Abstract
Histones control gene expression by regulating chromatin structure and function. The posttranslational modifications (PTMs) on the side chains of histones form the epigenetic landscape, which is tightly controlled by epigenetic modulator enzymes and further recognized by so-called reader domains. Histone microarrays have been widely applied to investigate histone–reader interactions, but not the transient interactions of Zn2+-dependent histone deacetylase (HDAC) eraser enzymes. Here, we synthesize hydroxamic acid-modified histone peptides and use them in femtomolar microarrays for the direct capture and detection of the four class I HDAC isozymes. Follow-up functional assays in solution provide insights into their suitability to discover HDAC substrates and inhibitors with nanomolar potency and activity in cellular assays. We conclude that similar hydroxamic acid-modified histone peptide microarrays and libraries could find broad application to identify class I HDAC isozyme-specific substrates and facilitate the development of isozyme-selective HDAC inhibitors and probes.
Similar content being viewed by others
Introduction
The human genetic code is stored in nucleosome particles, supramolecular assemblies of DNA wrapped around histone octamers, which comprise two copies of each of the core histone proteins H2A, H2B, H3, and H41. Epigenetic marks in the form of posttranslational modifications (PTMs) to histone amino acid side chains critically affect chromatin structure and ultimately gene expression2,3. Aberrant gene expression is directly associated with cancer, autoimmune deficiencies, and neurodegenerative diseases and may be modulated by altering the epigenetic state of the cell2,3,4,5. This epigenetic state is highly dynamic and responds to a variety of external stimuli of the cell such as nutrient composition and stress, but also at a systemic level to diet, lifestyle, environment, and, importantly, pharmaceutical intervention2,4. Therefore, a detailed understanding of the enzymes that shape the histone PTM landscape is of great interest. Peptide microarrays have been widely employed for the characterization of histone reader domains as well as histone-modifying enzymes and antibodies6,7,8,9,10,11,12,13. Also, chemical protein and peptide synthesis have facilitated the incorporation of modified amino acids to decipher how specific histone PTMs (e.g., acetylation, methylation, phosphorylation, ubiquitylation, and citrullination) affect recognition and chromatin structure at the molecular level14. One component of this complex, dynamic regulatory machinery is the reversible acetylation of the ε-amino group of lysine residues, which is regulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs 1–11; Fig. 1a)15,16. Further, the resulting ε-N-acetyllysine (Kac) mark is specifically recognized through bromodomains, constituting an important class of reader domains5. Peptide microarray strategies have proven valuable for providing binding affinities of large numbers of peptides17,18, and have especially proven efficient for interrogating the specificity of epigenetic reader domains6,7,9,11,12. Compared to HATs and bromodomains8,9,10,13, however, the application of peptide microarrays to the investigation of HDACs has remained more challenging19,20,21,22. The fast turnover of acetylated substrates and the resulting transient HDAC–substrate interaction necessitated coupling of the conventional peptide microarray approach with a secondary readout, such as radioactive isotopes23, fluorescence tags24,25, immunodetection of the resulting acetylation states21,22,26,27, or mass spectrometry19,20,28,29. Despite these efforts, the understanding of the inhibitor and substrate selectivity of the human Zn2+-dependent HDAC isoforms is still incomplete. This is particularly true for class I HDACs (HDACs 1‒3 and 8, Fig. 1a), which are the HDAC isozymes primarily targeted by clinically-used drugs30,31 and responsible for histone deacetylation16.
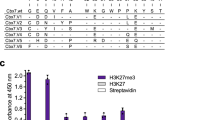
a Phylogenetic relationship of the 11 human Zn2+-dependent HDACs. b X-ray co-crystal structures showing the binding pocket of HDAC8 interacting with a Kac-containing peptide substrate or a hydroxamic acid. In contrast to Kac (left, PDB: 5DIC, Y306F mutation required for deactivation), hydroxamic acids (right, PDB: 3EW8) stall the transient interaction by chelating the Zn2+ ion in the active site. c Workflow. Peptide microarrays are prepared using the appropriate hydroxamic acid-containing aminosuberic acid analog (Asuha). Resulting arrays allow for the direct assessment of HDAC isoform-specific binding. Kme1, Kme2, Kme3: ε-N-mono-, di-, and trimethyllysine; pS: O-phosphoserine; pT: O-phosphothreonine; Kac: ε-N-acetyllysine.
Here, we report the synthesis and application of peptide microarrays that circumvent the aforementioned limitations by the introduction of hydroxamic acid-containing residues in place of the Kac residues (Fig. 1b)32,33,34, which thereby provide a platform to investigate HDAC binding preferences in high-throughput (Fig. 1c). Complementary functional assays evaluate the ability of this approach to predict HDAC binding, inhibition, and activity.
Results
Modified peptides enable interrogation of HDAC binding in microarray format
Conventional display of acetylated peptides in microarray format does not allow studying HDAC binding due to the fast turnover and resulting transient HDAC‒peptide interaction. We envisioned that modifying acetylated peptides similar to common competitive inhibitors of class I HDACs could enable the capture and detection of HDACs in microarray format. The cap group of such inhibitors interacts with the enzyme surface close to the catalytic pocket and the linker projects a zinc-binding group through a channel to the Zn2+ ion in the active site (Fig. 2a)31,32. Consequently, the turnover of Kac-containing peptides by Zn2+-dependent HDACs can be inhibited by entities where the Kac has been substituted with zinc-binding groups, and peptides containing the hydroxamic acid32,33,34 or the o-aminoanilide35 functionalities have been shown to bind and/or capture HDAC enzymes. Here, we explored the use of these moieties in histone tail peptide microarrays to interrogate the binding of class I HDACs. Modified peptide microarrays were produced using µSPOT, a variant of the widely applied SPOT method36,37,38. We found that replacing lysine residues in histone sequences by l-2-amino-8-(hydroxyamino)-8-oxooctanoic acid (Asuha) or l-2-amino-8-((2-aminophenyl)amino)-8-oxooctanoic acid (Asuapa) but not l-2-aminosuberic acid (Asu) or acetylated lysine (Kac) efficiently captured the HDAC–substrate interaction (Fig. 2b). Both Asuha and Asuapa provided sequence-dependent binding results, and Asuha was selected for further investigations due to the higher HDAC retention shown by the corresponding peptides. The length of the histone-based peptides was chosen to be 15 residues, because the synthetic efficiency of the µSPOT method decreases substantially above the 15–16-mer length. Our findings were in line with the observation that class I-selective o-aminoanilides, such as the approved drug chidamide and the clinical trial candidate entinostat, are among the most potent inhibitors, only surpassed by less selective hydroxamic acid-containing compounds such as panobinostat31.
a Approved drugs and examples of candidates in clinical trials that target class I HDACs31. Shared features include a cap group, a hydrophobic linker region, and zinc-binding group. b Display and probing of modified 15-mer histone peptides for capture of HDAC3 (10 nM) on microarray slides. Incorporation of a 2-aminosuberic acid hydroxamic acid derivative (Asuha) or a 2-aminosuberic acid o-aminoanilide derivative (Asuapa) but not the carboxylic acid (Asu) allow probing of HDAC binding in microarray format (n = 2 independent experiments). Source data are provided as a Source Data file. *Saturated chemiluminescence signal.
We next envisioned that hydroxamic acid-modified peptide microarrays would enable the direct and simple interrogation of HDAC isozyme binding and, potentially, facilitate the development of isozyme-selective inhibitors and probes (Fig. 1c). The implementation of this technique required an improved synthetic route for the Asuha building block. Modification of an acetamidomalonate alkylation/enzymatic resolution strategy39,40 provided an affordable and scalable synthetic route for Fmoc-Asuha(tBu)-OH (2 g, 20% overall yield, Supplementary Fig. 1) compared to previously reported syntheses33,34,41. Hydroxamic acid-modified µSPOT arrays were then produced and successful synthesis of the modified peptides was validated by LC-MS analysis of a subset of crude peptides prior to printing of the microarrays (Supplementary Table 1). Unfortunately, the crude peptides could not be triturated before chromatography due to the minute amounts produced and the resulting HPLC traces, therefore, included protecting group byproducts. Nevertheless, an average purity of 41%, corresponding to average coupling efficiency of at least 95%, was observed, which is in line with standard SPOT42,43 and µSPOT44 peptide synthesis. The detection of a dilution series of His-tagged HDAC2 spotted on a µSPOT slide indicated a linear correlation between signal and spotted amount (Supplementary Fig. 2, r2 = 0.96) when a horseradish peroxidase (HRP)-conjugated anti-His antibody was used. Thus, taken together, the µSPOT method allowed for economic production of hundreds of microarray copies from a single round of synthesis and thereby facilitated the semi-quantitative assessment of HDAC isozyme–peptide interactions (Fig. 1c)37,38.
Microarray-based screening and fine mapping of class I HDAC interactions
Next, we explored the use of hydroxamic acid-modified microarrays to study class I HDAC substrate recognition. We displayed a library of 291 histone-based peptides (16 amino acids in length and with a 3-residue frameshift) with the Asuha building block at the position of one lysine residue at a time, in order to comprehensively and systematically study recognition of the four core histones by HDACs 1‒3 and 8 (Supplementary Table 2). Microarray copies were titrated with recombinant human His-tagged class I HDACs and binding was detected with an anti-His antibody as before.
The recapitulation of known binding sites demonstrated the robustness of our platform. As expected, only peptides including Asuha residues were able to enrich HDACs. At the same time, we observed striking differences in isozyme affinity and selectivity (Fig. 3). HDACs 1, 2, and 8 displayed high sequence specificity, in agreement with previous substrate selectivity studies on HDAC828,45, and also similar to HDAC622. In contrast, HDAC3 bound more promiscuously and to sequences located both at the core and the tails of the histones. This might indicate that HDAC3 is less sensitive to the peptide sequence, as also reported for short peptide substrates46,47 and acetylated peptide microarrays20. Sequence logo plots were generated to highlight the amino acid preferences of each HDAC (Supplementary Fig. 3). These plots revealed an overall preference for Arg and Lys residues at the positions flanking the Asuha residue for all class I HDACs, which was previously observed for HDAC828 and, to some extent, also HDAC320. Interestingly, HDAC6 shows the opposite preference22. Further class I HDAC selectivity was observed for sequences containing Gly at the -2 and -3 positions, and Arg and Lys at the −4, +3, and +4 positions. Not surprisingly, histone tails are rich in Arg, Lys, and Gly residues, which could explain the obtained sequence preference. In this regard, we expect that future evaluation of potential isozyme-specific preferences would benefit from the use of randomized peptide arrays.
Heat map representation of relative HDAC binding affinity for Asuha (X)-containing histone peptides. The pEC50 values represent the concentration of HDAC enzyme required for half-maximal binding signal (n = 4 independent dose-response experiments). White squares represent weak binders (pEC50 < 7.6). N.D. = not determined (non-sigmoidal data). Source data are provided as a Source Data file. *HDAC3 tested in combination with the deacetylase activating domain (DAD) of NCoR2. †Matrix steric effects might impede protein binding to these peptides.
The microarrays were also applied for mapping binding sites with semi-quantitative estimation of binding affinities. This was outlined for histone H4 lysine 12 (H4K12) (Fig. 4a). Titration of single amino acid frame-shifted peptides and single amino acid truncation variants suggested exact binding sites and indicated that the C-terminus of the selected peptide is required for binding to HDACs 1, 2, and 8 (Supplementary Figs. 4‒7). Peptides 1a‒5a were then synthesized, purified, and tested as HDAC inhibitors to validate these observations. Peptides 4a and 5a showed 10-fold lower potency than the parent peptide (1a), as suggested by the data on HDACs 2 and 8, but not HDAC1 (Fig. 4c and Supplementary Fig. 8). More strikingly, peptides 2a and 3a with C-terminal truncations retained potency against all HDACs. This could indicate that particularly HDACs 1, 2, and 8 are sensitive to the steric effect of the microarray matrix and struggle to bind to Asuha residues closer than 5 amino acids to the C-terminal peptide conjugation site. On the other hand, HDAC3 was not affected by this effect, which could also explain the higher rate of positive results for this isozyme. Overall, the mapping of the H4K12 site revealed a complex sequence requirement for HDACs 1, 2, and 8, especially for residues within ±7 positions relative to the acetylated lysine. This mapping highlighted the importance of steric effects introduced by the immobilization and thus the importance to validate results with resynthesized peptides in solution.
a Fine mapping of relative HDAC1 binding affinities in microarray format (data represent mean ± SEM, n = 4 independent experiments). See Supplementary Figs. 4‒7 for complete data sets for HDACs 1‒3 and 8. X: Asuha. Source data are provided as a Source Data file. b Resynthesized peptide sequences including the parent peptide (1a), two frame-shifted sequences (2a, 4a), a C-truncated sequence (3a), and a N-truncated sequence (5a). c Comparison of the inhibitory curves of resynthesized peptides 1a‒5a with their corresponding microarray binding curves against HDAC1 (microarray data represent mean ± SEM, n = 4 independent experiments; inhibition data represent n = 2 independent experiments). See Supplementary Fig. 8 for data on HDACs 2, 3 and 8 and for half-maximal inhibition concentrations. Source data are provided as a Source Data file.
Microarray-based study of the binding effects of posttranslational modifications
The action of epigenetic modulators, including HDACs, is orchestrated by PTMs48. We next applied our microarray approach to resolve how specific PTM combinations control isozyme-specific class I HDAC function. To probe this, we selected two H3 N-terminal peptides for modification in a combinatorial manner according to reported histone PTMs (Supplementary Table 4)48,49. The included modifications were O-phosphoserine (pS), O-phosphothreonine (pT), ε-N-mono-, di-, and trimethyllysine (Kme1, Kme2, and Kme3, respectively), Kac, and ε-N-thioacetyllysine (Kthioac), which is a Kac analog with enhanced stability towards hydrolases50,51 that is only cleaved to some extent by HDACs 6 and 852 and class I sirtuins53. Remarkably, the microarray approach resolved complex and highly isozyme-specific binding profiles (Fig. 5a). Importantly, binding of HDACs to H3 peptides was diminished by phosphorylation of residues Ser10 or Thr11. Conversely, enrichment of HDACs 1, 2, and 8 was improved by modification of the Lys4 and Thr6 residues. Modifications at Lys23 also diminished HDAC8 binding, although its proximity to the C-terminus makes it difficult to interpret this trend due to the aforementioned matrix steric effects. Notably, the pronounced effect of phosphorylation of H3S10 in the microarray may mirror an altered recognition by epigenetic enzymes shown to play a pivotal role in the histone modification cross-talk54,55. To further validate the prediction of HDAC isozyme substrate recognition and selectivity, we studied deacetylation of resynthesized peptides modified at the Lys4 and Ser10 residues. Thus, H3 sequences were prepared as Kac-modified peptides with and without thioacetylation and phosphorylation by standard Fmoc/tBu SPPS (Fig. 5b) and were incubated with recombinant class I HDACs. Substrate conversion was followed by LC–MS analysis, which revealed that Ser10 phosphorylation, as predicted by the microarray, decreased the deacetylation by HDACs, whereas Lys4 thioacetylation did not promote conversion by HDACs 1, 2, and 8 (Fig. 5b and Supplementary Fig. 9). Taken together, our results highlight the applicability of hydroxamic acid-modified microarrays to interrogate changes in HDAC substrate that give rise to substantially altered recognition in high-throughput. In particular, we demonstrated that these microarrays may help identify individual positions and specific PTMs that critically alter peptide binding affinity and isozyme selectivity; thus, providing a useful tool to study HDAC function and its regulation.
a Heat map depicting effects of neighboring chemical modifications on the affinity of peptides H3(1‒16)K9Asuha and H3(10‒25)K18Asuha. p: phosphorylation, ac: acetylation, thioac: thioacetylation, me1: monomethylation, me2: demethylation, me3: trimethylation. Combinations of 4, 5, and 6 modifications abolished binding to H3(1‒16)K9Asuha peptides and have not been included. The pEC50 values represent the concentration of HDAC enzyme required for half-maximal binding signal, n = 4 independent experiments). White squares represent weak binders (pEC50 < 7.2). Source data are provided as a Source Data file. b Resynthesized peptide sequences with Kac at the position of study, and relative deacetylation by class I HDACs. Enzyme and substrate were incubated for 1 h at 37 °C, and relative conversion was determined by LCMS (see Supplementary Fig. 9 for sample assay traces). Microarray data represent mean ± SEM, n = 4 independent experiments; conversion data represent mean ± SD, n ≥ 2 independent experiments. *HDAC3 tested in combination with the DAD domain of NCoR2. †Matrix steric effects might contribute to the effects at these positions.
Identification of potent HDAC inhibitors with tailored isozyme selectivity
Multiple natural and synthetic inhibitors of class I HDACs (HDACs 1‒3 and 8) have been reported, some of which are approved for clinical use30,31. The majority are small molecules bearing a hydroxamic acid moiety, which exhibit limited isozyme selectivity. Microarray screening (Figs. 3 and 5) identified modified peptides with potential isozyme selectivity and suggested that even small chemical modifications can exert strong and isozyme-selective effects on HDAC recognition. To validate the reported selectivity and to quantify their inhibitory potency towards class I HDACs, we resynthesized additional nine hydroxamic acid-containing peptides (9a–17a) from the initial screen and evaluated HDAC inhibition in an in vitro fluorogenic assay56,57. All tested peptides displayed robust nanomolar potency against HDACs 1‒3, and also against HDAC8 in the case of peptides 6a, 13a, and 14a (Fig. 6a and Supplementary Fig. 10). As expected, control peptides with a lysine residue instead of Asuha did not inhibit any of the tested HDACs, even at high micromolar concentration.
a Dose-response HDAC inhibition curves for Asuha (X)-containing peptides (9a‒13a), the HDAC inhibitor SAHA and lysine-containing controls (9b‒13b). See Supplementary Fig. 10 for curves corresponding to 6a, 14a‒17a, and corresponding controls. b Sequence of resynthesized peptides. c Dose-response curves for peptides 11a, 13a, 14a, and 16a. d Summary of microarray screening data and the determined potency of resynthesized Asuha-containing peptides in solution. See Supplementary Fig. 12 for visual representation. *HDAC3 tested in combination with the DAD domain of NCoR2. All microarray data represent mean ± SEM, n = 4 independent experiments; all inhibition data represent n = 2 independent experiments. Source data are provided as a Source Data file.
The determined inhibitor profiles confirm the potency of multiple hits and validate the concept that peptide scaffolds can achieve isozyme selectivity, but they also reveal that potent inhibitors may be overlooked during microarray screening. Sequences 11a, 12a, and 13a exhibited selectivity for HDAC3 in the microarray (Fig. 3). For the resynthesized peptides, we found ~4-fold, ~2-fold, and >8-fold selectivity for HDAC3 vs. HDACs 1 and 2, respectively (Fig. 6c, d), which supports the use of our screening technique to discover selective lead inhibitors. Peptide 11a was also >60-fold selective for HDAC3 vs. HDAC8, whereas peptide 13a inhibited HDAC8 potently, which was not anticipated. Interestingly, peptides 9a and 16a displayed over 100-fold selectivity against HDACs 1‒3 versus HDAC8 (Fig. 6c, d), which is uncommon among hydroxamic acid-containing inhibitors57. Conversely, peptide 14a inhibited all four isozymes with similar potency, in agreement with the microarray data (Figs. 3 and 6d). The cytosolic isozyme HDAC6 was also included in the profiling because it is inhibited by several known hydroxamic acid-containing HDAC probes57. Peptides 1a and 15a were the most and least potent inhibitors against this isozyme, respectively (1a: Ki = 5.31 ± 0.03 nM, 15a: Ki = 114 ± 2 nM, Supplementary Fig. 11a). As expected, the inhibitors were highly selective over class IIa HDACs, represented by HDAC4, which was only partially inhibited at inhibitor concentrations of up to 50 µM (Supplementary Fig. 11b).
Though a number of the potencies recorded for resynthesized peptides recapitulated the microarray findings, the potencies of a number of the weaker binders were not estimated accurately in the microarray screening, thereby highlighting the need for detailed follow-up validation when applying this technology (Supplementary Fig. 12). Peptides 13a and 17a, which appeared to exhibit preference for HDACs 2 and 3 over HDAC1 in the array experiments, were both equipotent across all three recombinant HDACs 1–3, when tested in solution. This highlighted once more the importance of sufficient C-terminal spacing in microarray format, as both peptides included the Asuha residue within 5 amino acid positions of the linker. In addition, peptide 11a, which was among the most potent inhibitors of HDACs 1 and 2, did not show tight binding on the array for these two isozymes. Nevertheless, microarray screening enabled the discovery of potent peptide scaffolds with distinct profiles of HDAC inhibition. Additionally, our data confirmed a major contribution of the peptide sequence to substrate turnover, inhibitor potency, and isozyme selectivity (Fig. 6d and Supplementary Fig. 13), indicating that peptide scaffolds are an appropriate source for isozyme selective inhibitors and affinity probes.
Hydroxamic acid-modified peptides exhibit activity in cells
Next, we aimed to explore the cellular activity of selected HDAC inhibitors. We chose peptides 1a and 13a, which exhibit nanomolar potency against class I HDACs and HDAC6 and, at the same time, harbor multiple basic residues that could facilitate cell permeation. HEK293T cells were treated with the peptides for 5 h, and subsequent cell lysis and Western blot analysis revealed upregulation of acetylation of both tubulin and histones compared to DMSO control (Fig. 7a). These results strongly indicate that peptides 1a and 13a effectively inhibit the cytosolic HDAC6 and the nuclear HDACs 1‒3 in the cell, which is in agreement with their in vitro inhibition profiles against recombinant HDACs15. In particular, peptide 13a induced an 8-fold increase in tubulin acetylation and a 4-fold increase in H3K36 acetylation (Fig. 7b). Thus, the high-throughput screening and subsequent evaluation in solution delivered cellularly active HDAC probes.
a Sample Western blots on tubulin acetylation (Ac-α-tubulin) and histone acetylation (H3K27, H3K36) upon 5 h treatment with peptides 1a and 13a at 10 µM concentration, with SAHA as positive control (Ac-α-tubulin: 1 µM; H3K27ac, H3K36ac: 5 µM). See Supplementary Fig. 14 for full blots of all replicates. b Quantification of acetylation, normalized to vinculin loading control, and relative to DMSO (data represent mean ± SD, n = 4 biologically independent samples). Source data are provided as a Source Data file.
Discussion
Peptide microarrays are broadly applied to study writers and readers of histone modifications6,7,9,11,12, especially bromodomains8,9,10,13. Their application to erasers of histone acetylation, however, has remained limited to activity-based approaches, as microarray binding experiments required secondary readouts19,20,21,22,23,24,25,26,27,28,29. In addition, activity-based approaches rely on antibodies specific for a certain peptide modification, which can sometimes lead to biases induced by the sequence context. Here, we describe a facile, cost-effective, and scalable protocol for the preparation of the necessary hydroxamic acid building block and demonstrate its use to produce hydroxamic acid-modified peptide microarrays. These arrays allow the simple and direct profiling of HDAC binding and inhibition with minimal sample consumption and with the use of extensively validated tag-specific antibodies. When used systematically, similar arrays could provide large data sets and answer fundamental questions related to HDAC action and its regulation. Here, we demonstrated the use of this format to identify individual positions and modifications that determine isozyme-specific binding, and to the discovery of cellularly active peptides with tailored HDAC selectivity.
Hydroxamic acid-modified peptides have been used as substrate mimics and were shown to recapitulate important aspects of HDAC substrate recognition32,33,34,58. Here, we systematically explored the sequence and PTM requirements for the recognition of histone tails, as represented by Asuha-modified peptides, by HDACs 1, 2, 3, and 8. Our extensive dataset will enable future in-depth investigation of specific PTM effects on HDAC activity. These follow-up studies would benefit from the use of synthetic histones and reconstituted nucleosomes, which can reveal sequence effects beyond the length limitations of synthetic peptides45,58.
Selected inhibitor candidates displayed nanomolar potency and distinct selectivity profiles. Yet, the binding hierarchies in the screens did not represent the exact inhibitor selectivity profiles defined by more accurate in-solution assays. The potency of peptides with the Asuha residue close to the C-terminus was not well predicted, likely due to steric interference of the microarray matrix, and a number of additional potent inhibitors were overlooked by microarray screening. Kinetic insight from peptide macrocycles59 and an X-ray co-crystal structure of HDAC1 with a hydroxamate-containing H4-based peptide32 suggest that Asuha-containing peptides bind in the active site. Nonetheless, future studies should explore to what extent HDAC substrate turnover correlates with inhibitor potency or selectivity. This question could be addressed using the approach presented herein, together with alternative modifications such as the here described Asuapa building block. Such studies should take into account isozyme-specific effects, kinetic parameters, and contributions from additional binding partners60. Moreover, incubation with recombinant HDACs bearing active site mutations or co-incubation with validated HDAC probes could improve the selection of competitive inhibitor candidates from the array screening.
Selected microarray hits proved to penetrate the cell membrane and elicit hyperacetylation of class I and class IIb HDAC targets. Thus, combined with structural optimization such as stapling and/or N-methylation61,62, promising peptide-based HDAC probes could be afforded. The approach presented here could also be applied to the study class IIa HDACs, taking advantage of trifluoromethyl ketone-containing building blocks63, and deacetylase enzymes from other species64,65,66,67. The HDAC probes here reported may be the first of a new class of versatile tools for investigating class I HDACs and, importantly, also their native protein complexes33.
Further, the employed principle may be applied to alternative peptide microarray types including high-density lithography68 and laser-based on-chip synthesis approaches69,70, thereby further increasing the throughput. In addition, alternative binding detection methods could improve the dynamic range of the screening assay and potentially enhance the predictive value of the array18. Probing such microarrays with cellular lysates, using labeled HDACs and complex partners or highly specific antibodies, may provide a direct readout for the substrate-binding requirements of the native complexes16. Combination of such readout with different cells and tissues, as well as with perturbed HDAC levels, could greatly improve our current understanding of HDAC action in vivo.
Finally, we envision that the current approach can facilitate the development of selective chemoproteomic probes that would complement the already applied unspecific HDAC-targeting probes71,72,73. We expect that this work will enable the discovery, characterization, and selective targeting of cellularly active and, potentially, therapeutically relevant multiprotein complexes formed by the different HDAC isoforms16.
Methods
Chemical Synthesis
General synthetic methods, experimental procedures, and compound characterization can be found in the Supplementary Methods.
Assay Materials
Assays were performed in 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) buffer [50 mM HEPES/Na, 100 mM KCl, 0.001% (v/v) tween-20, 0.2 mM tris(2-carboxyethyl)phosphine (TCEP), 0.5 mg/mL bovine serum albumin (BSA), pH 7.4] unless otherwise stated. Recombinant HDAC enzymes employed in µSPOT, inhibitor, and substrate assays were from commercial sources: HDAC1 (full length, C-terminal His tag, C-terminal FLAG tag, BPS Bioscience, cat. #: 50051, lots 170105-1 and 181108-1, purity ≥ 79%, activity ≥ 460 pmol/min/µg), HDAC2 (full length, C-terminal His tag, BPS Bioscience, cat. #: 50002, lots 160701 and 160630, purity ≥ 84%, activity ≥ 675 pmol/min/µg), HDAC3/NCoR2 (full length, C-terminal His tag, NCoR2 N-terminal GST tag, BPS Bioscience, cat. #: 50003, lots 130819 and 190327, purity ≥ 80%, activity ≥ 3000 pmol/min/µg), HDAC8 (full length, C-terminal His tag, BPS Bioscience, cat. #: 50008, lots 150714 and 161216, purity ≥ 90%, activity ≥ 300 pmol/min/µg), HDAC4 (aa 627‒1084, BPS Bioscience, cat. #: 50004, lot 130828-G, purity ≥ 89%, activity ≥ 103225 pmol/min/µg), HDAC6 (full length, BPS Bioscience, cat. #: 50056, lot 151130-C, purity ≥ 88%, activity ≥ 150 pmol/min/µg). Antibodies: 6x-His tag monoclonal antibody (HIS.H8), HRP-conjugated (ThermoFisher, cat. #: MA1-21315-HRP).
µSPOT Microarray Preparation
Peptides were synthesized using a ResPep SL synthesis robot equipped with a CelluSPOT synthesis module (Intavis AG, Germany). Parallel synthesis was carried out on acid-soluble Fmoc-β-Ala-etherified cellulose disks (area 0.12 cm2, loading 1.0 μmol/cm2, Intavis AG), by Fmoc/tBu peptide synthesis on cellulose support (SPOT synthesis). The following protected amino acids were employed: Fmoc-Ala-OH, Fmoc-Arg(Pbf)-OH, Fmoc-Asu(tBu)-OH39, Fmoc-Asuapa(Boc)-OH (S8), Fmoc-Asuha(tBu)-OH (S5), Fmoc-Asn(Trt)-OH, Fmoc-Asp(OtBu)-OH, Fmoc-Cys(Trt)-OH, Fmoc-Gln(Trt)-OH, Fmoc-Glu(tBu)-OH, Fmoc-Gly-OH, Fmoc-His(Trt)-OH, Fmoc-Ile-OH, Fmoc-Leu-OH, Fmoc-Lys(Ac)-OH, Fmoc-Lys(Me,Boc)-OH, Fmoc-Lys(Me)2-OH, Fmoc-Lys(Me)3-OH, Fmoc-Lys(thioac)-OH, Fmoc-Lys(Boc)-OH, Fmoc-Pro-OH, Fmoc-Ser(tBu)-OH, Fmoc-Ser(PO3BzlH)-OH, Fmoc-Thr(tBu)-OH, Fmoc-Thr(PO3BzlH)-OH, Fmoc-Trp(Boc)-OH, Fmoc-Tyr(tBu)-OH and Fmoc-Val-OH. Peptide elongation was achieved by double coupling in DMF (20 and 15 min) with Fmoc-Xaa-OH/Oxyma/DIC (1.0:1.1:1.5 equiv) in at least 2-fold excess, followed by capping with acetic anhydride. Functionalized cellulose disks were transferred to 1.2 mL 96-tube racks (Intavis AG, cat. # 32.199) and shaken with TFA–CH2Cl2–H2O–iPr3SiH, [150 µL, 80:12:5:3 (v/v)] for 2 h at room temperature. Solutions were then removed, and disks were suspended in TFA–CF3SO3H–H2O–iPr3SiH [250 µL, 88.5:4:2.5:5 (v/v)] and shaken overnight at room temperature. Thereafter, solutions were triturated with ice-cold diethyl ether (700 µL), stored at −20 °C for 2 h and centrifuged at 1000 rcf (45 min, 4 °C). Supernatants were discarded, and pellets were rinsed with diethyl ether in the same manner. Then, 96-tube racks were ventilated for 2.5 h to allow for residual diethyl ether to evaporate, and pellets were dissolved in DMSO (500 µL) and stored at −20 °C. µSPOT microarrays were prepared by printing DMSO Stock/SSC buffer mixtures (30 nL, 2:1, saline sodium citrate buffer: 15 mM sodium citrate, 0.1 M NaCl, pH 7) in duplicate on polystyrene-coated microscope glass slides (Intavis AG, cat. # 54.112) with a Slide Spotter Robot (Intavis AG).
Analysis of Cleavable µSPOT Peptides
Disks containing Rink amide linkers were incubated in 1.5 mL plastic tubes with TFA–2,2′-(ethylenedioxy)diethanethiol (DODT)–H2O–iPr3SiH [250 µL, 92.5:2.5:2.5:2.5 (v/v)] for 2 h at room temperature. Solutions were transferred to new plastic tubes, triturated with ice-cold diethyl ether (750 µL), stored at –20 °C for 1 h and centrifuged at 17968 g (30 min, 4 °C). Thereafter, supernatants were discarded and pellets were rinsed again with ice-cold diethyl ether (750 µL). Plastic tubes were ventilated to allow for pellets to dry, and these were suspended in CH3CN/H2O [50 µL, 50:50 with 0.1% TFA (v/v)]. Analysis was performed by MALDI (1 µL), UPLC-MS (5 µL) and HPLC (15 µL).
Microarray Spotting of HDACs 2 and 3 and Assay Linearity Test
His-tagged HDAC2 and 3 were used at a concentration of 41.4 µM (in 40 mM Tris-HCl pH 8.0, 110 mM NaCl, 2.2 mM KCl, 0.04% (v/v) Tween-20, 20% (v/v) glycerol) and 7.5 µM (in 40 mM HEPES pH 8.0, 110 mM NaCl, 2.2 mM KCl, 20% (v/v) glycerol) respectively. Proteins were diluted to a concentration of 1 µM in a buffer consisting of 2/3 (v/v) DMSO and 1/3 (v/v) saline-sodium citrate (SSC) buffer (150 mM NaCl, 15 mM trisodium citrate, pH 7.0). A 1:2 (v/v) serial dilution of HDAC2 and 3 was prepared in a 384-well plate using a buffer of 2/3 DMSO and 1/3 SSC. Protein solutions were transferred to polystyrene-coated microscope glass slides (Intavis AG, cat. # 54.112) with a Slide Spotter Robot (Intavis AG). Slides were incubated in PBS with 2% (w/v) IGG-free BSA (2.5 mL; 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7.4) for 60 min at room temperature. Then, slides were incubated at room temperature with Anti-6x-His tag antibody (ThermoFisher Scientific, MA121315, Lot.: UF283577) in PBS with 0.1% BSA (1:5000, 2.5 mL, 15 min), washed with PBS (2.5 mL, 3 × 3 min), incubated with HRP-conjugated goat anti-mouse antibody (ThermoFisher Scientific, 31430, Lot: UJ293428) in PBS with 0.1% BSA (1:5000, 2.5 mL, 30 min), and washed with PBS (2.5 mL, 3 × 3 min). SuperSignal West Femto Maximum Sensitive Substrate was added to the slides (200 µL; ThermoFisher Scientific), and chemiluminescence signal was recorded in a c400 imaging system (Azure, Lowest Sensitivity, 5 min exposure). Binding intensities were evaluated using FIJI including the Microarray Profile addon (OptiNav). The error range and standard deviation were defined by comparing the intensities of two individual spots on the array.
Recombinant HDAC µSPOT Assays
µSPOT slides were incubated in HEPES buffer containing 10 mg/mL BSA (2 × 4 mL, 20 min, 25 °C) in 4-well microscope slide tray plates (MoBiTec GmbH, cat. #: 1145-40) and rinsed with standard HEPES buffer (3 × 2 mL). Then, slides were incubated with 6 HDAC concentrations (5-fold dilution; Fig. 3: HDAC1: 77‒0.025 nM, HDAC2: 57‒0.018 nM, HDAC3/NCoR2: 50‒0.016 nM, HDAC8: 56‒0.018 nM; Fig. 5a: HDAC1: 50‒0.016 nM, HDAC2: 250‒0.016 nM, HDAC3: 50‒0.016 nM, HDAC8: 150‒0.048 nM) in HEPES buffer (2 mL, 1 h, 25 °C), rinsed (4 × 2 mL), incubated with HRP-conjugated 6x-His tag antibody (1:10000 in HEPES buffer, 2 mL, 1 h, 25 °C), rinsed (4 × 2 mL), and developed. Detection was achieved using SuperSignal West femto maximum sensitivity substrate [ThermoFisher, cat. #: 34095, solution A and B (300 µL, 1:1)] and a chemiluminescence imaging system (PXi6, Syngene) with 30 s exposure time and no binning. The 6 slides incubated with different HDAC concentrations were imaged together in order to obtain data relative to top and bottom intensity levels. Experiments were performed twice, data were obtained with Array Analyze software (Active Motif), and data analysis and representation were performed with the GraphPad Prism 7 software. Outliers were curated manually and data were fitted to nonlinear regression [log(agonist) vs. response—variable slope (four parameters)]. Curves with top values outside of the 0.8‒1.2 range, pEC50 values below the studied enzyme concentration range, and incomplete curves were considered weak binders and color-coded with white squares. Non-sigmoidal curves were denoted as not determined (N.D.) The pEC50 values were determined as the concentration of HDAC enzyme required for half-maximal binding signal. Data presented in manuscript figures are reported with the number of significant figures allowed by the standard error of the mean (SEM) calculated for each experiment. The raw data in the Supporting Tables are not adjusted corresponding to standard error.
HDAC Inhibition Assays
Assays were performed in 96-well plates (Fischer Scientific, cat. #: 3686) in HEPES buffer (final volume: 25 µL/well). Asuha-containing peptide inhibitors (5-fold dilution series, or 50 µM and 5 µM concentration for HDAC4) were incubated with enzyme [HDAC1 (0.5‒4 nM), HDAC2 (0.5‒3 nM), HDAC3/NCoR2 (1 nM), HDAC8 (0.2‒0.5 nM), HDAC4 (0.01 nM) or HDAC6 (0.5‒1 nM)] and substrate [HDACs 1‒3 and 6: Ac-Leu-Gly-Lys(Ac)-AMC (20 µM), HDACs 4 and 8: Ac-Leu-Gly-Lys(Tfa)-AMC (20 µM and 100 µM, respectively)] for 30 min at 37 °C (AMC: 7-amino-4-methylcoumarin). A solution of trypsin (25 µL, 0.4 mg/mL; final concentration of 0.2 mg/mL) was subsequently added, and the assay was allowed to develop for 15 min at room temperature. Then, fluorescence was recorded and analyzed to afford residual enzymatic activity relative to control wells and, assuming a standard fast-on/fast-off mechanism, IC50 values were obtained by fitting the resulting data to the dose–response equation with variable Hill slope (Eq. 1). Inhibition Ki values were obtained by using the Cheng-Prusoff equation (Eq. 2) and Michaelis-Menten constants as reported (HDAC1: KM = 6 µM, HDAC2: KM = 3 µM, HDAC3: KM = 6 µM, HDAC6: KM = 16 µM, HDAC8: KM = 190 µM)57. Assays were performed twice with two internal replicates, and data was analyzed using GraphPad Prism 7 or 8.
HDAC Deacetylation Assays
Enzyme [HDAC1 (250 nM), HDAC2 (500 nM), HDAC3/NCoR2 (5 nM) or HDAC8 (250 nM)] and Kac peptide (HDACs 1‒3: 100 µM, HDAC8: 1 mM) were incubated in HEPES buffer (36 µL) in a 96-well plate for 1 h at 37 °C. Then, a sample from each well (25 µL) was quenched with CH3OH/HCOOH [94:6 (v/v) 12.5 µL] and analyzed by UPLC-MS. Gradients rising linearly 0‒10%, 5‒15%, 10‒20%, or 15‒25% during 4 or 9 minutes were employed for separation of starting material and deacetylated product. Mass spectra were used for identification of both species, and conversion was calculated by peak integration at 280 nm. Assays were performed at least twice.
Cell Treatment and Western Blot
HEK293T cells (ATCC), provided by the Pless lab at the University of Copenhagen, were cultured under a humidified 5% CO2 atmosphere in Dulbecco’s modified Eagle’s medium (DMEM, Thermo Scientific, cat. #: 11965118) supplemented with 10% (v/v) fetal bovine serum (FBS), 1% penicillin and 1% streptomycin. Cells were plated into 6-well plates and grown to 80–90% confluency. Treatment with compounds 1a (10 µM), 13a (10 µM), SAHA (1 µM), or DMSO was performed in the above-mentioned media for 5 h at 37 °C, after which the media was aspirated, cells were washed with PBS and treated with lysis buffer (1% Triton X-100, 0.2% SDS and cOmplete EDTA-free protease inhibitor cocktail, COEDTAF-RO Sigma-Aldrich, in PBS, 150 µL/well). Upon scraping, lysates were collected in microcentrifuge tubes, sonicated with a Bandelin Sonopuls mini20 (2 s on, 2 s off, 80% amplitude, 1 min), centrifuged (14000 g, 10 min, 4 °C), and protein concentrations of the supernatants were determined by the bicinchoninic acid assay (BCA1, Sigma-Aldrich). The concentrations of the samples were adjusted to the lowest concentration, and 33–54 µg of lysate were mixed with NuPAGE LDS sample buffer (ThermoFisher, NP0007) and sample reducing agent (ThermoFisher, NP0004) followed by heating to 95 °C for 10 min. Samples were then resolved by gel electrophoresis (SDS-PAGE) in NuPAGE gels (12 wells, 4‒12% Bis-Tris, 1.0 mm, ThermoFisher, NP0322BOX) with MES running buffer (ThermoFisher, NP000202). Proteins were transferred to PVDF membranes (ThermoFisher, IB24001) using the iBlot2 system and blocked with 5% skim milk in tris-buffered saline with 0.1% tween-20 (TBST) at 25 °C for 1 h. Membranes were then washed with TBST (3 × 5 min), incubated with primary antibody in TBST with 5% BSA (1:1000, 4 °C, overnight), washed with TBST (3 × 5 min), incubated with HRP-conjugated secondary antibody in TBST with 2% skim milk (1:10,000, 25 °C, 1 h), and washed with TBST (2 × 5 min) and TBS (1 × 5 min). Membranes were visualized using enhanced chemiluminescent reagents (Pierce ECL Western blotting substrate, ThermoFisher, 32106) on a syngene PXi4 image analysis system. Primary antibodies used: acetylated α-tubulin (6-11B-1, sc-23950, Santa Cruz Biotechnology, Inc), vinculin (E1E9V) XP (13901, Lot: 6, Cell Signaling Technology), acetyl-histone H3 (K36) (D9T5Q, 27683, Lot: 1, Cell Signaling Technology), acetyl-histone H3 (K27) (4353, Lot: 1, Cell Signaling Technology). Secondary antibodies used: Anti-rabbit IgG HRP-linked antibody (Cell Signaling Technology, CST-7074S), Anti-mouse IgG HRP-linked antibody (Cell Signaling Technology, CST-7076S). Protein marker used: Precision Plus Protein All Blue Standard (BioRAD, 161-0373).
Statistics and reproducibility
All measurements considered were taken from distinct samples. Repeated measurements of a single sample are considered n = 1.
Safety statement
No unexpected or unusually high safety hazards were encountered.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its supplementary information files. Source data are provided with this paper.
References
McGinty, R. K. & Tan, S. Nucleosome structure and function. Chem. Rev. 115, 2255–2273 (2014).
Ahuja, N., Sharma, A. R. & Baylin, S. B. Epigenetic therapeutics: a new weapon in the war against cancer. Annu. Rev. Med. 67, 73–89 (2016).
Tough, D. F., Tak, P. P., Tarakhovsky, A. & Prinjha, R. K. Epigenetic drug discovery: breaking through the immune barrier. Nat. Rev. Drug Discov. 12, 835–853 (2016).
Jakovcevski, M. & Akbarian, S. Epigenetic mechanisms in neurological disease. Nat. Med. 18, 1194–1204 (2012).
Arrowsmith, C. H., Bountra, C., Fish, P. V., Lee, K. & Schapira, M. Epigenetic protein families: a new frontier for drug discovery. Nat. Rev. Drug Discov. 11, 384–400 (2012).
Garske, A. L. et al. Combinatorial profiling of chromatin binding modules reveals multisite discrimination. Nat. Chem. Biol. 6, 283–290 (2010).
Bock, I. et al. Application of Celluspots peptide arrays for the analysis of the binding specificity of epigenetic reading domains to modified histone tails. BMC Biochem. 12, 48 (2011).
Fuchs, S. M., Krajewski, K., Baker, R. W., Miller, V. L. & Strahl, B. D. Influence of combinatorial histone modifications on antibody and effector protein recognition. Curr. Biol. 21, 53–58 (2011).
Filippakopoulos, P. et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell 149, 214–231 (2012).
Rothbart, S. B., Krajewski, K., Strahl, B. D., Fuchs, S. M. Peptide microarrays to interrogate the “histone code”. In Methods Enzymol., vol. 512 (Elsevier, 2012), 107–135.
Kungulovski, G. et al. Application of histone modification-specific interaction domains as an alternative to antibodies. Genome Res. 24, 1842–1853 (2014).
Su, Z. et al. ChIP-less analysis of chromatin states. Epigenet. Chromatin 7, 7 (2014).
Shanle, E. K. et al. Histone peptide microarray screen of chromo and Tudor domains defines new histone lysine methylation interactions. Epigenet. Chromatin 10, 12 (2017).
Müller, M. M. & Muir, T. W. Histones: at the crossroads of peptide and protein chemistry. Chem. Rev. 115, 2296–2349 (2014).
Yang, X.-J. & Seto, E. The Rpd3/Hda1 family of lysine deacetylases: from bacteria and yeast to mice and men. Nat. Rev. Mol. Cell Biol. 9, 206–218 (2008).
Millard, C. J., Watson, P. J., Fairall, L. & Schwabe, J. W. R. Targeting class I histone deacetylases in a “complex” environment. Trends Pharmacol. Sci. 38, 363–377 (2017).
Nady, N. et al. A spot on the chromatin landscape? Histone peptide arrays as a tool for epigenetic research. Trends Biochem. Sci. 33, 305–313 (2008).
Szymczak, L., Kuo, H.-Y. & Mrksich, M. Peptide arrays: development and application. Anal. Chem. 90, 266–282 (2017).
Gurard‐Levin, Z. A., Kim, J. & Mrksich, M. Combining mass spectrometry and peptide arrays to profile the specificities of histone deacetylases. ChemBioChem 10, 2159–2161 (2009).
Gurard-Levin, Z. A., Kilian, K. A., Kim, J., Bähr, K. & Mrksich, M. Peptide arrays identify isoform-selective substrates for profiling endogenous lysine deacetylase activity. ACS Chem. Biol. 5, 863–873 (2010).
Lee, J.-H. et al. Identifying dysregulated epigenetic enzyme activity in castrate-resistant prostate cancer development. ACS Chem. Biol. 12, 2804–2814 (2017).
Kutil, Z. et al. The unraveling of substrate specificity of histone deacetylase 6 domains using acetylome peptide microarrays and peptide libraries. FASEB J. 33, 4035–4045 (2019).
Clemente, S., Franco, L. & López-Rodas, G. Distinct site specificity of two pea histone deacetylase complexes. Biochemistry 40, 10671–10676 (2001).
Garske, A. L. & Denu, J. M. SIRT1 top 40 hits: use of one-bead, one-compound acetyl-peptide libraries and quantum dots to probe deacetylase specificity. Biochemistry 45, 94–101 (2006).
Riester, D., Hildmann, C., Schwienhorst, A. & Meyer-Almes, F.-J. Histone deacetylase inhibitor assay based on fluorescence resonance energy transfer. Anal. Biochem. 362, 136–141 (2007).
Smith, B. C., Settles, B., Hallows, W. C., Craven, M. W. & Denu, J. M. SIRT3 substrate specificity determined by peptide arrays and machine learning. ACS Chem. Biol. 6, 146–157 (2011).
Rauh, D. et al. An acetylome peptide microarray reveals specificities and deacetylation substrates for all human sirtuin isoforms. Nat. Commun. 4, 2327 (2013).
Gurard-Levin, Z. A. & Mrksich, M. The activity of HDAC8 depends on local and distal sequences of its peptide substrates. Biochemistry 47, 6242–6250 (2008).
Kuo, H.-Y., DeLuca, T. A., Miller, W. M. & Mrksich, M. Profiling deacetylase activities in cell lysates with peptide arrays and SAMDI mass spectrometry. Anal. Chem. 85, 10635–10642 (2013).
Falkenberg, K. J. & Johnstone, R. W. Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nat. Rev. Drug Discov. 13, 673–691 (2014).
Kristensen, H. M. E., Madsen, A. S. & Olsen, C. A. Inhibitors of the Zinc‐dependent histone deacetylases. In Epigenetic Drug Discovery, (eds Sippl, W. & Jung, M.) 155–184 (John Wiley & Sons, 2018).
Watson, P. J. et al. Insights into the activation mechanism of class I HDAC complexes by inositol phosphates. Nat. Commun. 7, 11262 (2016).
Dose, A. et al. Interrogating substrate selectivity and composition of endogenous histone deacetylase complexes with chemical probes. Angew. Chem. Int. Ed. 55, 1192–1195 (2016).
Mahindra, A. et al. Synthesis of HDAC substrate peptidomimetic inhibitors using fmoc amino acids incorporating zinc-binding groups. Org. Lett. 21, 3179–3182. (2019).
Seidel, J. et al. Peptide-based 2-aminophenylamide probes for targeting endogenous class I histone deacetylase complexes. ChemBioChem 20, 3001–3005 (2019).
Frank, R. Spot-synthesis: an easy technique for the positionally addressable, parallel chemical synthesis on a membrane support. Tetrahedron 48, 9217–9232 (1992).
Winkler, D. F. H.; Hilpert, K.; Brandt, O.; Hancock, R. E. W., Synthesis of peptide arrays using SPOT-technology and the CelluSpots-method. (eds. Cretich, M., Chiari, M.) In Peptide Microarrays, 157–174, 2009.
Wu, C., Li, S. S.-C. CelluSpots™: a reproducible means of making peptide arrays for the determination of SH2 domain binding specificity. In Peptide Microarrays, (Springer, 2009) 197–202.
Kahnberg, P. et al. Design, synthesis, potency, and cytoselectivity of anticancer agents derived by parallel synthesis from α-aminosuberic acid. J. Med. Chem. 49, 7611–7622 (2006).
Montero, A., Beierle, J. M., Olsen, C. A. & Ghadiri, M. R. Design, synthesis, biological evaluation, and structural characterization of potent histone deacetylase inhibitors based on cyclic α/β-tetrapeptide architectures. J. Am. Chem. Soc. 131, 3033–3041 (2009).
Wilson, D. M., Silverman, L. N., Bergauer, M. & Keshari, K. R. Solid phase synthesis of hydroxamate peptides for histone deacetylase inhibition. Tetrahedron Lett. 54, 151–153 (2013).
Katz, C. et al. Studying protein–protein interactions using peptide arrays. Chem. Soc. Rev. 40, 2131–2145 (2011).
Barnash, K. D., Lamb, K. N., James, L. I. & Frye, S. V. Peptide technologies in the development of chemical tools for chromatin‐associated machinery. Drug Dev. Res. 78, 300–312 (2017).
Sereikaite, V. et al. Targeting the gamma-Aminobutyric Acid Type B (GABAB) Receptor Complex: Development of Inhibitors Targeting the K(+) Channel Tetramerization Domain (KCTD) Containing Proteins/GABAB Receptor Protein-Protein Interaction. J. Med. Chem. 62, 8819–8830 (2019).
Castañeda, C. A. et al. HDAC8 substrate selectivity is determined by long-and short-range interactions leading to enhanced reactivity for full-length histone substrates compared with peptides. J. Biol. Chem. 292, 21568–21577 (2017).
Madsen, A. S. & Olsen, C. A. Profiling of substrates for zinc‐dependent lysine deacylase enzymes: HDAC3 exhibits decrotonylase activity in vitro. Angew. Chem. 124, 9217–9221 (2012).
McClure, J. J. et al. Comparison of the deacylase and deacetylase activity of zinc-dependent HDACs. ACS Chem. Biol. 12, 1644–1655 (2017).
Lee, J.-S., Smith, E. & Shilatifard, A. The language of histone crosstalk. Cell 142, 682–685 (2010).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Fatkins, D. G. & Zheng, W. Substituting Nε-thioacetyl-lysine for Nε-acetyl-lysine in peptide substrates as a general approach to inhibiting human NAD+-dependent protein deacetylases. Int. J. Mol. Sci. 9, 1–11 (2008).
Fatkins, D. G. & Zheng, W. A spectrophotometric assay for histone deacetylase 8. Anal. Biochem. 372, 82–88 (2008).
Zessin, M. et al. One-atom substitution enables direct and continuous monitoring of histone deacylase activity. Biochemistry 58, 4777–4789 (2019).
Rajabi, N.; Nielsen, A. L.; Olsen, C. A., Dethioacylation by sirtuins 1–3: considerations for drug design using mechanism-based sirtuin inhibition. ACS Med. Chem. Lett. 11, 1886–1892 (2020).
Hirota, T., Lipp, J. J., Toh, B.-H. & Peters, J.-M. Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438, 1176–1180 (2005).
Liokatis, S. et al. Phosphorylation of histone H3 Ser10 establishes a hierarchy for subsequent intramolecular modification events. Nat. Struct. Mol. Biol. 19, 819–823 (2012).
Bowers, A. et al. Total synthesis and biological mode of action of largazole: a potent class I histone deacetylase inhibitor. J. Am. Chem. Soc. 130, 11219–11222 (2008).
Bradner, J. E. et al. Chemical phylogenetics of histone deacetylases. Nat. Chem. Biol. 6, 238–243 (2010).
Wu, M. et al. Lysine-14 acetylation of histone H3 in chromatin confers resistance to the deacetylase and demethylase activities of an epigenetic silencing complex. eLife 7, e37231 (2018).
Kitir, B. L. et al. Chemical editing of macrocyclic natural products and kinetic profiling reveal slow, tight-binding histone deacetylase inhibitors with picomolar affinities. Biochemistry 56, 5134–5146 (2017).
Maolanon, A. R., Madsen, A. S. & Olsen, C. A. Innovative strategies for selective inhibition of histone deacetylases. Cell Chem. Biol. 23, 759–768 (2016).
Chatterjee, J., Rechenmacher, F. & Kessler, H. N‐methylation of peptides and proteins: an important element for modulating biological functions. Angew. Chem. Int. Ed. 52, 254–269 (2013).
Lau, Y. H., De Andrade, P., Wu, Y. & Spring, D. R. Peptide stapling techniques based on different macrocyclisation chemistries. Chem. Soc. Rev. 44, 91–102 (2015).
Moreno-Yruela, C. & Olsen, C. A. Synthesis of trifluoromethyl ketone containing amino acid building blocks for the preparation of peptide-based histone deacetylase (HDAC) inhibitors. Synthesis 50, 4037–4046 (2018).
Gregoretti, I., Lee, Y.-M. & Goodson, H. V. Molecular evolution of the histone deacetylase family: functional implications of phylogenetic analysis. J. Mol. Biol. 338, 17–31 (2004).
Marek, M. et al. Structural basis for the inhibition of histone deacetylase 8 (HDAC8), a key epigenetic player in the blood fluke Schistosoma mansoni. PLoS Pathog. 9, e1003645 (2013).
Hailu, G. S. et al. Lysine deacetylase inhibitors in parasites: past, present, and future perspectives. J. Med. Chem. 60, 4780–4804 (2017).
Diedrich, D. et al. One-pot, multi-component synthesis and structure-activity relationships of peptoid-based histone deacetylase (HDAC) inhibitors targeting malaria parasites. Eur. J. Med. Chem. 158, 801–813 (2018).
Loeffler, F. F. et al. High-flexibility combinatorial peptide synthesis with laser-based transfer of monomers in solid matrix material. Nat. Commun. 7, 11844 (2016).
Forsström, B. et al. Proteome-wide epitope mapping of antibodies using ultra-dense peptide arrays. Mol. Cell. Proteom. 13, 1585–1597 (2014).
Hansen, L. B., Buus, S. & Schafer-Nielsen, C. Identification and mapping of linear antibody epitopes in human serum albumin using high-density peptide arrays. PLoS ONE 8, e68902 (2013).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Becher, I. et al. Chemoproteomics reveals time-dependent binding of histone deacetylase inhibitors to endogenous repressor complexes. ACS Chem. Biol. 9, 1736–1746 (2014).
Weigt, D., Hopf, C. & Médard, G. Studying epigenetic complexes and their inhibitors with the proteomics toolbox. Clin. Epigenetics 8, 76 (2016).
Acknowledgements
We thank A.S. Madsen for assistance with data analysis, K. Strømgaard for access to the peptide microarray synthesis and printing equipment, V. Sereikaite for assistance with peptide microarray synthesis, and N. Rajabi for donation of the Fmoc-Asu(tBu)-OH building block. This work was supported by the University of Copenhagen (PhD fellowship to C.M.-Y.), the Carlsberg Foundation (2013-01-0333 and CF18-0442; C.A.O.), the Lundbeck Foundation (Running Cost grant R289-2018-2074), and the European Research Council (ERC-CoG-725172 – SIRFUNCT; C.A.O.).
Author information
Authors and Affiliations
Contributions
Conceptualization, H.M.M. and C.A.O.; Methodology, C.M.-Y., H.M.M., and C.A.O.; Investigation, C.M.-Y., M.B., A.-E.V., and C.S.; Writing—Original Draft, C.M.-Y., H.M.M., and C.A.O.; Writing ‒ Review & Editing, C.M.-Y., M.B., A.-E.V., C.S., H.M.M., and C.A.O.; Visualization, C.M.-Y.; Supervision, H.M.M. and C.A.O.; Project Administration, C.A.O.; Funding Acquisition, C.A.O.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moreno-Yruela, C., Bæk, M., Vrsanova, AE. et al. Hydroxamic acid-modified peptide microarrays for profiling isozyme-selective interactions and inhibition of histone deacetylases. Nat Commun 12, 62 (2021). https://doi.org/10.1038/s41467-020-20250-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-20250-9
This article is cited by
-
Interrogating epigenetic mechanisms with chemically customized chromatin
Nature Reviews Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.