Abstract
Atomistic modeling of chemically reactive systems has so far relied on either expensive ab initio methods or bond-order force fields requiring arduous parametrization. Here, we describe a Bayesian active learning framework for autonomous “on-the-fly” training of fast and accurate reactive many-body force fields during molecular dynamics simulations. At each time-step, predictive uncertainties of a sparse Gaussian process are evaluated to automatically determine whether additional ab initio training data are needed. We introduce a general method for mapping trained kernel models onto equivalent polynomial models whose prediction cost is much lower and independent of the training set size. As a demonstration, we perform direct two-phase simulations of heterogeneous H2 turnover on the Pt(111) catalyst surface at chemical accuracy. The model trains itself in three days and performs at twice the speed of a ReaxFF model, while maintaining much higher fidelity to DFT and excellent agreement with experiment.
Similar content being viewed by others
Introduction
Accurate modeling of chemical reactions is a central challenge in computational physics, chemistry, and biology, lying at the heart of in silico design of covalent drugs1 and next-generation catalysts with higher activity and selectivity2. Reactive molecular dynamics (MD) simulation is an essential tool in advancing such rational design efforts3. By directly simulating the motion of individual atoms without fixing any chemical bonds, reactive MD enables unbiased discovery of reaction mechanisms at atomic resolution as well as prediction of reaction rates complementary to experimental studies4.
Reactive MD requires a flexible model of the potential energy surface (PES) of the system that is both (i) chemically accurate in describing bond breaking and formation, and (ii) computationally affordable to be able to access long timescales necessary to capture rare reactive events. Accurate evaluation of the PES at each time-step can be achieved with ab initio methods such as density functional theory (DFT) and post-Hartree-Fock techniques. However, these methods are limited to small systems due to nonlinear scaling with the number of electrons, precluding their use for dynamical simulation of realistic systems beyond a few hundred atoms spanning tens of picoseconds.
For many nonreactive systems, a viable alternative to ab initio MD is to parameterize a force field whose evaluation cost scales linearly with the number of atoms and is often orders of magnitude cheaper than ab initio methods. However, many traditional force fields, such as the AMBER and CHARMM models extensively used in biomolecular simulations5,6, explicitly fix certain chemical bonds in the system, making them unsuitable for describing chemical reactions. The more flexible ReaxFF model is capable of describing bond breaking and formation and as such has been applied to a wide range of reactive systems in the past two decades3,7. However, ReaxFF models require expert fine-tuning for each system and can significantly deviate from the ab initio PES in many cases8 due to their limited parametric functional form.
Machine-learned (ML) force fields have emerged in the past decade as a powerful tool for building linear-scaling models of the PES at near-DFT accuracy9,10,11,12. These ML models map atomic configurations onto potential energies, atomic forces, and virial stress tensors, using ab initio calculations as the ground truth for the regression. Typically, training these models involves manual construction of the ab initio database targeting the system of interest. This manual approach has been used to describe a range of simple bulk materials8,13,14 and organic molecules15,16,17. However, manual training often requires considerable time, expertise, and computing resources, and is particularly challenging for reactive systems where relevant transition state pathways and their sampling requirement can be difficult to gauge in advance. As a result, ML-driven MD simulations of reactive processes remain scarce in the literature18,19,20,21,22.
A powerful emerging alternative is to generate the training set autonomously using active learning23,24,25,26,27,28. In this approach, model errors or uncertainties are used to decide if a candidate test structure is reliably predicted or should be added to the training set, in which case an ab initio calculation is performed and the model is updated. A particularly promising approach involves training the model "on the fly” during an MD simulation, where the ML model is used to propagate atomic motion and is updated only when the model uncertainties exceed a chosen threshold29,30,31,32. Active learning has been applied successfully in the past year to a range of nonreactive systems and phenomena, including phase transitions in hybrid perovskites33, melting points of solids34, superionic transport in silver iodide32, surface restructuring of palladium deposited on silver35, the 2D-to-3D phase transition of stanene36, and lithium-ion diffusion in solid electrolytes37. It has also been extended recently to chemical reactions within the Gaussian approximation potential (GAP) framework21. The kernel-based GAP force field, although highly accurate, has a prediction cost that scales linearly with the size of the sparse set, making it several orders of magnitude more expensive than traditional force fields such as ReaxFF38.
Chemically reactive systems pose a particular challenge for machine-learned force fields. Owing to its central importance in numerous catalytic processes such as selective hydrogenation39 and hydrogen storage40, H2 reactivity on transition metal surfaces has been a subject of extensive computational investigation, including DFT calculations of H2 activation and diffusion41,42,43, as well as MD simulations using DFT44 and parametric models such as ReaxFF45, embedded-atom method46, tight-binding47, and low-dimensional models48. We note that all previous models only consider either a bare surface interacting with a single H2 molecule or H-covered surface coupled to an implicit reservoir representing the gas phase. These approaches do not explicitly treat the two interacting phases due to either high cost or limited expressiveness insufficient for capturing many-body interactions in a heterogeneous setting. This limits their transferability to realistic systems involving multiple gas-phase H2 molecules and chemisorbed H atoms.
Here, we develop an autonomous method for training reactive many-body force fields on the fly and accelerating the resulting ML models by over an order of magnitude. We apply our method to large-scale reactive MD simulations of a prototypical system in the field of heterogeneous catalysis: reactive turnover of H2 on the (111) surface of platinum, including dissociative adsorption, diffusion, exchange, and recombinative desorption. For the H/Pt system, our prediction speed exceeds that of ReaxFF by more than a factor of two while achieving much higher near-quantum accuracy. Importantly, the training process takes only a few days compared to the months previously required with manual approaches. We accomplish this by using Bayesian uncertainties of a sparse Gaussian process (SGP) model to automatically decide which structures to include in the training set. Once the model is trained, the prediction speed is significantly increased via lossless mapping of the SGP model onto an equivalent parametric model whose cost is independent of the training set size. Our method builds on the sparse Gaussian process force fields introduced in refs. 10,49, and the SGP-based on-the-fly training methods of refs. 33, 34, extending these methods to a canonical chemically reactive system and establishing their equivalence to a simpler class of polynomial models. Our method also builds on our own previous active learning workflow, which relied on a significantly more expensive exact Gaussian process32 and was limited to two- and three-body interactions that are insufficiently descriptive for chemically reactive systems. Our active learning approach overcomes these limitations through efficient data-driven construction of the PES for the entire H/Pt system, at high fidelity to the chosen ab initio method. To our best knowledge, we perform the first large-scale direct MD simulations of reactive H2 turnover on Pt(111), capturing the explicit two-phase boundary across the molecular gas phase and fully thermalized substrate with varying H coverages. Importantly, the procedure requires no simplifications or prior assumptions about the reaction mechanisms, proceeding entirely autonomously and providing direct computational measurement of the catalytic reaction kinetics. The resulting apparent activation energy for H2 turnover is found to be in excellent agreement with surface science experiments.
Results
Active learning and acceleration of many-body force fields
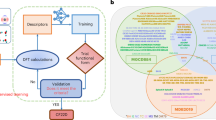
Figure 1 shows an overview of our method for autonomous training and acceleration of many-body Bayesian force fields. The on-the-fly training loop (Fig. 1a) is driven by an SGP that provides Bayesian uncertainties of model predictions. The objective is to automatically construct both the training set of the SGP and the sparse set, which is a collection of representative local atomic environments ρt that are summed over at test time to make predictions and evaluate uncertainties (see Methods). The training loop begins with a DFT calculation on an initial atomic structure, initializing the training set and serving as the first frame of the MD simulation.
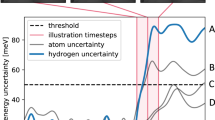
a At each time-step of the MD simulation, local energies, forces, and stresses are computed with the SGP model. If the uncertainty on a local energy exceeds the chosen threshold ΔDFT, a DFT calculation is performed and the model is updated. b Mapping of local environments ρi onto multielement descriptors derived from the atomic cluster expansion. The environment is first mapped onto an equivariant descriptor ci, products of which are used to compute the rotationally invariant descriptor di that serves as an input to the model. c Mapping of a ξ = 2 SGP force field onto an equivalent quadratic model. The prediction cost of the SGP scales linearly with the number of sparse environments NS, while the cost of the corresponding polynomial model is independent of NS.
At each time-step of the MD simulation, the SGP predicts the potential energy, forces, and stresses of the current structure and assigns Bayesian uncertainties to local energy predictions ε. These uncertainties take the form of a scaled predictive variance \({\widetilde{V}}_{\varepsilon }\) valued between 0 and 1 and are defined to be independent of the hyperparameters of the SGP kernel function (see Eq. (17), Methods). This formulation provides a robust measure of the distance between the local atomic environments ρi observed in the training simulation and the environments ρt stored in the sparse set of the SGP. If the uncertainty is below a chosen "prediction threshold” ΔDFT, the predictions of the SGP are accepted and an MD step is taken using the model forces.
When the uncertainty exceeds the prediction threshold, the simulation is halted and a DFT calculation is performed on the current structure. The computed DFT energy, forces, and stresses are then added to the training set of the SGP, and local environments ρi with uncertainty above an "update threshold” Δsparse ≤ ΔDFT are added to the sparse set. This active selection of sparse points reduces redundancy in the model by ensuring that only sufficiently novel local environments are added to the sparse set. It also helps to reduce the cost of SGP prediction, which scales linearly with the size of the sparse set (see Eq. (9), Methods). The training simulation is terminated when calls to DFT become infrequent, typically after 3–10 ps of dynamics.
The accuracy of the learned force field is in large part determined by the expressiveness of the descriptor vectors assigned to local atomic environments. Our procedure for mapping local environments ρi onto symmetry-preserving many-body descriptors di draws on the atomic cluster expansion (ACE) introduced by Drautz50 and is sketched in Fig. 1b. A rotationally equivariant descriptor ci (see Eq. (4), Methods) is first computed by passing interatomic distance vectors through a basis set of radial functions and spherical harmonics and summing over all neighboring atoms of a particular species. Rotationally invariant contractions of the tensor product ci ⊗ ci are then collected in an array di of many-body invariants, which serves as input to the SGP model. The vector di corresponds to the B2 term in the multielement atomic cluster expansion and is closely related to the SOAP descriptor51. In each of these approaches, the number of elements of di scales quadratically with the number of chemical species in the system.
Crucially, once sufficient training data have been collected, we map the trained SGP model onto an equivalent and much faster model whose prediction cost is independent of the size of the sparse set (Fig. 1c). This mapping draws on the duality in machine learning between kernel-based models on the one hand, which make comparisons with the training set at test time and are therefore “memory-based,” and linear parametric models on the other, which are polynomials of the features and do not depend explicitly on the training set once the model is trained52.
As shown in the Methods section, mean predictions of an SGP trained with a normalized dot product kernel raised to an integer power ξ can be evaluated as a polynomial of order ξ in the descriptor di. The integer ξ determines the body-order of the learned force field, which we define following ref. 53 to be the smallest integer n at which the derivative of the local energy with respect to the coordinates of any n distinct neighboring atoms vanishes. The simplest case, ξ = 1, corresponds to a model that is linear in di, and since the elements of di are sums of three-body contributions, the resulting model is formally three-body. ξ = 2 models are quadratic in di and thus five-body, with general ξ corresponding to a (2ξ + 1)-body model when using the B2 term of ACE.
We evaluate the performance of kernels with different ξ values by comparing the log marginal likelihood \({{{{{{{\mathcal{L}}}}}}}}({{{{{{{\bf{y}}}}}}}}|\xi )\) of SGP models trained on the same structures. \({{{{{{{\mathcal{L}}}}}}}}\) quantifies the probability of the training labels y given a particular choice of hyperparameters and can be used to identify hyperparameters that optimally balance model accuracy and complexity (see Eq. (19), Methods). For H/Pt(111), we find that the likelihood \({{{{{{{\mathcal{L}}}}}}}}({{{{{{{\bf{y}}}}}}}}|\xi )\) for ξ = 2 is considerably higher than for ξ = 1 but nearly the same as for ξ = 3, and decreases for ξ > 3 (Supplementary Fig. 1). We therefore choose to train five-body ξ = 2 models, allowing local energies to be evaluated as a simple vector-matrix-vector product ε(ρi) = diβdi that can be rapidly computed (see Eq. (20) for the definition of β). We have implemented this quadratic model as a custom pair-style in the LAMMPS code, which exhibits a dramatic acceleration over standard SGP mean prediction (Supplementary Fig. 4). Remarkably, the ` resulting LAMMPS model is more than twice as fast as a recent H/Pt ReaxFF model45, opening up pathways to accurate ML-driven reactive MD simulations that are more efficient than their classical counterparts.
On-the-fly training of a reactive H/Pt force field
Figure 2 presents our procedure for training a reactive H/Pt force field on the fly. We performed four independent training simulations targeting gas-phase H2, bulk and surface Pt, and H2 interaction with the Pt(111) slab (Fig. 2a). Each training was performed from scratch, without any data initially stored in the training set of the SGP. As reported in Table 1, the single-element training simulations were each completed in less than two hours of wall time on 32 CPUs, with 24 DFT calls made during the gas-phase H2 simulation and only 4 and 6 calls made during the surface and bulk Pt simulations, respectively. The majority of DFT calls were made in the reactive H/Pt(111) simulation, with 216 calls made in total during the 3.7 ps simulation, resulting in nearly 50,000 training labels in total (consisting of the potential energy, atomic forces, and six independent stress tensor components from each DFT calculation). The order-of-magnitude increase in the number of accumulated sparse environments during the H/Pt training simulation reflects the greater diversity of chemical environments present in this two-phase simulation, encompassing gaseous H2, H surface and sub-surface diffusion, and Pt surface and bulk dynamics. Potential energy predictions made during the run are plotted in Fig. 2b, showing excellent agreement between the SGP and DFT to within 1 meV/atom (see Supplementary Fig. 2 for the corresponding plots of the single-element training simulations).
a Snapshots of four independent training simulations (left to right): gas-phase H2; bulk Pt; (111) facet of the Pt surface; and H2 interaction with the Pt(111) slab. b Energy versus time during the H/Pt(111) training simulation, with the SGP predictions in green and DFT evaluations shown as black dots. The inset is a zoom-in of the model predictions during the first recombination event observed at t ≈ 0.3 ps. c Snapshots of the first recombination event. Atoms are colored by the uncertainty of the local energy prediction, with red corresponding to the prediction threshold ΔDFT.
The H/Pt(111) training was initialized with five randomly oriented H2 molecules in the gas phase and with one side of the slab at full H coverage. The temperature was set at 1500 K to facilitate sampling of rare recombinative desorption events on the surface. The first recombination event occurred at t ≈ 0.3 ps, shown as a sequence of MD snapshots in Fig. 2c. Here, each atom is colored by the uncertainty of its local energy, ranging from blue for negligible uncertainty to red corresponding to uncertainty near the prediction threshold ΔDFT. The formation of the H–H bond triggers two DFT calls (frames ii and iii), demonstrating the model’s ability to automatically detect and incorporate novel atomic environments into the training set without any prior information about the nature of the reaction.
Force field model validation and comparisons with ReaxFF
We pool together all the structures and sparse environments collected in the four independent training simulations to construct the final SGP model, which we validate extensively on a range of properties against DFT. Our objective is to obtain a model that achieves accurate prediction of not only energy, forces, and stresses during MD simulations—quantities that the model was directly trained on—but also fundamental properties of Pt, H2, and H/Pt that were not explicitly included in the training set. For bulk Pt, we predict the lattice constant to within 0.1% of the DFT value, as well as the bulk modulus and elastic constants to within 6% (see Table 2). The latter considerably improves on the recent ReaxFF force field for H/Pt45, which overestimates the C44 elastic constant by nearly 200%.
We also consider a more stringent test of model performance by forcing the model to extrapolate on structures that are significantly different from those encountered during training. In Fig. 3, we plot model predictions of bulk Pt energies as a function of volume, gas-phase H2 dissociation and dimer interaction profiles, surface energies of several Pt facets, and H adsorption energies at different binding sites. In each case, we present the 99% confidence region associated with each prediction, which we compute under the Deterministic Training Conditional approximation of the GP predictive variance (see Methods).
In each plot, shaded regions indicate 99% confidence regions of the SGP model. a Energy versus volume of bulk Pt. b Energy versus H–H bond distance of a single H2 molecule. c Energy versus intermolecular distance of two H2 molecules oriented perpendicular (blue) and parallel (green) to each other. d Surface energies of Pt for (111), (110), (100), and (211) facets. e H adsorption energies at different binding sites and Pt facets.
In general, we observe low uncertainties and excellent agreement with DFT for configurations that are well-represented in the training set. For bulk Pt, the uncertainties nearly vanish close to the equilibrium volume (Fig. 3a), which was extensively sampled in the 0 GPa training simulation of bulk Pt. The model also gives confident and accurate predictions for H2 bond lengths between ~ 0.5 and 1.2 Å (Fig. 3b) and for dimer separations above 1.8 Å (Fig. 3c). The confidence region expectedly grows for extreme bond lengths and dimer separations that were not encountered during training (see Supplementary Fig. 3 for the radial distribution function of the H2 training simulation averaged over all frames). For surface energies and H adsorption energies (Fig. 3d, e), the model agrees well with DFT for Pt(111). Surprisingly, the model is able to generalize and provide robust predictions for other surface facets that have not been included in training. The largest uncertainties are observed for H adsorption energies at (110) and (100) hollow sites, most likely due to geometric differences from the (111) binding site configurations.
Next, we validate model predictions of potential energies, atomic forces, and virial stress tensor components against DFT and compare with the ReaxFF model45 (see Supplementary Figs. 7–9 for the parity plots). The models are tested on 50 regularly spaced frames from two 500-ps trajectories that were generated independently at 1200 K using the SGP and ReaxFF models. The structure was initialized with one side of the slab at full H coverage. We first highlight that qualitatively different behaviors are generated by the two force field models. While our SGP model displays only adsorption and desorption events, the ReaxFF model gives rise to non-physical surface sublimation of isolated Pt hydride species, even at temperatures as low as 600 K (see Supplementary Fig. 14). To assess the fidelity of both models to DFT, we examine the accuracy of SGP and ReaxFF predictions on structures drawn from both MD trajectories.
Supplementary Table 1 summarizes the mean absolute errors (MAE) with respect to DFT for the three classes of properties. Overall, the SGP predictions demonstrate significantly higher fidelity to DFT than the ReaxFF predictions for both trajectories. In particular, even though the surface-evaporated structures encountered in the ReaxFF trajectory were not included in our training set, the SGP model is capable of extrapolating on these structures closer to the DFT values than the ReaxFF model. This improvement is most apparent in the potential energy predictions on the ReaxFF trajectory. ReaxFF predicts the high-energy evaporated structures to be low-energy (MAE = 93 meV/atom); the SGP model does so as well, but to a much lesser extent (MAE = 26 meV/atom), as evidenced by the cluster of data points being closer to the parity line in Supplementary Fig. 8.
We now examine full transition state pathways for H2 dissociative adsorption and atomic H diffusion on Pt(111) at the low-coverage limit (Supplementary Fig. 11). Both DFT and our SGP model predict the three binding sites (FCC hollow, HCP hollow, top) to be nearly isoenergetic (within ~ 0.05 eV of each other), with the FCC hollow site being the most stable (Table 3). In contrast, ReaxFF predicts the top site to be the most stable (by 0.16 eV compared to the HCP hollow site) and as such it was precluded from the analysis here. According to the minimum energy pathway obtained from DFT (Supplementary Fig. 10), gas-phase H2 molecule first approaches the top site, followed by dissociation into the nearest FCC hollow sites. Molecular adsorption is not favored, and as such the structure spontaneously relaxes to the fully dissociated state. The chemisorption process is barrierless and overall exothermic by 1.02 eV. Subsequent diffusion from FCC to HCP hollow site has the smallest energy barrier (0.07 eV), followed by HCP hollow to top (0.09 eV) and FCC hollow to top (0.15 eV). As summarized in Table 3, the SGP predictions of these barriers (and the associated pathways; see Supplementary Fig. 10) are in excellent agreement with DFT.
Lastly, we examine atomic H diffusion using SGP to perform MD simulations in the temperature range of 300–900 K (for details see Supplementary Figs. 12–14 in SI). The Arrhenius analysis yields an apparent activation energy of 92 meV and a diffusion prefactor of 3.53 × 1013 Å2/s (Supplementary Fig. 13), in good agreement with the experimental values of 68 meV and 1 × 1013 Å2/s from helium atom scattering measurements54. This dynamical estimate of the diffusion barrier is consistent with the static values from the transition state pathways (Table 3), in the range of 50–150 meV (SGP) and 65-149 meV (DFT).
Large-scale reactive MD simulations
The trained SGP model was mapped onto a fast polynomial model, as described in Methods, and used to perform explicit two-phase reactive MD simulations of H2 turnover on Pt(111). These large-scale simulations allow for a direct estimation of the reaction rates of H2 dissociative adsorption and recombinative desorption, resembling a realistic experimental measurement. The system was initialized with both sides of the slab at full H coverage and 80 randomly oriented H2 molecules in the gas phase, giving 864 Pt atoms and 448 H atoms in total (see Fig. 4a for an example MD snapshot). The simulations were performed at four temperatures—450, 600, 750, and 900 K—with the vertical dimension of the box frozen at 6.8 nm and with the pressure along the transverse axes set to 0 GPa. 500 ps of dynamics were simulated in total with a time-step of 0.1 fs. Reactive events were counted by monitoring the coordination number of each H atom within a 1 Å shell, which is zero for atomic H adsorbed on the surface, and one for molecular H2 in the gas phase.
a Snapshot from a 500 ps simulation at 900 K. The system is a six-layer slab model of a 12 × 12 unit cell of Pt(111) with H2 reactive events occurring at both sides of the slab. b Cumulative number of dissociation and recombination events observed at 450 K (dark blue), 600 K (light blue), 750 K (orange), and 900 K (red). The reaction rate is estimated from the slope of each curve after an initial equilibration period of 200ps. c Arrhenius plot of the reaction rate versus inverse temperature. An apparent activation energy of 0.25(2) eV is obtained, in good agreement with the experimental values of 0.21–0.24 eV55,56.
During the first ~ 100–200 ps, the recombination rate exceeds the dissociation rate as the system approaches an equilibrium surface H coverage. Higher temperatures are seen to be associated with lower equilibrium coverage values (see Supplementary Fig. 5 for a plot of the surface coverage as a function of time). Once the slab reaches an equilibrium coverage, the rates of dissociation and recombination become roughly equal (Fig. 4b). The reaction rates are estimated by performing a linear fit of the cumulative number of reactive events for the final 300 ps of each simulation. The Arrhenius plot of these rates against inverse temperature provides an apparent activation energy of 0.25(2) eV (Fig. 4c). To check the effect of pressure, the simulations were repeated with double the size of the vacuum, giving a consistent estimate of 0.20(3)eV for the apparent activation energy (Supplementary Fig. 6). Both estimates are in good agreement with the experimental values reported in the range of 0.21–0.24 eV from surface science experiments conducted at ultrahigh vacuum55, as well as ambient pressures56.
Discussion
We have developed an active learning method for autonomous, on-the-fly training of reactive force fields, achieving excellent accuracy relative to DFT and computational efficiency surpassing that of the primary traditional force field used for reactive simulations, ReaxFF. The method presented here is intended to reduce the time, effort, and expertise required to train accurate reactive force fields, making it possible to obtain a high-quality model in a matter of days with minimal human supervision. In particular, our method has enabled direct and unbiased MD simulations of H2 turnover on Pt(111) including all degrees of freedom with explicit molecular gas phase and chemisorbed H atoms, thereby overcoming the limitations of previous studies constrained to either an implicit gas phase or a single H2 molecule.
Our method bridges two of the leading approaches to ML force fields, combining the principled Bayesian uncertainties available in kernel-based approaches like GAP with the computational efficiency of parametric force fields such as SNAP11, qSNAP57, and MTP12. By simplifying the training procedure and reducing the computational cost of reactive force fields at the same time, this unified approach will help extend the range of applicability and predictive power of reactive MD as a modeling tool, providing new insights into complex systems that have so far remained out of reach. Identifying more compact descriptors for many-element systems and distributing automated training protocols across many processors are promising directions that would help extend the method presented here to systems of even greater complexity. Of particular interest are applications to biochemical reactions and more complex heterogeneous catalysis.
Methods
Here, we present our implementation of SGP force fields, our procedure for mapping them onto accelerated polynomial models, and the computational details of our training workflow, DFT calculations, MD simulations, and transition state modeling.
Sparse Gaussian process (SGP) force fields
Our SGP force fields are defined in three key steps: (i) Fixing the energy model, which involves expressing the total potential energy as a sum over local energies assigned to local atomic environments ρi, (ii) mapping these local environments onto descriptor vectors di that serve as input to the model, and (iii) choosing a kernel function k(di, dj) that quantifies the similarity between two local environments. In this section, we summarize these steps and then present our approach to making predictions, evaluating Bayesian uncertainties, and optimizing hyperparameters with the SGP.
Defining the energy model
As in the Gaussian Approximation Potential formalism10, we express the total potential energy E(r1, …, rN; s1, …sN) of a structure of N atoms as a sum over local energies ε assigned to atom-centered local environments,
The decomposition of the total energy into a sum over local, atom-centered contributions gives the model the key property of scaling linearly with the number of atoms in the system. Here ri is the position of atom i, si is its chemical species, and we define the local environment ρi of atom i to be the set of chemical species and interatomic distance vectors connecting atom i to neighboring atoms j ≠ i within a cutoff sphere of radius \({r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}\),
Note that we allow the cutoff radius \({r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}\) to depend on the central and environment species si and sj. This additional flexibility was found to be important in the H/Pt system, with H–H and H–Pt interactions requiring shorter cutoffs than Pt-Pt.
Describing local environments
To train an SGP model, local environments ρi must be mapped onto fixed-dimension descriptor vectors di that respect the physical symmetries of the potential energy surface. Different from SOAP descriptors used in GAP, we use the multielement atomic cluster expansion of Drautz50,58 to efficiently compute many-body descriptors that satisfy rotational, permutational, translational, and mirror symmetry. To satisfy rotational invariance, the descriptors are computed in a two-step process: the first step is to compute a rotationally equivariant descriptor ci of atom i by looping over the neighbors of i and passing each interatomic distance vector rij through a basis set of radial basis functions multiplied by spherical harmonics, and the second step is to transform ci into a rotationally invariant descriptor di by forming a tensor product of ci with itself and keeping only the rotationally invariant elements of the resulting tensor. In the first step, the interatomic distance vectors rij are passed through basis functions ϕnℓm of the form
where \({\tilde{r}}_{ij}=\frac{{r}_{ij}}{{r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}}\) is a scaled interatomic distance, Rn are radial basis functions defined on the interval [0, 1], Yℓm are the real spherical harmonics, and c is a cutoff function that smoothly goes to zero as the interatomic distance rij approaches the cutoff radius \({r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}\). The equivariant descriptor ci (called the "atomic base” in ref. 50) is a tensor indexed by species s, radial number n and angular numbers ℓ and m, and is computed by summing the basis functions over all neighboring atoms of a particular species s,
where \({\delta }_{s,{s}_{j}}=1\) if s = sj and 0 otherwise. Finally, in the second step, the rotationally invariant vector di is computed by invoking the sum rule of spherical harmonics,
To eliminate redundancy in the invariant descriptor, we notice that interchanging the s and n indices leaves the descriptor unchanged,
In practice we keep only the unique values, which can be visualized as the upper- or lower-triangular portion of the matrix formed from invariant contractions of the tensor product ci ⊗ ci, as shown schematically in Fig. 1(b). This gives a descriptor vector of dimension
where Nspecies is the number of species, Nrad is the number of radial functions, and \({\ell }_{\max }\) is the maximum value of ℓ in the spherical harmonics expansion. When computing descriptor values, we also compute gradients with respect to the Cartesian coordinates of each neighbor in the local environment, which are needed to evaluate forces and stresses.
The cutoff radii \({r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}\) and radial and angular basis set sizes Nrad and \({\ell }_{\max }\) are hyperparameters of the model that can be tuned to improve accuracy. We chose the Chebyshev polynomials of the first kind for the radial basis set and a simple quadratic for the cutoff,
We set the Pt-Pt cutoff to 4.25 Å and the H–H and H–Pt cutoffs to 3.0 Å, and truncated the basis expansion at Nrad = 8, \({\ell }_{\max }=3\). These hyperparameters performed favorably on a restricted training set of H/Pt structures, as determined by the log marginal likelihood of the SGP (see Supplementary Fig. 1).
Making model predictions
The SGP prediction of the local energy ε assigned to environment ρi is evaluated by performing a weighted sum of kernels between ρi and a set of representative sparse environments S,
where NS is the number of sparse environments, k is a kernel function quantifying the similarity of two local environments, and α is a vector of training coefficients. For the kernel function, we use a normalized dot product kernel raised to an integer power ξ, similar to the SOAP kernel51:
Here, d1 = ∥d1∥ and d2 = ∥d2∥. The hyperparameter σ quantifies variation in the learned local energy, and in our final trained model is set to 3.84 eV.
The training coefficients α are given by
where \({{{{{{{\boldsymbol{\Sigma }}}}}}}}={({{{{{{{{\bf{K}}}}}}}}}_{SF}{{{{{{{{\boldsymbol{\Lambda }}}}}}}}}^{-1}{{{{{{{{\bf{K}}}}}}}}}_{FS}+{{{{{{{{\bf{K}}}}}}}}}_{SS})}^{-1}\), KSF is the matrix of kernel values between the sparse set S and the training set F (with \({{{{{{{{\bf{K}}}}}}}}}_{FS}={{{{{{{{\bf{K}}}}}}}}}_{SF}^{\top }\)), KSS is the matrix of kernel values between the sparse set S and itself, Λ is a diagonal matrix of noise values quantifying the expected error associated with each training label, and y is the vector of training labels consisting of potential energies, forces, and virial stresses. For the noise values in Λ, we chose for the final trained model a force noise of σF = 0.1 eV/Å, an energy noise of σE = 50 meV, and a stress noise of σS = 0.1 GPa. In practice, we found that performing direct matrix inversion to compute Σ in Eq. (11) was numerically unstable, so we instead compute α with QR decomposition, as proposed in ref. 59.
Evaluating uncertainties
To evaluate uncertainties on total potential energies E, we compute the GP predictive variance VE under the Deterministic Training Conditional (DTC) approximation60,
Here kEE is the GP covariance between E and itself, which is computed as a sum of local energy kernels
with i and j ranging over all atoms in the structure. The row vector kES stores the GP covariances between the potential energy E and the local energies of the sparse environments, with \({{{{{{{{\bf{k}}}}}}}}}_{SE}={{{{{{{{\bf{k}}}}}}}}}_{ES}^{\top }\).
Surface energies and binding energies are linear combinations of potential energies, and their uncertainties can be obtained from a straightforward generalization of Eq. (12). Consider a quantity Q of the form Q = aE1 + bE2, where a and b are scalars and E1 and E2 are potential energies. GP covariances are bilinear, so that for instance
and as a consequence the GP predictive variance assigned to Q is obtained by replacing kEE and kES in Eq. (12) with kQQ and kQS, respectively, where
and
We use these expressions to assign confidence regions to the surface and binding energies reported in Fig. 3.
To evaluate uncertainties on local energies ε, we first compute a simplified predictive variance
Formally, Vε is the predictive variance of an exact GP trained on the local energies of the sparse environments, and it has two convenient properties: (i) it is independent of the noise hyperparameters Λ, and (ii) it is proportional to but otherwise independent of the signal variance σ2. This allows us to rescale the variance to obtain a unitless measure of uncertainty \({\widetilde{V}}_{\varepsilon }\),
Notice that \({\widetilde{V}}_{\varepsilon }\) lies between 0 and 1 and is independent of the kernel hyperparameters, providing a robust uncertainty measure on local environments that we use to guide our active learning protocol.
Optimizing hyperparameters
To optimize the hyperparameters of the SGP, we evaluate the DTC log marginal likelihood
where Nlabels is the total number of lables in the training set. Eq. (19) quantifies the likelihood of the training labels y given a particular choice of hyperparameters. The first term penalizes model complexity while the second measures the quality of the fit, and hence hyperparameters that maximize \({{{{{{{\mathcal{L}}}}}}}}\) tend to achieve a favorable balance of complexity and accuracy61. During on-the-fly runs, after each of the first ten updates to the SGP, the kernel hyperparameters σ, σE, σF, and σS are optimized with the L-BFGS algorithm by evaluating the gradient of \({{{{{{{\mathcal{L}}}}}}}}\). We also use the log marginal likelihood to evaluate different descriptor hyperparameters Nrad, \({\ell }_{\max }\), and \({r}_{{{{{{{{\rm{cut}}}}}}}}}^{({s}_{i},{s}_{j})}\) and the discrete kernel hyperparameter ξ (see Supplementary Fig. 1 for a comparison).
Mapping to an equivalent polynomial model
Letting \({\tilde{{{{{{{{\boldsymbol{d}}}}}}}}}}_{i}=\frac{{{{{{{{{\bf{d}}}}}}}}}_{i}}{{d}_{i}}\) denote the normalized descriptor of local environment ρi, we observe that with the dot product kernel defined in Eq. (10), local energy prediction can be rewritten as
where in the final two lines we have gathered all terms involving the sparse set into a symmetric tensor β of rank ξ. Once β is computed, mean predictions of the SGP can be evaluated without performing a loop over sparse points, which can considerably accelerate model predictions for small ξ. For ξ = 1, corresponding to a simple dot product kernel, mean predictions become linear in the descriptor,
Evaluating the local energy with Eq. (21) requires a single dot product rather than a dot product for each sparse environment, accelerating local energy prediction with the SGP by a factor of NS. For ξ = 2, mean predictions are quadratic in the descriptor and can be evaluated with a vector-matrix-vector product,
The cost of the matrix-vector product \({{{{{{{\boldsymbol{\beta }}}}}}}}\tilde{{{{{{{{\boldsymbol{d}}}}}}}}}\) scales quadratically with the descriptor dimension ndesc and becomes the principal bottleneck when the descriptor dimension is large, as shown in Supplementary Figs. 15 and 16.
The mapping in Eq. (22) is exact and provides an acceleration of SGP local energy prediction if the number of sparse environments exceeds the descriptor dimension, NS > ndesc, with quadratic prediction expected to be faster by a factor of \(\frac{{N}_{S}}{{n}_{{{{{{{{\rm{desc}}}}}}}}}}\). For general ξ, this ratio of efficiencies is equal to \(\frac{{N}_{S}}{{n}_{{{{{{{{\rm{desc}}}}}}}}}^{\xi }}\) and diminishes rapidly with ξ. In our H/Pt models, for which NS = 2424 and ndesc = 544, we found ξ = 2 models to give considerable improvement over ξ = 1, but found no benefit for ξ ≥ 3 (see Supplementary Table I). We therefore selected ξ = 2 for our final trained model and used quadratic prediction when performing production MD simulations, giving a theoretical speed up over SGP mean prediction of \(\frac{{N}_{S}}{{n}_{{{{{{{{\rm{desc}}}}}}}}}}\,\approx\, 4.5\). In practice we observe a speed up of greater than 10x due to better optimization of the quadratic LAMMPS model (see Supplementary Fig. 4).
Computational details
Training workflow
To calculate the ACE descriptors, train SGP models, and map SGPs onto accelerated quadratic models, we have developed the FLARE++ code, available at https://github.com/mir-group/flare_pp. On-the-fly training simulations were performed with the FLARE code32, available at https://github.com/mir-group/flare, which is coupled to the MD engine as implemented in the Atomic Simulation Environment (ASE)62 and the DFT engine as implemented in the Vienna Ab Initio Simulation Package (VASP)63.
Density functional theory (DFT)
We perform DFT calculations using plane-wave basis sets and the projector augmented-wave (PAW) method as implemented in the Vienna Ab Initio Simulation Package (VASP)63. The plane-wave kinetic energy cutoff is set at 450 eV. The Methfessel-Paxton smearing scheme is employed with a broadening value of 0.2 eV. The total energy is converged to 10−5 eV. Gas-phase H2 is optimized in a 14 × 15 × 16 Å3 cell at the Γ-point. The lattice constant of bulk face-centered cubic Pt is optimized according to the third-order Birch-Murnaghan equation of state, using a 19 × 19 × 19 k-point grid. For training and benchmark, we use a six-layer slab model of a 3 × 3 unit cell of Pt(111). The slab is spaced with 16 Å of vacuum along the direction normal to the surface in order to avoid spurious interactions between adjacent unit cells. The Brillouin zone is sampled using a Γ-centered 7 × 7 × 1 k-point grid. For force field validation (Supplementary Figs. 7–9), we use a slightly larger 5 × 5 unit cell with a 5 × 5 × 1 k-point grid.
We employ the Perdew-Burke-Ernzerhof (PBE) parametrization64 of the generalized gradient approximation (GGA) of the exchange-correlation functional. PBE provides Pt lattice constant of 3.97 Å, in good agreement with the experimental benchmark of 3.92 Å65. We also examine the dissociative adsorption energy of H2, defined as the change in energy from an isolated slab and a gas phase H2 to a combined system of atomic H adsorbed on the slab:
PBE provides H adsorption energy of − 0.52 eV at the dilute limit, within 0.16 eV of the experimental benchmark of − 0.36 eV66. Based on these observations, we conclude that PBE is an appropriate reference functional that can provide reasonable comparison with experiments for H2 chemisorption on Pt(111)67.
Molecular dynamics (MD)
We perform MD simulations using the Large-scale Atomic/Molecular Massively Parallel Simulator (LAMMPS)68 with our custom pair-style for mapped SGP force fields available in the FLARE++ repository. We also make various comparisons with an available H/Pt ReaxFF model45,69 for force field validation (Supplementary Figs. 7–9 and 14). For simulations of H2 reactivity, we use a six-layer slab model of a 12 × 12 unit cell of Pt(111). Periodic boundary condition is enforced in all Cartesian directions. The box dimension normal to the surface is fixed to retain the vacuum. A velocity-Verlet integrator is used with a time-step of δt = 0.1 fs to evolve the equations of motion. Production simulations are run for 500 ps within the isothermal-isobaric (NPT) ensemble at a temperature range of 300–900 K. Pressure and temperature are enforced on the system using a Nos\(\acute{{{{{{{{\rm{e}}}}}}}}}\)-Hoover barostat (1000 δt = 100 fs coupling) and thermostat (100 δt = 10 fs coupling), respectively.
For Arrhenius analysis of atomic H diffusion (Supplementary Figs. 12 and 13), we use a 10 × 10 unit cell of Pt(111). The bottommost layer is immobilized (velocities and forces set to zero) to prevent it from acting as a surface. All simulations consist of 10 ps equilibration within the isothermal-isobaric (NPT) ensemble, followed by 200 ps production within the canonical (NVT) ensemble at the same temperature range of 300–900 K. We take the second half of the production as a linear diffusive regime, where the mean-squared displacement (MSD) behaves linearly as a function of time. For each temperature, the MSD is averaged over 30 parallel replicas to ensure sufficient noise reduction.
Transition state modeling
We perform ab initio transition state modeling using the VASP Transition State Tools (VTST) (Supplementary Figs. 10 and 11). The bottom three layers are fixed at their bulk positions to mimic bulk properties. All initial and final states are optimized via ionic relaxation, with the total energy and forces converged to 10−5 eV and 0.02 eV/Å, respectively. Transition state pathways are first optimized via the climbing-image nudged elastic band (CI-NEB) method, using three intermediate images generated by linear interpolation with a spring constant of 5 eV/Å2. The total forces, defined as the sum of the spring force along the chain and the true force orthogonal to the chain, are converged to 0.05 eV/Å. Then, the image with the highest energy is fully optimized to a first-order saddle point via the dimer method, this time converging the total energy and forces to 10−7 eV and 0.01 eV/Å, respectively. We confirm that the normal modes of all transition states contain only one imaginary frequency by calculating the Hessian matrix within the harmonic approximation, using central differences of 0.01 Å at the same level of accuracy as the dimer method.
For comparison against DFT, the transition state pathways are also optimized with our SGP model via the CI-NEB method as implemented in LAMMPS, this time using ten intermediate images generated by linear interpolation.
Data availability
Input and output files of the molecular dynamics simulations described in this study are available at https://archive.materialscloud.org/record/2022.92.
Code availability
An open-source implementation of the FLARE++ code is available at https://github.com/mir-group/flare_pp.
References
Singh, J., Petter, R. C., Baillie, T. A. & Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 10, 307–317 (2011).
Friend, C. M. & Xu, B. Heterogeneous catalysis: a central science for a sustainable future. Acc. Chem. Res. 50, 517–521 (2017).
Van Duin, A. C., Dasgupta, S., Lorant, F. & Goddard, W. A. Reaxff: a reactive force field for hydrocarbons. J. Phys. Chem. A 105, 9396–9409 (2001).
Mihalovits, L. M., Ferenczy, G. G. & Keserű, G. M. Affinity and selectivity assessment of covalent inhibitors by free energy calculations. J. Chem. Inf. Model. 60, 6579–6594(2020).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Vanommeslaeghe, K. et al. Charmm general force field: a force field for drug-like molecules compatible with the charmm all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Senftle, T. P. et al. The reaxff reactive force-field: development, applications and future directions. NPJ Comput. Mater. 2, 1–14 (2016).
Bartók, A. P., Kermode, J., Bernstein, N. & Csányi, G. Machine learning a general-purpose interatomic potential for silicon. Phys. Rev. X. 8, 041048 (2018).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Bartók, A. P., Payne, M. C., Kondor, R. & Csányi, G. Gaussian approximation potentials: The accuracy of quantum mechanics, without the electrons. Phys. Rev. Lett. 104, 136403 (2010).
Thompson, A. P., Swiler, L. P., Trott, C. R., Foiles, S. M. & Tucker, G. J. Spectral neighbor analysis method for automated generation of quantum-accurate interatomic potentials. J. Comput. Phys. 285, 316–330 (2015).
Shapeev, A. V. Moment tensor potentials: a class of systematically improvable interatomic potentials. Multiscale Model. Simul. 14, 1153–1173 (2016).
Deringer, V. L. & Csányi, G. Machine learning based interatomic potential for amorphous carbon. Phys. Rev. B 95, 094203 (2017).
Behler, J. First principles neural network potentials for reactive simulations of large molecular and condensed systems. Angew. Chem. Int. Ed. 56, 12828–12840 (2017).
Schütt, K. T., Sauceda, H. E., Kindermans, P.-J., Tkatchenko, A. & Müller, K.-R. Schnet–a deep learning architecture for molecules and materials. J. Chem. Phys. 148, 241722 (2018).
Chmiela, S., Sauceda, H. E., Poltavsky, I., Müller, K.-R. & Tkatchenko, A. sgdml: Constructing accurate and data efficient molecular force fields using machine learning. Comput. Phys. Commun. 240, 38–45 (2019).
Deringer, V. L. et al. Origins of structural and electronic transitions in disordered silicon. Nature 589, 59–64 (2021).
Mailoa, J. P. et al. A fast neural network approach for direct covariant forces prediction in complex multi-element extended systems. Nat. Mach. Intell. 1, 471–479 (2019).
Zeng, J., Cao, L., Xu, M., Zhu, T. & Zhang, J. Z. Complex reaction processes in combustion unraveled by neural network-based molecular dynamics simulation. Nat. Commun. 11, 1–9 (2020).
Yang, M., Bonati, L., Polino, D. & Parrinello, M. Using metadynamics to build neural network potentials for reactive events: the case of urea decomposition in water. Catal. Today 387, 143–149 (2021).
Young, T. A., Johnston-Wood, T., Deringer, V. L. & Duarte, F. A transferable active-learning strategy for reactive molecular force fields. Chem. Sci. 12, 10944–10955 (2021).
Park, C. W. et al. Accurate and scalable graph neural network force field and molecular dynamics with direct force architecture. NPJ Comput. Mater. 7, 73 (2021).
Artrith, N. & Behler, J. High-dimensional neural network potentials for metal surfaces: A prototype study for copper. Phys. Rev. B 85, 045439 (2012).
Rupp, M. et al. Machine learning estimates of natural product conformational energies. PLoS Comput. Biol. 10, e1003400 (2014).
Smith, J. S., Nebgen, B., Lubbers, N., Isayev, O. & Roitberg, A. E. Less is more: Sampling chemical space with active learning. J. Chem. Phys. 148, 241733 (2018).
Zhang, L., Lin, D.-Y., Wang, H., Car, R. & Weinan, E. Active learning of uniformly accurate interatomic potentials for materials simulation. Phys. Rev. Mater. 3, 023804 (2019).
Bernstein, N., Csányi, G. & Deringer, V. L. De novo exploration and self-guided learning of potential-energy surfaces. Npj Comput. Mater. 5, 1–9 (2019).
Sivaraman, G. et al. Machine-learned interatomic potentials by active learning: amorphous and liquid hafnium dioxide. NPJ Comput. Mater. 6, 1–8 (2020).
Li, Z., Kermode, J. R. & De Vita, A. Molecular dynamics with on-the-fly machine learning of quantum-mechanical forces. Phys. Rev. Lett. 114, 096405 (2015).
Podryabinkin, E. V. & Shapeev, A. V. Active learning of linearly parametrized interatomic potentials. Comput. Mater. Sci. 140, 171–180 (2017).
Jinnouchi, R., Miwa, K., Karsai, F., Kresse, G. & Asahi, R. On-the-fly active learning of interatomic potentials for large-scale atomistic simulations. J. Phys. Chem. Lett. 11, 6946–6955 (2020).
Vandermause, J. et al. On-the-fly active learning of interpretable bayesian force fields for atomistic rare events. NPJ Comput. Mater. 6, 1–11 (2020).
Jinnouchi, R., Lahnsteiner, J., Karsai, F., Kresse, G. & Bokdam, M. Phase transitions of hybrid perovskites simulated by machine-learning force fields trained on the fly with bayesian inference. Phys. Rev. Lett. 122, 225701 (2019).
Jinnouchi, R., Karsai, F. & Kresse, G. On-the-fly machine learning force field generation: Application to melting points. Phys. Rev. B 100, 014105 (2019).
Lim, J. S. et al. Evolution of metastable structures at bimetallic surfaces from microscopy and machine-learning molecular dynamics. J. Am. Chem. Soc. 142, 15907–15916 (2020).
Xie, Y., Vandermause, J., Sun, L., Cepellotti, A. & Kozinsky, B. Bayesian force fields from active learning for simulation of inter-dimensional transformation of stanene. NPJ Comput. Mater. 7, 1–10 (2021).
Hajibabaei, A., Myung, C. W. & Kim, K. S. Towards universal sparse gaussian process potentials: Application to lithium diffusivity in superionic conducting solid electrolytes. Phys. Rev. B 103, 214102 (2021).
Plimpton, S. J. & Thompson, A. P. Computational aspects of many-body potentials. MRS Bull. 37, 513–521 (2012).
Luneau, M. et al. Guidelines to achieving high selectivity for the hydrogenation of alpha,beta-unsaturated aldehydes with bimetallic and dilute alloy catalysts: a review. Chem. Rev. 120, 12834–12872 (2020).
Pyle, D. S., Gray, E. M. & Webb, C. J. Hydrogen storage in carbon nanostructures via spillover. Int. J. Hydrog. Energy 41, 19098–19113 (2016).
Chen, B. W. J. & Mavrikakis, M. How coverage influences thermodynamic and kinetic isotope effects for H-2/D-2 dissociative adsorption on transition metals. Catal. Sci. Technol. 10, 671–689 (2020).
Kristinsdóttir, L. & Skúlason, E. A systematic DFT study of hydrogen diffusion on transition metal surfaces. Surf. Sci. 606, 1400–1404 (2012).
Greeley, J. & Mavrikakis, M. Surface and subsurface hydrogen: Adsorption properties on transition metals and near-surface alloys. J. Phys. Chem. B 109, 3460–3471 (2005).
Kraus, P. & Frank, I. On the dynamics of H-2 adsorption on the Pt(111) surface. Int. J. Quantum Chem. 117, 25407 (2017).
Gai, L., Shin, Y. K., Raju, M., van Duin, A. C. & Raman, S. Atomistic adsorption of oxygen and hydrogen on platinum catalysts by hybrid grand canonical monte carlo/reactive molecular dynamics. J. Phys. Chem. C 120, 9780–9793 (2016).
Tokumasu, T. A. Molecular dynamics study for dissociation of h2 molecule on pt (111) surface. Adv. Mat. Res. 452, 1144–1148.
Ahmed, F. et al. Comparison of reactivity on step and terrace sites of Pd(3 3 2) surface for the dissociative adsorption of hydrogen: a quantum chemical molecular dynamics study. Appl. Surf. Sci. 257, 10503–10513 (2011).
Kroes, G.-J. & Diaz, C. Quantum and classical dynamics of reactive scattering of H-2 from metal surfaces. Chem. Soc. Rev. 45, 3658–3700 (2016).
Handley, C. M., Hawe, G. I., Kell, D. B. & Popelier, P. L. Optimal construction of a fast and accurate polarisable water potential based on multipole moments trained by machine learning. Phys. Chem. Chem. Phys. 11, 6365–6376 (2009).
Drautz, R. Atomic cluster expansion for accurate and transferable interatomic potentials. Phys. Rev. B 99, 014104 (2019).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B 87, 184115 (2013).
Bishop, C. M. Pattern Recognition and Machine Learning (Springer, 2006).
Glielmo, A., Zeni, C. & De Vita, A. Efficient nonparametric n-body force fields from machine learning. Phys. Rev. B 97, 184307 (2018).
Graham, A., Menzel, A. & Toennies, J. Quasielastic helium atom scattering measurements of microscopic diffusional dynamics of H and D on the Pt(111) surface. J. Chem. Phys. 111, 1676–1685 (1999).
Christmann, K., Ertl, G. & Pignet, T. Adsorption of hydrogen on a pt (111) surface. Surf. Sci. 54, 365–392 (1976).
Montano, M., Bratlie, K., Salmeron, M. & Somorjai, G. A. Hydrogen and deuterium exchange on pt (111) and its poisoning by carbon monoxide studied by surface sensitive high-pressure techniques. J. Am. Chem. Soc. 128, 13229–13234 (2006).
Wood, M. A. & Thompson, A. P. Extending the accuracy of the snap interatomic potential form. J. Chem. Phys. 148, 241721 (2018).
Dusson, G. et al. Atomic cluster expansion: Completeness, efficiency and stability. J. Comput. Phys. 454, 110946 (2022).
Foster, L. et al. Stable and efficient gaussian process calculations. J. Mach. Learn. Res. 10, 857–882 (2009).
Quinonero-Candela, J. & Rasmussen, C. E. A unifying view of sparse approximate gaussian process regression. J. Mach. Learn. Res. 6, 1939–1959 (2005).
Bauer, M., van der Wilk, M. & Rasmussen, C. E. Understanding probabilistic sparse gaussian process approximations. (eds. Lee, D., Sugiyama, M., Luxburg, U., Guyon, I. & Garnett, R.) 30th Conference on Neural Information Processing Systems, Vol. 29 (Curran Associates, Inc., 2016).
Larsen, A. H. et al. The atomic simulation environment-a python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Kresse, G. Ab-initio molecular-dynamics for liquid-metals. J. Non-Cryst. Solids 193, 222–229 (1995).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Csonka, G. I. et al. Assessing the performance of recent density functionals for bulk solids. Phys. Rev. B 79, 155107 (2009).
Wellendorff, J. et al. A benchmark database for adsorption bond energies to transition metal surfaces and comparison to selected DFT functionals. Surf. Sci. 640, 36–44 (2015).
Gautier, S., Steinmann, S. N., Michel, C., Fleurat-Lessard, P. & Sautet, P. Molecular adsorption at Pt(111). How accurate are DFT functionals? Phys. Chem. Chem. Phys. 17, 28921–28930 (2015).
Plimpton, S. Fast parallel algorithms for short-range molecular dynamics. J. Comput. Phys. 117, 1–19 (1995).
Aktulga, H. M., Fogarty, J. C., Pandit, S. A. & Grama, A. Y. Parallel reactive molecular dynamics: numerical methods and algorithmic techniques. Parallel Comput. 38, 245–259 (2012).
Acknowledgements
We acknowledge enlightening discussions with Albert P. Bartók, Steven B. Torrisi, Lixin Sun, Matthew M. Montemore, and Philippe Sautet. We also thank David Clark, Anders Johansson, and Claudio Zeni for their contributions to the FLARE++ code. J.V. was supported by Bosch Research LLC and the National Science Foundation under Grant No. 1808162 and 2003725. Y.X. was supported by the US Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences (BES) under Award No. DE-SC0020128 (Design & Assembly of Atomically Precise Quantum Materials & Devices). J.S.L. was supported by the Integrated Mesoscale Architectures for Sustainable Catalysis (IMASC), an Energy Frontier Research Center funded by the US Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences (BES) under Award No. DE-SC0012573. C.J.O. was supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. DGE1745303. J.V., J.S.L., and C.J.O used the Cannon cluster, FAS Division of Science, Research Computing Group at Harvard University. J.S.L. and C.J.O. used the National Energy Research Scientific Computing Center (NERSC), a DOE Office of Science User Facility supported under Contract No. DE-AC02-05CH11231, through allocation m3275.
Author information
Authors and Affiliations
Contributions
J.V. designed the method, performed simulations, led development of the FLARE++ code, and implemented the mapping of the SGP model and the LAMMPS pair-style with contributions from Y.X. Y.X. implemented the on-the-fly training workflow coupled with ASE. J.S.L. performed force field validation, transition state modeling, and comparisons with ReaxFF. C.J.O. performed the Arrhenius analysis of atomic H diffusion. B.K. conceived the reactive application, contributed to algorithm development, and supervised the work. J.V. wrote the manuscript. All authors contributed to manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vandermause, J., Xie, Y., Lim, J.S. et al. Active learning of reactive Bayesian force fields applied to heterogeneous catalysis dynamics of H/Pt. Nat Commun 13, 5183 (2022). https://doi.org/10.1038/s41467-022-32294-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-32294-0
This article is cited by
-
Uncertainty driven active learning of coarse grained free energy models
npj Computational Materials (2024)
-
Machine-learning driven global optimization of surface adsorbate geometries
npj Computational Materials (2023)
-
Hyperactive learning for data-driven interatomic potentials
npj Computational Materials (2023)
-
Accurate energy barriers for catalytic reaction pathways: an automatic training protocol for machine learning force fields
npj Computational Materials (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.