Abstract
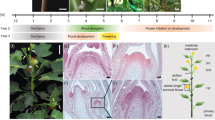
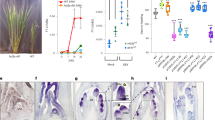
DELLA proteins associate with transcription factors to control plant growth in response to gibberellin1. Semi-dwarf DELLA mutants with improved harvest index and decreased lodging greatly improved global food security during the ‘green revolution’ in the 1960–1970s2. However, DELLA mutants are pleiotropic and the developmental basis for their effects on plant architecture remains poorly understood. Here, we show that DELLA proteins have genetically separable roles in controlling stem growth and the size of the inflorescence meristem, where flowers initiate. Quantitative three-dimensional image analysis, combined with a genome-wide screen for DELLA-bound loci in the inflorescence tip, revealed that DELLAs limit meristem size in Arabidopsis by directly upregulating the cell-cycle inhibitor KRP2 in the underlying rib meristem, without affecting the canonical WUSCHEL-CLAVATA meristem size regulators3. Mutation of KRP2 in a DELLA semi-dwarf background restored meristem size, but not stem growth, and accelerated flower production. In barley, secondary mutations in the DELLA gain-of-function mutant Sln1d 4 also uncoupled meristem and inflorescence size from plant height. Our work reveals an unexpected and conserved role for DELLA genes in controlling shoot meristem function and suggests how dissection of pleiotropic DELLA functions could unlock further yield gains in semi-dwarf mutants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Daviere, J.-M. & Achard, P. Gibberellin signaling in plants. Development 140, 1147–1151 (2013).
Pingali, P. L. Green revolution: impacts, limits, and the path ahead. Proc. Natl Acad. Sci. USA 109, 12302–12308 (2012).
Holt, A. L., van Haperen, J. M., Groot, E. P. & Laux, T. Signaling in shoot and flower meristems of Arabidopsis thaliana. Curr. Opin. Plant Biol. 17, 96–102 (2014).
Chandler, P. M. & Harding, C. A. ‘Overgrowth’ mutants in barley and wheat: new alleles and phenotypes of the ‘Green Revolution’ Della gene. J. Exp. Bot 64, 1603–1613 (2013).
King, K. E., Moritz, T. & Harberd, N. P. Gibberellins are not required for normal stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics 159, 767–776 (2001).
Daviere, J. M. et al. Class I TCP-DELLA interactions in inflorescence shoot apex determine plant height. Curr. Biol. 24, 1923–1928 (2014).
Peng, J. et al. The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev. 11, 3194–3205 (1997).
Dill, A. & Sun, T. P. Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 159, 777–785 (2001).
Ubeda-Tomas, S. et al. Gibberellin signaling in the endodermis controls Arabidopsis root meristem size. Curr. Biol. 19, 1194–1199 (2009).
Koornneef, M. & van der Veen, J. H. Induction and analysis of gibberellin sensitive mutants in Arabidopsis thaliana (L.) heynh. Theor. Appl. Genet. 58, 257–263 (1980).
Koini, M. A. et al. High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Curr. Biol. 19, 408–413 (2009).
Jasinski, S. et al. KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr. Biol. 15, 1560–1565 (2005).
Achard, P. et al. Gibberellin signaling controls cell proliferation rate in Arabidopsis. Curr. Biol. 19, 1188–1193 (2009).
Marín-de la Rosa, N. et al. Genome wide binding site analysis reveals transcriptional coactivation of cytokinin-responsive genes by DELLA proteins. PLoS Genet. 11, e1005337 (2015).
Achard, P. et al. Integration of plant responses to environmentally activated phytohormonal signals. Science 311, 91–94 (2006).
De Veylder, L. et al. Functional analysis of cyclin-dependent kinase inhibitors of Arabidopsis. Plant Cell 13, 1653–1668 (2001).
Schiessl, K., Muino, J. M. & Sablowski, R. Arabidopsis JAGGED links floral organ patterning to tissue growth by repressing Kip-related cell cycle inhibitors. Proc. Natl Acad. Sci. USA 111, 2830–2835 (2014).
Rast, M. I. & Simon, R. The meristem-to-organ boundary: more than an extremity of anything. Curr. Opin. Genet. Dev 18, 287–294 (2008).
Sanz, L. et al. The Arabidopsis D-type cyclin CYCD2;1 and the inhibitor ICK2/KRP2 modulate auxin-induced lateral root formation. Plant Cell 23, 641–660 (2011).
Mayer, K. F. et al. Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95, 805–815 (1998).
Yadav, R. K. et al. WUSCHEL protein movement mediates stem cell homeostasis in the Arabidopsis shoot apex. Genes Dev. 25, 2025–2030 (2011).
Savaldi-Goldstein, S., Peto, C. & Chory, J. The epidermis both drives and restricts plant shoot growth. Nature 446, 199 (2007).
Je, B. I. et al. Signaling from maize organ primordia via FASCIATED EAR3 regulates stem cell proliferation and yield traits. Nat. Genet. 48, 785–791 (2016).
Leyser, H. O. & Furner, I. Characterisation of three shoot apical meristem mutants of Arabidopsis thaliana. Development 116, 397–403 (1992).
Hensel, L. L., Nelson, M. A., Richmond, T. A. & Bleecker, A. B. The fate of inflorescence meristems is controlled by developing fruits in Arabidopsis. Plant Physiol. 106, 863–876 (1994).
Dill, A., Jung, H. S. & Sun, T. P. The DELLA motif is essential for gibberellin-induced degradation of RGA. Proc. Natl Acad. Sci. USA 98, 14162–14167 (2001).
Huang, S. et al. Transgenic studies on the involvement of cytokinin and gibberellin in male development. Plant Physiol. 131, 1270–1282 (2003).
Waddington, S. R., Cartwright, P. M. & Wall, P. C. A quantitative scale of spike initial and pistil development in barley and wheat. Ann. Bot. 51, 119–130 (1983).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Xu, C. et al. A cascade of arabinosyltransferases controls shoot meristem size in tomato. Nat. Genet. 47, 784–792 (2015).
Koornneef, M. et al. A gibberellin insensitive mutant of Arabidopsis thaliana. Physiol. Plant. 65, 33–39 (1985).
Gallego-Giraldo, C. et al. Role of the gibberellin receptors GID1 during fruit-set in Arabidopsis. Plant J. 79, 1020–1032 (2014).
Silverstone, A. L. et al. Repressing a repressor: gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell 13, 1555–1565 (2001).
Reddy, G. V. & Meyerowitz, E. M. Stem-cell homeostasis and growth dynamics can be uncoupled in the Arabidopsis shoot apex. Science 310, 663–667 (2005).
Gallois, J. L., Woodward, C., Reddy, G. V. & Sablowski, R. Combined SHOOT MERISTEMLESS and WUSCHEL trigger ectopic organogenesis in Arabidopsis. Development 129, 3207–3217 (2002).
Sieburth, L. E., Drews, G. N. & Meyerowitz, E. M. Non-autonomy of AGAMOUS function in flower development: use of a Cre/loxP method for mosaic analysis in Arabidopsis. Development 125, 4303–4312 (1998).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Smyth, D. R., Bowman, J. L. & Meyerowitz, E. M. Early flower development in Arabidopsis. Plant Cell 2, 755–767 (1990).
Bencivenga, S., Serrano-Mislata, A., Bush, M., Fox, S. & Sablowski, R. Control of oriented tissue growth through repression of organ boundary genes promotes stem morphogenesis. Dev. Cell 39, 198–208 (2016).
Serrano-Mislata, A., Schiessl, K. & Sablowski, R. Active control of cell size generates spatial detail during plant organogenesis. Curr. Biol. 25, 2991–2996 (2015).
Kurihara, D., Mizuta, Y., Sato, Y. & Higashiyama, T. ClearSee: a rapid optical clearing reagent for whole-plant fluorescence imaging. Development 142, 4168–4179 (2015).
Bailey, T. L. et al. MEME suite: tools for motif discovery and searching. Nucleic Acids Res 37, W202 (2009).
Edgar, R., Domrachev, M. & Lash, A. E. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the \({2}^{-\Delta \Delta {{\rm{C}}}_{{\rm{T}}}}\) method. Methods 25, 402–408 (2001).
Acknowledgements
We thank P. Chandler, L. Østergaard, T.-P. Sun, N. Harberd, P. Doerner, V. Reddy, Miguel Á. Pérez Amador and the European Arabidopsis Stock Centre for plasmids and seeds, G. Calder for advice with confocal microscopy and B. Shi for help with vibratome sections. The work was supported by BBSRC grants BB/J007056/1, BB/J004588/1 and BB/M003825/1, and by a grant from the Ministerio de Educación, Cultura y Deporte, Spain (EX-2010-0491) to A.S.-M.
Author information
Authors and Affiliations
Contributions
Conceptualization, A.S.-M., S.Bo. and R.S.; investigation, A.S.-M., S.Be., M.B., S.Bo. and K.S.; software, R.S.; formal analysis and data curation, A.S.-M. and R.S.; writing — original draft, A.S.-M. and R.S.; writing — review and editing, A.S.-M., S.Be., M.B., S.Bo., K.S. and R.S.; funding acquisition, R.S. and A.S.-M.; supervision, R.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Supplementary Methods, Supplementary References, Supplementary Figures 1-6, Supplementary Table legends.
Table 1
High-confidence targets of RGAp:GFP-rga_17 identified by ChIP-seq in inflorescence apices.
Table 2
Manually annotated functional categories of selected RGAp:GFP-rgaD17 target genes identified by ChIP-seq in inflorescence apices.
Table 3
Source images and processing steps for data presented in each figure.
Table 4
Imaging data — filtering parameters, source data and statistical analysis for each figure.
Table 5
ChIP–PCR source data and statistics (related to Fig. 3c).
Rights and permissions
About this article
Cite this article
Serrano-Mislata, A., Bencivenga, S., Bush, M. et al. DELLA genes restrict inflorescence meristem function independently of plant height. Nature Plants 3, 749–754 (2017). https://doi.org/10.1038/s41477-017-0003-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-017-0003-y
This article is cited by
-
DELLA proteins and CBL-CIPK signaling pathway are emerging players in plant biotechnology
Theoretical and Experimental Plant Physiology (2024)
-
Uncovering the involvement of DoDELLA1-interacting proteins in development by characterizing the DoDELLA gene family in Dendrobium officinale
BMC Plant Biology (2023)
-
The master growth regulator DELLA binding to histone H2A is essential for DELLA-mediated global transcription regulation
Nature Plants (2023)
-
Wheat genomic study for genetic improvement of traits in China
Science China Life Sciences (2022)
-
Improving architectural traits of maize inflorescences
Molecular Breeding (2021)