Abstract
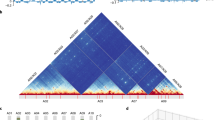
The formation of polyploids significantly increases the complexity of transcriptional regulation, which is expected to be reflected in sophisticated higher-order chromatin structures. However, knowledge of three-dimensional (3D) genome structure and its dynamics during polyploidization remains poor. Here, we characterize 3D genome architectures for diploid and tetraploid cotton, and find the existence of A/B compartments and topologically associated domains (TADs). By comparing each subgenome in tetraploids with its extant diploid progenitor, we find that genome allopolyploidization has contributed to the switching of A/B compartments and the reorganization of TADs in both subgenomes. We also show that the formation of TAD boundaries during polyploidization preferentially occurs in open chromatin, coinciding with the deposition of active chromatin modification. Furthermore, analysis of inter-subgenomic chromatin interactions has revealed the spatial proximity of homoeologous genes, possibly associated with their coordinated expression. This study advances our understanding of chromatin organization in plants and sheds new light on the relationship between 3D genome evolution and transcriptional regulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lieberman-Aiden, E. Comprehensive mapping of long-range interactions reveals folding principles ofthe human genome. Science 326, 289–293 (2009).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Zhang, Y. B. et al. Chromatin connectivity maps reveal dynamic promoter-enhancer long-range associations. Nature 504, 306–310 (2013).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Feng, S. et al. Genome-wide Hi-C analyses in wild-type and mutants reveal high-resolution chromatin interactions in Arabidopsis. Mol. Cell 55, 694–707 (2014).
Grob, S., Schmid, Marc., W. & Grossniklaus, U. Hi-C analysis in Arabidopsis identifies the KNOT, a structure with similarities to the flamenco locus of Drosophila. Mol. Cell 55, 678–693 (2014).
Wang, C. et al. Genome-wide analysis of local chromatin packing in Arabidopsis thaliana. Genome Res. 25, 246–256 (2015).
Liu, C. et al. Genome-wide analysis of chromatin packing in Arabidopsis thaliana at single-gene resolution. Genome Res. 26, 1057–1068 (2016).
Wang, M. et al. Asymmetric subgenome selection and cis-regulatory divergence during cotton domestication. Nat. Genet. 49, 579–587 (2017).
Senchina, D. S. et al. Rate variation among nuclear genes and the age of polyploidy in Gossypium. Mol. Biol. Evol. 20, 633–643 (2003).
Zhang, T. et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537 (2015).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–249 (2015).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228 (2017).
Tjong, H. et al. Population-based 3D genome structure analysis reveals driving forces in spatial genome organization. Proc. Natl Acad. Sci. USA 113, E1663–E1672 (2016).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Li, F. et al. Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 33, 524–530 (2015).
Schmidt, D. et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages.Cell 148, 335–348 (2012).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Merkenschlager, M. & Odom, D. T. CTCF and cohesin: linking gene regulatory elements with their targets. Cell 152, 1285–1297 (2013).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381 (2017).
Thevenin, A., Ein-Dor, L., Ozery-Flato, M. & Shamir, R. Functional gene groups are concentrated within chromosomes, among chromosomes and in the nuclear space of the human genome. Nucleic Acids Res. 42, 9854–9861 (2014).
Xie, T., Yang, Q. Y., Wang, X. T., McLysaght, A. & Zhang, H. Y. Spatial colocalization of human ohnolog pairs acts to maintain dosage-balance. Mol. Biol. Evol. 33, 2368–2375 (2016).
Wang, M. et al. Multi-omics maps of cotton fibre reveal epigenetic basis for staged single-cell differentiation. Nucleic Acids Res. 44, 4067–4079 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Cronn, R. C., Small, R. L. & Wendel, J. F. Duplicated genes evolve independently after polyploid formation in cotton. Proc. Natl Acad. Sci. USA 96, 14406–14411 (1999).
Page, J. T. et al. DNA sequence evolution and rare homoeologous conversion in tetraploid cotton. PLoS Genet. 12, e1006012 (2016).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Servant, N. et al. HiTC: exploration of high-throughput ‘C’ experiments. Bioinformatics 28, 2843–2844 (2012).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Shin, H. et al. TopDom: an efficient and deterministic method for identifying topological domains in genomes. Nucleic Acids Res. 44, e70 (2016).
Boyle, A. P., Guinney, J., Crawford, G. E. & Furey, T. S. F-Seq: a feature density estimator for high-throughput sequence tags. Bioinformatics 24, 2537–2538 (2008).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Sun, Q. & Zhou, D. X. Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. Proc. Natl Acad. Sci. USA 105, 13679–13684 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
We thank K. Lindsey (Durham University) for revising this manuscript. This work was funded by the National Natural Science Foundation of China (31230056, 31301005), China Postdoctoral Science Foundation (2015M572169) and State Key Laboratory of Cotton Biology Open Fund (CB2016A08).
Author information
Authors and Affiliations
Contributions
X.Z., M.W. and Q.Y. conceived and designed the project. M.L., P.W. and Z.Y. performed the experiments and managed sequencing. M.W., C.S. and J.L. analysed the data. G.L. and L.T. contributed to manuscript discussion. M.W. wrote the manuscript draft. X.Z., Q.Y. and G.L. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Tables 1–4
Supplementary Table 5
Identification of intergenic DHSs and enhancers in cotton
Supplementary Table 6
Summary of enhancer-associated lincRNAs in cotton
Supplementary Table 7
Summary of TADs in cotton
Supplementary Table 8
Summary of compartment switching in cotton
Supplementary Table 9
Summary of DEGs associated with changes of chromatin structures
Supplementary Table 10
Inter-chromosomal interactions in G. hirsutum
Supplementary Table 11
Inter-chromosomal interactions in G. barbadense
Supplementary Table 12
Homologous gene pairs with chromatin interactions in cotton
Rights and permissions
About this article
Cite this article
Wang, M., Wang, P., Lin, M. et al. Evolutionary dynamics of 3D genome architecture following polyploidization in cotton. Nature Plants 4, 90–97 (2018). https://doi.org/10.1038/s41477-017-0096-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-017-0096-3
This article is cited by
-
Genomic rearrangements and evolutionary changes in 3D chromatin topologies in the cotton tribe (Gossypieae)
BMC Biology (2023)
-
Dynamic chromatin architectures provide insights into the genetics of cattle myogenesis
Journal of Animal Science and Biotechnology (2023)
-
Regulation of chromatin organization during animal regeneration
Cell Regeneration (2023)
-
Cotton pedigree genome reveals restriction of cultivar-driven strategy in cotton breeding
Genome Biology (2023)
-
3D organization of regulatory elements for transcriptional regulation in Arabidopsis
Genome Biology (2023)