Abstract
NifB is a radical S-adenosyl-L-methionine (SAM) enzyme that is essential for nitrogenase cofactor assembly. Previously, a nitrogen ligand was shown to be involved in coupling a pair of [Fe4S4] clusters (designated K1 and K2) concomitant with carbide insertion into an [Fe8S9C] cofactor core (designated L) on NifB. However, the identity and function of this ligand remain elusive. Here, we use combined mutagenesis and pulse electron paramagnetic resonance analyses to establish histidine-43 of Methanosarcina acetivorans NifB (MaNifB) as the nitrogen ligand for K1. Biochemical and continuous wave electron paramagnetic resonance data demonstrate the inability of MaNifB to serve as a source for cofactor maturation upon substitution of histidine-43 with alanine; whereas x-ray absorption spectroscopy/extended x-ray fine structure experiments further suggest formation of an intermediate that lacks the cofactor core arrangement in this MaNifB variant. These results point to dual functions of histidine-43 in structurally assisting the proper coupling between K1 and K2 and concurrently facilitating carbide formation via deprotonation of the initial carbon radical.
Similar content being viewed by others
Introduction
NifB plays a key role in the biosynthesis of the active site of Mo-nitrogenase, an [(R-homocitrate)-MoFe8S9C] metallocofactor (designated M-cluster) that catalyzes the ambient conversion of dinitrogen (N2) to ammonia (NH3)1,2,3,4,5,6. A member of the radical S-adenosyl-L-methionine (SAM) enzyme family7,8, NifB is a monomeric protein carrying three [Fe4S4] clusters:9,10,11,12 one, designated the SAM-cluster, is ligated by a canonical CxxxCxxC motif and coordinates the SAM-binding [Fe4S4] cluster; the other two, designated the K-cluster, are ligated by a number of conserved ligands and supply the full complement of iron (Fe) atoms for the biosynthesis of an 8Fe precursor to the mature cofactor. Previous studies of the NifB proteins from Azotobacter vinelandii and Methanosarcina acetivorans have led to the proposal that NifB utilizes a unique, radical SAM-dependent mechanism for carbide insertion concomitant with the transformation of the K-cluster to an [Fe8S9C] cluster (designated the L-cluster), which represents both an 8Fe precursor and an all-Fe core of the M-cluster13,14,15,16. This process begins with an SN2-type transfer of a methyl group from a SAM molecule to the K-cluster (Supplementary Fig. 1a, ➀), and it is followed by hydrogen abstraction of the methyl group by a 5′-deoxyadenosyl (5′-dA•) radical that is derived from the homolytic cleavage of a second SAM molecule (Supplementary Fig. 1a, ➁). The resultant methylene radical then undergoes further deprotonation/dehydrogenation to yield a μ6-coordinated interstitial carbide concomitant with the radical-chemistry-based coupling and rearrangement of the two [Fe4S4] modules of the K-cluster (designated K1- and K2-cluster, respectively) into an [Fe8S9C] L-cluster, which represents an all-Fe core of the M-cluster except for the substitution of one Fe atom for Mo/homocitrate at one end of the cluster (Supplementary Fig. 1a, ➂). Once generated, the L-cluster is passed from NifB onto NifEN, the next biosynthetic scaffold, where it is matured into a fully complemented M-cluster upon insertion of Mo and homocitrate by NifH prior to the delivery of M-cluster to its final binding site in NifDK, the catalytic component of Mo-nitrogenase (Supplementary Fig. 1a, ➃)17,18,19,20,21,22.
The unique radical chemistry that occurs on NifB during the nitrogenase cofactor core assembly makes this protein an attractive subject of investigation, as knowledge in this regard will shed light on the unusual synthetic route to a high-nuclearity metallocofactor that is biologically important and chemically elusive. Previously, we have identified three sets of Cys ligands—three Cys per set—for the three [Fe4S4] clusters in M. acetivorans NifB (designated MaNifB), namely, the SAM- cluster (Cys50, Cys54, and Cys57), the K1-cluster (Cys30, Cys63, and Cys129) and the K2-cluster (Cys264, Cys274, and Cys277) (Supplementary Fig. 1b, c)11. Further, we have established the coexistence of SAM- and K2-clusters as a prerequisite for methyltransfer and hydrogen abstraction to occur and pinpointed the K2-cluster as the site for methyl attachment and the subsequent hydrogen abstraction from the methyl group by a 5′-dA• radical11. Perhaps even more excitingly, using pulse EPR techniques, we have identified a nitrogen atom from a His residue as the fourth ligand for the K1-cluster that is lost upon coupling between the K1- and K2-clusters into an L-cluster11. This observation suggests an important role of the His/nitrogen ligand in generating the nitrogenase cofactor core (i.e., the L-cluster), particularly given the ability of His to undergo protonation/deprotonation, which either provides a release mechanism for the L-cluster onto the next biosynthetic apparatus or allows this residue to participate in the further deprotonation/dehydrogenation of the carbon radical to give rise to a central carbide. As such, it is crucial to identify this nitrogen ligand and probe its function in the process of nitrogenase cofactor core formation.
Here we use a combination of site-directed mutagenesis and pulse EPR spectroscopy to show that the His43 residue is the specific nitrogen ligand for the K1-cluster of MaNifB. Our biochemical and EPR analyses demonstrate the essential role of His43 in the formation of the nitrogenase cofactor core structure, although substitution of this residue with Ala does not impact early steps of carbide insertion leading to the initial hydrogen atom abstraction of the K2-associated methyl group. Our XAS/EXAFS analysis further reveals a shortened distance between the K1- and K2-clusters upon substitution of His43 with Ala, as well as a further processing of these clusters into an intermediate between the K- and L-clusters upon incubation with SAM. These observations point to a dual function of His43 in positioning/orienting the K1-cluster relative to the K2-cluster for proper coupling and deprotonating the initial carbon radical for carbide formation.
Results
The identity of the histidine ligand of the K1-cluster
Sequence analysis of MaNifB revealed the presence of three highly conserved His residues, His28, His43 and His219, which could potentially serve as the nitrogen ligand for the K1-cluster on NifBMa (Supplementary Fig. 1b). On the basis of this analysis, three MaNifB variants were heterologously expressed in Escherichia coli. Designated MaNifBH28A, MaNifBH43A and MaNifB219A, respectively, each variant has one of the three conserved His residues substituted with Ala. Purified MaNifB variants, like their wildtype counterpart (designated MaNifBwt), are monomers of ~38 kDa (Supplementary Fig. 2a). Moreover, upon FeS reconstitution, MaNifBH28A, MaNifBH43A, and MaNifBH219A have Fe contents comparable to that of MaNifBwt, all of which carry three [Fe4S4] clusters (i.e., the SAM-, K1- and K2-clusters) per protein (Supplementary Fig. 2b). Thus, a loss of the His ligand does not seem to impact the ability of MaNifB to ligate any of the three [Fe4S4] clusters, likely due to the 3-Cys-coordination of these clusters that is sufficient to secure them in place.
To assess whether a nitrogen ligand is still present in these MaNifB variants, pulse EPR spectroscopy—specifically, three-pulse electron spin echo envelope modulation (3P-ESEEM)—was used to observe modulations to the time domain spectrum and corresponding peaks in the fast Fourier transformed (FFT) spectrum that can be assigned to the coordinating nuclei (Fig. 1; also see Supplementary Fig. 3). Consistent with our previous observation11, MaNifBwt shows deep modulations in the time domain of the ESEEM spectrum (Fig. 1a, trace 1) and corresponding intensity between 0 and 6 MHz in the FFT (Fig. 1b, trace 1), which have been previously assigned to the hyperfine and quadrupole couplings of a cluster-ligated 14N nucleus11. Similar modulations and intensities are present in the ESEEM spectrum and FFT of MaNifBK1, a variant carrying only the K1-cluster but no SAM- and K2-clusters (because of substitutions of the Cys ligands of the SAM- and K2-clusters with Ala11), affirming the previous assignment of the nitrogen ligand to the K1-cluster (Fig. 1a, b, trace 5). While similarly deep modulations and intensities are observed in the ESEEM spectra and FFTs of MaNifBH28A and MaNifBH219A (Fig. 1a, b, traces 2, 3), these features are clearly absent from the ESEEM spectrum and FFT of MaNifBH43A (Fig. 1a, b, trace 4), suggesting His43 as the nitrogen ligand for the K1-cluster. To seek further support for this assignment, MaNifBK1-H43A—another MaNifB variant carrying only the Cys ligands of K1 along with a substitution of His43 with Ala—was heterologously expressed in E. coli. The purified MaNifBK1-H43A demonstrates the same subunit composition as MaNifBK1, as well as the same Fe content that is consistent with the presence of one [Fe4S4] cluster (i.e., the K1-cluster) per protein upon FeS reconstitution (Supplementary Fig. 2a, b). However, contrary to MaNifBK1 (Fig. 1a, b, trace 5), MaNifBK1-H43A does not show deep modulations and intensities in its ESEEM spectrum and FFT (Fig. 1a, b, trace 6), firmly establishing His43 as the nitrogen ligand that specifically coordinates the K1-cluster.
a Time domain and (b) fast Fourier transformed (FFT) spectra of MaNifBwt (1), MaNifBH28A (2), MaNifBH219A (3), MaNifBH43A (4), MaNifBK1 (5) and MaNifBK1-H43A (6). The time domain spectra of the His-ligand containing samples (i.e., 1, 2, 3, and 5) have modulations from 14N that appear as peaks in the fast Fourier transformed (FFT) spectra between 1 and 6 MHz. The sharp modulations between 250 and 500 ns in the time domain and the resulting broad peak near 14 MHz in the FFT are from nearby weakly coupled protons. The ESEEM spectra were recorded at 10 K, τ = 128–144 ns, π/2 = 12 ns, and 9.3366 GHz. The experiment was performed three times independently (n = 3 independent samples), and representative results are shown in the figure. All protein samples have a concentration of 15 mg mL−1. The CW EPR spectra of these dithionite-reduced MaNifB proteins are shown in Supplementary Fig. 3.
The role of histidine ligand in L-cluster maturation
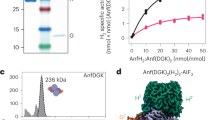
Continuous wave (CW) EPR analysis provided the first insights into the function of His43 in the cofactor core assembly process. Consistent with the coupling and rearrangement of the K1- and K2-clusters into an L-cluster, an L-cluster-specific, g = 1.94 signal9,10,15 is observed in the spectra of MaNifBwt, MaNifBH28A, and MaNifBH219A upon incubation of these proteins with SAM (Fig. 2a, traces 1–3). In contrast, the g = 1.94 signal is absent from the spectrum of MaNifBH43A following the same treatment, suggesting a lack of K- to L-cluster transformation on this protein after incubation with SAM (Fig. 2a, trace 4). In support of this assignment, the SAM-treated MaNifBwt, MaNifBH28A, and MaNifBH219A can be used as M-cluster sources for the subsequent reconstitution and activation of apo-NifDK in an in vitro assay; whereas the SAM-treated MaNifBH43A cannot support the reconstitution and activation of apo-NifDK in the same assay (Fig. 2b). Interestingly, the activity of MaNifBH28A in this assay is ~41% less than those of MaNifBwt and MaNifBH219A, suggesting a possible involvement of His28 in the K- to L-cluster conversion due to the extremely close location of this residue to one of the Cys ligands (Cys30) of the K1-cluster in the primary sequence (see Supplementary Fig. 1b) and, consequently, the tertiary structure of NifB. More importantly, the abolished activity of in MaNifBH43A in the K- to L-cluster transformation points to a critical role of His43 in coupling the K1- and K2-clusters into an 8Fe L-cluster.
a EPR spectra of IDS-oxidized MaNifBwt (1), MaNifBH28A (2), MaNifBH219A (3), and MaNifBH43A (4) upon addition of SAM. Formation of the L-cluster was monitored by the appearance of an L-cluster-specific S = 1/2 signal at g = 1.94 (dashed vertical line). All protein samples have a concentration of 15 mg mL−1. The EPR spectra were recorded at 5 mW and 20 K. b M-cluster maturation activity of MaNifBwt (1), MaNifBH28A (2), MaNifBH219A (3), and MaNifBH43A (4). The activity of M-cluster maturation was determined on the basis of the C2H2-reducing activity of reconstituted NifDK, using SAM-treated MaNifB proteins as the M-cluster sources. The EPR analysis was performed three times independently (n = 3 independent samples), and representative results are shown in (a). The maturation assay was performed five times independently (n = 5 independent samples), and data are presented as mean ± S.D. (b). See Methods for the detailed composition of maturation assays.
To further explore the role of His43 in this process, high performance liquid chromatography (HPLC) was performed to examine the products generated upon incubation of MaNifBH43A with SAM. Like MaNifBwt, MaNifBH28A, and MaNifBH219A (Fig. 3a, b, traces 1–3), MaNifBH43A (Fig. 3a, b, trace 4) can cleave SAM into S-adenosyl-L-homocysteine (SAH) and 5′-deoxyadenosine (5′-dAH). In addition, as observed in the cases of MaNifBwt, MaNifBH28A, and MaNifBH219A (Fig. 3c, traces 1–3), formation of methanethiol is detected upon acid quench of an incubation mixture of MaNifBH43A and SAM (Fig. 3c, trace 4). This observation is not particularly surprising because our previous study demonstrates that SAH, 5′-dAH and methanethiol can be generated as long as both SAM- and K2-clusters are present, even in absence of the K1-cluster. The fact that substitution of the K1-specific ligand, His43, with Ala does not impact the reactivities associated with the SAM- and K2-clusters is consistent with this observation and places the perturbation of the K- to L-cluster conversion by this substitution after the hydrogen atom abstraction from the K2-associated methyl group (see Supplementary Fig. 1a).
a, b HPLC elution profiles of SAH (a) and 5′-dAH (b) upon incubation of SAM with MaNifBwt (1), MaNifBH28A (2), MaNifBH219A (3), and MaNifBH43A (4). c GC analyses of methanethiol (Me-SH) formation upon acid quenching of incubation mixtures containing SAM and MaNifBwt (1), MaNifBH28A (2), MaNifBH219A (3), and MaNifBH43A (4). All products were identified using standards13,14. The HPLC (a, b) and GC (c) experiments were each performed three times independently (n = 3 independent samples), and representative results are shown in the figure. The protein and SAM concentrations are 0.4 mM and 4 mM, respectively, in (a, b); and 40 μM and 0.3 mM, respectively, in (c). See Methods for detailed compositions of these assays.
Dual actions of histidine ligand in L-cluster maturation
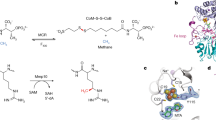
X-ray absorption spectroscopy (XAS)/extended x-ray absorption fine structure (EXAFS) analysis provided further insights into the role of His43 in L-cluster maturation. XAS/EXAFS analysis has proven to be a valuable tool for obtaining structural information of the cluster species related to the function and assembly of nitrogenase, and previous studies of the wildtype and variant MaNifB proteins have established XAS/EXAFS parameters that can be used in combination with the EPR and biochemical data to conclusively assign cluster species and monitor cluster transformation in this protein. The x-ray absorption near edge structure (XANES) data reveal a K-edge energy for MaNifBH43A similar to that for MaNifBwt before or after incubation with SAM (Fig. 4a), suggesting a similar sulfur-rich environment in all these protein species16. The pre-edge feature of MaNifBH43A is also similar in intensity to that of MaNifBwt before incubation with SAM; however, its amplitude does not increase as much as that of MaNifBwt upon incubation with SAM (Table 1), suggesting that the transition metal center in SAM-treated MaNifBH43A (designated MaNifBH43A/SAM) is less distorted away from centrosymmetry23,24 than that in SAM-treated MaNifBwt (designated MaNifBwt/SAM). Consistent with this observation, the smoothed second derivative of the pre-edge data of MaNifBwt/SAM transitions from a single inverted peak at ~7112.6 eV to two inverted peaks at ~7112.6 eV and ~7114.5 eV, respectively (Fig. 4b). Such a change has been attributed to the conversion of the K-cluster (with typical tetrahedral Fe-site geometries25) to an L-cluster (with an unusual intermediate geometry between tetrahedral and trigonal pyramidal25) in MaNifBwt upon incubation with SAM16. In the case of MaNifBH43A, while a similar peak at ~7112.6 eV is observed in the second derivative before and after incubation with SAM, the line-shapes of these plots beyond 7113 eV are different than those of the corresponding MaNifBwt species (Fig. 4b). More importantly, the second peak at ~7114.5 eV is absent from the plot of MaNifBH43A/SAM, although MaNifBH43A/SAM seems to undergo a transition analogous to that of MaNifBwt/SAM on the basis of the similar line-shapes of their second derivative plots (Fig. 4b).
a Pre-edge regions of the normalized fluorescence spectra and (b) smoothed second derivatives of the pre-edge regions, (c) Fourier transforms of the EXAFS data (dotted) and the best fits of data (solid), and (d) k3-weighted EXAFS data (dotted) and the best fits of data (solid). Spectra are shown for MaNifBwt before (1) and after (2, designated MaNifBwt/SAM) incubation with SAM, and MaNifBH43A before (3) and after (4, designated MaNifBH43A/SAM) incubation with SAM. Note that MaNifBwt carries the K-cluster (a [Fe4S4] cluster pair); whereas MaNifBwt/SAM carries the L-cluster ([Fe8S9C]). The peaks at ~7112.6 eV and ~7114.5 eV of the pre-edge regions are indicated by dashed vertical lines to illustrate the transition from a single peak at ~7112.6 eV to two peaks at ~7112.6 eV and ~7114.5 eV in the spectrum of MaNifBwt (b, 1 vs. a, 2), which corresponds to conversion of the K-cluster to an L-cluster upon incubation of MaNifBwt with SAM. Such a change is not observed in the case of MaNifBH43A following the same treatment with SAM (b, 3 vs. a, 4). The XAS analysis was performed three times independently (n = 3 independent samples), and representative results are shown in the figure. All protein samples have a concentration of 50 mg mL−1. All scans were taken at ~10 K. See Supporting Information for more details of EXAFS fits.
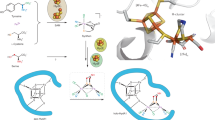
Extended X-ray absorption fine structure (EXAFS) analysis of the Fe K-edges of MaNifBH43A provided important insights into the structural metrics of its associated cluster species. Prior to SAM treatment, MaNifBH43A and MaNifBwt display two similar features at R + Δ ~1.7 and 2.4 Å, respectively, in the Fourier transforms (FT; Fig. 4c) of their EXAFS data (Fig. 4d), although the feature of MaNifBH43A at R + Δ ~2.4 Å is much more prominent than that of MaNifBwt. For MaNifBH43A, these FT features can be best fit with Fe–S and Fe∙∙∙Fe scatterers at 2.29 and 2.71 Å, respectively; whereas for MaNifBwt, they are best fit with Fe–S scatterers at 2.29 Å and Fe∙∙∙Fe scatterers at 2.51 and 2.69 Å, respectively (Table 2). Upon incubation with SAM, MaNifBH43A/SAM and MaNifBwt/SAM seemingly undergo similar changes, both showing extra FT features at R + Δ ~3.0 and ~3.5 Å; yet, while MaNifBwt/SAM displays a substantially increased intensity and a clear shift of its FT feature at R + Δ ~2.4 Å, the corresponding FT feature of MaNifBH43A/SAM remains largely unchanged (Fig. 4c). The differences between the two SAM-treated MaNifB species are clearly illustrated in the best fits of their EXAFS data: MaNifBH43A/SAM is best modeled with two types of Fe–S scatterers at 2.27 and 3.88 Å, respectively, and one type of Fe∙∙∙Fe scatterers at 2.69 Å; whereas MaNifBwt/SAM is best modeled with one type of Fe–S scatterers at 2.23 Å and two types of Fe∙∙∙Fe scatterers at 2.64 and 3.70 Å, respectively (Table 2). Most notably, the long-range Fe∙∙∙Fe distance at 3.70 Å, which originates from the intercubane scattering between the six carbide-coordinated Fe atoms at the cofactor core25, is present only in MaNifBwt/SAM but absent from MaNifBH43A/SAM. This observation suggests that unlike MaNifBwt, MaNifBH43A does not enable the formation of an L-cluster with a μ6-coordinated central carbide in place upon incubation with SAM. In support of this argument, MaNifBH43A carries clusters with short-range Fe∙∙∙Fe distances that are characteristic of the [Fe4S4] clusters before and after incubation with SAM (Table 2). However, modeling of the cluster species on MaNifBH43A/SAM, contrary to that of the cluster species on MaNifBH43A, requires the inclusion of an extra Fe–S distance at 3.88 Å (Table 2) that corresponds to the distance from a sulfide to a Fe at the opposite vertex of a [Fe4S4] cluster. The appearance of such a distance is consistent with an increased order of the two K-cluster units (i.e., K1 and K2), or a further processing of these [Fe4S4] units into a cluster intermediate between the K- and L-clusters on MaNifBH43A/SAM; more importantly, it highlights an overall homogeneity of the cluster species on MaNifBH43A/SAM, as this Fe–S distance can only be observed when the clusters are well-ordered and, therefore, uniform in nature. Overall, the mean squared deviations (σ2) of the S and Fe scatterers are rather small (<5 × 10−3 Å2; see Supplementary Tables 1 and 2), which further supports the homogeneity of the cluster species in the MaNifB proteins. Three most likely configurations can be proposed for this cluster intermediate: (i) a partial [Fe4S3] cluster pair bridged by two sulfide (S2-) atoms and a carbon (CHx) species, (ii) a face-on [Fe4S4] cluster pair bridged by a CHx species, and (iii) a vertex-on [Fe4S4] cluster pair bridged by a CHx species, all of which lack the characteristic cofactor core structure that is defined by an interstitial carbide coordinated with six Fe atoms (Supplementary Fig. 4). Given the homogeneity of the cluster species on MaNifBH43A/SAM, it is likely that one of these proposed models will be identified through future structural characterization of this protein.
Discussion
Formation of a cluster intermediate on the SAM-treated MaNifBH43A that is distinct from both the K- and L-clusters implies that His43 serves as a key structural element and/or reaction component during the process of cofactor core formation on MaNifB. The substantially increased intensity of the FT feature of MaNifBH43A at R + Δ ~2.4 Å relative to that in MaNifBwt prior to incubation with SAM (see Fig. 4c) points to a much stronger Fe∙∙∙Fe scattering that results from a much closer distance between the two [Fe4S4] units of the K-cluster in MaNifBH43A than those in MaNifBwt. Such a change in the distance and/or orientation of the K1-cluster relative to that of the K2-cluster apparently presents a challenge for MaNifBH43A to initiate a proper coupling between the two K-cluster units. The His43 ligand, therefore, could play a steric role in keeping the two K-cluster units in the correct distance/orientation to each other by either indirectly pulling the K1-cluster away from the K2-cluster via its ligand capacity or directly separating the K1- and K2-clusters with its bulky imidazole ring (Supplementary Fig. 4, ➀). Additionally, given our observation of a loss of the nitrogen ligand upon conversion of the K-cluster to an L-cluster on MaNifB11, His43 may lose its coordination to the K1-cluster via protonation, thereby freeing up K1 for the subsequent coupling with K2 into an L-cluster, and facilitating the release of the completed L-cluster to the next biosynthetic apparatus for further maturation. As such, His43 likely functions as a molecular switch via reversible protonation and deprotonation events, securing the cluster in place in its deprotonated state while giving the cluster certain structural flexibility in its protonated state to accommodate the different states required for cluster conversion. A similarly labile nitrogen ligand is also found in mitoNEET, where protonation of a His ligand to a [Fe2S2] cluster permits transfer of the cluster to downstream acceptor proteins26,27.
In light of this proposal, it is interesting to consider a coupling of the function of His43 as a molecular switch with another role of this residue in cofactor core formation, one that is involved in the further deprotonation/dehydrogenation of the initial methylene radical to yield the interstitial carbide at the center of the L-cluster. The observation that the two K-cluster units in SAM-treated MaNifBH43A become more aligned with each other but remain largely separate FeS cubanes in character (Supplementary Fig. 4, ➁) is consistent with an interruption of the carbide formation/insertion process that is required for the coupling/rearrangement of K1 and K2 into the geometry of a cofactor core. Consequently, the cluster is rendered in an unfinished state with the carbon intermediate (CHx) not fully deprotonated or dehydrogenated and attached to one or both of the K-cluster units (Supplementary Fig. 4, ➂). It is important to note that, other than His43, additional residues may also be involved in processing the carbon intermediate to an interstitial carbide, as substitution of His28 with Ala apparently reduces the efficiency of L-cluster formation on NifB by 41% (see Fig. 2b). A proton relay mechanism involving multiple histidine (or equivalent) residues may be employed in this case to facilitate efficient deprotonation/dehydrogenation of the initial methylene radical to eventually yield a carbide in the center of the L-cluster. While the role of His43 and other relevant players in the cofactor core formation process is yet to be elucidated, the results of this study provide an important framework for further investigations into the unique radical chemistry underlying the formation of the core structure of the nitrogenase cofactor. Knowledge obtained from these studies will contribute to a better understanding of the mechanism of nitrogenase and shed important light on the mechanisms of analogous biological systems.
Methods
General information
Unless otherwise specified, all chemicals were purchased from Sigma-Aldrich (St. Louis, MO) and Thermo Fisher Scientific (Waltham, MA), and all experiments were performed under an Ar atmosphere using Schlenk techniques and a glove box operating at <3 ppm O2.
Cell growth and protein purification
E. coli strains expressing His-tagged MaNifBwt (strain YM114EE), MaNifBH28A (strain YM242EE), MaNifBH43A (strain YM244EE), MaNifBH219A (strain YM246EE), MaNifBK1 (strain YM165EE) and MaNifBK1-H43A (strain YM307EE) were grown in 10-L batches in Difco LB medium containing 100 mg L-1 ampicillin (BD Biosciences) in a BIOFLO 415 fermenter (New Brunswick Scientific) at 37 °C, with 200 rpm agitation and 10 L˄min-1 airflow. When OD600 reached 0.5, the temperature was lowered to 25 °C before expression of the wildtype and variant MaNifB proteins was induced by addition of 25 µM IPTG. Expression of proteins was allowed to continue for 16 h before cells were harvested by centrifugation using a Thermo Fisher Scientific Legend XTR centrifuge. Subsequently, His-tagged MaNifB proteins were purified by immobilized metal affinity chromatography (IMAC)9,10.
SDS-PAGE analysis
The purified wildtype and variant MaNifB proteins were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis on a 4-20% precast Tris-glycine gel (Bio-Rad).
Iron determination
The iron contents of the FeS-reconstituted wildtype and variant MaNifB proteins were determined by inductively coupled plasma optical emission spectroscopy (ICP-OES) using a Thermo Scientific iCAP7000. Stock solutions of elemental iron (1 mg mL−1, Inorganic Ventures) were diluted to make standard solutions for calibration. Each protein sample was mixed with 100 µL concentrated sulfuric acid (H2SO4) and 100 µL concentrated nitric acid (HNO3) and heated at 250 °C for 30 min. This procedure was repeated until the solutions became colorless. The solution was then cooled to room temperature and diluted to a total volume of 10 mL with 2% HNO3 prior to sample analysis.
FeS cluster reconstitution
The purified wildtype or variant MaNifB was treated with 20 mM bathophenanthroline disulfonate, an iron chelator, in a buffer containing 2 mM dithionite (DT; Na2S2O4), 50 mM Tris-HCl (pH 8.0) and 500 mM NaCl, followed by incubation at room temperature for 1 h to remove the endogenous FeS clusters associated with the protein. Subsequently, this mixture was diluted with a buffer containing 50 mM Tris-HCl (pH 8.0) and loaded on a Q Sepharose column (GE Healthcare). The column was then washed with a buffer containing 2 mM DT, 50 mM Tris-HCl (pH 8.0) and 50 mM NaCl prior to elution of the MaNifB protein with a buffer containing 50 mM Tris-HCl (pH 8.0) and 500 mM NaCl. Dithionite was removed by running protein through a Sephadex G-25 (GE Healthcare) column equilibrated with 50 mM Tris-HCl (pH 8.0) and 10% glycerol. Reconstitution of the wildtype or variant MaNifB protein with synthetic [Fe4S4] clusters15 was carried out by adding a dimethylformamide (DMF) solution of synthetic [Fe4S4] cluster dropwise at a molar ratio of 5:1 to the MaNifB protein in a buffer containing 20 mM β-mercaptoethanol and 50 mM Tris-HCl (pH 8.0), with continuous stirring on ice. After incubation on ice for 1 h, the reaction mixture was diluted with a buffer containing 2 mM DT and 50 mM Tris-HCl (pH 8.0) and loaded on a Q Sepharose column. The column was then washed with a buffer containing 2 mM DT, 50 mM Tris-HCl and 50 mM NaCl prior to elution of the reconstituted MaNifB with a buffer containing 2 mM DT, 50 mM Tris-HCl (pH 8.0) and 500 mM NaCl. Reconstituted wildtype and variant MaNifB proteins were subjected to metal determination, activity assays and EPR analysis.
Cofactor maturation assays
The cofactor maturation assay contained, in a total volume of 1.0 mL, 25 mM Tris-HCl (pH 8.0), 20 mM DT, 3.5 mg FeS-reconstituted wildtype or variant MaNifB, 10 mM SAM, 2 mg ΔnifB AvNifEN, 1.4 mg NifH, 0.8 mM ATP, 1.6 mM MgCl2, 10 mM creatine phosphate, 8 units creatine phosphokinase, 0.3 mM homocitrate, 0.3 mM Na2MoO4 and 0.5 mg ΔnifB AvNifDK. This mixture was incubated at 30 °C for 30 min before it was examined for enzymatic activities9,10.
S-adenosyl-L-methionine (SAM) cleavage assays
The SAM cleavage assay contained, in a total volume of 0.3 mL, 25 mM Tris-HCl (pH 8.0), 5% glycerol (v/v), 40 µM wildtype or variant MaNifB, and 0.3 mM SAM. Assays were incubated at 25 °C for 60 min with intermittent mixing, before they were terminated by filtration through Amicon Ultra 30,000 MWCO centrifugal filters. Samples were then supplemented by trifluoroacetic acid (TFA) to a concentration of 0.14% before being analyzed by a Thermo Scientific Dionex Ultimate 3000 UHPLC system equipped with an Acclaim 120 C18 column (4.6 × 100 mm, 5-µm particle size). The flow rate of buffer was 0.5 mL˄min-1, and the column was kept at 30 °C. The column was equilibrated with 98% buffer A (50 mM KH2PO4, pH 6.6) and 2% buffer B (100% methanol) before each injection of a 100-µL sample. After sample injection, a linear gradient of 2–60% buffer B was applied over 20 min, followed by 8 min of isocratic flow with 60% buffer B and a linear gradient of 60–2% buffer B over 4 min. Elution of products was monitored at a UV wavelength of 254 nm. After each run, the column was equilibrated for 5 min with 2% buffer B before the injection of the next sample.
Acid quench experiments
Detection of MaNifB-dependent production of methanethiol was performed14. First, excess DT was removed from wildtype and variant MaNifB via gel filtration with Sephadex G-25 fine resin that was equilibrated with a buffer containing 25 mM Tris-HCl (pH 8.0). Immediately following the removal of excess reductant, 40 nmol of MaNifB was added to a sealed 300-μL glass vial that contained 400 nmol SAM in a total volume of 100 μL. These 100-μL reactions were then incubated for 30 min at 25 °C before being quenched by 25 μL of 1 M HCl. To observe the formation of the volatile methanethiol, the acid-quenched samples were incubated at 60 °C for 15 min and equilibrated to room temperature for 10 min before the entire headspace was injected by a gas-tight syringe onto a GC–MS (Thermo-Fisher Scientific Trace 1300 GC connected to a Thermo-Fisher Scientific ISQ QD single quadrupole mass spectrometry) with a Restek Rxi-1ms column (30 m, 0.32 mm ID, 4.0 μm df). The GC inlet and oven temperatures were maintained at 30 °C, while the mass spectrometry transfer line and ion source were maintained at 250 °C. Total ion chromatograms were generated under SIM conditions in electron ionization mode, and methanethiol was detected at an m/z ratio of 47. The base peaks were selected on the basis of the characterization of standard samples (Sigma-Aldrich) under full scan conditions and comparison to those reported in the National Institute of Standards and Technology database.
Electron paramagnetic resonance (EPR) analysis
Sample preparation was carried out in a Vacuum Atmospheres glove box with less than 1 ppm O2 and flash frozen in liquid nitrogen before analysis. The dithionite-reduced samples were prepared by incubating 50 μM wildtype or variant MaNifB with 40 mM SAM for 15 min, followed by re-isolation of MaNifB into a buffer containing 50 mM Tris-HCl (pH 8.0), 500 mM NaCl, and 2 mM dithionite (DT). The indigo disulfonate (IDS)-oxidized samples were prepared by incubating the re-isolated MaNifB with excess IDS for 5 min, followed by removal of excess IDS was using a Sephadex G-25 desalting column. The DT-reduced or IDS-oxidized MaNifB sample was then concentrated to 15 mg mL−1, followed by transfer of the sample into EPR tubes. The samples were then flash frozen and stored in liquid nitrogen. CW EPR spectra were recorded by an ESP 300 Ez spectrophotometer (Bruker) interfaced with an ESR-9002 liquid-helium continuous-flow cryostat (Oxford Instruments) using a microwave power of 5 mW (IDS-oxidized samples) or 50 mW (IDT-reduced samples), a gain of 5 × 104, a modulation frequency of 100 kHz, and a modulation amplitude of 5 G. Five scans was recorded for each sample in perpendicular mode at 20 K (IDS-oxidized samples) or 10 K (DT-reduced samples) using a microwave frequency of 9.62 GHz.
All pulse EPR studies were carried out at the UC Davis CalEPR center, using a Bruker EleXsys E580 pulse EPR spectrometer equipped with an Oxford-CF935 liquid-helium cryostat and an ITC-503 temperature controller. Pulse data were collected using a Bruker MS5 probe (X-band) or an R.A. Isaacson-designed cylindrical TE011 resonator (Q-band)28 adapted for pulse EPR in an Oxford Instruments CF935 cryostat. Three-pulse ESEEM spectra were collected using the pulse sequence π/2-τ-π/2-T-π/2-τ-echo where the delay time, T, was increased by 16 ns steps. ESEEM spectra were recorded at 10 K, τ = 128-144 ns (values chosen to minimize proton modulations to the spectra), π/2 = 12 ns, and a microwave frequency of 9.3366 GHz. Spectral processing (subtraction of a background exponential, application of a hamming window, and FFT) were performed using the EasySpin 5.2.27 toolbox in Matlab R2019a29.
X-ray absorption spectroscopy (XAS) analysis
The XAS samples were prepared the same way as the EPR samples (see above). The sample concentrations were 50 mg mL−1. Fe K-edge X-ray absorption spectra were collected on SSRL beam line 7-3 using a 30-element solid state Ge detector (Canberra) with a SPEAR3 storage ring current of ~500 mA at an energy of 3.0 GeV. The BL7-3 optics consists of a flat, bent, harmonic rejection vertically collimating Rh-coated Si M0 mirror and a liquid nitrogen cooled double crystal Si(220) monochromator. A total of 7 and 6 scans, respectively, were collected for MaNifBH43A before and after incubation with SAM (designated MaNifBH43A and MaNifBH43A/SAM). All scans were taken between 6882 and 8000 eV at ~10 K using an Oxford Instruments CF1208 continuous flow liquid-helium cryostat using a closed-cycle cooled He gas loop. An iron foil was placed in the beam pathway prior to the ionization chamber I0 and scanned concomitantly for an energy calibration, with the first inflection point of the edge assigned to 7112.0 eV. A Soller slit with a 3 μm Mn filter was used to increase the signal-to-noise ratio of the spectra. Photoreduction was monitored by scanning the same spot on the sample twice and comparing the first derivative peaks associated with the edge energy during data collection.
The detector channels from the scans were examined, calibrated and averaged using EXAFSPAK30 and then processed for EXAFS analysis using PYSPLINE31 to extract χ(k). PYSPLINE was used to subtract a second-order background from the entire range of data and subsequently generate a spline function to model background absorption through the EXAFS region. A four-region spline was chosen with 2, 3, 3 order polynomials over the post edge region, and the data were normalized to have an edge jump of 1.0 at 7130 eV. Following a modified data analysis protocol25, the Fe K-edge EXAFS data for the clusters associated with MaNifBH43A and MaNifBH43A/SAM were generated by subtracting the k-weighted EXAFS data, χ(k), of MaNifBSAM (an MaNifB variant carrying only the SAM-cluster11) from the χ(k) of the samples in a 1:2 ratio on the basis of the proportionate iron quantity for each cluster species (i.e., 4 Fe for the SAM-cluster and 8 Fe for the K-clusters). Theoretical phase and amplitude parameters for a given absorber–scatterer pair were calculated using FEFF 8.4032 and subsequently applied to the nonlinear least squares Opt fitting program of the EXAFSPAK package during curve fitting. Parameters for each species were calculated using an appropriate model derived from either the crystal structure of the M-cluster in NifDK (PDB code 3U7Q)5, where the Mo atom was exchanged for an Fe atom, or from the [Fe4S4] cluster in NifH (PDB code 1G5P)33 because there are no available crystal structures of MaNifB. In all analyses, the coordination number of a given shell (N) was a fixed parameter and was varied iteratively in integer steps, whereas the bond lengths (R) and mean-square deviation (σ2) were allowed to freely float. The estimated uncertainties in R, σ2, and N are 0.02 Å, 0.1 × 10-3 Å2, and 20%, respectively. The amplitude reduction factor S0 was fixed at 1.0 for the Fe K-edge data, whereas the edge-shift parameter ΔE0 was allowed to float as a single value for all shells. Thus, in any given fit, the number of floating parameters was typically equal to 2 × number of shells + 1. The goodness of fit (GOF) parameters were calculated as follows:
The Fe K-edge data were analyzed with a k range of 2–11.2 Å−1 (ΔR = 0.17 Å) to allow comparison between previously reported data16, although the data could be analyzed with higher resolution with a k range of 2–14 Å−1 for MaNifBH43A and MaNifBH43A/SAM. Pre-edge analysis was performed on the Fe K-edge fluorescence data normalized to have an edge jump of 1.0 at 7130 eV in PYSPLINE. The pre-edge features were fit as described elsewhere23 between 7108 and 7117 eV using the Fityk34 program with pseudo-Voigt functions composed of 50:50 Gaussian/Lorentzian functions.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availibility
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Information files and from the corresponding authors upon reasonable request. The National Institute of Standards and Technology database is available at https://www.nist.gov/data. Parameters used for cluster modeling are available at https://www.rcsb.org/ using PDB IDs 3U7Q and 1G5P.
References
Burgess, B. K. & Lowe, D. J. Mechanism of molybdenum nitrogenase. Chem. Rev. 96, 2983–3012 (1996).
Rees, D. C. et al. Structural basis of biological nitrogen fixation. Philos. Trans. A: Math. Phys. Eng. Sci. 363, 971–984 (2005).
Hoffman, B. M., Lukoyanov, D., Yang, Z. Y., Dean, D. R. & Seefeldt, L. C. Mechanism of nitrogen fixation by nitrogenase: the next stage. Chem. Rev. 114, 4041–4062 (2014).
Einsle, O. et al. Nitrogenase MoFe-protein at 1.16 Å resolution: a central ligand in the FeMo-cofactor. Science 297, 1696–1700 (2002).
Spatzal, T. et al. Evidence for interstitial carbon in nitrogenase FeMo cofactor. Science 334, 940 (2011).
Lancaster, K. M. et al. X-ray emission spectroscopy evidences a central carbon in the nitrogenase iron-molybdenum cofactor. Science 334, 974–977 (2011).
Frey, P. A., Hegeman, A. D. & Ruzicka, F. J. The radical SAM superfamily. Crit. Rev. Biochem. Mol. Biol. 43, 63–88 (2008).
Broderick, J. B., Duffus, B. R., Duschene, K. S. & Shepard, E. M. Radical S-adenosylmethionine enzymes. Chem. Rev. 114, 4229–4317 (2014).
Wiig, J. A., Hu, Y. & Ribbe, M. W. NifEN-B complex of Azotobacter vinelandii is fully functional in nitrogenase FeMo cofactor assembly. Proc. Natl Acad. Sci. U.S.A. 108, 8623–8627 (2011).
Fay, A. W., Wiig, J. A., Lee, C. C. & Hu, Y. Identification and characterization of functional homologs of nitrogenase cofactor biosynthesis protein NifB from methanogens. Proc. Natl Acad. Sci. U.S.A. 112, 14829–14833 (2015).
Rettberg, L. A. et al. Probing the coordination and function of Fe4S4 modules in nitrogenase assembly protein NifB. Nat. Commun. 9, 2824 (2018).
Wilcoxen, J. et al. Electron paramagnetic resonance characterization of three iron-sulfur clusters present in the nitrogenase cofactor maturase NifB from Methanocaldococcus infernus. J. Am. Chem. Soc. 138, 7468–7471 (2016).
Wiig, J. A., Hu, Y., Lee, C. C. & Ribbe, M. W. Radical SAM-dependent carbon insertion into the nitrogenase M-cluster. Science 337, 1672–1675 (2012).
Wiig, J. A., Hu, Y. & Ribbe, M. W. Refining the pathway of carbide insertion into the nitrogenase M-cluster. Nat. Commun. 6, 8034 (2015).
Tanifuji, K. et al. Tracing the ‘9th sulfur’ of the nitrogenase cofactor via a semi-synthetic approach. Nat. Chem. 10, 568–572 (2018).
Jasniewski, A. J. et al. Spectroscopic characterization of an eight-iron nitrogenase cofactor precursor that lacks the ‘9th sulfur’. Angew. Chem. Int. Ed. Engl. 58, 14703–14707 (2019).
Hu, Y. et al. FeMo cofactor maturation on NifEN. Proc. Natl Acad. Sci. U.S.A. 103, 17119–17124 (2006).
Yoshizawa, J. M. et al. Optimization of FeMoco maturation on NifEN. J. Am. Chem. Soc. 131, 9321–9325 (2009).
Kaiser, J. T., Hu, Y., Wiig, J. A., Rees, D. C. & Ribbe, M. W. Structure of precursor-bound NifEN: a nitrogenase FeMo cofactor maturase/insertase. Science 331, 91–94 (2011).
Fay, A. W. et al. Spectroscopic characterization of the isolated iron-molybdenum cofactor (FeMoco) precursor from the protein NifEN. Angew. Chem. Int. Ed. 50, 7787–7790 (2011).
Hu, Y. et al. Nitrogenase Fe protein: a molybdate/homocitrate insertase. Proc. Natl Acad. Sci. U.S.A. 103, 17125–17130 (2006).
Schmid, B. et al. Structure of a cofactor-deficient nitrogenase MoFe protein. Science 296, 352–356 (2002).
Westre, T. E. et al. A multiplet analysis of Fe K-edge 1s → 3d pre-edge features of iron complexes. J. Am. Chem. Soc. 119, 6297–6314 (1997).
Sarangi, R. X-ray absorption near-edge spectroscopy in bioinorganic chemistry: application to M–O2 systems. Coord. Chem. Rev. 257, 459–472 (2013).
Corbett, M. C. et al. Structural insights into a protein-bound iron-molybdenum cofactor precursor. Proc. Natl Acad. Sci. U.S.A. 103, 1238–1243 (2006).
Karmi, O.et al. The unique fold and lability of the[2Fe-2S] clusters of NEET proteins mediate their key functions in health anddisease. J. Biol. Inorg. Chem. 23, 599–612 (2018).
Tamir, S. et al. Structure–function analysis of NEET proteins uncovers their role as key regulators of iron and ROS homeostasis in health and disease. Biochim. Biophys. Acta 1853, 1294–1315 (2015).
Flores, M. et al. Probing hydrogen bonding to quinone anion radicals by 1H and 2H ENDOR spectroscopy at 35 GHz. J. Chem. Phys. 294, 401–413 (2003).
Stoll, S. & Schweiger, A. EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J. Magn. Reson. 178, 42–55 (2006).
George, G. N. in EXAFSPAK: A Suite of Computer Programs for Analysis of X-ray Absorption Spectra, https://www-ssrl.slac.stanford.edu/exafspak.html (Stanford Synchrotron Radiation Laboratory, Stanford Linear Accelerator Center: Menlo Park, CA, 1990).
Tenderholt, A., Hedman, B. & Hodgson, K. O. PySpline: a modern, cross‐platform program for the processing of raw averaged XAS edge and EXAFS data. AIP Conf. Proc. 882, 105–107 (2007).
Ankudinov, A. L., Ravel, B., Rehr, J. J. & Conradson, S. D. Real-space multiple-scattering calculation and interpretation of X-ray-absorption near-edge structure. Phys. Rev. B 58, 7565–7576 (1998).
Strop, P. et al. Crystal structure of the all-ferrous [4Fe-4S]0 form of the nitrogenase iron protein from Azotobacter vinelandii. Biochemistry 40, 651–656 (2001).
Wojdyr, M. Fityk: a general-purpose peak fitting program. J. Appl. Crystallogr. 43, 1126–1128 (2010).
Acknowledgements
This work was supported by NIH-NIGMS grant GM67626 (to M.W.R. and Y.H.) and NIH-NIHGMS grant 1R35GM126961-01 (to R.D.B.).
Author information
Authors and Affiliations
Contributions
L.A.R., J.W., A.J.J., C.C.L., and K.T. performed experiments and analyzed data; J.W., Y.H., R.D.B., and M.W.R. designed experiments, analyzed data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rettberg, L.A., Wilcoxen, J., Jasniewski, A.J. et al. Identity and function of an essential nitrogen ligand of the nitrogenase cofactor biosynthesis protein NifB. Nat Commun 11, 1757 (2020). https://doi.org/10.1038/s41467-020-15627-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-15627-9
This article is cited by
-
Tracing the incorporation of the “ninth sulfur” into the nitrogenase cofactor precursor with selenite and tellurite
Nature Chemistry (2021)
-
Bioassembly of complex iron–sulfur enzymes: hydrogenases and nitrogenases
Nature Reviews Chemistry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.