Abstract
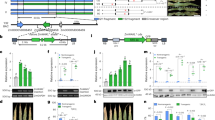
Localized control of cell death is crucial for the resistance of plants to pathogens. Papain-like cysteine proteases (PLCPs) regulate plant defence to drive cell death and protection against biotrophic pathogens. In maize (Zea mays), PLCPs are crucial in the orchestration of salicylic acid (SA)-dependent defence signalling. Despite this central role in immunity, it remains unknown how PLCPs are activated, and which downstream signals they induce to trigger plant immunity. Here, we discover an immune signalling peptide, Z. mays immune signalling peptide 1 (Zip1), which is produced after salicylic acid (SA) treatment. In vitro studies demonstrate that PLCPs are required to release bioactive Zip1 from its propeptide precursor. Conversely, Zip1 treatment strongly elicits SA accumulation in leaves. Moreover, transcriptome analyses revealed that Zip1 and SA induce highly overlapping transcriptional changes. Consequently, Zip1 promotes the infection of the necrotrophic fungus Botrytis cinerea, while it reduces virulence of the biotrophic fungus Ustilago maydis. Thus, Zip1 represents the previously missing signal that is released by PLCPs to activate SA defence signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
van der Hoorn, R. A. L. Plant proteases: from phenotypes to molecular mechanisms. Annu Rev. Plant Biol. 59, 191–223 (2008).
Xia, Y. et al. An extracellular aspartic protease functions in Arabidopsis disease resistance signaling. EMBO J. 23, 980–988 (2004).
Tian, M., Huitema, E., da Cunha, L., Torto-Alalibo, T. & Kamoun, S. A kazal-like extracellular serine protease inhibitor from Phytophthora infestans targets the tomato pathogenesis-related protease P69B. J. Biol. Chem. 279, 26370–26377 (2004).
Tornero, P., Conejero, V. & Vera, P. Primary structure and expression of a pathogen-induced protease (PR-P69) in tomato plants: similarity of functional domains to subtilisin-like endoproteases. Proc. Natl Acad. Sci. USA 93, 6332–6337 (1996).
Stegmann, M. et al. The receptor kinase FER is a RALF-regulated scaffold controlling plant immune signaling. Science 355, 287–289 (2017).
Misas-Villamil, J. C., van der Hoorn, R. A. L. & Doehlemann, G. Papain-like cysteine proteases as hubs in plant immunity. New Phytol. 212, 902–907 (2016).
Rooney, H. C. et al. Cladosporium Avr2 inhibits tomato Rcr3 protease required for Cf-2-dependent disease resistance. Science 308, 1783–1786 (2005).
Song, J. et al. Apoplastic effectors secreted by two unrelated eukaryotic plant pathogens target the tomato defense protease Rcr3. Proc. Natl Acad. Sci. USA 106, 1654–1659 (2009).
Lozano-Torres, J. L. et al. Dual disease resistance mediated by the immune receptor Cf-2 in tomato requires a common virulence target of a fungus and a nematode. Proc. Natl Acad. Sci. USA 109, 10119–10124 (2012).
Glazebrook, J. Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu. Rev. Phytopathol. 43, 205–227 (2005).
Pieterse, C. M., Leon-Reyes, A., Van der Ent, S. & Van Wees, S. C. Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 5, 308–316 (2009).
Yan, S. & Dong, X. Perception of the plant immune signal salicylic acid. Curr. Opin. Plant Biol. 20, 64–68 (2014).
Doherty, H. M., Selvendran, R. R. & Bowles, D. J. The wound response of tomato plants can be inhibited by aspirin and related hydroxy-benzoic acids. Phys. Mol. Plant Pathol. 33, 377–384 (1988).
Pefia-Cortes, H., Albrecht, T., Prat, S., Weiler, E. W. & Willmitzer, L. Aspirin prevents wound-induced gene expression in tomato leaves by blocking jasmonic acid biosynthesis. Planta 191, 123–128 (1993).
Thomma, B. P. et al. Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc. Natl Acad. Sci. USA 95, 15107–15111 (1998).
Huffaker, A. et al. Plant elicitor peptides are conserved signals regulating direct and indirect antiherbivore defense. Proc. Natl Acad. Sci. USA 110, 5707–5712 (2013).
Huffaker, A., Dafoe, N. J. & Schmelz, E. A. ZmPep1, an ortholog of Arabidopsis elicitor petide 1, regulates maize innate immunity and enhances disease resistance. Plant Physiol. 155, 1325–1338 (2011).
Huffaker, A., Pearce, G. & Ryan, C. A. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. Proc. Natl Acad. Sci. USA 103, 10098–10103 (2006).
Brefort, T. et al. Ustilago maydis as a pathogen. Annu. Rev. Phytopathol. 47, 423–445 (2009).
Doehlemann, G. et al. Reprogramming a maize plant: transcriptional and metabolic changes induced by the fungal biotroph Ustilago maydis. Plant J. 56, 181–195 (2008).
Hof, A., Zechmann, B., Schwammbach, D., Huckelhoven, R. & Doehlemann, G. Alternative cell death mechanisms determine epidermal resistance in incompatible barley-ustilago interactions. Mol. Plant Microbe. Interact. 27, 403–414 (2014).
Doehlemann, G. et al. Pep1, a secreted effector protein of Ustilago maydis, is required for successful invasion of plant cells. PLoS Pathog. 5, e1000290 (2009).
van der Linde, K. et al. A maize cystatin suppresses host immunity by inhibiting apoplastic cysteine proteases. Plant Cell 24, 1285–1300 (2012).
Mueller, A. N., Ziemann, S., Treitschke, S., Assmann, D. & Doehlemann, G. Compatibility in the Ustilago maydis-maize interaction requires inhibition of host cysteine proteases by the fungal effector Pit2. PLoS Pathog. 9, (2013).
Dolezal, A. L. et al. Aspergillus flavus infection induces transcriptional and physical changes in developing maize kernels. Front Microbiol. 5, 384 (2014).
Ray, S. et al. Turnabout is fair play: herbivory-induced plant chitinases excreted in fall armyworm frass suppress herbivore defenses in maize. Plant Physiol. 171, 694–706 (2016).
Barrett, A. J., Kembhavi, A. A. & Hanada, K. E-64 [L-trans-epoxysuccinyl-leucyl-amido(4-guanidino)butane] and related epoxides as inhibitors of cysteine proteinases. Acta Biol. Med Ger. 40, 1513–1517 (1981).
Cravatt, B. F., Wright, A. T. & Kozarich, J. W. Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu Rev. Biochem 77, 383–414 (2008).
Greenbaum, D., Medzihradszky, K. F., Burlingame, A. & Bogyo, M. Epoxide electrophiles as activity-dependent cysteine protease profiling and discovery tools. Chem. Biol. 7, 569–581 (2000).
Paireder, M. et al. The death enzyme CP14 is a unique papain-like cysteine proteinase with a pronounced S2 subsite selectivity. Arch. Biochem Biophys. 603, 110–117 (2016).
Choe, Y. et al. Substrate profiling of cysteine proteases using a combinatorial peptide library identifies functionally unique specificities. J. Biol. Chem. 281, 12824–12832 (2006).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Zipfel, C. et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428, 764–767 (2004).
Monaghan, J. et al. The calcium-dependent protein kinase CPK28 buffers plant immunity and regulates BIK1 turnover. Cell Host Microbe 16, 605–615 (2014).
Miya, A. et al. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl Acad. Sci. 104, 19613–19618 (2007).
Jabaiah, A. M., Getz, J. A., Witkowski, W. A., Hardy, J. A. & Daugherty, P. S. Identification of protease exosite-interacting peptides that enhance substrate cleavage kinetics. Biol. Chem. 393, 933–941 (2012).
Lori, M. et al. Evolutionary divergence of the plant elicitor peptides (Peps) and their receptors: interfamily incompatibility of perception but compatibility of downstream signalling. J. Exp. Bot. 66, 5315–5325 (2015).
Bartels, S. & Boller, T. Quo vadis, Pep? Plant elicitor peptides at the crossroads of immunity, stress, and development. J. Exp. Bot. 66, 5183–5193 (2015).
van der Linde, K. et al. Pathogen Trojan horse delivers bioactive host protein to alter maize (Zea mays) anther cell behavior in situ. Plant Cell. https://doi.org/10.1105/tpc.17.00238 (2018).
Cirino, G. & Vergnolle, N. Proteinase-activated receptors (PARs): crossroads between innate immunity and coagulation. Curr. Opin. Pharmacol. 6, 428–434 (2006).
Schmidlin, F. & Bunnett, N. W. Protease-activated receptors: how proteases signal to cells. Curr. Opin. Pharmacol. 1, 575–582 (2001).
Srivastava, R., Liu, J. X., Guo, H., Yin, Y. & Howell, S. H. Regulation and processing of a plant peptide hormone, AtRALF23, in Arabidopsis. Plant J. 59, 930–939 (2009).
Srivastava, R., Liu, J. X. & Howell, S. H. Proteolytic processing of a precursor protein for a growth-promoting peptide by a subtilisin serine protease in Arabidopsis. Plant J. 56, 219–227 (2008).
Paireder, M. et al. The papain-like cysteine proteinases NbCysP6 and NbCysP7 are highly processive enzymes with substrate specificities complementary to Nicotiana benthamiana cathepsin B. Biochim Biophys Acta 1865, 444–452 (2017).
Chen, Z., Zheng, Z., Huang, J., Lai, Z. & Fan, B. Biosynthesis of salicylic acid in plants. Plant Signal. Behav. 4, 493–496 (2009).
Djamei, A. et al. Metabolic priming by a secreted fungal effector. Nature 478, 395–398 (2011).
Acosta, I. et al. Tasselseed1 is a lipoxygenase affecting jasmonic acid signaling in sex determination of maize. Science 323, 262–265 (2009).
Huffaker, A. et al. Novel acidic sesquiterpenoids constitute a dominant class of pathogen-induced phytoalexins in maize. Plant Physiol. 156, 2082–2097 (2011).
van der Linde, K., Kastner, C., Kumlehn, J., Kahmann, R. & Doehlemann, G. Systemic virus-induced gene silencing allows functional characterization of maize genes during biotrophic interaction with Ustilago maydis. New Phytol. 189, 471–483 (2011).
Pearce, G. et al. Isolation and characterization of hydroxyproline-rich glycopeptide signals in black nightshade leaves. Plant Physiol. 150, 1422–1433 (2009).
Pearce, G., Strydom, D., Johnson, S. & Ryan, C. A. A polypeptide from tomato leaves induces wound-inducible proteinase inhibitor proteins. Science 253, 895–897 (1991).
Ryan, C., Pearce, G., Scheer, J. & Moura, D. Polypeptide hormones. Plant Cell Online, 251–264 (2002).
Chen, Y. C., Siems, W. F., Pearce, G. & Ryan, C. A. Six peptide wound signals derived from a single precursor protein in Ipomoea batatas leaves activate the expression of the defense gene sporamin. J. Biol. Chem. 283, 11469–11476 (2008).
Acknowledgements
This work is funded by the German Research Foundation (DFG) via grant DO 1421/5-1 (GD). Mass spectrometry work was financially supported by an ERC starting grant (M.K., grant No. 258413) and the Deutsche Forschungsgemeinschaft (M.K., grant no. INST 20876/127-1 FUGG). Research in the Zipfel laboratory is supported by the Gatsby Charitable Foundation. We are very grateful to R. Kahmann for helpful discussions and the Max Planck Institute for Terrestrial Microbiology, Marburg, Germany, for continuous support and access to laboratory facilities. We are also very thankful to A. Matei for meaningful discussions and for critical reading the manuscript. We thank R. van der Hoorn (Oxford University) for generously providing us with ABPP probes.
Author contributions
S.Z., K.L. and G.D. designed the experiments and analysed the data. S.Z., K.L. and B.A. performed the functional analysis of Zip1/PROZIP1. N.H. and C.Z. designed and analysed ROS and MAPK assays. Y.D., A.H. and E.S. designed, performed and analysed salicylic acid measurements. U.L. analysed the transcriptome data. F.K., T.C. and M.K. performed MS experiments and MS-related data analysis. S.Z. and G.D. wrote the manuscript with input from all authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Materials and Methods, Supplementary References, Supplementary Figures 1–7, Supplementary Table 2, Supplementary Table 3.
Supplementary Table 1
Complete gene list of RNAseq analyses with differentially expressed genes in response to SA and Zip1 compared to mock samples, respectively
Rights and permissions
About this article
Cite this article
Ziemann, S., van der Linde, K., Lahrmann, U. et al. An apoplastic peptide activates salicylic acid signalling in maize. Nature Plants 4, 172–180 (2018). https://doi.org/10.1038/s41477-018-0116-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-018-0116-y
This article is cited by
-
Enhancing tomato resistance by exploring early defense events against Fusarium wilt disease
Phytopathology Research (2023)
-
A papain-like cysteine protease-released small signal peptide confers wheat resistance to wheat yellow mosaic virus
Nature Communications (2023)
-
Ustilago maydis PR-1-like protein has evolved two distinct domains for dual virulence activities
Nature Communications (2023)
-
Planthopper salivary sheath protein LsSP1 contributes to manipulation of rice plant defenses
Nature Communications (2023)
-
Mechanisms controlling plant proteases and their substrates
Cell Death & Differentiation (2023)