Abstract
Predicting effects of gene regulatory elements (GREs) is a longstanding challenge in biology. Machine learning may address this, but requires large datasets linking GREs to their quantitative function. However, experimental methods to generate such datasets are either application-specific or technically complex and error-prone. Here, we introduce DNA-based phenotypic recording as a widely applicable, practicable approach to generate large-scale sequence-function datasets. We use a site-specific recombinase to directly record a GRE’s effect in DNA, enabling readout of both sequence and quantitative function for extremely large GRE-sets via next-generation sequencing. We record translation kinetics of over 300,000 bacterial ribosome binding sites (RBSs) in >2.7 million sequence-function pairs in a single experiment. Further, we introduce a deep learning approach employing ensembling and uncertainty modelling that predicts RBS function with high accuracy, outperforming state-of-the-art methods. DNA-based phenotypic recording combined with deep learning represents a major advance in our ability to predict function from genetic sequence.
Similar content being viewed by others
Introduction
Recent progress in DNA sequencing and synthesis has facilitated reading and (re-)writing of the genetic makeup of biological systems on a massive scale1,2. Despite this progress, the relationship between a genetic sequence and its functional properties is poorly understood, and thus the question what to write remains largely unanswered3,4. As the number of possible sequences scales exponentially with their length, the theoretical sequence space cannot be exhaustively explored by experiments, even for small GREs5,6,7. Therefore, innovative high-throughput (HTP) approaches are required that allow to collect a quantitative functional readout for large numbers of genetic sequences7,8. At the same time, novel methods are required that identify statistical patterns and dependencies in the resulting data sets to generate models that accurately predict the properties of untested sequences. Deep learning maximizes the benefit of big data collection owing to its ability to capture complex, non-linear dependencies and to its computational scalability9, which led to several successful applications in computational biology, from genomics to proteomics10,11,12,13,14,15. These methods promise to be able to model sequence–function dependencies with minimal prior assumptions, provided that large experimental training data sets that link sequence to quantitative measure of function16,17 are available.
Although next-generation sequencing (NGS) allows obtaining sequence information at extremely large scale, our ability to assign a quantitative functional readout to each sequence has not kept pace. In previous efforts to alleviate this experimental bottleneck3,18,19,20,21, the functional readout is performed in a separate technical step, and retroactively mapped back to the corresponding sequence by statistical inference. This introduces errors and limits data quality21,22 impairing prediction accuracy. Furthermore, ribosome loading23, DNA methylation7,24, and enrichment by growth selection25 have been suggested in combination with NGS as alternative approaches. In a particularly noteworthy recent study, Yus and coworkers have used dam methylase to facilitate a functional readout quantifiable by NGS with high throughput24. However, these approaches either require elaborate sample processing procedures, which are prone to introduce bias, or are restricted to specific functional readouts. RNA sequencing techniques avoid some of these limitations but are restricted to transcriptional effects and can be greatly biased due to variability in reverse transcription, barcode-induced bias, and DNA amplification efficiencies26,27. Therefore, the need for widely applicable, technically simple and yet accurate high-throughput approaches to ascribe functional (or phenotypic) readouts to genetic sequences persists.
Here, we introduce a method that relies on DNA-based phenotypic recording to address the limitations enumerated above. Its core innovation is a three-component genetic architecture that combines on the same DNA molecule the gene of a site-specific DNA recombinase, a GRE controlling its expression, and the recombinase substrate. Thus, a physical link between GRE and the recombinase substrate is established, and the latter serves as stable, heritable record of the GRE’s effect on gene expression. Each DNA molecule embodying this architecture contains information about both the GRE sequence and a measure of its function (i.e. the modified vs. unmodified state of the substrate), both of which can be read in a single sequencing read and thus unambiguously linked. Relying on this principle, large libraries of GREs can be assessed solely relying on NGS rendering separate functional experimentation obsolete. This greatly simplifies experimental procedures, enables measurements at high kinetic resolution, eliminates technical bias associated with sample processing, and avoids the need to infer the functional readout. Importantly, while any single DNA molecule generates a binary functional record, the resolution of the readout can be arbitrarily increased by sequencing multiple DNA copies to obtain a frequency of modified substrates for each individual candidate GRE.
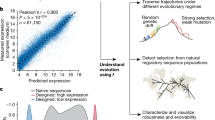
We use this approach termed uASPIre (ultradeep Acquisition of Sequence-Phenotype Interrelations) to record more than 2.7 million sequence–function pairs in a single experiment to kinetically measure translation from 303,503 RBSs in Escherichia coli. Further, we exploit the resulting high-resolution kinetic data to train a residual convolutional neural network ensemble (SAPIENs: Sequence-Activity Prediction In Ensemble of Networks) that quantitatively predicts RBS activities and quantifies reliably the uncertainty of prediction. Crucially, the combination of uASPIre and SAPIENs leads to hitherto unmatched prediction accuracy for RBSs as reflected by a coefficient of determination R2 of 0.927 and mean absolute error MAE of 0.039, notably without requirement for prior mechanistic knowledge about the translation process.
Results
The uASPIre principle
In its broadest sense, uASPIre relies on a three-component DNA architecture comprising a genetic sequence to be investigated (diversifier), the gene of a DNA-modifying enzyme (modifier), and the cognate DNA substrate of this enzyme (discriminator), all located on the same DNA molecule (Fig. 1a). The modifier can alter the discriminator sequence, which can thus appear in at least two discrete states corresponding to modified and unmodified DNA substrate, respectively. The diversifier is placed in a genetic context that allows it to either directly or indirectly affect the activity of the modifier through gene regulation (e.g. if the diversifier is a GRE) or otherwise. The more a diversifier activates (or inactivates) the modifier, the higher (or lower) is the likelihood of discriminator modification. Hence, the discriminator serves as a DNA record of functional information about the diversifier’s activity, a concept we term DNA-based phenotypic recording. We define the term phenotypic as any observable, functional characteristic arising from a genetic sequence. Consequently, both sequence and function of the diversifier can be determined concomitantly in a single sequencing read. Although binary on the level of a single DNA molecule, the fraction of modified discriminators amongst all DNA copies that share the same diversifier constitutes a direct, quantitative, and internally normalized readout of diversifier function that can be precisely tracked over time. If only a single diversifier variant is present per cell or compartment, an unambiguous link between a diversifier’s sequence and its function is stably and heritably established and maintained on the level of single DNA molecules. Therefore, NGS can be used to assess extremely large diversifier libraries. Crucially, dynamic range, resolution of the functional readout (i.e. fraction of modified discriminators), and overall throughput of the method can be arbitrarily increased by adapting sequencing depth (i.e. number of reads per diversifier variant) and number of total reads. Herein, we establish of a proof of concept for the described approach of DNA-based phenotypic recording by demonstrating the assessment of large numbers of RBSs as exemplary diversifiers.
a Generalized genetic architecture underlying the uASPIre approach. A genetic sequence of interest (diversifier; e.g. a GRE) controls, either positively (+) or negatively (−), the activity of a DNA-modifying enzyme (modifier), which can modify its cognate substrate DNA (discriminator). If placed on the same DNA molecule, diversifier sequence and discriminator state can be both determined by sequencing, for instance by NGS using forward (seqfwd) and reverse (seqrev) primers. b Readout of the uASPIre method. Under monoclonal conditions (i.e., only one diversifier variant per compartment/cell), sequencing of multiple DNA copies that share the same diversifier allows to determine the fraction of modified discriminators, which can be used as a continuous, normalized readout for diversifier function. c Prototype plasmid employing recombinase Bxb1 as a modifier controlled by the rhamnose-inducible promoter Prha. Bxb1 inverts an mCherry CDS into the correct orientation relative to a constitutive promoter Pconst, thus activating mCherry expression. attB/P and attL/R: Bxb1 attachment sites before and after recombination. d, e Kinetics of Bxb1-mediated discriminator modification in shake flask cultivations of E. coli. Recombination is detected by d direct fluorescent measurement (open diamonds, mean and s.d. of n = 3 technical replicates) and counting of red colonies after plasmid extraction and retransformation (closed diamonds, n = 1, minimum of 275 total colonies per data point) as well as e bulk Sanger sequencing of the discriminator sequence. Source data for d are available as a Source Data file.
uASPIre for GRE assessment
Although several DNA-modifying enzymes are available, we chose site-specific recombinases as a modifier for practical realization of uASPIre in this study. Recombinases have been used to record cellular events, for example by inducing reporter genes in certain cell types28,29, or to discover cell- and tissue-specific promoters30. Diversification of a recombinase coding sequence (CDS) was performed to discover variants with altered specificity31. We selected the well-characterized integrase from bacteriophage Bxb1 (bxb1/Bxb1) because it is self-sufficient in catalysing irreversible recombination, active in Pro- and Eukarya, and highly specific due to its long attachment sites attB and attP (50 and 53 bp) making off-target effects unlikely32,33,34. This orthogonality is an important advantage compared to other DNA-modifying enzymes, such as methylases, whose expression affects transcription, plasmid copy number, and cell cycle control35, often accompanied by toxic effects36. The two-state discriminator used in conjunction with Bxb1 is a short DNA sequence flanked by attB and attP in an orientation leading to irreversible sequence inversion by the recombinase (referred to as flipping hereafter).
We anticipated two critical prerequisites for the use of recombinases such as Bxb1 for uASPIre. First, recombinase expression must be tightly regulated to ensure precise control over discriminator modification. Second, modification of discriminators should occur within a practical time window of a few hours to make use of the full dynamic measurement range. To evaluate technical feasibility of these requirements, we constructed an E. coli prototype plasmid (pASPIre1) to track the activity of Bxb1 using fluorescence measurements as a proxy. In this construct, Bxb1-mediated recombination inverts an mCherry CDS into sense orientation relative to its promoter converting a non-fluorescent discriminator state into a fluorescent one (Fig. 1c). The requirement for tight regulation of recombinase expression prompted us to use the l-rhamnose-inducible promoter Prha37 to control bxb1 transcription. Using pASPIre1, we assessed Bxb1 recombination relying on bulk fluorescence measurements, counting of red colonies after retransformation of plasmid isolates from the culture (Fig. 1d), and by Sanger sequencing (Fig. 1e). These experiments collectively showed that Bxb1 is tightly controlled by addition of rhamnose to the culture without substantial recombination occurring beforehand. Moreover, the reaction proceeds for several hours allowing to flexibly sample the dynamic measurement range of discriminator inversion.
Next, we adapted the prototype architecture (Fig. 1a) to enable large-scale assessment of GREs, specifically RBSs, directly by NGS (Fig. 2a). First, superfolder green fluorescent protein (sfGFP) was fused to the Bxb1 C-terminus for later recording of calibration curves (see below). This Bxb1-sfGFP fusion retained activity exhibiting similar reaction dynamics as the sfGFP-less variant (plasmid pASPIre2, Supplementary Fig. 1). Moreover, we replaced mCherry in the discriminator with 150 bp of non-coding DNA (plasmid pASPIre3, Fig. 2a; Supplementary Fig. 2), and constructed a rhamnose utilization-deficient strain to avoid inducer consumption and ensure stable induction throughout the cultivation (E. coli TOP10ΔrhaA, Supplementary Fig. 3). Next, we used this system to characterize libraries of RBSs at high throughput relying on the uASPIre principle. As a part of the 5′-untranslated region (5′-UTR) of bacterial mRNAs, RBSs dictate the rate-limiting initiation of translation38. Because few mutations in this region can lead to orders-of-magnitude differences in protein expression, RBSs have become proven targets for optimization of cellular protein levels, in particular in multi-protein systems such as metabolic pathways39,40. This trend has been largely fueled by models that predict the relative strength of RBSs41,42,43,44 and tools for smart RBS library design45,46. However, current models are insufficiently accurate to reliably allow accurate prediction and rational forward engineering46,47,48, mainly due to the fact that they are based on small data sets of experimental endpoint measurements (<103 RBS variants), which do not cover a representative fraction of the vast number of possible RBSs and disregard the highly dynamic nature of translation. We hypothesized that time-resolved activity data for much larger RBS populations could be used to develop predictive models with greatly improved accuracy.
a DNA architecture to facilitate simultaneous sequence–function assessment using NGS. As discriminator, a stretch of non-coding DNA flanked by attB/P sites is used. A Bxb1-sfGFP fusion is used as a modifier and controlled by different RBSs as target for diversification (plasmid pASPIre3). b Experimental workflow for the uASPIre of RBSs. An RBS library with 17 randomized bases upstream of the bxb1 start codon (shown upside down) is used to transform E. coli. Monoclonal transformants each carry a different RBS variant (indicated by cells in different shades of blue). [IN1/2]: sample-specific indices. c Kinetic behavior of 10,427 RBSs (gray lines) recorded by uASPIre. The fraction of flipped discriminators is plotted over time after induction. Flipping profiles for 100 randomly selected RBSs are highlighted in blue for clarity. Data retrieved from a single Illumina NextSeq run (~400 million paired-end reads, average of ~18,700 reads per RBS). d Optimization of sampling schedule. Left panel: Effect of reducing the number of sampling time points on the approximation error between reduced (2 to 17 time points) and initial (18 time points) flipping profiles for n = 9334 RBSs from the library (see “Methods” section). Boxes range from first (lower line) to third (upper line) quartile and contain the median (center line). Whiskers contain 1.5-fold interquartile range and outliers are displayed as circles. Right panel: Initial (18 time points) and optimized (nine time points) sampling schedule. e Throughput optimization by adjustment of NGS loading. The effect of the total library size on throughput (i.e. number of variants above read-count threshold) of uASPIre for the optimized sampling schedule is shown for different read-count thresholds indicated by differentially colored circles. Source data for c–e are available as a Source Data file.
We established the following experimental workflow for the uASPIre of RBSs in E. coli (Fig. 2b; see “Methods” section). An RBS library with diversified 5′-UTR is used to transform E. coli and monoclonal transformants are co-cultivated. After induction with rhamnose, the culture is sampled at different time points and plasmid DNA is extracted, followed by agarose gel purification of target DNA fragments spanning both RBS and discriminator. Next, duplex DNA adapters containing time-sample-specific indices are ligated to the target fragments, samples are pooled, and the library pool is subjected to NGS. RBS sequence and discriminator state (here unflipped or flipped) are determined using paired-end reading. Finally, NGS raw data are processed (see “Methods” section) to obtain the dynamics of translation for each RBS, as reflected by the fraction of flipped discriminators among all sequencing reads obtained for this RBS (fraction flipped hereafter) over time (flipping profiles hereafter).
We used this workflow to analyze a library of ~10,500 variants with 17 randomized bases (N17) in the 5′-UTR of the bxb1-sfGFP mRNA over 18 time points (Fig. 2c; see “Methods” section). Importantly, a single NGS run (Illumina NextSeq, ~400 million reads) was used at an excessively high coverage (on average ~18,700 reads per variant). While the library size for this proof of concept was chosen conservatively small, it pointed towards important features of our approach: (i) the achievable throughput is very high (here 187,686 sequence–function pairs distributed across 18 time points), and it could be significantly increased by optimizing read coverage and number of time samples; (ii) The RBS activity represented by the flipping profiles is directly and quantitatively assessed at high resolution for the functional readout (fraction flipped) and with high technical reproducibility (Supplementary Fig. 4); (iii) Measurements can be performed in short intervals down to a few minutes or less facilitating acquisition of precise kinetic data. Notably, this is required to properly resolve the diversity of translation rates in the library, since endpoint measurements would either lead to underestimation of strong variants (late sampling) or low resolution for weak RBSs (early sampling); (iv) PCR-amplification is avoided during the entire workflow, which we have identified as a major source of non-systematic bias (Supplementary Fig. 5) that is absent for the PCR-free workflow (Supplementary Fig. 6).
Using this proof-of-concept data set, we optimized critical experimental parameters to increase the uASPIre throughput. First, we evaluated the effect of reducing the number of sampling time points on the ability to reconstruct the full 18 time-point flipping profiles from the proof-of-concept experiment (see “Methods” section). This analysis indicated that the number of sampling time points can be significantly reduced without major deviation from the full flipping profiles (Fig. 2d) to save NGS capacity and increase the overall throughput. We selected an optimized schedule with nine sampling time points for the following experiments as a compromise between throughput increase and accuracy of flipping profiles (98% of sequences below 5% approximation error). Afterwards, we simulated how the total library size affects the throughput by estimating the number of variants above different read-count thresholds (Fig. 2e; see “Methods” section). Here, the read-count threshold is defined as the minimum number of NGS reads per variant and time point above which the functional readout (fraction flipped) is considered statistically robust. The throughput is the number of variants above this threshold. At a given limit of obtainable reads per NGS run, the total library size (i.e. number of variants subjected to NGS) represents the main experimental parameter that can be tuned to adjust the overall throughput of our method. Increasing total library size is expected to increase the throughput but also to lead to a higher relative fraction of RBSs below threshold. Our analysis indicated that a library of ~250,000–500,000 RBS variants would be optimal to robustly retrieve high quality (i.e., above-threshold) data for a maximized number of variants (Fig. 2e).
Ultrahigh-throughput characterization of RBSs
We created a second, larger RBS library diversifying the 17 bases directly upstream of the bxb1 start codon (N17). Such libraries are known to be prone to strong skew towards weak RBSs46, which we also observed for the first RBS library (Supplementary Fig. 7a). Initial efforts on training a machine learning (ML) model on these data indicated a systematic underestimation of translation activity particularly for strong variants (see “Methods” section; Supplementary Fig. 7b). This observation, which we attributed to the skew in the initial library, prompted us to construct three additional libraries (High1-3) likely enriched for intermediate and strong RBSs. Libraries High1-3 were designed based on the first data set and added to an approximate total of one fifth to the N17 library (Fig. 3a; Supplementary Fig. 8; see “Methods” section). The composite library (~350,000 pooled transformants) spiked with a set of 31 internal-standard RBSs spanning a wide range of activities (see “Methods” section) was subjected to the uASPIre workflow. This experiment yielded the fraction flipped for 303,503 RBSs over nine time points constituting over 2.7 million sequence–function pairs (Fig. 3b). The applied threshold of at least 20 reads per RBS and time point corresponds to a robust minimum coverage of 180-fold for each variant with the average coverage amounting to 587-fold. This threshold resulted from comparing the predictive performance of ML models (see below) trained on data sets with different thresholds and tested on a validation set (Supplementary Fig. 9; see “Methods” section). Notably, while the same NGS platform was used, the throughput was increased about 29-fold compared to the proof-of-concept experiment due to optimized sampling and library size. This experiment was done in three independent biological replicates with low variability substantiating high reproducibility of uASPIre (Supplementary Fig. 10).
a Diversification strategy for RBSs. Four RBS sub-libraries were designed randomizing 17 bases in the 5′-UTR with full (N17) and partial (High1-3) degeneracy. High1-3 are designed to achieve a predicted reduced skew towards weak RBSs (see “Methods” section). Sequences are displayed in reverse complement. b Kinetic behavior of 303,503 RBS variants from the composite RBS library. Horizontal lines are the 9-time-point flipping profiles with fraction flipped shown in color code, and variants are ranked according to their IFP0–480 min. Data correspond to more than 2.7 million sequence–function pairs recorded in a single Illumina NextSeq run. c Schematic representation of IFP0–480 min (blue shaded area). d Correlation of IFP0–480 min and cellular Bxb1-sfGFP fluorescence as shown for 31 internal-standard RBSs spanning a wide range of activities (see “Methods” section). R2 and Rv2 are coefficient of determination and coefficient of determination after leave-one-out cross validation for the logistic fit (dashed line), respectively. Mean and s.d. of the slope of the cell-specific GFP signal of n = 3 independent shake flask cultures (biological replicates) are displayed (see “Methods” section). e Distribution of IFP0–480 min in the four sub-libraries. Violins contain percentiles 0.5–99.5 of variants with median and outliers as white and blue dots, respectively. Black bars contain the 25th to 75th percentiles. Source data for b, d, and e are available as a Source Data file.
To correlate the functional readout (fraction flipped) obtained by NGS with the cellular Bxb1-sfGFP concentration, we recorded cell-specific fluorescence of the Bxb1-sfGFP fusion for the aforementioned 31 internal-standard RBSs in individual shake flask cultivations (Supplementary Fig. 11; see “Methods” section). We compared the resulting curves with the corresponding flipping profiles obtained in NGS analyzing pairs of integral- and slope-based summary statistics (i.e., quantitative curve representations) for the two measurement types (see “Methods” section). Discriminator inversion strongly correlated with the prevailing cellular Bxb1-sfGFP concentration as indicated by high R2 values (0.85–0.98) for all tested combinations (Supplementary Fig. 12a, b). We selected the integral of the flipping profile between 0 and 480 min. after induction (IFP0–480 min, Fig. 3c) for further steps due to its high correlation with different summary statistics for Bxb1-sfGFP fluorescence and the high degree of diversity for IFP0–480 min in the library (Fig. 3d; Supplementary Fig. 12c). Notably, compared to the GFP measurements, the functional readout obtained by NGS exhibited a larger dynamic range and higher sensitivity at the lower (and to a lesser extent higher) range of the RBS activity spectrum (Fig. 3d). Relying on IFP0–480 min, we assessed different sub-libraries (Fig. 3e). As expected for fully degenerate RBS diversification, we found the N17 sub-library to be strongly skewed towards low activity. By contrast, all three designed sub-libraries (High1-3) were enriched for intermediate and strong RBSs, indicating that our design goal was met.
Deep learning for RBS prediction through SAPIENs
We developed an ML approach to exploit the data sets obtainable by uASPIre for quantitative prediction of RBS strength from sequence. Our deep-learning model SAPIENs uses the RBS sequence as an input in a binary matrix representation (Fig. 4a, see “Methods” section). The SAPIENs architecture is an ensemble of ten residual neural networks (ResNets)49,50, each consisting of three residual blocks of two convolutional layers51 each. The last convolutional layer’s output is fed into two sets of fully connected layers, which integrate information across all positions of the RBS. These two layer sets provide the final output of each ResNet of the ensemble, which is two shape parameters of a probability distribution for RBS activity (beta distribution). The ten ResNet models were independently trained with different randomly initialized parameters, random hyperparameters and batches of sequences. In this way, SAPIENs models the predicted distribution of IFP0–480 min as a uniformly weighted mixture of ten beta distributions, parametrized by ten independent sets of shape parameters. By outputting a distribution, we are able to characterize both the predicted value of IFP0–480 min and the uncertainty of prediction, which we quantify by the mean and standard deviation of the predictive distribution, respectively (see “Methods” section). Such a sequence-by-sequence characterization of predictive uncertainty would not be feasible with pointwise predictions. Crucially, this does not only provide quantitative predictions of the RBS activity but also a well-calibrated confidence score. A detailed description of all components of SAPIENs is available in the ML Annex.
a Schematic architecture of a single ResNet. One-hot encoded 17-bp RBS sequences are fed into three residual blocks, composed of two convolutional layers (conv1/2), and two sets of two fully connected layers (FC1α/β, FC2α/β). Yellow and purple boxes represent the output of the convolutional and fully connected layers, respectively. The gray box represents the output of the flattening operation. The model yields a probability distribution of IFP0–480 min for each sequence from which the predicted IFP0–480 min value (mean µ) and an uncertainty estimate (s.d. σ) are calculated. SAPIENs is a combination of ten individually parametrized ResNets. b Comparison of predictive performance of single ResNet and SAPIENs with classical ML models trained on the same set of 248,451 RBSs (see “Methods” section). MAE: mean absolute error, RMSE: root-mean-square error. c Comparison of IFP0–480 min values predicted by SAPIENs with experimental values measured by uASPIre. Test-set sequences were binned (bin size: 0.05) according to measured IFP0–480 min. Violins comprise percentiles 0.5–99.5 of sequences with median and outliers shown as white circles and blue dots. Black bars contain the 25th to 75th percentiles. d Cellular Bxb1-sfGFP concentrations are reliably predicted from experimentally determined (light green circles) and predicted (dark green circles) IFP0–480 min values as shown for the 31 internal-standard RBSs. IFP0–480 min values (n = 1) were converted into the slope of the cell-specific Bxb1-sfGFP signal between 0 and 290 min after induction relying on the logistic fit parameters determined earlier (Fig. 2e). e Dependence of predictive performance of different ML models on the size of the training data set (see “Methods” section). Predictive performance is evaluated with four metrics: percentage of predicted IFP0–480 min values within 2-fold error, MAE, RMSE and R2. The color scheme indicates performance, from dark blue (best) to dark red (worst). f The confidence intervals of the predicted probability distributions (horizontal axis) fully assess the uncertainty of the prediction values (vertical axis), i.e. x% of the predicted values lie within x% confidence interval. The obtained values (blue circles, n = 1) are well aligned with a theoretical perfect uncertainty assessment (dotted line). Source data for b–f are available as a Source Data file.
We trained SAPIENs and several classical linear and non-linear ML models on the same 248,451 RBS sequences chosen at random from the larger uASPIre data set. Hyperparameters were optimized exclusively on a validation set (27,398 sequences) and afterwards all models were evaluated on a held-out test set (27,654 sequences, see “Methods” section). The linear model Ridge Regression52 (R2 = 0.678) was clearly outperformed by non-linear models k-nearest neighbors53 (k-NN, R2 = 0.738), Random Forest54 (R2 = 0.835) and gradient tree boosting55 (GTB, R2 = 0.893), which highlights the importance of interactions between nucleotides in the RBS (Fig. 4b, c). Notably, SAPIENs outperformed all other approaches reaching an R2 of 0.927 and MAE of 0.039. It exhibited consistently high predictive performance across the entire range of RBS activities including the 31 internal-standard RBSs that were excluded from training (Fig. 4c; Supplementary Fig. 13). Moreover, the systematic inaccuracy in predicting strong RBSs was eliminated as a result of the addition of the designed sub-libraries High1-3 (Supplementary Fig. 14). The predicted IFP0–480 min values were converted into summary statistics for cellular Bxb1-sfGFP concentrations relying on calibration curves (Fig. 3d) and the resulting predicted Bxb1-sfGFP values correlated well with their experimentally determined counterparts (Fig. 4d). This indicates that our model reliably predicts cellular protein levels even for unseen sequences. Importantly, except for the overall weakest-performing Ridge Regression, prediction accuracy increased with training set size for all models reflected by rising confidence (R2, percentage of sequences within 2-fold error) and decreasing errors (RMSE, MAE; Fig. 4e). While a general trend towards saturation was observed, no plateau is reached even for the largest training set of 248,451 sequences. This points to the high value of large-scale sequence–function data and emphasizes the potential of uASPIre. Moreover, to go beyond global metrics of accuracy (R2, MAE), SAPIENs produces a well-calibrated confidence score for each prediction56, which can be used to guide forward engineering of RBSs (Fig. 4f, Supplementary Fig. 15, see “Methods” section). Lastly, we observed that our high prediction accuracy is reproducible across the three biological replicates (Supplementary Fig. 16, ML Annex).
Identification of sequence determinants of RBSs
We analyzed the impact of different factors known to influence RBS activity in our data. We did not find a significant overall correlation between mRNA folding energy and RBS activity (Supplementary Fig. 17a). Notably, sequences with particularly strong secondary structures (i.e. free folding energy below −15 kcal mol−1) showed a tendency to be weak RBSs (Supplementary Fig. 17b). However, such strong secondary structures were underrepresented in the library, likely due to the full randomization of only a short part of the mRNA rendering strong base pairing unlikely. Therefore, in order to allow for a proper analysis of the impact of mRNA folding or particular secondary structures, an alternative diversification strategy would be required. We then assessed the impact of Shine–Dalgarno (SD)-like motifs (i.e. AGGAGG and subsequences thereof) and additional start codons in the 5′-UTR within the N17 library. Clearly, SD-like motifs exhibit a strong positive effect on translation, which is lost (or even slightly inverted) if the motif is too close to the translational start (Fig. 5a). Similarly, a positive effect was observed for additional in-frame AUG codons (Fig. 5b) and, to a lesser extent, for GUG and UUG (Supplementary Fig. 17c, d). By contrast, out-of-frame start codons showed no globally consistent tendency but overall favored translation, in particular for positions −17 to −8. This is likely due to Gs in the start codons facilitating 16S-rRNA binding, which expectedly is most prevalent for GUG (Supplementary Fig. 17c) and difficult to disentangle from a genuine start codon effect.
a, b Influence of Shine–Dalgarno-like motifs (a) and AUG codons (b) in the 5′-UTR on the RBS activity of N17 library members. Black horizontal lines indicate the median IFP0–480 min in the data set. Boxplots (a) contain a variable number of RBSs (n between 24 and 1246) depending on the occurrence of the respective motif, and boxes range from first (lower line) to third (upper line) quartile with median (red center line) and percentiles 20/80 (whiskers). Circles (b) represent median IFP0–480 min with percentiles 20/80 (shaded areas) and in-frame positions (highlighted red). c Importance of ResNet filters for the prediction. Pearson correlation between filter activation and RBS activities of all held-out sequences is displayed per filter and position for the first convolutional layer of one randomly selected ResNet. Five filter stacks with apparent high significance are framed in bold and the average weight per base and position of the corresponding centroid filter is shown (right). d Visualization of integrated gradients scores of SAPIENs in a low-dimensional space. T-distributed stochastic neighbor embedding (t-SNE) is applied to the integrated gradient scores of test set RBSs. t-SNE dim1/2 are the two dimensions resulting from the t-SNE algorithm. e Impact of 5′-UTR bases and positions on RBS activity. Using an all-zeroes input as baseline, the average attribution score per base and position is displayed as determined for the test-set sequences. Letter size corresponds to the importance score and orientation to the direction of effect (i.e. upward/downward corresponding to a tendency to increase/decrease IFP0–480 min). f Attribution of bases and positions to strong RBSs. The strongest 5% of sequences in the test set were distributed into five clusters using k-means algorithm. The displayed motifs are the medoids of each cluster (i.e. the sequences closest to the respective cluster centroid). g, h In silico evolution of RBSs. Starting from the sequence with the lowest (g) and highest (h) predicted IFP0–480 min in the test set, pairwise mutations (underlined) are greedily applied until no further increase (g) or decrease (h) in IFP0–480 min is observed (total of 10 and 8 rounds for g and h, respectively). Source data for a–c, e, and f are available as a Source Data file.
We analyzed one ResNet model to gain an understanding about the relative importance of RBS bases and positions by clustering filters of the first convolutional layer according to the correlation between their output features and the RBS activity (see “Methods” section). We found that the model had captured translation-promoting (A, G) and translation-reducing (C) effects of bases (Fig. 5c). Moreover, a positioning effect was observable: filters with large positive weight for Gs or negative weight for Us/Cs correlated positively with RBS activity when scanning upstream regions but negatively when closer to the translational start (centroids 1–4). By contrast, filters promoting Us/Cs correlated negatively with RBS strength for most positions (centroid 5).
For further interpretation, we used integrated gradients57, an attribution method commonly used for deep-learning models (see “Methods” section). For the test set, a low-dimensional embedding of SAPIENs integrated gradients scores indicated a clear structure with strong and weak sequences clustering in almost linearly separable fashion (Fig. 5d). Global analysis of the integrated gradients scores revealed that specific positions and bases are particularly indicative for RBS activity (Fig. 5e). Substantiating the observation from Fig. 5c, Gs strongly promote translation while Cs appear to be consistently adverse. The translation-promoting effect for Gs is only observable if the distance from the start codon is at least seven bp, while a neutral or even unfavorable effect prevails for other regions. However, no distinct SD-like motif appeared because this global analysis only represents per-base and -position averages. A more targeted analysis obtained by clustering of strong and weak sequences (Fig. 5f; Supplementary Fig. 18; see “Methods” section) revealed SD-like motifs with most impactful positions ranging from −13 to −6 and invariance or slight preference for weakly pairing bases (A, U) outside the motif. Hence, our model successfully reconstructed SD-like patterns, notably without any prior knowledge about the process of translation.
Finally, we performed in silico evolution with uASPIre by greedily applying pairwise mutations to the weakest and strongest sequence in the test set to maximize and minimize predicted IFP0–480 min, respectively (see “Methods” section). Confirming our previous findings, the model systematically mutated U or C to A or G to form SD-like motifs or create in-frame start codons upon increasing RBS strength (Fig. 5g), while removing Gs and adding Cs when decreasing it (Fig. 5h). Moreover, evolving a strong sequence (gain of function) required more steps than diminishing RBS activity (loss of function) due to the sparsity of strong sequences within the search space.
Discussion
Herein, we introduce a method termed uASPIre, which relies on phenotypic recording in DNA to enable experimental generation of large-scale sequence–function data of high quality while significantly reducing experimental effort and minimizing error. Since uASPIre solely relies on NGS, extremely high throughputs are achievable, which scale linearly with the number of obtained sequencing reads and are independent of other technical parameters such as sorting speed and efficiency. For instance, transferring the experimental setup used in this study to currently available benchmark NGS systems (e.g. Illumina NovaSeq 6000) would allow recording of 108 or more sequence–function pairs per experiment, with further increases expectable in the future due to the ongoing development of NGS technology.
Notably, uASPIre requires no sophisticated instruments such as cell sorters or specialized facilities (except for NGS which can be outsourced to service providers) and only standard methods for sample preparation (DNA purification, restriction, ligation). Determination of library composition and functional characterization are performed concomitantly in a single device. Compared to previously available methods, this grants experimental practicability and, more importantly, avoids bias and error from multiple devices and processing steps. To this end, neither barcoding nor DNA amplification after the actual experiment (e.g. via PCR or clonal expansion by growth) are required, both of which are known sources of bias (compare Supplementary Figs. 5 and 6). Variant treatment is fully parallelized throughout the workflow from library generation to final readout, which avoids introduction of bias due to long and/or differential processing times, constituting a major advantage over sequential approaches. Importantly, functional information is recorded directly and does not have to be statistically inferred from read distributions, which is a well-known source of error22.
The functional readout (i.e. fraction of modified discriminators) is a quantitative, internally normalized metric for variant comparison. It exhibits high sensitivity and large dynamic range as can be appreciated from its superior ability to resolve differences between variants at the low and high end of the activity range compared to fluorescence measurements (Fig. 2d). Its resolution can be arbitrarily adjusted by adapting the sequencing depth (i.e. number of reads per variant) and could be further enhanced using systems that allow more than two discriminator states58,59. The instantaneous and continuous recording of the functional readout in situ avoids the need for immediate measurements during or directly after cultivation, which are for instance required for transient reporters such as fluorescent proteins. Therefore, the kinetic resolution of uASPIre is only limited by the time required for sampling of the culture, which can be performed in intervals of one minute or less. This is a key feature, since most biological phenomena are highly dynamic and therefore inappropriately depicted by endpoint measurements as our data on RBSs show (Figs. 2c and 3b). Crucially, such high-resolution kinetics cannot be achieved in approaches relying on elaborate and lengthy procedures such as cell sorting.
In this study, we capitalize on the advantages of uASPIre enumerated above and demonstrate its utility by recording the effect on translation of more than 313,000 RBSs from two libraries in a total of over 2.9 million sequence–function pairs. Furthermore, we exploit the high-quality data sets by deep learning to quantitatively predict RBS behavior. Notably, only the combination of big data obtained through uASPIre and the model SAPIENs facilitated the high predictive performance achieved in this study (Fig. 4e), emphasizing the potential of high-throughput experimentation combined with state-of-the-art deep learning. SAPIENs accurately quantifies the uncertainty of its predictions, which is a useful practical criterion to pick sequences the most reliable predictions. In addition, interpretation of the uASPIre data and SAPIENs revealed position-specific sequence motifs in a fully data-driven fashion without requirement for prior knowledge about RBSs. It should be noted that RBS activity is known to depend on the respective sequence context, which includes mRNA regions upstream of the RBS and the 5′-part of the CDS60. These regions have to be likewise diversified in order to obtain a truly generalizable RBS prediction. Similarly, additional modifications to the setup could be considered to enable generalization across different experimental conditions, species and sequences with low similarity to the ones currently used for training. In this context, ML techniques to improve robustness for out-of-distribution data, such as transfer learning and domain adaptation, could be used (see ML Annex).
Importantly, uASPIre is not restricted to specific functional traits of interest and the approach introduced herein can be repurposed to address a wide range of biologically relevant questions. To this end, RBS library characterization should be viewed as an application example only, and the approach can be used to interrogate different types of GREs and mechanisms of gene regulation on all levels of the central dogma. This is of high significance since gene regulation is of utmost importance for cellular function and impaired regulation of genes is frequently associated with disease. Moreover, we anticipate utility of uASPIre also beyond the realm of gene regulation arguing that, in principle, any trait of interest, which can be coupled to a gene expression output, could be accessed with the method. For instance, transcriptional or translational biosensors may be used to drive modifier expression in response to certain stimuli or small molecules of interest rendering a plethora of alternative applications accessible. Lastly, we expect uASPIre to be applicable in a wide range of host organisms, since Bxb1 is functional in pro- and eukaryotes including human cells33,34 and a variety of well-characterized alternative recombinases and other DNA-modifying enzymes are readily available. We therefore envision a wide applicability of the uASPIre approach in several research domains including metabolic engineering, genetic circuit design, and microbiome research, to name but a few.
Methods
Chemicals and reagents
Unless stated otherwise, all chemicals and reagents were obtained from Sigma Aldrich (Buchs, Switzerland). Enzymes were obtained from New England Biolabs (Ipswich, MA, USA). Oligonucleotides (Supplementary Table 1), custom duplex DNA adapters, synthetic genes, and gene fragments were obtained from Integrated DNA Technologies (Leuven, Belgium).
Cultivation of E. coli
E. coli strains were commonly cultivated in lysogeny broth (LB) supplemented with 10 g L−1 d-glucose, 50 mg L−1 kanamycin, 50 mg L−1 streptomycin, and 15 g L−1 agar, where appropriate. Rhamnose-utilization deficiency was assessed by cultivation of strains in defined mineral medium61 supplemented with 0.1 g L−1 l-leucine, 0.03 g L−1 l-isoleucine, 0.15 g L−1 l-valine, and 10 g L−1 of either d-glucose or l-rhamnose as major carbon source. Shake flask cultures (LB) were inoculated from monoclonal pre-cultures to an initial OD600 of 0.05 and cultivated in a shaking incubator (37 °C, 200 rpm). Expression was induced at an OD600 of ~0.5 by addition of 2 g L−1 l-rhamnose to the cultures. Microtiter plate cultivations were performed in sterile 96-well plates (flat bottom Nunclon™ Delta Surface, ThermoFisher Scientific, Waltham, MA, US) containing 200 μL LB per well. Wells were inoculated from monoclonal pre-cultures to an initial OD600 of 0.05 and plates were incubated in an Infinite® M1000 PRO plate reader (Tecan Group, Männedorf, Switzerland) at 37 °C without lid (orbital shaking mode, 6 mm amplitude).
Construction of plasmids
Plasmids used are listed in Supplementary Table 2. All plasmids constructed in the course of this study are based on pSEVA29162. Inserts were created relying on synthetic genes or gene fragments, which were inserted into the vector backbone by conventional restriction-ligation cloning. Maps and sequences of plasmids created in this study can be found in the Supplementary Information (Supplementary Figs. 19–23). A detailed description of library construction procedures is provided below.
Assessment of Bxb1 recombination
Bxb1-mediated inversion of the discriminator was assessed by direct fluorescent measurement, counting of red- and non-fluorescent colonies after retransformation of isolated plasmid and Sanger sequencing of the discriminator region. For direct measurement of mCherry fluorescence, 1 mL samples were collected from shake flask cultivations, spun down in a tabletop centrifuge (1 min, 8000 rcf) and pellets were re-suspended in 1 mL of ice-cold, sterile-filtered phosphate-buffered saline (PBS). To ensure full chromophore maturation, samples were incubated at 4 °C overnight before measurement in an Infinite® M1000 PRO plate reader (Tecan Group, Männedorf, Switzerland) in 96-well plates (flat bottom Nunclon™ Delta Surface, ThermoFisher Scientific, 200 μL per well). Cell-specific mCherry fluorescence was determined by dividing the red fluorescence signal (excitation at λEx = 587 nm, emission at λEm = 610 nm) by the OD600 for each sample. For microtiter plate cultivations, fluorescence of mCherry and sfGFP (λEx = 485 nm, λEm = 535 nm) was directly measured in the culture broth and normalized for OD600. To assess the dynamics of Bxb1-mediated recombination directly on the DNA level, plasmid DNA was extracted from shake flask culture samples and the state of the discriminator was determined by Sanger sequencing. Furthermore, 50 ng of extracted plasmid DNA were used to re-transform E. coli TOP10 and the transformation mixture was plated on LB agar containing 50 mg L−1 kanamycin and 10 g L−1 d-glucose to shut down transcription from PRha. After overnight incubation at 37 °C, plates were stored at 4 °C for maturation of the mCherry chromophore. Afterwards, colonies (at least 275) were manually counted to determine the ratio of clones that had received a plasmid copy of pASPIre1 with a flipped (red colonies) or unflipped (white colonies) discriminator upon transformation, respectively.
Construction of knockout strains
All E. coli strains used in this study are listed in Supplementary Table 2. Knockout of the genomic rhaA gene (l-rhamnose isomerase) in parent strain E. coli TOP10 was achieved using the method described by Datsenko and Wanner63. Primers 1 and 2 were used to generate the required linear DNA fragment containing the kanamycin resistance gene (kanR) from plasmid pKD1363 flanked by sequences homologous to the target locus.
Library generation
RBS libraries were generated via PCR on template plasmid pASPIre3 using forward primer 3 and degenerate reverse primers 4, 5, 6, and 7 to diversify the respective RBS region. The resulting PCR products were digested with PstI and SacI (37 °C, 4 h) and ligated into pASPIre3 pre-treated with the same restriction enzymes. The ligation mixtures were purified and used for electroporation of E. coli TOP10 ΔrhaA. Transformants were plated on several plates of LB agar (10 g L−1 d-glucose, 50 mg L−1 kanamycin, 50 mg L−1 streptomycin) in order to estimate library size by colony counting and facilitate pooling of the different libraries in defined ratios. After overnight incubation (37 °C), 5 mL of LB were added to the plates and colonies were scraped off. The resulting cell suspensions were pooled and sterile glycerol was added to a final concentration of 15% (v/v). Last, OD600 of the cell suspensions was determined before library stocks were snap-frozen and stored at −80 °C until further use.
Library cultivation and sampling
Library stocks were thawed on ice and used to inoculate 600 mL pre-warmed LB (50 mg L−1 kanamycin) to an initial OD600 of 0.05 in 5 L baffled cultivation flasks. Cultures were incubated (37 °C, 200 rpm) until an OD600 of ~0.5 was reached and 2 g L−1 l-rhamnose were added to induce Bxb1-mediated recombination. Samples taken throughout the cultivation were immediately mixed with an excess of ice-cold, sterile PBS for rapid cooling and then centrifuged (10 min, 4000 rcf, 4 °C), and cell pellets were frozen on dry ice until extraction of plasmid DNA was performed using a commercial kit (ZymoPURE Miniprep Kit, Zymo Research) and stored at −20 °C until further use.
NGS sample preparation
Plasmid DNA isolated from culture samples was digested with NcoI and SacI (37 °C, 4 h). After, target fragments (308 bp) containing both RBS and the attP/R site were purified by agarose gel electrophoresis and sample-specific combinations of customized, indexed DNA duplexes (Supplementary Table 3) were ligated to the sticky ends of the target fragment. For the PCR-amplified sample (Supplementary Fig. 4), NGS fragments were generated using primers 8 and 9 to specifically amplify the target region and add required overhangs for Illumina sequencing to both ends. The resulting linear DNA fragments were purified by agarose gel electrophoresis and concentration of the target was determined using capillary electrophoresis (12-capillary Fragment Analyzer, Advanced Analytical/Agilent). Afterwards, indexed samples were pooled according to their determined concentrations to adjust equal molarity for all samples and the pooled sample was subjected to NGS.
NGS
NGS was performed using an Illumina NextSeq 500 platform and a High Output kit v2.5 (75 cycles, PE 33/51) using ~20% genomic PhiX library as spike-in to increase sequence diversity. Primary data analyses were done with Illumina RTA version 2.4.11 and bcl2fastq v2.20.0.422.
Computational scripts and data sets
An annotated script for the processing of NGS data (see below) as well as the pre-processed data sets used in this study are available under: github.com/JeschekLab/uASPIre. A detailed description of the ML models is provided in the separate ML Annex, and code describing how to define and fit the SAPIENs model as well as the resulting parameters of the fitted model can be obtained under: github.com/BorgwardtLab/SAPIENs.
Processing of NGS data
The algorithms for processing of NGS data for this project were written in bash and python and are available under: http://github.com/JeschekLab/uASPIre. Briefly, forward and reverse reads retrieved from fastq files were paired and all reads with more than six consecutive unidentified nucleotides were removed. Afterwards, target fragments were selected by a 10-bp constant region (GAGCTCGCAT, max. 3 mismatches) and sequences from different samples were deconvoluted by their unique combination of two 6-bp indices (Supplementary Table 3). Next, the discriminator state was determined by searching for the presence of an attP or attR site corresponding to the sequences GGGTTTGTACCGTACAC or GCCCGGATGATCCTGAC, respectively (max. 3 mismatches, differential bases highlighted in bold). RBS sequences were determined by retrieving the 17 nucleotides upstream of the bxb1 start codon. Finally, variants with mismatches in the bxb1 CDS in more than 8% of reads were removed to exclude off-target mutations.
Internal-standard RBSs and recording of calibration curves
The internal-standard RBSs used in this study are listed in Supplementary Table 4. RBSs R1-R22 were selected from the proof-of-concept library with the goal to span the entire range of observed RBS activities. First, kinetic profiles were cropped at 720 min and only RBSs with at least 100 reads per sample were used which resulted in a set of high-quality profiles for ~9500 RBSs. Afterwards, profiles were grouped according to their dynamic behavior relying on k-medoid clustering52 (k = 25) and the RBS corresponding to the centre of each cluster was selected as representative internal-standard RBS (Supplementary Fig. 24). In addition, three weak (R23–R25) and four strong (R26–R29) RBSs handpicked from the initial library as well as two strong RBSs (R30, R31) designed using the RBS calculator41 were included. These RBSs were individually introduced into pASPIre3 by conventional cloning procedures to obtain derivatives that carry the respective RBS sequence controlling Bxb1-sfGFP translation. Activity of these RBSs was assessed by recording of the cell-specific fluorescence of three biological replicates of each variant in individual shake flask cultivations. 100 mL of pre-warmed LB (50 mg L−1 kanamycin) in 1 L baffled shake flasks were inoculated from overnight pre-cultures to an initial OD600 of 0.05. Bxb1-sfGFP expression was induced at an OD600 of ~0.5 by adding 2 g L−1 l-rhamnose. Samples taken throughout the cultivation were immediately mixed with an excess of ice-cold, sterile PBS for rapid cooling and then centrifuged (10 min, 4000 rcf, 4 °C). Cell pellets were re-suspended in PBS and suspensions were stored overnight in micro centrifuge tubes at 4 °C for sfGFP maturation. Fluorescence (λEx = 485 nm, λEm = 535 nm) and OD600 were measured in technical triplicates in 96-well plates (Corning 96-well Clear Bottom Black Polystyrol, 200 µL per well) in a TECAN Infinite® M1000 PRO plate reader. Curves of cell-specific sfGFP fluorescence were obtained by normalizing the blanked fluorescence signal for the blanked OD600 measurements and subtracting the cell-specific background fluorescence of an sfGFP-less variant (empty vector control), which was included for every cultivation batch. Furthermore, a dilution series of fluorescein was included in every 96-well plate to compensate for variations of the fluorescent readout over time.
Correlation of Bxb1 recombination with Bxb1-sfGFP levels
In order to convert Bxb1-catalyzed discriminator flipping into cellular Bxb1 concentrations, we compared the recorded cellular fluorescence profiles for the 31 internal-standard RBSs with their corresponding flipping profiles as recorded by NGS. To this end, we sought to (i) establish a combination of summary statistics which exhibit a high degree of correlation between the two measured quantities across the entire range of RBS strengths, (ii) identify the best (potentially non-linear) fit between the two summary statistics, and (iii) ensure that a high degree of diversity is maintained for the representation of the discriminator flipping across the entire set of sequences in the data set. We used integral-based (i.e. area under the curve) summary statistics for the flipping profiles and slope-based representations (i.e. slope of the linear fit) for the fluorescence profiles (Supplementary Fig. 12a, b). For the flipping statistics, we also quantified the diversity of each representation by estimating the differential entropy64 of its probability density (Supplementary Fig. 12c). For each type of summary statistic, we additionally treated the time ranges over which both the fluorescence and flipping summaries are computed as additional hyperparameters to be optimized. Moreover, for each pair of candidate summary statistics, we evaluated linear, log-linear and generalized logistic fits. We quantified the quality of each pair of summary statistics using the resulting R2 of the fit as evaluated using leave-one-out cross validation on the pool of 31 internal-standard RBSs in order to compensate for potential effects of overfitting in the analysis. Moreover, the standard deviation of each summary statistic for fluorescence was computed for all internal-standard RBSs relying on the three biological replicates. Further details on the evaluation of summary statistics are provided in the ML Annex.
Optimization of sampling time points
Sampling times were optimized using the high-quality kinetic profiles obtained from the proof-of-concept RBS library (see previous section for definition of high-quality profiles). To avoid biases towards the initial sampling schedule, we first represented the profile p of each RBS by an approximation with a logistic function \(\hat p\) imputed at 5 min intervals, which was fixed as the minimal time difference between two samples (Supplementary Fig. 25a). In cases where logistic approximation was not possible (i.e. failed parameter optimization), an exponential decay function was used. Formally, RBS i is represented by its approximation\(\hat p^i = (\hat p^i_0,\hat p^i_5, \ldots ,\hat p^i_{720})\). Afterwards, optimal sampling times were greedily selected while fixing the first and last sample at 0 and 720 min after induction. The goal of the optimization was to find a set S of optimal sampling times (initially S = (0, 720)) which allows to reconstruct \(\hat p^i\) such that a linear approximation using time points in S is as close to \(\hat p^i\) as possible. Given the set of possible sampling times T = {0, 5, 10, …, 720} and the subset I of RBS profiles on which the sampling times should be inferred, the greedy optimization finds the next optimal sampling time point s* from T as follows:
where \(\hat l^{i,\,S}\) corresponds to the linear approximation of \(\hat p^i\) using only sampling times in S (Supplementary Fig. 25b). In other words, s* is the time point that (i) is not part of the sampling schedule S yet and (ii) results in the smallest cumulative reconstruction error over all RBS profiles in I. Subsequently, S is augmented by s*, and Eq. (1) is evaluated to find the next optimal s*, until S contains the desired number of sampling times. Finally, the quality of the optimal sampling schedule for every RBS i is evaluated by computing the approximation error ri between the observed profile pi, and its linear interpolation at the optimal time points S, termed li,S (Supplementary Fig. 25c):
This optimization was performed for different sets of RBS profiles I: we first sorted RBSs by their observed strength (i.e. difference in the fraction of flipped discriminators between first and last sample) and optimized sampling for the top 5%, 10%, 25%, 50%, and 100%, respectively. This strategy was chosen to compensate for the strong bias towards weak RBSs in the initial library. Afterwards, we computed the cumulative approximation error on the entire library (top 100%) for the sampling schedules optimized on the different subsets. We found that for seven or more samples the difference in approximation error between subsets became indistinguishable and hence chose the sampling times inferred on the top 10% of profiles for the following experiments.
Optimization of NGS loading
To increase the throughput of uASPIre, we analyzed the data from the proof-of-concept (poc) experiment, which contained kinetic data of ~10,000 RBS variants. We sought to estimate an optimal number of variants to be loaded into NGS in order to retrieve a maximized number of variants with high-quality data (i.e. above different minimal read-count thresholds θ). For this simulation, we assumed that the limiting factor is the NGS throughput and that the maximal number of valid reads (i.e. reads that pass the pre-processing pipeline quality constraints) retrieved by NGS is constant across experiments under the same experimental conditions. This simulation is based on the idea that increasing the number of RBS variants reduces the coverage and vice versa, as the maximum number of valid reads is constant. For the distribution of read counts, we assumed that it follows a log-normal distribution and that its variance is independent of the coverage. The proof-of-concept data set is composed of ~2 × 108 valid reads, which are spread among nt = 18 time points and npoc = 10,427 variants with an average coverage of cov ~1000 reads per variant per time point. If the coverage of the small data set is reduced by a factor of rc > 1, and the number of time points by a factor of rt > 1, the total number of variants that could be loaded into NGS without loss would be ninput (rc, rt) = npoc × rc × rt, by conservation of the maximal number of valid reads. However, out of these ninput (rc, rt) variants, only noutput (θ, rc, rt) < ninput (rc, rt) would pass the quality control as enforced by the minimal read threshold θ. To simulate the effect of the minimal read threshold, we downsampled the read counts of the proof-of-concept data set by a factor rc and applied to it the minimal read threshold θ resulting in a number of variants above-threshold nsimul (θ, rc) < npoc. The estimated final number of variants is therefore noutput (θ, rc, rt) = nsimul (θ, rc) × rc × rt. Figure 2e illustrates the estimation of the number of variants for rt = nt / 9, several minimal read thresholds θ and several downsampling factors rc.
RBS library design
Initial efforts for training a convolutional neural network (CNN)51 based on the proof-of-concept data set resulted in a systematic underestimation of RBS strength, in particular for strong RBSs. This is likely due to the library being skewed towards weak sequences as a result of the full randomization of the 17 bases upstream of the Bxb1 start codon (Supplementary Fig. 7). To overcome this, three libraries (High1-3) presumably enriched in moderate-to-strong RBSs were designed in silico based on the proof-of-concept data set and added to a fully randomized library (N17). Libraries High1 and High2 were designed using position probability matrices (PPMs), 2D matrices in which each element represents the proportion of times a nucleotide occurs at a given position in the sequence. To this end, RBSs from the proof-of-concept data set were grouped into 10 linearly distributed bins according to a proxy for the normalized integral of their flipping profile (IFPtrz, contained in [0, 1]), for each of which a PPM was computed. The IFPtrz was computed using the trapezoidal rule on the flipping profiles. Degenerate RBS sequences for High1 and High2 were designed with the goal to obtain PPMs that most closely resemble (minimal mean-squared error) the PPMs of the highest and second highest bin, respectively. Library High3 was designed using a genetic algorithm on the basis of predictions from an initial CNN trained on the proof-of-concept data set. The RBS sequences from the three highest IFPtrz bins were randomly mutated for 200 iterations (1–2 mutations per sequence and iteration). Only sequences for which the predicted IFPtrz was increased due to the mutations were propagated to the next iteration. At the end of this process, we calculated the PPM of the resulting pool of sequences with high predicted IFPtrz, randomly selected 20,000 sequences from this PPM, and computed the predicted IFPtrz distribution for this sub-sample. Finally, the degenerate RBS sequence of High3 was obtained by greedily minimizing the Kolmogorov–Smirnov distance between the predicted IFPtrz distribution of the sub-sample and the predicted IFPtrz distribution for the respective degenerate candidate RBS sequence. For further details regarding the computation of the IFPtrz, the CNN and the genetic algorithm please refer to the ML Annex.
Normalization of biological replicates
In order to facilitate comparison of biological replicates, we capitalized on the 31 internal-standard RBSs. These serve as internal references spanning a large range of RBS activities and allow to compensate for potential batch effects and other systematic biases between replicates. Formally, for each of the 31 internal-standard RBSs, we denote by x and y the measured normalized integral of the flipping profile (IFP) for the biological replicate to be normalized and the reference replicate, respectively. We fit either a polynomial function of degree two, \(f:\left[ {0,\,1} \right] \to {\Bbb R}\) with f(x) = I + Ax + Bx2, or its inverse f(x) = g−1(x) with g(z) = I + Az + Bz2, such that the mean-squared error between f(x) and y is minimized across the 31 measurement pairs. Moreover, we impose the following constraints on the parameters of f: first, RBSs that show no activity in one replicate should remain inactive in the other replicates (f(0) = 0). Second, RBSs whose discriminators are entirely flipped before induction in one replicate should exhibit that behavior in the other replicates (f(1) = 1). Third, the ranking of RBSs according to their strength should be preserved across replicates (f is monotonically non-decreasing in [0, 1]). It should be noted that, empirically, these assumptions appear to hold across the three biological replicates in this study. Imposing the first two constraints above reduces the number of free parameters of the polynomial function from three to one, resulting in the family of functions parametrized by A: f(x) = Ax + (1 − A)x2. Moreover, the third constraint translates into the following bounds on the set of allowed values for the free parameter A: 0 ≤ A ≤ 2. This procedure was carried out for each pair of biological replicates. The quality of the resulting fits was then evaluated on the full data sets, excluding the 31 internal-standard RBSs that were used to optimize A.
Machine learning core model
We fitted the flipping profile of each RBS with a generalized logistic function (ML Annex), integrating the fitted kinetic curves between the time points at 0 and 480 min. and normalized the integral value by dividing by 480 (min). The resulting normalized integral value (range between 0 and 1; IFP0–480 min) was used as a descriptor of RBS behavior and was selected as an exemplary target for prediction since it exhibits high correlation with cellular Bxb1-sfGFP levels and a high diversity across the RBS libraries (Supplementary Fig. 12). Initially, we defined a set of preliminary candidate deep-learning architectures for a predictive model according to standard practices12,65. These included convolutional neural networks (CNNs) with and without residual blocks, as well as multilayer perceptrons. These architectures were assessed as part of the hyperparameter selection process, which indicated superior performance of the CNN with residual blocks (ResNet)49,50 for this particular application, resulting in a model with three residual blocks of two convolutional layers and two sets of two fully connected layers. We applied three main variations to the ResNet model in order to improve predictive accuracy and additionally provide a measure for predictive uncertainty. First, we chose the negative log-likelihood, which is a proper scoring rule, as the training criterion to achieve better uncertainty estimates56. The predicted IFP0–480 min was modeled using a beta distribution, as it provides a flexible distribution with support in the interval [0, 1]. Second, the last two fully connected layers in the network were modified to output two values instead of one, thereby allowing to independently parametrize the two shape parameters of the predictive beta distribution for each input sequence. Equivalently, as the first two moments of the beta distribution are functions of the shape parameters, we were able to retrieve the mean and the standard deviation of the predictive distribution for each input sequence. Third, we used an ensemble of N = 2 × 5 ResNet models56, each trained separately with a different random initialization of network parameters, a random order of training sequences during stochastic gradient-based optimization and different architecture and optimizer hyperparameters. This third variation helped increase predictive accuracy and capture epistemic uncertainty. The final model, SAPIENs, is an ensemble composed of five ResNet models with three residual blocks of two convolutional layers, composed of 64 filters of sizes 9 and 1, respectively, followed by two sets of two fully connected layers with 64 and 1 units, respectively (weight decay parameter: 10−6, learning rate: 0.01) and five ResNet models with three residual blocks of two convolutional layers, composed of 512 filters of sizes 10 and 1, respectively, followed by two sets of two fully connected layers with 64 and 1 units, respectively (weight decay parameter: 10−6, learning rate: 0.001). In all cases, we kept a held-out test set and split the remaining data set into a training and a validation set while keeping the same proportion of strong RBSs as defined by the 15th percentile of the IFP0–480 min distribution and softplus activation functions for the two output layers. We used batch-normalization66 followed by LeakyReLU activation functions between each layer. For optimization, we used the Adam optimizer67. The model was implemented in Keras with the Tensorflow68 backend. All hyperparameters (number of filters and layers, filters sizes, number of units of the fully connected layers, weight decay, learning rate, batch size) were selected with random search69 on the basis of their performance on the validation set. Additional details about the neural network can be found in the ML Annex.
Uncertainty estimation
The measured IFP0–480 min for each RBS was modeled as a draw from a beta distribution. The mean and variance of this distribution estimated by the ResNet model (see above) correspond to the predicted IFP0–480 min value and an indication of the aleatoric uncertainty of prediction, respectively. To complement this aleatoric estimate with an estimate of epistemic uncertainty, we first used an ensemble of N = 5 ResNet models with identical architecture and optimizer hyperparameters but different random parameter initialization and ordering of the input sequences. The uncertainty estimate is therefore given by the standard deviation of the mixture of N = 5 beta distributions (Supplementary Fig. 15a–c). Furthermore, we extended this ensemble strategy at a later stage by also including M different configurations for the higher level hyperparameters, such as architecture and optimizer hyperparameters, with five ResNet models per configuration, resulting in a total of N = M × 5 ResNet models in the ensemble. Finally, a number of configurations M = 2 was fixed as a trade-off between predictive performance and computational complexity (Supplementary Fig. 15d). The reliability diagram for this final ResNet ensemble (SAPIENs, N = 2 × 5) showed well-calibrated uncertainty estimates (Fig. 4f) indicating that the uncertainty of each predicted target value seems to be accounted for. This is confirmed by the fact that the mean absolute error is positively correlated with the predicted standard deviations (Supplementary Fig. 15e). Both these results suggest that the predicted standard deviations can be used as scores to evaluate the quality of each individual prediction.
Minimal read number threshold
A minimal threshold for the number of NGS reads per RBS was determined as a quality control criterion for both training and test sets. Increasing this threshold is expected to trade off two opposite effects since it increases the average quality of the data leading to a decrease in the underlying aleatoric uncertainty but at the same time reduces the data set size available for training, which generally lowers predictive performance. To this end, we first defined six filtered data sets obtained by keeping only sequences with at least 10, 15, 20, 30, 40, or 50 reads per sampling time point. Then, we randomly split each filtered data set into training, validation and test sets as described above and made sure that for each split the high-quality training, validation and test sets were contained in the lower quality training, validation and test sets, respectively. Moreover, a test set was held out for the following prediction experiments. In order to identify an optimal lower read-count threshold, we trained a single ResNet model for 150 epochs. We randomized the search for hyperparameters69 (see above) used the same 150 sets of hyperparameters for each filtered training data set and determined the coefficient of determination on the validation set. Hence, the minimal threshold was effectively treated as a hyperparameter. This analysis indicated that a minimal threshold of 20 reads per time point was optimal for predictive performance, which saturated for lower thresholds despite the increase in overall data set size (Supplementary Fig. 9a). We kept this training/validation/test split (Split0) for the following prediction experiments. Finally, we confirmed that these conclusions were not an artifact of the random split of the original data set by repeating this analysis using five different training, validation and test-set splits (Supplementary Fig. 9b).
Evaluation and benchmarking of the prediction model
Using Split0, we evaluated our model in more detail. Importantly, this implies that the test set had not been used in previous experiments in order to avoid overfitting. First, we used random search for selecting the best combination among 150 sets of hyperparameters on the validation set (see above), let SAPIENs run for 300 epochs and used an early stopping criterion on the validation set to avoid overfitting by selecting the epoch with the best validation R2 (Fig. 4c). To compare our single ResNet models and SAPIENs to different available ML approaches, we trained different models (Fig. 4b, e) on the training set and tuned their hyperparameters by optimizing predictive performance on a subset of the validation set of Split0. The single ResNet and SAPIENs models were trained for a maximum of 150 epochs, using early stopping. A total of 100 randomly generated model architectures with 1–3 residual blocks were considered. Hyperparameters tuned for the other models were regularization strength for Ridge Regression52, number of neighbors K for k-Nearest Neighbors53, number of trees for Random Forests54, and maximum depth and learning rate for Gradient Tree Boosting55, the later also benefited from early stopping in the validation set. The impact of the training set size on predictive performance (Fig. 4e) was evaluated by training the different models on different smaller data sets, while ensuring that the training and validation sets were contained in the training and validation sets of higher sample size experiments (i.e. nested training and validation sets). Hyperparameters for all models were optimized independently for each training set size on the corresponding validation set. The effect of adding designed sub-libraries to increase the fraction of stronger RBSs in the bulk library (Fig. 3e) was further analyzed to evaluate a potential gain in predictive performance for the intermediate and strong sequences (Supplementary Fig. 14). To this end, we performed cross-analyses with the fully degenerate sub-library (N17) and the bulk library (N17+High1-3). We trained on N17 and predicted on unseen subsets of N17 and N17+High1-3, and trained N17+High1-3 and predicted on unseen subsets of N17 and N17+High1-3 (Supplementary Fig. 14a). In another set of analyses, we omitted each of the enriched sub-libraries while training by moving them to the test sets and evaluated the corresponding effect (Supplementary Fig. 14b). We trained a single ResNet model for 300 epochs for computational considerations and we used early stopping in the validation set. Hyperparameters were tuned independently for each data set and selected from 150 random configurations in the corresponding validation set. All analyses were done with the same training and validation set sizes. Comparative analyses were performed with the same test set.
Evaluation of sequence motifs and model interpretation
We analyzed the fully degenerate sub-library (N17) in order to measure the impact of the position of known motifs of influence on the RBS activity, such as start codons (AUG, UUG, GUG) or the consensus Shine–Dalgarno sequence (AGGAGG and subsequences). To this end, for each position, for each group of RBSs that presented the motif of interest at the given position, we calculated simple statistics (median, interquartile ranges, 20/80 percentiles) on the target IFP0–480 min of the sequences in the group (Fig. 5a, b; Supplementary Fig. 17c, d). We excluded from these groups RBSs that contained at least one start codon other than the one at the position of interest. We also analyzed the filters of the first convolutional layer (excluding the first skip connection) of a ResNet model of the ensemble chosen at random (Fig. 5c). To this end, the effect of each filter was evaluated by calculating Pearson’s correlation coefficient between the filter activations at each position and the flipping integral for all sequences in the test set. Consequently, each filter is represented by a vector of correlations of size 17, which corresponds to the number of positions at which the filter influence is estimated. Finally, the filter representations are then clustered in 12 groups with a complete linkage clustering method using Hamming distance as the underlying metric for comparing individual sequences in order to group filters of similar influence. The integrated gradients57 method assigns attribution scores to each base and position by computing the linear path integral between the sequence of interest and a baseline sequence chosen a priori. The attribution scores measure the effect of individual bases on the predicted IFP0–480 min, relative to a baseline. We applied the integrated gradients method to SAPIENs and chose a “blank” one-hot encoded sequence as a neutral baseline (i.e. an all-zeros array). We first used a dimension reduction method, the t-distributed stochastic neighbor embedding (tSNE) method, to visualize how sequences behave in a low-dimensional space (perplexity = 12, early exaggeration = 30) (Fig. 5d). We also averaged the attribution scores of all sequences in the test set, per base and per position, to get a better understanding of the important positions and bases, which contribute either to a high RBS activity or to a low one (Fig. 5e). Finally, in order to account for non-linearities between positions and to understand the drivers of very strong or very weak sequences, we selected the top 5% and the bottom 5% sequences in the test set after removing outlier sequences and clustered each pool with k-means according to their attribution score profiles into five clusters. The medoids of these five clusters are displayed for the strong (Fig. 5f) and weak RBSs (Supplementary Fig. 18). For in silico evolution, we selected the weakest (respectively strongest) sequence in the test set and aimed to mutate it progressively to a sequence presenting a maximum (respectively minimum) attainable RBS activity as predicted by SAPIENs (Fig. 5g, h). To do so, we considered all sequences that could result from applying one or two mutations to the current sequence and kept the strongest (respectively weakest) one in each round until no candidate exhibited a change in predicted IFP0–480 min in the desired direction.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Plasmids pASPIre1, pASPIre2, and pASPIre3 are available from Addgene under identification numbers ID154842, ID154843, and ID154843, respectively. Source data for Figs. 1d, 2c–e, 3b, d, e, 4b–f, and 5a–c, e, f and Supplementary Figs. 1b, 3a, b, 4a, b, 5, 6b, 7a, b, 9a, b, 10, 11, 12a–c, 13, 14a, b, 15d, e, 16, 17a–d, 18, and 25 are provided in the Source Data file. The raw NGS data for the three biological replicate experiments for the large RBS library are provided in the NCBI SRA under accession codes SAMN15215026, SAMN15215027, and SAMN15215027. Source data are provided with this paper.
Code availability
The processed NGS data sets are available under: github.com/JeschekLab/uASPIre. The code generated and used in this study is available under: github.com/JeschekLab/uASPIre (NGS data processing) and github.com/BorgwardtLab/SAPIENs (code for ML). Source data are provided with this paper.
References
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Kosuri, S. & Church, G. M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Sharon, E. et al. Inferring gene regulatory logic from high-throughput measurements of thousands of systematically designed promoters. Nat. Biotechnol. 30, 521–530 (2012).
Mutalik, V. K. et al. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat. Methods 10, 354–360 (2013).
Fowler, D. M. et al. High-resolution mapping of protein sequence-function relationships. Nat. Methods 7, 741–746 (2010).
Atwal, G. S. & Kinney, J. B. Learning quantitative sequence-function relationships from massively parallel experiments. J. Stat. Phys. 162, 1203–1243 (2016).
Raad, M., Modavi, C., Sukovich, D. J. & Anderson, J. C. Observing biosynthetic activity utilizing next generation sequencing and the DNA linked enzyme coupled assay. ACS Chem. Biol. 12, 191–199 (2017).
Hertzberg, R. P. & Pope, A. J. High-throughput screening: new technology for the 21st century. Curr. Opin. Chem. Biol. 4, 445–451 (2000).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, Cambridge, MA, 2016.
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Park, Y. & Kellis, M. Deep learning for regulatory genomics. Nat. Biotechnol. 33, 825–826 (2015).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Zou, J. et al. A primer on deep learning in genomics. Nat. Genet. 51, 12–18 (2019).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Camacho, D. M., Collins, K. M., Powers, R. K., Costello, J. C. & Collins, J. J. Next-generation machine learning for biological networks. Cell 173, 1581–1592 (2018).
Kinney, J. B., Murugan, A., Callan, C. G. & Cox, E. C. Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence. Proc. Natl Acad. Sci. USA 107, 9158–9163 (2010).
Kosuri, S. et al. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl Acad. Sci. USA 110, 14024–14029 (2013).
Cambray, G., Guimaraes, J. C. & Arkin, A. P. Evaluation of 244,000 synthetic sequences reveals design principles to optimize translation in Escherichia coli. Nat. Biotechnol. 36, 1005–1015 (2018).
de Boer, C. G. et al. Deciphering eukaryotic gene-regulatory logic with 100 million random promoters. Nat. Biotechnol. 38, 56–65 (2019).
Peterman, N. & Levine, E. Sort-seq under the hood: implications of design choices on large-scale characterization of sequence-function relations. BMC Genomics 17, 206 (2016).
Sample, P. J. et al. Human 5′ UTR design and variant effect prediction from a massively parallel translation assay. Nat. Biotechnol. 37, 803–809 (2019).
Yus, E., Yang, J. S., Sogues, A. & Serrano, L. A reporter system coupled with high-throughput sequencing unveils key bacterial transcription and translation determinants. Nat. Commun. 8, 368 (2017).
Cuperus, J. T. et al. Deep learning of the regulatory grammar of yeast 5′ untranslated regions from 500,000 random sequences. Genome Res. 27, 2015–2024 (2017).
Parekh, S., Ziegenhain, C., Vieth, B., Enard, W. & Hellmann, I. The impact of amplification on differential expression analyses by RNA-seq. Sci. Rep. 6, 25533 (2016).
Katayama, S. et al. Guide for library design and bias correction for large-scale transcriptome studies using highly multiplexed RNAseq methods. BMC Bioinformatics 20, 418 (2019).
Orban, P. C., Chui, D. & Marth, J. D. Tissue- and site-specific DNA recombination in transgenic mice. Proc. Natl Acad. Sci. USA 89, 6861–6865 (1992).
Kaczmarczyk, S. J. & Green, J. E. A single vector containing modified cre recombinase and LOX recombination sequences for inducible tissue-specific amplification of gene expression. Nucleic Acids Res. 29, E56–E56 (2001).
Altier, C. & Suyemoto, M. A recombinase-based selection of differentially expressed bacterial genes. Gene 240, 99–106 (1999).
Buchholz, F. & Stewart, A. F. Alteration of Cre recombinase site specificity by substrate-linked protein evolution. Nat. Biotechnol. 19, 1047–1052 (2001).
Kim, A. I. et al. Mycobacteriophage Bxb1 integrates into the Mycobacterium smegmatis groEL1 gene. Mol. Microbiol. 50, 463–473 (2003).
Xu, Z. Y. et al. Accuracy and efficiency define Bxb1 integrase as the best of fifteen candidate serine recombinases for the integration of DNA into the human genome. BMC Biotechnol. 13, 78 (2013).
Jusiak, B. et al. Comparison of integrases identifies Bxb1-GA mutant as the most efficient site-specific integrase system in mammalian cells. ACS Synth. Biol. 8, 16–24 (2019).
Lobner-Olesen, A., Skovgaard, O. & Marinus, M. G. Dam methylation: coordinating cellular processes. Curr. Opin. Microbiol. 8, 154–160 (2005).
Southall, T. D. et al. Cell-type-specific profiling of gene expression and chromatin binding without cell isolation: assaying RNA Pol II occupancy in neural stem cells. Dev. Cell 26, 101–112 (2013).
Egan, S. M. & Schleif, R. F. A regulatory cascade in the induction of rhaBAD. J. Mol. Biol. 234, 87–98 (1993).
Laursen, B. S., Sorensen, H. P., Mortensen, K. K. & Sperling-Petersen, H. U. Initiation of protein synthesis in bacteria. Microbiol. Mol. Biol. Rev. 69, 101–123 (2005).
Jeschek, M., Gerngross, D. & Panke, S. Combinatorial pathway optimization for streamlined metabolic engineering. Curr. Opin. Biotechnol. 47, 142–151 (2017).
Jervis, A. J. et al. Machine learning of designed translational control allows predictive pathway optimization in Escherichia coli. ACS Synth. Biol. 8, 127–136 (2019).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Na, D. & Lee, D. RBSDesigner: software for designing synthetic ribosome binding sites that yields a desired level of protein expression. Bioinformatics 26, 2633–2634 (2010).
Seo, S. W. et al. Predictive design of mRNA translation initiation region to control prokaryotic translation efficiency. Metab. Eng. 15, 67–74 (2013).
Borujeni, A. E., Channarasappa, A. S. & Salis, H. M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res. 42, 2646–2659 (2014).
Farasat, I. et al. Efficient search, mapping, and optimization of multi-protein genetic systems in diverse bacteria. Mol. Syst. Biol. 10, 731 (2014).
Jeschek, M., Gerngross, D. & Panke, S. Rationally reduced libraries for combinatorial pathway optimization minimizing experimental effort. Nat. Commun. 7, 11163 (2016).
Reeve, B., Hargest, T., Gilbert, C. & Ellis, T. Predicting translation initiation rates for designing synthetic biology. Front. Bioeng. Biotechnol. 2, 1–6 (2014).
Vigar, J. R. J. & Wieden, H. J. Engineering bacterial translation initiation—do we have all the tools we need? Biochim. Biophys. Acta 1861, 3060–3069 (2017).
He, K. M., Zhang, X. Y., Ren, S. Q. & Sun, J. in Proc. IEEE Conference on Computer Vision and Pattern Recognition 770–778 (2016).
Xie, S., Girshick, R., Dollár, P., Tu, Z. & He, K. in Proc. IEEE Conference on Computer Vision and Pattern Recognition, 1492–1500 (2017).
LeCun, Y. et al. Backpropagation applied to handwritten zip code recognition. Neural Comput. 1, 541–551 (1989).
Hastie, T., Tibshirani, R. & Friedman, J. H. The Elements of Statistical Learning: Data Mining, Inference, and Prediction 2 (Springer, New York, 2009).
Altam, N. S. An introduction to kernel and nearest-neighbor nonparametric regression. Am. Stat. 46, 175–185 (1992).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Friedman, J. H. Greedy function approximation: a gradient boosting machine. Ann. Stat. 29, 1189–1232 (2001).
Lakshminarayanan, B., Pritzel, A. & Blundell, C. Simple and scalable predictive uncertainty estimation using deep ensembles. Adv. Neur. 30, 6402–6413 (2017).
Sundararajan, M., Taly, A. & Yan, Q. Axiomatic attribution for deep networks. Proc. 34th Int. Conf. Mach. Learn. 70, 3319–3328 (2017).
Lapique, N. & Benenson, Y. Genetic programs can be compressed and autonomously decompressed in live cells. Nat. Nanotechnol. 13, 309–315 (2018).
Roquet, N., Soleimany, A. P., Ferris, A. C., Aaronson, S. & Lu, T. K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
Kudla, G., Murray, A. W., Tollervey, D. & Plotkin, J. B. Coding-sequence determinants of gene expression in Escherichia coli. Science 324, 255–258 (2009).
Jeschek, M. et al. Biotin-independent strains of Escherichia coli for enhanced streptavidin production. Metab. Eng. 40, 33–40 (2017).
Martinez-Garcia, E., Aparicio, T., Goni-Moreno, A., Fraile, S. & de Lorenzo, V. SEVA 2.0: an update of the Standard European Vector Architecture for de-/re-construction of bacterial functionalities. Nucleic Acids Res. 43, D1183–D1189 (2015).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Perez-Cruz, F. Estimation of information theoretic measures for continuous random variables. Adv. Neural Inform. Process. Syst. 21, 1257–1264 (2009).
Kelley, D. R. et al. Sequential regulatory activity prediction across chromosomes with convolutional neural networks. Genome Res. 28, 739–750 (2018).
Ioffe, S. & S., C. Batch normalization: accelerating deep network training by reducing internal covariate shift. Proc. 32nd Int. Conf. Mach. Learn. 37, 448–456 (2015).
Kingma, D. P. & Ba, J. Adam: A Method for Stochastic Optimization. CoRR abs/1412.6980 (2015).
Abadi, M. et al. in Proc. 12th USENIX Symposium on Operating Systems Design and Implementation (2016). https://www.usenix.org/system/files/conference/osdi16/osdi16-abadi.pdf
Bergstra, J. & Bengio, Y. Random search for hyper-parameter optimization. J. Mach. Learn. Res. 13, 281–305 (2012).
Acknowledgements
We kindly thank all members of the Genomics Facility Basel for support with NGS experiments, Prof. Michael C. Jewett for helpful scientific discussions, and Dr. Tania Roberts for experimental advice. This work was supported in part by the European Commission (project: Madonna; grant no. 766975), the SNSF Starting Grant “Significant Pattern Mining” (grant no. 155913; K.B.) and the NCCR “Molecular Systems Engineering” (Y.B.).
Author information
Authors and Affiliations
Contributions
M.J. and Y.B. conceived the project. M.J., Y.B., and K.B. coordinated the study. K.B. conceived and supervised machine learning analyses. M.J. supervised experimental work. S.H., K.F., and M.J. performed experiments. S.H., K.F., L.P., A.C.G., and M.J. analyzed data. A.C.G., L.P., and M.J. developed measures to increase throughput. S.H. and L.P. developed the algorithm for processing of NGS data. L.P. conceived, developed, and analyzed machine learning models. C.B. advised design of DNA adapters and NGS. M.J., L.P., and Y.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Koji Tsuda, Luis Serrano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Höllerer, S., Papaxanthos, L., Gumpinger, A.C. et al. Large-scale DNA-based phenotypic recording and deep learning enable highly accurate sequence-function mapping. Nat Commun 11, 3551 (2020). https://doi.org/10.1038/s41467-020-17222-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-17222-4
This article is cited by
-
Accuracy and data efficiency in deep learning models of protein expression
Nature Communications (2022)
-
Towards next-generation cell factories by rational genome-scale engineering
Nature Catalysis (2022)
-
pGlycoQuant with a deep residual network for quantitative glycoproteomics at intact glycopeptide level
Nature Communications (2022)
-
Feasibility of predicting allele specific expression from DNA sequencing using machine learning
Scientific Reports (2021)
-
A deep learning approach to programmable RNA switches
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.