Abstract
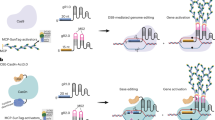
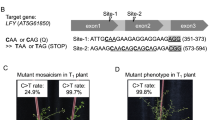
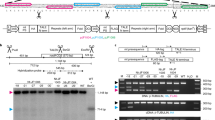
The recent development of adenine base editors (ABEs) has enabled efficient and precise A-to-G base conversions in higher eukaryotic cells. Here, we show that plant-compatible ABE systems can be successfully applied to protoplasts of Arabidopsis thaliana and Brassica napus through transient transfection, and to individual plants through Agrobacterium-mediated transformation to obtain organisms with desired phenotypes. Targeted, precise A-to-G substitutions generated a single amino acid change in the FT protein or mis-splicing of the PDS3 RNA transcript, and we could thereby obtain transgenic plants with late-flowering and albino phenotypes, respectively. Our results provide ‘proof of concept’ for in planta ABE applications that can lead to induced neo-functionalization or altered mRNA splicing, opening up new avenues for plant genome engineering and biotechnology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
23 August 2018
In Supplementary Fig. 1b originally published with this Brief Communication, the DNA sequence of nickase Cas9 was incorrect; this has now been amended.
References
Yin, K., Gao, C. & Qiu, J.-L. Nat. Plants 3, 17107 (2017).
Woo, J. W. et al. Nat. Biotechnol. 33, 1162–1164 (2015).
Alonso-Blanco, C. et al. Plant Cell 21, 1877–1896 (2009).
Slade, A. J., Fuerstenberg, S. I., Loeffler, D., Steine, M. N. & Facciotti, D. Nat. Biotechnol. 23, 75–81 (2005).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Nature 533, 420–424 (2016).
Nishida, K. et al. Science 353, 1248–1257 (2016).
Zong, Y. et al. Nat. Biotechnol. 35, 438–440 (2017).
Shimatani, Z. et al. Nat. Biotechnol. 35, 441–443 (2017).
Gaudelli, N. M. et al. Nature 551, 464–471 (2017).
Bae, S., Park, J. & Kim, J. S. Bioinformatics 30, 1473–1475 (2014).
Kim, D. et al. Nat. Methods 12, 237–243 (2015).
Yan, L. et al. Mol. Plant 8, 1820–1823 (2015).
Tsutsui, H. & Higashiyama, T. Plant Cell Physiol. 58, 46–56 (2017).
Hanzawa, Y., Money, T. & Bradley, D. Proc. Natl Acad. Sci. USA 102, 7748–7753 (2005).
Qin, G. et al. Cell Res. 17, 471–482 (2007).
Hua, K., Tao, X., Yuan, F., Wang, D. & Zhu, J.-K. Mol. Plant 11, 627–630 (2018).
Yan, F. et al. Mol. Plant 11, 631–634 (2018).
Kim, H. et al. J. Integr. Plant Biol. 58, 705–712 (2016).
Zhang, X., Henriques, R., Lin, S. S., Niu, Q. W. & Chua, N. H. Nat. Protoc. 1, 641–646 (2006).
Park, J., Lim, K., Kim, J.-S. & Bae, S. Bioinformatics 33, 286–288 (2017).
Acknowledgements
We thank all members of the Center for Genome Engineering for their support. We thank J. Suh for comments on the manuscript and technical assistance. This work was supported by the research fund from the Institute for Basic Science, Republic of Korea (IBS-R021-D1).
Author information
Authors and Affiliations
Contributions
J.-Y.Y., B.-C.K., S.-T.K. and J.W.W. designed the experiments. B.-C.K., J.-Y.Y., J.R., Y.S. and M.C. generated all the constructs. B.-C.K. performed the transient assay in protoplasts. J.-Y.Y., Y.S. and J.R. generated the transgenic Arabidopsis and analysed the plants. J.-Y.Y., S.-T.K., B.-C.K. and J.-S.K. wrote the manuscript with the help of all other authors. J.-S.K. supervised the project.
Corresponding author
Ethics declarations
Competing interests
J.-S.K. is a co-founder of and holds stocks in ToolGen, Inc. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8, Supplementary Table 1
Rights and permissions
About this article
Cite this article
Kang, BC., Yun, JY., Kim, ST. et al. Precision genome engineering through adenine base editing in plants. Nature Plants 4, 427–431 (2018). https://doi.org/10.1038/s41477-018-0178-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0178-x
This article is cited by
-
Recent advances in therapeutic CRISPR-Cas9 genome editing: mechanisms and applications
Molecular Biomedicine (2023)
-
Single-cell resolution analysis reveals the preparation for reprogramming the fate of stem cell niche in cotton lateral meristem
Genome Biology (2023)
-
A streamlined guide RNA screening system for genome editing in Sorghum bicolor
Plant Methods (2023)
-
Recent advances in the plant epitranscriptome
Genome Biology (2023)
-
Split complementation of base editors to minimize off-target edits
Nature Plants (2023)