytic properties of hybrid Rubisco comprising tobacco small and sunflower large subunits mirror the kinetically equivalent source Rubiscos and can support tobacco growth. Plant Physiol. 146, 83–96 (2008).
Ashton, A. R., Burnell, J. N., Furbank, R. T., Jenkins, C. L. D. & Hatch, M. D. in Methods in Plant Biochemistry Vol 3 (ed. Lea, P. J.) 39–71 (Academic, London, 1999).
Sharwood, R. E., Sonawane, B. V. & Ghannoum, O. Photosynthetic flexibility in maize exposed to salinity andshade. J. Exp. Bot. 65, 3715–3724 (2014).
Kromdijk, J., Griffiths, H. & Schepers, H. E. Can the progressive increase of C4 bundle sheath leakiness at low PFD be explained by incomplete suppression of photorespiration? Plant Cell Environ. 33, 1935–1948 (2010).
Evans, J., Sharkey, T., Berry, J. & Farquhar, G. Carbon isotope discrimination measured concurrently with gas exchange to investigate CO2 diffusion in leaves of higher plants. Funct. Plant Biol. 13, 281–292 (1986).
Barbour, M. M., Evans, J. R., Simonin, K. A. & Caemmerer, S. Online CO2 and H2O oxygen isotope fractionation allows estimation of mesophyll conductance in C4 plants, and reveals that mesophyll conductance decreases as leaves age in both C4 and C3 plants. New Phytol. 210, 875–889 (2016).
Kromdijk, J., Ubierna, N., Cousins, A. B. & Griffiths, H. Bundle-sheath leakiness in C4 photosynthesis: a careful balancing act between CO2 concentration and assimilation. J. Exp. Bot. 65, 3443–3457 (2014).
Acknowledgements
We acknowledge T. Clemente and S. Sato (University of Nebraska–Lincoln) for assembling the final transformation constructs, performing maize transformations, and providing seed from T0 lines. T. Nelson is acknowledged for sharing ME antibody. This research was supported by the Agriculture and Food Research Initiative from the National Institute of Food and Agriculture, US Department of Agriculture, under award number 2016-67013-24464. Travel to the Australian National University was supported by the Mario Einaudi Center for International Studies, International Research Travel Grant at Cornell University. We thank S. Long (University of Illinois) for helpful discussions and manuscript suggestions. R.E.S. is funded by the ARC Centre of Excellence for Translational Photosynthesis (CE140100015) and ARC DECRA (DE13010760).
Author information
Authors and Affiliations
Contributions
C.S. participated in all experiments and drafted the manuscript. R.S. participated in experiments shown in Figs. 1, 2, 3, 5 and Table 1. F.A.B. participated in experiments shown in Fig. 3 and Table 1. J.K. participated in some experiments presented in Fig. 5 and Table 1. V.B. participated in experiments shown in Supplementary Figs. 3 and 4. D.S. was responsible for project management, finalization of data analysis and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–4.
Rights and permissions
About this article
Cite this article
Salesse-Smith, C.E., Sharwood, R.E., Busch, F.A. et al. Overexpression of Rubisco subunits with RAF1 increases Rubisco content in maize. Nature Plants 4, 802–810 (2018). https://doi.org/10.1038/s41477-018-0252-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0252-4
This article is cited by
-
Improvement of photosynthesis in changing environment: approaches, achievements and prospects
Plant Biotechnology Reports (2024)
-
From leaf to multiscale models of photosynthesis: applications and challenges for crop improvement
Photosynthesis Research (2024)
-
Optimization of photosynthesis for sustainable crop production
CABI Agriculture and Bioscience (2022)
-
CRISPR gene editing of major domestication traits accelerating breeding for Solanaceae crops improvement
Plant Molecular Biology (2022)
-
Transcriptome dynamic landscape underlying the improvement of maize lodging resistance under coronatine treatment
BMC Plant Biology (2021)