Abstract
The phytohormone auxin is the information carrier in a plethora of developmental and physiological processes in plants1. It has been firmly established that canonical, nuclear auxin signalling acts through regulation of gene transcription2. Here, we combined microfluidics, live imaging, genetic engineering and computational modelling to reanalyse the classical case of root growth inhibition3 by auxin. We show that Arabidopsis roots react to addition and removal of auxin by extremely rapid adaptation of growth rate. This process requires intracellular auxin perception but not transcriptional reprogramming. The formation of the canonical TIR1/AFB–Aux/IAA co-receptor complex is required for the growth regulation, hinting to a novel, non-transcriptional branch of this signalling pathway. Our results challenge the current understanding of root growth regulation by auxin and suggest another, presumably non-transcriptional, signalling output of the canonical auxin pathway.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Leyser, O. Auxin signaling. Plant Physiol. 176, 465–479 (2018).
Gray, W. M., Kepinski, S., Rouse, D., Leyser, O. & Estelle, M. Auxin regulates SCFTIR1-dependent degradation of AUX/IAA proteins. Nature 414, 271–276 (2001).
Evans, M. L., Ishikawa, H. & Estelle, M. A. Responses of Arabidopsis roots to auxin studied with high temporal resolution—Comparison of wild-type and auxin-response mutants. Planta 194, 215–222 (1994).
Knight, T. A. On the direction of the radicle and germen during the vegetation of seeds. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 96, 99–108 (1806).
Darwin, C. The Power of Movement in Plants (John Murray, London, 1880).
Nemec, B. Ueber die art der wahrnehmung des schwek—Raftreizes bei den pflanzen. Ber. Dtsch. Bot. Ges. 18, 241–245 (1900).
Adamowski, M. & Friml, J. PIN-dependent auxin transport: Action, regulation, and evolution. Plant Cell 27, 20–32 (2015).
Went, F. & Thimann, K. Phytohormones (Macmillan, New York, 1937).
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007).
Fendrych, M., Leung, J. & Friml, J. TIR1/AFB–Aux/IAA auxin perception mediates rapid cell wall acidification and growth of Arabidopsis hypocotyls. eLife 5, e19048 (2016).
Uchida, N. et al. Chemical hijacking of auxin signaling with an engineered auxin-TIR1 pair. Nat. Chem. Biol. 14, 299–305 (2018).
Leyser, H. M. O., Pickett, F. B., Dharmasiri, S. & Estelle, M. Mutations in the AXR3 gene of Arabidopsis result in altered auxin response including ectopic expression from the SAUR-AC1 promoter. Plant J. 10, 403–413 (1996).
Ruegger, M. et al. The TIR1 protein of Arabidopsis functions in auxin response and is related to human SKP2 and yeast grr1p. Genes Dev. 12, 198–207 (1998).
Scheitz, K., Lüthen, H. & Schenck, D. Rapid auxin-induced root growth inhibition requires the TIR and AFB auxin receptors. Planta 238, 1171–1176 (2013).
Monshausen, G. B., Miller, N. D., Murphy, A. S. & Gilroy, S. Dynamics of auxin-dependent Ca2 + and pH signaling in root growth revealed by integrating high-resolution imaging with automated computer vision-based analysis. Plant J. 65, 309–318 (2011).
Shih, H. W., Depew, C. L., Miller, N. D. & Monshausen, G. B. The cyclic nucleotide-gated channel CNGC14 regulates root gravitropism in Arabidopsis thaliana. Curr. Biol. 25, 3119–3125 (2015).
Grossmann, G. et al. The RootChip: An integrated microfluidic chip for plant science. Plant Cell 23, 4234–4240 (2011).
von Wangenheim, D. et al. Live tracking of moving samples in confocal microscopy for vertically grown roots. eLife 6, e26792 (2017).
Band, L. R. R. et al. Root gravitropism is regulated by a transient lateral auxin gradient controlled by a tipping-point mechanism. Proc. Natl Acad. Sci. USA 109, 4668–4673 (2012).
Brunoud, G. et al. A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482, 103–106 (2012).
Hoyle, N. P. & Ish-Horowicz, D. Transcript processing and export kinetics are rate-limiting steps in expressing vertebrate segmentation clock genes. Proc. Natl Acad. Sci. USA 110, E4316–E4324 (2013).
McClure, B. A., Hagen, G., Brown, C. S., Gee, M. A. & Guilfoyle, T. J. Transcription, organization, and sequence of an auxin-regulated gene cluster in soybean. Plant Cell 1, 229–39 (1989).
Moreno-Risueno, M. A. et al. Oscillating gene expression determines competence for periodic Arabidopsis root branching. Science 329, 1306–1311 (2010).
Macdonald, I. R. & Ellis, R. J. Does cycloheximide inhibit protein synthesis specifically in plant tissues?. Nature 222, 791–792 (1969).
Okushima, Y. et al. Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana. Plant Cell 17, 444–463 (2005).
Swarup, R. et al. Root gravitropism requires lateral root cap and epidermal cells for transport and response to a mobile auxin signal. Nat. Cell Biol. 7, 1057–1065 (2005).
Bennett, M. J. et al. Arabidopsis AUX1 gene: A permease-like regulator of root gravitropism. Science 273, 948–950 (1996).
Delbarre, A., Muller, P., Imhoff, V. & Guern, J. Comparison of mechanisms controlling uptake and accumulation of 2,4-dichlorophenoxy acetic acid, naphthalene-1-acetic acid, and indole-3-acetic acid in suspension-cultured tobacco cells. Planta 198, 532–541 (1996).
Dindas, J. et al. AUX1-mediated root hair auxin influx governs SCFTIR1/AFB-type Ca2 + signaling. Nat. Commun. 9, 1174 (2018).
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119 (2005).
Hayashi, K. et al. Rational design of an auxin antagonist of the SCF(TIR1) auxin receptor complex. ACS Chem. Biol. 7, 590–598 (2012).
Swarup, R. et al. Structure-function analysis of the presumptive Arabidopsis auxin permease AUX1. Plant Cell 16, 3069–3083 (2004).
Grossmann, G. et al. Time-lapse fluorescence imaging of Arabidopsis root growth with rapid manipulation of the root environment using the RootChip. J. Vis. Exp. 65, e4290 (2012).
Li, L., Krens, S. F. G., Fendrych, M. & Friml, J. Real-time analysis of auxin response, cell wall pH and elongation in Arabidopsis thaliana hypocotyls. Bio-Protoc. 7, 1–10 (2018).
Thévenaz, P., Ruttimann, U. E., U., M., Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Schneider, Ca, Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
The authors thank R. Hauschild for help with image analysis. M.F. was supported by the Austrian Science Fund (FWF) (M 2128-B21), and in the final stages of the project by the Czech Science Foundation (18-10116Y). M.A. was supported by the Austrian Science Fund (FWF) (M2379-B28). Additional funding for M.G. was received from the Ministry of Education of the Czech Republic/MŠMT project NPUI-LO1417. N.U. received MEXT/JSPS KAKENHI JP16H01462, S.H. received JP17H06350, K.T. received JP26440140 and K.U.T. received JP16H01237. K.U.T. is supported by GBMF Grant No. 3035. S.H. is a JST PRESTO Investigator and K.U.T. is an HHMI-GBMF Investigator. J.F. acknowledges the ERC Advanced Grants (ETAP).
Author information
Authors and Affiliations
Contributions
M.F. initiated the project, acquired funding, performed the experiments, analysed the data and wrote the manuscript. M.A. created the computational model of auxin fluxes and wrote the manuscript. J.M. designed, optimized and fabricated the microfluidic devices. M.G. performed experiments and edited the manuscript. S.H., K.T., N.U. and K.U.T. designed, synthesized and developed the engineered cvxIAA–ccvTIR1 system and edited the manuscript. J.F. initiated the project, acquired funding and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Text, Supplementary Figures 1–5, Supplementary Tables 1–4 and Supplementary References
Supplementary Data 1
Contains data supporting the results in the paper
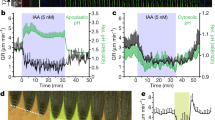
Supplementary Video 1
The reaction of a root to addition and removal of 10 nM IAA. This movie is a spectacular demonstration of the transient nature of the root growth inhibition by auxin. The root almost instantaneously limits growth when IAA is added (marked by the magenta dye), and the growth is resumed following IAA removal
Supplementary Video 2
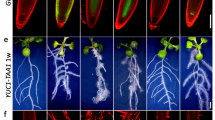
aux1 mutant compared to control after addition of 5 nM IAA auxin (indicated by OFF and ON)
Supplementary Video 3
Modelled temporal dynamics of IAA intracellular accumulation in control and aux1 modelled roots
Supplementary Video 4
The reaction of ccvTIR1 and controlTIR1 roots to the addition and removal of 50 nM cvxIAA (false coloured in blue)
Rights and permissions
About this article
Cite this article
Fendrych, M., Akhmanova, M., Merrin, J. et al. Rapid and reversible root growth inhibition by TIR1 auxin signalling. Nature Plants 4, 453–459 (2018). https://doi.org/10.1038/s41477-018-0190-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0190-1
This article is cited by
-
New fluorescent auxin derivatives: anti-auxin activity and accumulation patterns in Arabidopsis thaliana
Plant Growth Regulation (2024)
-
A biostimulant yeast, Hanseniaspora opuntiae, modifies Arabidopsis thaliana root architecture and improves the plant defense response against Botrytis cinerea
Planta (2024)
-
Plant adenylate cyclases have come full circle
Nature Plants (2023)
-
Structure and growth of plant cell walls
Nature Reviews Molecular Cell Biology (2023)
-
Antigravitropic PIN polarization maintains non-vertical growth in lateral roots
Nature Plants (2023)