Abstract
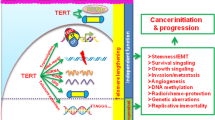
Cancers require telomere maintenance mechanisms for unlimited replicative potential. They achieve this through TERT activation or alternative telomere lengthening associated with ATRX or DAXX loss. Here, as part of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, we dissect whole-genome sequencing data of over 2500 matched tumor-control samples from 36 different tumor types aggregated within the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium to characterize the genomic footprints of these mechanisms. While the telomere content of tumors with ATRX or DAXX mutations (ATRX/DAXXtrunc) is increased, tumors with TERT modifications show a moderate decrease of telomere content. One quarter of all tumor samples contain somatic integrations of telomeric sequences into non-telomeric DNA. This fraction is increased to 80% prevalence in ATRX/DAXXtrunc tumors, which carry an aberrant telomere variant repeat (TVR) distribution as another genomic marker. The latter feature includes enrichment or depletion of the previously undescribed singleton TVRs TTCGGG and TTTGGG, respectively. Our systematic analysis provides new insight into the recurrent genomic alterations associated with telomere maintenance mechanisms in cancer.
Similar content being viewed by others
Introduction
Telomeres are nucleoprotein complexes at the ends of chromosomes that prevent DNA degradation and genome instability1. The typically 10–15 kb long chromosome termini are composed of long stretches of TTAGGG (t-type) repeat arrays with an increasing number of variants toward proximal, subtelomeric regions, the most common being TGAGGG (g-type), TCAGGG (c-type), and TTGGGG (j-type) repeats2,3.
Telomeres play an important role in cellular aging, as they are shortened with each cell division and finally trigger a DNA damage response resulting in senescence4,5. To avoid this permanent growth arrest, cells with unlimited proliferative potential need to extend their telomeres. In humans, telomeric DNA is synthesized by telomerase, an enzyme that is composed of the reverse transcriptase TERT and the RNA template TERC. This complex is active in the germline and stem cells, but absent in most somatic cells6. Telomerase is upregulated in ~85% of human cancers by different genetic aberrations, including TERT amplifications7, rearrangements8, or mutations in the TERT promoter9,10. The remaining tumors employ an alternative lengthening of telomeres (ALT) pathway, which is based on DNA recombination of telomeric sequences11. Details on the ALT mechanism remain elusive, but it has been associated with loss-of-function mutations in the chromatin remodeling genes ATRX (α-thalassaemia/mental retardation syndrome X-linked) and DAXX (death domain-associated protein)12. Telomeres of ALT cells characteristically have heterogeneous lengths and contain a range of telomere variant repeats (TVRs)13,14,15. Other hallmarks of ALT include ALT-associated promyelocytic leukemia nuclear bodies, abundance of extrachromosomal telomeric repeats of various forms (such as C-circles), and genome instability11,16.
While normally located at the chromosome termini, telomere sequences are also found within chromosomes. As such, interstitial telomeric sequences with large blocks of telomere repeats exist in humans and other species, which probably arose from ancestral genome rearrangements or other evolutionary events17. Recently, also ALT-specific, targeted telomere insertions into chromosomes have been described that lead to genomic instability18. Another source for unexpected telomere repeat occurrence is the stabilizing function of telomeres at broken chromosomes. After a double-strand break, telomeres can be added de novo to the unprotected break sites (“telomere healing”)19,20 or acquired from other chromosomal positions (“telomere capture”)21,22.
The here presented study was conducted within the scope of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, which aggregated whole-genome sequencing (WGS) data from 2658 cancers across 38 tumor types generated by the ICGC and TCGA projects. This data was reanalyzed with standardized, high-accuracy pipelines to align to the human genome (reference build hs37d5), and identify germline variants and somatically acquired mutations, as described in ref. 23.
Here, we characterize the telomere landscape of 2519 tumor samples from 36 different tumor types using the WGS alignments, somatic mutation, and chromothripsis calls provided by the PCAWG Consortium23,24. Besides determining telomere content and searching for mutations associated with different telomere maintenance mechanisms (TMMs), we systematically detect 2683 somatic telomere insertions and show that different TMMs are associated with the enrichment of previously undescribed singleton TVRs.
Results
Telomere content across cohorts
Due to the repetitive nature of telomere sequences, short sequencing reads from telomeres cannot be uniquely aligned to individual chromosomes. However, a mean telomere content for the tumor as a whole can be estimated from the number of reads containing telomere sequences25. Here, we extracted reads containing at least six telomere repeats per 100 bases, allowing the canonical telomere repeat TTAGGG and the three most common TVRs TCAGGG, TGAGGG, and TTGGGG. The telomere content was defined as the number of unaligned telomere reads normalized by sequencing coverage and GC content. Of the 2583 high-quality tumor samples available in PCAWG, we selected those from donors with a single tumor sample. From each donor, a control sample was available. In most cases this consisted of a blood sample, but could also stem from tumor-adjacent or other tissue23. The telomere content was determined for the remaining 2519 tumor samples and matched controls from 36 different tumor types. Several of these tumor types were not covered in a recent pan-cancer overview of telomere lengths26, including medulloblastoma, pilocytic astrocytoma, chronic lymphocytic leukemia, pancreatic endocrine cancers, benign bone cancer, and osteosarcoma. All relevant donor information and results used in this study are summarized in Supplementary Data 1.
Telomere content of the controls anticorrelated with age (r = −0.36, Spearman correlation; Supplementary Fig. 1a). However, this age effect only has a low contribution to the strong correlation between the telomere content of the tumor and control samples (r = 0.47 and rpartial = 0.46 given the patient age, Spearman correlation, Supplementary Fig. 1b). Thus, the association of tumor and control telomere content must mainly be caused by other genetic27,28, environmental29, or technical factors26. We normalized for these contributions by computing the ratio of tumor and control telomere content per individual.
Most tumor samples had a lower telomere content than the matched control (Fig. 1a). However, there were systematic differences between the different tumor types. Among those with the highest telomere content increase were osteosarcomas and leiomyosarcomas (median telomere content tumor/control log2 ratios = 0.7 and 0.6, respectively). A particularly low telomere content was found in colorectal adenocarcinoma and medulloblastoma (median telomere content tumor/control log2 ratios = −1.0).
a Overview of the telomere content distribution of all analyzed tumor types. The number of samples in each tumor type is indicated. Cohorts with sample sizes <15 are not shown. b TMM-associated mutations in different tumor types. c Telomere content in samples with different TMM-associated mutations. d TERT expression in samples with different TMM-associated mutations. The center lines of the boxplots are the medians, the bounds of the boxes represent the first and third quartiles, the upper and lower whiskers extend from the hinge to the largest or smallest value, respectively, no further than 1.5 × IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). ****p < 0.0001, Wilcoxon rank-sum tests.
Prevalence of TMM-associated mutations
Different types of mutations in ATRX or DAXX, and at the TERT locus have been associated with ALT and telomerase activation, respectively. We therefore searched for these types of somatic mutations to infer the active TMM in a given tumor. Somatic mutations in ATRX, DAXX, or TERT were found in 16% of tumor samples. In total, 64 tumor samples had truncating ATRX (n = 53) or DAXX alterations (n = 11), and are referred to as ATRX/DAXXtrunc in the following analysis. Of note, 10 of the 11 DAXX alterations were found in pancreatic endocrine tumors, while ATRX mutations were seen in a wider variety of entities. An additional 46 samples had nontruncating ATRX/DAXX simple nucleotide variants. TERT alterations (TERTmod) were detected in 270 of the 2519 tumor samples (11%). The latter group comprised 198 activating C228T or C250T promoter mutations (of which 132 were obtained from the PCAWG simple nucleotide variant consensus calls and the remaining were detected with a targeted approach), 11 amplifications leading to at least six additional TERT copies, 55 structural variations within 20 kb upstream of TERT (TERTpSV), and 6 samples with more than one of these modifications. Additionally, 18 tumor samples had both ATRX/DAXX truncating or other missense mutations and TERT alterations.
“Enhancer hijacking” near the TERT transcription start site (TSS) has been described in neuroblastoma8 and has recently been indicated in further cancer types26. In our data set, the strikingly focal distribution of structural variations upstream of the TERT TSS also points to this phenomenon (Supplementary Fig. 2). Further evidence is given by the direct overlap of 40% (n = 25/62) of the juxtaposed positions within 20 kb upstream of the TERT TSS with enhancers from the dbSUPER database30. In contrast, only 13% (n = 9/69) of the juxtaposed positions between 20 and 1000 kb corresponded to a predicted super-enhancer. Besides in melanoma (13% prevalence), chromophobe renal cell (9%), hepatocellular (9%), bladder transitional cell (4%), biliary (3%), and renal clear cell carcinoma (3%), previously undescribed TERTpSV in osteosarcoma (9%) and stomach adenocarcinoma (1%) were found. Moreover, the subtype-specific histological classification available in this study showed that TERTpSV were more frequent in liposarcoma (32%) than in leiomyosarcoma (7%).
The tumor types with the highest prevalence of ATRX/DAXXtrunc mutations were liposarcomas (32%), adult lower grade gliomas (28%), pancreatic endocrine tumors (23%), and osteosarcoma (17%; Fig. 1b and Supplementary Fig. 3a, b), all of which have previously been associated with ALT12,31. TERTmod were most prevalent in transitional cell bladder cancer (70%), glioblastoma (67%), lower grade gliomas (61%), and melanoma (51%).
The telomere content in TERTmod samples differed significantly from that in ATRX/DAXXtrunc samples (p = 1.1 × 10−9, Wilcoxon rank-sum test; Fig. 1c, a detailed overview is shown in Supplementary Fig. 4a). On average, telomere content was gained in ATRX/DAXXtrunc (mean telomere content tumor/control log2 ratio = 0.3), while telomere sequences were lost in TERTmod samples (mean telomere content tumor/control log2 ratio = −0.4). Samples with nontruncating ATRX/DAXX simple nucleotide variants had a similar telomere content as TERTmod samples (p > 0.05, Wilcoxon rank-sum test), suggesting that most of the nontruncating ATRX/DAXX mutations are passenger events. In TERTmod samples and samples with unknown TMM, the telomere content correlated with TERT expression (r = 0.20, Pearson correlation; p = 4.1 × 10−10, significance of fitted linear regression model) and TERT expression was significantly higher in TERTmod samples than in ATRX/DAXXtrunc samples (p = 1.3 × 10−9, Wilcoxon rank-sum test; Fig. 1d, detailed overviews are shown in Supplementary Figs. 3c and 4b).
High amount of telomere insertions in ATRX/DAXXtrunc tumors
To find insertions of telomeres into nontelomeric regions of the genome, we searched for tumor-specific discordant paired-end reads, where one end maps to the chromosome and the other end is telomeric. Exact positions of the insertions were determined from reads spanning the junction site and visual inspection (Fig. 2).
a One-sided telomere insertion in liposarcoma sample SP121774. Blue tracks show the sequencing coverage; light blue represents clipped sequences. Individual reads are grey and clipped bases are colored. Dark grey reads represent the nontelomeric end of a discordant read pair. b Two-sided telomere insertion in breast adenocarcinoma sample SP5636. Nontelomeric clipped bases are transparent. c One-sided telomere insertion accompanied by copy number loss of the adjacent chromosome end in glioblastoma sample SP29559. Arches represent structural variations. d Multiple telomere insertions in a chromosome that underwent chromothripsis in melanoma sample SP124441. e One-sided telomere insertion accompanied by a translocation of the adjacent chromosome segment in pancreatic adenocarcinoma sample SP125764.
Overall, 2683 telomere insertions were detected. These were distributed unevenly between samples and different tumor types (Fig. 3a). Telomere insertions were found in 27% of the tumor samples, with counts ranging between 1 and 228 telomere insertion events. The tumor types with the highest amount of telomere insertions per tumor sample were liposarcoma, leiomyosarcoma, and osteosarcoma, all of which also had a relatively high mean telomere content. In fact, the number of telomere insertions positively correlated with the telomere content (r = 0.19, Spearman correlation). Moreover, the number of telomere insertions was associated with the number of genomic breakpoints in the sample (r = 0.38, Spearman correlation). To test for a synergistic effect, linear models that predict telomere insertions from telomere content and breakpoint abundance with and without an interaction term were computed. The models with the interaction term (p = 8.8 × 10−234) performed substantially better than purely additive models (p = 5.8 × 10−90).
a Number of telomere insertions in samples of different tumor types. The tumor types are sorted by mean telomere content tumor/control log2 ratios. Cohorts with sample sizes <15 are not shown. b Number of telomere insertions in samples with different TMM-associated mutations. c Number of breakpoints in samples with different TMM-associated mutations. d Percent of breakpoints coinciding with telomere insertions in samples with different TMM-associated mutations. The center line of the boxplot is the median, the bounds of the box represent the first and third quartiles, the upper and lower whiskers extend from the hinge to the largest or smallest value, respectively, no further than 1.5 × IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). ****p < 0.0001, Wilcoxon rank-sum test. e Copy number changes of adjacent segments accompanying telomere insertions. “Complex” means that the copy numbers between segments differ in more than four copies. Overlaps with regions of chromothripsis are indicated. For telomere insertions that did not overlap with regions of chromothripsis, structural variations, or additional telomere insertions within 10 kb are indicated.
There was clearly a higher percentage of samples with telomere insertions in ATRX/DAXXtrunc tumors (80%) than TERTmod tumors (28%; Fig. 3b). As expected, ATRX/DAXXtrunc samples also had a higher number of breakpoints (mean = 733) than TERTmod samples (mean = 291; Fig. 3c). Overall, the fraction of genomic breakpoints overlapping with telomere insertion sites was significantly higher in ATRX/DAXXtrunc than TERTmod samples (p = 1.7 × 10−20, Wilcoxon rank-sum test; Fig. 3d). In agreement with the high breakpoint frequency, chromothripsis (numerous chromosomal rearrangements occurring in a single event)32 was more prevalent in the ATRX/DAXXtrunc samples (59%) compared to TERTmod samples (34%) and samples without ATRX/DAXXtrunc and TERTmod mutations (29%). Similarly, ATRX/DAXXtrunc samples were more likely to have an autosomal breakage-fusion-bridge (BFB) event (44%) than the remaining samples (TERTmod: 31%, other: 32%). In ATRX/DAXXtrunc samples, autosomal chromosome arms that showed evidence for BFB cycles and chromothripsis had the highest incidence of telomere insertions (Supplementary Fig. 5).
Correlation analysis of telomere insertions and mutations in telomere maintenance-associated genes from the TelNet database33 [http://www.cancertelsys.org/telnet] revealed significant association with TP53 (q = 1.9 × 10−42), ATRX (q = 2.6 × 10−6), PLCB2 (q = 7.8 × 10−4), MEN1 (q = 0.017), TSSC4 (q = 0.017), RB1 (q = 0.018), DAXX (q = 0.019), and ABCC8 mutations (q = 0.04, Wilcoxon rank-sum tests after Benjamini–Hochberg correction). Most of these genes have been implicated in the maintenance of telomere length or structure in humans (Supplementary Table 1). The exceptions are PLCB2 and ABCC8, whose homologues have so far only been reported in association with telomere length regulation in yeast34,35.
The detected telomere insertions were scattered across different chromosomes and regions within the chromosome (Supplementary Fig. 6). No clear preferential insertion sites were identified, but several de novo telomere junctions occurred at the chromosome ends (5% within 50 kb of the first or last chromosomal segment). A total of 44% of the telomere insertions were in genes, and 8% of these disrupted exons. Several tumor suppressor genes were affected, e.g., CHEK1 encoding for a protein involved in cell cycle arrest upon DNA damage36 (Fig. 2a).
Of note, patterns of microhomology were observed in 79% of telomere insertions with t-type repeats at the junction site (Supplementary Fig. 7).
Frequent copy number losses at telomere insertion sites
Most of the telomere insertions were one-sided (98%), i.e., telomere sequences were only attached to one side of the breakpoint (Fig. 2a). Telomere insertions were defined as two-sided, if there was a second telomere insertion event downstream in the opposite orientation (Fig. 2b). Two-sided telomere insertions can occur via a telomere sequence that bridges two chromosome fragments or, alternatively, telomere sequences are independently fused to both ends of the chromosome break. Reads supporting the first scenario were found in 14 of the 25 two-sided telomere insertions pairings. For the other cases, the inserted repeat sequence was too long to distinguish between the two scenarios.
Because so many breakpoints were one-sided, we investigated the fate of the corresponding broken fragment using complementary information from copy number changes and structural variation annotation (Fig. 3e). As expected, one-sided telomere insertions coincided most frequently with copy number loss of the adjacent segment (46%, Fig. 2c). In contrast, copy number gains of the fragment were rare (6%). Surprisingly, telomere insertions were frequently located at copy-number neutral sites (42%). Overlaps with regions of chromothripsis were found for 25% and structural variations without chromothripsis overlap (including telomere insertions) were detected near the insertion site for 28% of the copy-number neutral cases (Fig. 2d, e). The remaining telomere insertions at copy-number neutral sites are likely to be subclonal (Supplementary Fig. 8a) or have undetected structural variations nearby (Supplementary Fig. 8b).
Occasional TERRA expression at telomere insertions
ALT-positive tumors have been associated with elevated levels of long noncoding telomeric repeat-containing RNA (TERRA)16. We searched for TERRA expression in the RNA-sequencing data of 867 tumor samples. In line with the results of Barthel et al.26, TERRA levels were higher in ATRX/DAXXtrunc compared to TERTmod samples (p = 5.0 × 10−7, Wilcoxon rank-sum test, Supplementary Fig. 9). In 16 samples, evidence for TERRA expression at telomere insertion sites was found. For most of these, the number of split reads supporting TERRA expression was low (between 1 and 8 reads). However, 146 TERRA reads expressed from only two telomere insertion sites were detected in an ATRX/DAXXtrunc liposarcoma sample, making up almost 6% of its total TERRA read count. This percentage is likely to be notably higher, as the short read length does not allow assignment of the total number of TERRA reads stemming from these telomere insertions. Thus, TERRA is not exclusively transcribed from TSSs in the subtelomeric region but can also arise from telomere insertions. Of note, these telomere insertion transcripts do not always contain the canonical UUAGGG repeat but can also be composed of the reverse complement CCCUAA.
Enrichment of singleton TVRs in ATRX/DAXXtrunc samples
It has previously been shown that ALT leads to an increased integration of TVRs into telomeres, the most common ones being hexamers of the type NNNGGG15. To detect differences in the telomere composition of ATRX/DAXXtrunc and TERTmod tumors, we therefore searched for NNNGGG repeats in telomere reads. The most frequent TVRs across all tumor samples were TGAGGG, TCAGGG, and TTGGGG (Supplementary Fig. 10), which are known to be enriched in proximal telomeric regions2,3.
These and the seven other most frequent TVRs (TAAGGG, GTAGGG, CATGGG, TTCGGG, CTAGGG, TTTGGG, and ATAGGG) were chosen to search for common telomere repeat combinations. For this, the neighboring 18 base pairs on either side of the TVRs were determined (Supplementary Data 2). Most TVRs were surrounded by many different pattern combinations (e.g., TTGGGG). Others were dominated by a certain repeat context, which was similar in ATRX/DAXXtrunc and TERTmod tumors (e.g., CATGGG or ATAGGG). However, TTCGGG stood out, as 41% of the TVRs in ATRX/DAXXtrunc samples were surrounded by canonical t-type repeats, whereas this context was observed for only 4% of TTCGGG TVRs in TERTmod tumors.
Following up on this observation, we compared variant hexamers surrounded by at least three t-type repeats on either side (“singletons”) to TVRs in an arbitrary sequence context. This revealed that singletons are generally well suited to distinguish ATRX/DAXXtrunc from TERTmod samples (Fig. 4a, an overview of all patterns is shown in Supplementary Fig. 11). The remaining variant analysis therefore focused on such TVR singletons. CATGGG was excluded as it did not occur as singletons. For the other TVRs, the median of absolute counts varied between 12 and 100, but counts in individual tumor samples reached >10,000 (Supplementary Fig. 12).
a Receiver operating characteristic for the classification of samples with ALT-associated mutations from telomere variant repeats. Red: no specific sequence context required. Blue: singletons ((TTAGGG)3-NNNGGG-(TTAGGG)3). The more the area under the curve (AUC) deviates from 0.5, the better the repeat occurrence distinguishes ATRX/DAXXtrunc from TERTmod samples. b Pattern count tumor/control log2 ratios of all patients plotted against telomere content tumor/control log2 ratios for selected singletons. The regression line through the TERTmod samples is shown in green and is defined as the expected pattern count in the following. c Distance to the expected singleton repeat count in ATRX/DAXXtrunc and TERTmod samples. The center line of the boxplot is the median, the bounds of the box represent the first and third quartiles, the upper and lower whiskers extend from the hinge to the largest or smallest value, respectively, no further than 1.5 × IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). ****p < 0.0001; ***p < 0.001, Wilcoxon rank-sum tests after Bonferroni correction. The profiles of all analyzed patterns are shown in Supplementary Figs. 11 and 13.
As expected, normalized singleton repeat counts generally rose with increasing telomere content (Fig. 4b, an overview of all patterns is shown in Supplementary Fig. 13). However, TGAGGG, TCAGGG, TTGGGG, and TTCGGG singletons had significantly higher counts than expected in ATRX/DAXXtrunc compared to TERTmod samples (p = 7.3 × 10−11, 7.9 × 10−6, 5.6 × 10−4, and 5.8 × 10−13, respectively, Wilcoxon rank-sum test after Bonferroni correction; Fig. 4c). Especially, TGAGGG and TTCGGG seemed to be highly interspersed in a subset of ATRX/DAXXtrunc tumors. In contrast, TTTGGG singletons were observed less frequently in ATRX/DAXXtrunc tumors (p = 3.8 × 10−12, Wilcoxon rank-sum test after Bonferroni correction).
This seemingly ALT-specific TVR enrichment or depletion occurred in different tumor types, with the highest prevalence in leiomyosarcomas (60%), pancreatic endocrine tumors (42%), liposarcomas (37%), osteosarcomas (29%), and lower grade gliomas (28%; Supplementary Table 2). In the ATRX/DAXXtrunc samples, singleton TVR occurrences correlated with each other (Supplementary Fig. 14). The strongest correlations were between TGAGGG occurrence and TTCGGG and TTGGGG singletons (r = 0.57 and r = 0.58, respectively, Spearman correlation).
ALT prediction
ALT has several different hallmarks with which it can be reliably identified11. However, most of these are not detectable in short-read WGS data. Using ATRX/DAXXtrunc as indicators of ALT, we have shown several possible TMM classification features based on WGS. Most ATRX/DAXXtrunc samples are already separated well from TERTmod samples by nonsupervised clustering of normalized TGAGGG, TCAGGG, TTGGGG, TTCGGG, and TTTGGG singleton repeat counts (Supplementary Fig. 15). As expected, the clusters of ATRX/DAXXtrunc samples had a high telomere content and a high number of telomere insertions relative to the total number of breakpoints. These features were further used to build a random forest classifier distinguishing ATRX/DAXXtrunc from TERTmod samples (area under the curve: 0.95; sensitivity: 0.73; specificity: 0.99; all after 10-fold cross-validation). The variables with the highest importance for the classification were the divergence of observed TTTGGG and TTCGGG singleton TVRs from the expected count, the number of breakpoints and the number of telomere insertions (Supplementary Table 3). It may be pivotal for further understanding of this mechanism to determine the causal relationship between these features and the ALT phenotype.
The scores resulting from the classifier can be interpreted as an ALT probability. As expected, ATRX/DAXXtrunc had a high ALT probability (mean = 0.91), while TERTmod samples had a low ALT probability (mean = 0.13, Supplementary Fig. 16). A total of 17 samples without ATRX/DAXXtrunc mutations had an ALT probability of over 0.9, of which three had nontruncating ATRX/DAXX mutations and one sample had a frameshift insertion in ATRX and a TERT amplification (11 TERT copies, triploid). Across the entire dataset, most samples had a low ALT probability (Supplementary Fig. 17), suggesting that their TMM is telomerase based. This included some samples with ATRX/DAXX missense mutations, suggesting that the mutations in those samples may be more of a passenger event than functionally relevant. Tumor types with a high ALT probability were leiomyosarcoma, osteosarcoma, pancreatic endocrine tumors, and liposarcomas, in keeping with the known high prevalence of ALT in these entities37,38,39.
Discussion
In this study, we have shown that the presence of ALT-associated mutations in tumors correlates with increased telomere content, enrichment of isolated TVRs in t-type context (singletons), a higher number of genomic breakpoints, and intrachromosomal telomere insertions (Fig. 5). In contrast, tumors with mutations associated with a possible telomerase activation showed moderate decrease of telomere content and increased TERT expression. Hence, TERT reactivation may not suffice to fully counteract the telomere loss associated with high proliferation and/or occur when advanced telomere attrition increases the selective pressure to activate telomere maintenance. The observed telomere content increase in ATRX/DAXXtrunc versus the decrease in TERTmod samples is in agreement with the recent findings of Barthel et al.26. The higher telomere content in ATRX/DAXXtrunc tumors indicates that the negative feedback loop that constrains telomere elongation to a physiological level in healthy telomerase-expressing cells40,41 is bypassed by the ALT process, while it seems to remain intact in telomerase-positive tumors. In addition to telomere elongation, the increase of telomere content in ALT-positive tumors detected by sequencing-based methods may partly stem from aberrant intrachromosomal telomere insertions18 or extrachromosomal telomeric DNA42. Although almost all tumors must maintain their telomeres43, we only detected somatic mutations highly associated with ALT or telomerase activation in a subset of the samples. In tumors arising from tissues with high rates of self-renewal, telomerase is likely to already be epigenetically activated in the cell of origin44,45. Thus, telomere maintenance activating mutations occur more frequently in tumors derived from slowly replicating cells46. In line with this assumption, we observed high rates of TMM-associated mutations in brain, liver, bladder, and kidney tumors and TERT expression despite lack of TMM-associated mutations in lymphomas, tumors of the gastrointestinal tract, and female reproductive system. The exceptions were pilocytic astrocytoma, pancreatic, and prostate adenocarcinoma, which all originate from slowly replicating tissues, but had almost no TMM-associated alterations. In pancreatic and prostate cancer, TERT activity has been detected47,48, suggesting other means of telomerase activation. In pilocytic astrocytoma, neither telomerase expression/activity nor ALT was observed, but preALT characteristics have been reported49,50. Therefore, a TMM may only be fully activated upon progression of this slow-growing tumor type. Medulloblastoma samples only had a TERTmod frequency of 14% and one of the lowest average telomere contents in our study. While TERTpmut tend to occur in older patients of the SHH subgroup, the TERT promoter is frequently methylated in younger SHH patients and other medulloblastoma subgroups51. Interestingly, SHH medulloblastomas are thought to arise from granule neuron precursor cells52. This is a cell type with an extremely high rate of turnover during development and infancy, which may explain the TERT promoter methylation in younger SHH patients. In agreement with data suggesting that TERT expression is higher in TERTpmut than in TERT promoter methylated medulloblastomas51, we found that the telomere content was significantly higher in medulloblastomas with TERTmod than in those without (p = 0.0045, Wilcoxon rank-sum test).
It is known that telomeres elongated by telomerase have a homologous length with few TVRs in distal telomeric regions (left), while ALT telomeres have heterogeneous lengths with an increased amount of TVRs (right). Moreover, ALT cells have abundant extrachromosomal telomeric sequences. From this study, we conclude that the chromosomes of ALT cells have a higher number of aberrant interstitial telomere insertions, most of which are one-sided and accompanied by a loss of the adjacent chromosomal segment. We also showed that several TVRs occurring as singletons are more abundant in ALT telomeres, while one singleton (TTTGGG) was more abundant in telomerase-elongated telomeres. Please note that it is currently undetermined whether the different types of singletons are located in proximal or distal telomeric regions.
In our study, we systematically mapped telomere insertions into nontelomeric genomic regions using WGS data. They were most frequently accompanied by a loss of the adjacent chromosomal segment or located at copy-number neutral sites. Surprisingly, the latter telomere insertions were rarely two-sided and chromothripsis or other structural variations in the adjacent genomic regions occurred only in about half of the cases. As broken chromosome ends are highly unstable, the remaining segments must have undetected structural rearrangements, such as subclonal copy number changes or undetected DNA fusions. Taken together, the results suggest that we observe telomere healing or capture20,21 rather than telomere insertions followed by chromosomal instabilities18,53. As microhomology around telomere insertion sites was frequent, the sequences were probably inserted by nonhomologous end-joining54 or a microhomology-mediated mechanism55.
Telomere insertions were particularly frequent in ATRX/DAXXtrunc tumors, in which the abundant extrachromosomal telomeric DNA expands the telomere template pool for microhomology-mediated double-strand repair. We speculate that in this cellular environment, a high load of genomic breakpoints subsequently leads to the observed disproportionately increased number of telomere capture-like events. Due to the stochastic nature of ALT, the likelihood of telomere crisis is elevated, an event that can induce BFB cycles56,57 and chromothripsis58. Nevertheless, ALT can also stabilize telomeres, which has been shown to counteract genomic instability in certain instances59. Either scenario may account for the higher prevalence of chromothripsis and BFB events in ATRX/DAXXtrunc cases observed in this study. Together with the correlation of telomere insertions and mutations in TP53, RB1, MEN1, ATRX, and DAXX, these findings suggest that genome instability and the ALT phenotype are prerequisites for a high number of telomere insertions. Mutations in TP53, RB1, and MEN1 have been associated with impaired DNA damage response and repair60,61,62. This may make telomere crisis and genomic rearrangements more likely, while at the same time preventing apoptosis or senescence. Supporting all of these associations, an increased incidence of ALT in combination with chromothripsis was observed in SHH medulloblastomas with TP53 germline or somatic mutations59.
Telomere elongation by ALT or telomerase enriches distinct TVRs15. Here, we report a stronger association of singleton TVRs with ATRX/DAXXtrunc mutations than TVRs in an arbitrary context. The increase of TVRs has been attributed to the inclusion of subtelomeric regions during ALT via homologous recombination14. Whether telomeric sequences with lower TVR density are under positive selection or regions with higher TVR density are under negative selection remains to be clarified.
A possible function for TVRs has been reported in ALT-positive cell lines, where TCAGGG repeats that recruit nuclear receptors were enriched14,18. This enrichment was confirmed in a subset of primary ATRX/DAXXtrunc tumor samples in our study. However, we found a more pronounced enrichment of TTCGGG or TGAGGG. While TGAGGG has previously been associated with ALT14, the high prevalence of TTCGGG singletons in ALT is a novel discovery. No proteins with strong affinity to these two TVRs are currently known. This may indicate a more passive mode of action, for instance deprotection of telomeres by shelterin displacement14, and/or alteration of the telomeric G-quadruplex conformation15. Notably, we report for the first time that TTTGGG singletons but not TTTGGG in arbitrary context were depleted in ATRX/DAXXtrunc samples. This finding underlines the necessity to consider the sequence context of TVRs. None of the current models of ALT provide an explanation for this specific TVR depletion.
The methodologies presented here expand the established telomere content estimation from genomic sequencing by the context-dependent analysis of TVRs and telomere insertions. By applying them within a large-scale pan-cancer study, we provide a valuable resource for the further characterization of different TMMs in cancer cells based on WGS data.
Methods
Sequencing data
WGS and expression data were obtained from the ICGC/TCGA PCAWG project23. The WGS reads of tumor and control samples were aligned with bwa-mem by the PCAWG-tech group. Control samples were usually blood. In a small number of cases, the controls were obtained from tumor-adjacent or other tissue23. Tumors with multiple samples were excluded from this study, as well as one sample pair with reads shorter than 30 bp. Expression data was in the format of normalized RNA read counts per gene and only available for 1033 of 2519 patients. RNA sequencing BAM files aligned with STAR by the PCAWG-tech group were available for 867 tumor samples. To avoid confusion, we used the name “(central nervous system) CNS-LGG” (lower grade glioma, i.e., grades II and III) for the “CNS-Oligo” tumor type, because several samples in this cohort did not have the genetic markers (i.e., 1p/19q co-deletions) for oligodendroglioma required by the WHO63. A detailed overview of tumor type abbreviations with the included subtypes is given in Supplementary Table 4.
Mutation data
Somatic simple nucleotide, structural variations, and copy numbers were obtained from the PCAWG consensus calls (Synapse IDs syn7364923, syn7596712, and syn8042992, respectively). Structural variations were not available for 24 tumor samples.
Telomere read extraction and computational telomere content estimation
The telomere content of WGS samples was determined using the software tool TelomereHunter64. In short, telomeric reads containing six nonconsecutive instances of the four most common telomeric repeat types (TTAGGG, TCAGGG, TGAGGG, and TTGGGG) were extracted. For the further analysis, only unmapped reads or reads with a very low alignment confidence (mapping quality lower than 8) were considered. The telomere content was determined by normalizing the telomere read count by all reads in the sample with a GC content of 48–52%.
Determining TMM-associated mutations
Samples with a truncating ATRX or DAXX alteration (frame-shift insertion/deletion, stop-codon gain, or structural variation breakpoint within the gene) were defined as ATRX/DAXXtrunc, samples with other simple nucleotide variants were defined as ATRX/DAXXnon-trunc. Deletions that only affected intronic regions of ATRX were not considered. Of note, a frame-shift deletion called in the tumor sample of donor SP112201 was excluded as a false positive after visual inspection in the Integrative Genomics Viewer (IGV)65,66. Samples with a structural variation breakpoint on the plus strand 20 kb upstream of TERT the TSS were defined as TERTpSV. TERTamp samples had at least six additional copies of the TERT gene compared to the mean ploidy of the sample. Tumor samples with a C228T or C250T TERT promoter mutation were defined as TERTpmut. Due to the low sequencing coverage at the TERT promoter, these mutations were called using less stringent criteria (at least two reads with the mutated base, mutational frequency of at least 20%) in addition to the PCAWG consensus single nucleotide variant (SNV) calls (Synapse ID syn7364923). If multiple of these TERT modifications were present, the sample was defined as TERTmult. Samples with these TERT alterations were summarized as TERTmod. Samples without any of these alterations were defined as “wild type”. If a sample had both a TERTmod alteration and an ATRX/DAXX alteration, it was defined as “mixed”. For some analyses, ATRX/DAXXnon-trunc, mixed and wild-type samples were summarized as “other”.
Overlap of juxtaposed positions upstream of TERT and predicted super-enhancers
For the closest structural variation (SV) of each tumor sample to the TERT TSS, the juxtaposed genomic coordinates were compared to 65,950 predicted super-enhancers from the dbSUPER database30. Only SVs on the plus strand and within 1 mb of the TERT TSS were considered. Overlaps of juxtaposed positions with super-enhancer sites were defined as direct overlaps. Super-enhancer sites within 1 mb of the juxtaposed position were defined as indirect overlaps.
Telomere insertion detection
To find insertions of telomeric sequences into nontelomeric regions in the genome, we searched for tumor-specific discordant paired-end reads, where one end was an extracted telomere read and the other end was nontelomeric and uniquely mapped to a chromosome (mapping quality > 30). In 1 kb regions containing at least three discordant reads in the tumor sample and none in the matching control, exact positions of telomere insertions were defined by at least three split reads spanning the insertion site. The split reads had to contain at least one TTAGGG repeat. Regions with discordant read pairs in at least 15 control samples were excluded. Finally, the insertion sites were visualized using IGV65,66 to identify and remove remaining false positives. A telomere insertion was defined as two-sided if another telomere insertion in opposite orientation was found in the downstream 10 kb of the reference genome with the same repeat on the forward strand. Otherwise it was defined as one-sided.
Breakpoint detection
Breakpoints were obtained from the consensus breakpoint list of structural and copy number variation calls (Synapse ID syn8042992). In short, six copy number detection tools were run on all samples, including the consensus structural variations breakpoints. From the obtained chromosomal segments of the individual callers another set of consensus breakpoints was calculated.
Chromothripsis detection
To identify chromothripsis events, we extended the set of statistical criteria proposed by Korbel and Campbell67. The basic idea is to determine whether there is a statistically significant number of interleaved structural variants in a contiguous genomic region. We did this by constructing a graph whose nodes correspond to SVs and whose edges connect interleaved SVs. The identified clusters of SVs were also tested for the presence of alternating copy number and loss-of-heterozygosity patterns. The resulting chromothripsis calls were validated visually. The full description of the methodology and the detailed patterns of chromothripsis events in the genomes are described in a separate study24. Only high-confidence chromothripsis calls were included in this analysis.
BFB detection
At least two fold-back inversions on the same chromosome arm were defined as BFB events. Fold-back inversions had to fulfill the following requirements adapted from Cheng et al.68: (1) the two breakpoints of the inversion are less than 20 kb apart; (2) the inversion does not have a reciprocal partner, such that inversion1_start < inversion2_start < inversion1_end < inversion2_end; and (3) there is a copy number change at the inversion site.
Copy number changes at telomere insertion sites
Copy numbers of chromosomal segments were obtained from the PCAWG consensus calls (Synapse ID syn8042992). Copy numbers reveal gains or losses of chromosomal segments based on coverage and B-allele frequency, but were here limited to segments of at least 10 kb. The breakpoint estimations could differ from the actual site by up to 50 kb. Therefore, telomere insertions were assigned to the closest breakpoint within 50 kb. If there was no breakpoint within 50 kb or the copy numbers at either side of the telomere insertion were the same, the copy number change at the telomere insertion was defined as neutral.
Structural variations near telomere insertion sites
Structural variation annotation was obtained from the PCAWG consensus calls (Synapse ID syn7596712), which was based on discordant mate pairs and split reads, providing exact breakpoints. Because copy number variations smaller than 10 kb were not detected by copy number callers, small deletions next to the telomere insertion site may be missed. We therefore searched for structural variations within 10 kb of a telomere insertion to detect these cases.
Candidate gene selection for correlation analysis
A list of 1725 telomere maintenance associated human genes was obtained from TelNet33 on February 20, 2017. After removing genes without a unique Ensembl IDs in the GENCODE69 v19 HAVANA annotation, the remaining 1686 genes were used for correlation of telomere insertions and simple nucleotide variants.
TERRA detection
TelomereHunter was run on RNA-Seq BAM files to count reads containing at least k nonconsecutive instances of the four most common telomeric repeat types (TTAGGG, TCAGGG, TGAGGG, and TTGGGG). The repeat threshold k was chosen depending on the read length: k = 7 for 45–50 bp, k = 10 for 75–76 bp, and k = 14 for 99–101 bp. The resulting TERRA read counts were normalized by the total number of reads in the sample. For better readability, this number was multiplied by 1 Mio.
Detection of TVRs
TVRs were detected by searching for hexamers of the type NNNGGG in the extracted telomere reads. Each base was required to have a base quality of at least 20. The neighboring 18 bp on either side of the TVR were determined. For further analysis, NNNGGG TVRs were once computed for arbitrary context and once for t-type context ((TTAGGG)3-NNNGGG-(TTAGGG)3, also called “singletons”). The absolute counts were normalized to the total number of reads in the sample. The expected pattern counts in arbitrary context were calculated as: telomere content tumor/control log2 ratio. The expected singleton counts at different telomere content tumor/control log2 ratios were taken from the regression line through TERTmod samples. The singleton occurrence heatmap was generated using the R package ComplexHeatmap70.
Classifier for predicting active TMMs
A random forest classifier to distinguish ATRX/DAXXtrunc and TERTmod samples was built using the R packages “randomForest”71 and “caret”72 with the following eight features: telomere content tumor/control log2 ratio, number of telomere insertions, number of breakpoints, and the distance of TGAGGG, TCAGGG, TTGGGG, TTCGGG, and TTTGGG singletons (i.e., repeats in a t-type context) to their expected occurrence. To deal with the imbalance in the data set (i.e., 266 TERTmod samples versus 63 ATRX/DAXXtrunc samples without missing data), the model was trained with a down-sampled training set. The performance was determined using 10-fold cross-validation.
Statistics
Differences between ATRX/DAXXtrunc and TERTmod samples in terms of telomere content, percent breakpoints with telomere insertions, and singleton repeat abundance were tested using two-sided Wilcoxon rank-sum tests. Singleton repeat abundance p-values were corrected for multiple testing using the Bonferroni method. To reduce the influence of outliers, correlation coefficients were calculated with the Spearman method. Correlation between control telomere content and age as well as tumor and control telomere content was tested with linear regression. All statistical analyses were carried out using R (R Foundation for Statistical Computing).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Somatic and germline variant calls, mutational signatures, subclonal reconstructions, transcript abundance, splice calls, and other core data generated by the ICGC/TCGA PCAWG Consortium is described here23 and available for download at [https://dcc.icgc.org/releases/PCAWG]. Additional information on accessing the data, including raw read files, can be found at [https://docs.icgc.org/pcawg/data/]. In accordance with the data access policies of the ICGC and TCGA projects, most molecular, clinical and specimen data are in an open tier which does not require access approval. To access potentially identification information, such as germline alleles and underlying sequencing data, researchers will need to apply to the TCGA Data Access Committee (DAC) via dbGaP for access to the TCGA portion of the dataset, and to the ICGC Data Access Compliance Office (DACO) for the ICGC portion. In addition, to access somatic SNVs derived from TCGA donors, researchers will also need to obtain dbGaP authorization.
Derived data sets described specifically used in this manuscript are catalogued on Synapse. Data access is possible via the ICGC data portal (DCC), were the original files are split into ICGC and TCGA subsets due to different access regulations:
somatic simple nucleotide calls (syn7364923): and [https://dcc.icgc.org/releases/PCAWG/consensus_snv_indel],
structural variation calls (syn7596712): and [https://dcc.icgc.org/releases/PCAWG/consensus_sv],
copy number calls (syn8042992): and [https://dcc.icgc.org/releases/PCAWG/consensus_cnv].
Code availability
The software TelomereHunter used for the in silico analysis of the genomic footprints of activated TMMs is available from [https://www.dkfz.de/en/applied-bioinformatics/telomerehunter/telomerehunter.html]. The core computational pipelines used by the PCAWG Consortium for alignment, quality control, and variant calling are available to the public at https://dockstore.org/search?search=pcawg under the GNU General Public License v3.0, which allows for reuse and distribution.
Change history
08 December 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41467-022-32328-7
References
O’Sullivan, R. J. & Karlseder, J. Telomeres: protecting chromosomes against genome instability. Nat. Rev. Mol. Cell Biol. 11, 171–181 (2010).
Allshire, R. C., Dempster, M. & Hastie, N. D. Human telomeres contain at least three types of G-rich repeat distributed non-randomly. Nucleic Acids Res. 17, 4611–4627 (1989).
Baird, D. M., Jeffreys, A. J. & Royle, N. J. Mechanisms underlying telomere repeat turnover, revealed by hypervariable variant repeat distribution patterns in the human Xp/Yp telomere. EMBO J. 14, 5433–5443 (1995).
Harley, C. B., Futcher, A. B. & Greider, C. W. Telomeres shorten during ageing of human fibroblasts. Nature 345, 458–460 (1990).
d’Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198 (2003).
Wright, W. E., Piatyszek, M. A., Rainey, W. E., Byrd, W. & Shay, J. W. Telomerase activity in human germline and embryonic tissues and cells. Dev. Genet. 18, 173–179 (1996).
Zhang, A. et al. Frequent amplification of the telomerase reverse transcriptase gene in human tumors. Cancer Res. 60, 6230–6235 (2000).
Peifer, M. et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature 526, 700–704 (2015).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Huang, F. W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Cesare, A. J. & Reddel, R. R. Alternative lengthening of telomeres: models, mechanisms and implications. Nat. Rev. Genet. 11, 319–330 (2010).
Heaphy, C. M. et al. Altered telomeres in tumors with ATRX and DAXX mutations. Science 333, 425 (2011).
Varley, H., Pickett, H. A., Foxon, J. L., Reddel, R. R. & Royle, N. J. Molecular characterization of inter-telomere and intra-telomere mutations in human ALT cells. Nat. Genet. 30, 301–305 (2002).
Conomos, D. et al. Variant repeats are interspersed throughout the telomeres and recruit nuclear receptors in ALT cells. J. Cell Biol. 199, 893–906 (2012).
Lee, M. et al. Telomere extension by telomerase and ALT generates variant repeats by mechanistically distinct processes. Nucleic Acids Res. 42, 1733–1746 (2014).
Lovejoy, C. A. et al. Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet. 8, e1002772 (2012).
Lin, K. W. & Yan, J. Endings in the middle: current knowledge of interstitial telomeric sequences. Mutat. Res. 658, 95–110 (2008).
Marzec, P. et al. Nuclear-receptor-mediated telomere insertion leads to genome instability in ALT cancers. Cell 160, 913–927 (2015).
Wilkie, A. O., Lamb, J., Harris, P. C., Finney, R. D. & Higgs, D. R. A truncated human chromosome 16 associated with alpha thalassaemia is stabilized by addition of telomeric repeat (TTAGGG)n. Nature 346, 868–871 (1990).
Flint, J. et al. Healing of broken human chromosomes by the addition of telomeric repeats. Am. J. Hum. Genet. 55, 505–512 (1994).
Meltzer, P. S., Guan, X. Y. & Trent, J. M. Telomere capture stabilizes chromosome breakage. Nat. Genet. 4, 252–255 (1993).
Slijepcevic, P. & Bryant, P. E. Chromosome healing, telomere capture and mechanisms of radiation-induced chromosome breakage. Int. J. Radiat. Biol. 73, 1–13 (1998).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature https://doi.org/10.1038/s41586-020-1969-6 (2020).
Cortes-Ciriano, I. et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. https://doi.org/10.1038/s41588-019-0576-7 (2020).
Lee, M. et al. Comparative analysis of whole genome sequencing-based telomere length measurement techniques. Methods 114, 4–15 (2017).
Barthel, F. P. et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat. Genet. 49, 349–357 (2017).
Slagboom, P. E., Droog, S. & Boomsma, D. I. Genetic determination of telomere size in humans: a twin study of three age groups. Am. J. Hum. Genet. 55, 876–882 (1994).
Codd, V. et al. Identification of seven loci affecting mean telomere length and their association with disease. Nat. Genet. 45, 422–427, 427e1–2(2013).
Shammas, M. A. Telomeres, lifestyle, cancer, and aging. Curr. Opin. Clin. Nutr. Metab. Care 14, 28–34 (2011).
Khan, A. & Zhang, X. dbSUPER: a database of super-enhancers in mouse and human genome. Nucleic Acids Res. 44, D164–D171 (2016).
Heaphy, C. M. et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am. J. Pathol. 179, 1608–1615 (2011).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Braun, D. M., Chung, I., Kepper, N., Deeg, K. I. & Rippe, K. TelNet - a database for human and yeast genes involved in telomere maintenance. BMC Genet. 19, 32 (2018).
Hu, Y. et al. Telomerase-null survivor screening identifies novel telomere recombination regulators. PLoS Genet. 9, e1003208 (2013).
Askree, S. H. et al. A genome-wide screen for Saccharomyces cerevisiae deletion mutants that affect telomere length. Proc. Natl Acad. Sci. USA 101, 8658–8663 (2004).
Patil, M., Pabla, N. & Dong, Z. Checkpoint kinase 1 in DNA damage response and cell cycle regulation. Cell Mol. Life Sci. 70, 4009–4021 (2013).
Amorim, J. P., Santos, G., Vinagre, J. & Soares, P. The role of ATRX in the alternative lengthening of telomeres (ALT) phenotype. Genes (Basel) 7, 66 (2016).
de Wilde, R. F. et al. Loss of ATRX or DAXX expression and concomitant acquisition of the alternative lengthening of telomeres phenotype are late events in a small subset of MEN-1 syndrome pancreatic neuroendocrine tumors. Mod. Pathol. 25, 1033–1039 (2012).
Chudasama, P. et al. Integrative genomic and transcriptomic analysis of leiomyosarcoma. Nat. Commun. 9, 144 (2018).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Hockemeyer, D. & Collins, K. Control of telomerase action at human telomeres. Nat. Struct. Mol. Biol. 22, 848–852 (2015).
Nabetani, A. & Ishikawa, F. Unusual telomeric DNAs in human telomerase-negative immortalized cells. Mol. Cell Biol. 29, 703–713 (2009).
Reddel, R. R. Telomere maintenance mechanisms in cancer: clinical implications. Curr. Pharm. Des. 20, 6361–6374 (2014).
Guilleret, I. et al. Hypermethylation of the human telomerase catalytic subunit (hTERT) gene correlates with telomerase activity. Int. J. Cancer 101, 335–341 (2002).
Takasawa, K. et al. DNA hypermethylation enhanced telomerase reverse transcriptase expression in human-induced pluripotent stem cells. Hum. Cell 31, 78–86 (2018).
Killela, P. J. et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc. Natl Acad. Sci. USA 110, 6021–6026 (2013).
Hiyama, E. et al. Telomerase activity is detected in pancreatic cancer but not in benign tumors. Cancer Res. 57, 326–331 (1997).
Sommerfeld, H. J. et al. Telomerase activity: a prevalent marker of malignant human prostate tissue. Cancer Res. 56, 218–222 (1996).
Chong, E. Y., Lam, P. Y., Poon, W. S. & Ng, H. K. Telomerase expression in gliomas including the nonastrocytic tumors. Hum. Pathol. 29, 599–603 (1998).
Slatter, T. et al. Pilocytic astrocytomas have telomere-associated promyelocytic leukemia bodies without alternatively lengthened telomeres. Am. J. Pathol. 177, 2694–2700 (2010).
Lindsey, J. C. et al. TERT promoter mutation and aberrant hypermethylation are associated with elevated expression in medulloblastoma and characterise the majority of non-infant SHH subgroup tumours. Acta Neuropathol. 127, 307–309 (2014).
Schuller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008).
Jia, P., Chastain, M., Zou, Y., Her, C. & Chai, W. Human MLH1 suppresses the insertion of telomeric sequences at intra-chromosomal sites in telomerase-expressing cells. Nucleic Acids Res. 45, 1219–1232 (2016).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Ottaviani, D., LeCain, M. & Sheer, D. The role of microhomology in genomic structural variation. Trends Genet. 30, 85–94 (2014).
McClintock, B. The stability of broken ends of chromosomes in Zea mays. Genetics 26, 234–282 (1941).
Lo, A. W. et al. DNA amplification by breakage/fusion/bridge cycles initiated by spontaneous telomere loss in a human cancer cell line. Neoplasia 4, 531–538 (2002).
Maciejowski, J., Li, Y., Bosco, N., Campbell, P. J. & de Lange, T. Chromothripsis and kataegis induced by telomere crisis. Cell 163, 1641–1654 (2015).
Ernst, A. et al. Telomere dysfunction and chromothripsis. Int. J. Cancer 138, 2905–2914 (2016).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Huang, P. H., Cook, R. & Mittnacht, S. RB in DNA repair. Oncotarget 6, 20746–20747 (2015).
Gallo, A., Agnese, S., Esposito, I., Galgani, M. & Avvedimento, V. E. Menin stimulates homology-directed DNA repair. FEBS Lett. 584, 4531–4536 (2010).
Louis, D. N. et al. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 131, 803–820 (2016).
Feuerbach, L. et al. TelomereHunter - in silico estimation of telomere content and composition from cancer genomes. BMC Bioinforma. 20, 272 (2019).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013).
Cheng, C. et al. Whole-genome sequencing reveals diverse models of structural variations in esophageal squamous cell carcinoma. Am. J. Hum. Genet. 98, 256–274 (2016).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Liaw, A. W. M. Classification and regression by randomForest. R. News 2, 18–22 (2002).
Kuhn, M. Building Predictive Models in R Using the caret Package. J. Stat. Softw. 28 (2008).
Acknowledgements
We thank J. Kerssemakers, M. Prinz, and M. Heinold for their help in data processing. We thank I. Buchhalter for annotation of simple nucleotide variations. We thank D. Hübschmann for his support in correlation analysis. We thank all TelNet curators for sharing the results of their extensive literature research. We thank K.I. Deeg, P. Lichter, and S.M. Pfister for their support in early stages of the study. We thank I. Chung for comments and discussion. The work was supported by grants from the German Federal Ministry of Education and Research (BMBF) within the e:Med program (project CancerTelSys, 01ZX1302 to K.R.) and the program for medical genome research (01KU1001A, -B, -C, and -D; 01KU1505A). S.D.K. received funding from the German Research Foundation (DFG) in research priority program SPP1463 (grant no. Br3535/1-2). I.C.C. has received funding from the European Union’s Framework Programme For Research and Innovation Horizon 2020 (2014–2020) under the Marie Sklodowska-Curie Grant Agreement No. 703543. We acknowledge the contributions of the many clinical networks across ICGC and TCGA who provided samples and data to the PCAWG Consortium, and the contributions of the Technical Working Group and the Germline Working Group of the PCAWG Consortium for collation, realignment, and harmonised variant calling of the cancer genomes used in this study. We thank the patients and their families for their participation in the individual ICGC and TCGA projects.
Author information
Authors and Affiliations
Consortia
Contributions
L.S. was involved in all bioinformatical analyses. C.H. performed structural variation annotation, principal component analysis and classification. L.F. was responsible for correlation analysis. S.D.K. analyzed gene expression data and was involved in visualization. L.S., P.G. and L.F. were involved in method development. K.K. and M.S. were involved in copy number analysis. R.K. was involved in data preprocessing. D.M.B. developed and curated the TelNet database. I.C.C., R.X. and P.J.P. provided regions of chromothripsis. B.H. was involved in early stages of experimental design. R.E. was responsible for data management and creating the IT-infrastructure. K.R. and D.T.W.J. provided insights into telomere biology. D.T.W.J. and L.F. conceived the study. B.B. and L.F. oversaw the experimental design and execution. L.S. and L.F. wrote the manuscript with contributions by K.K., D.M.B., M.S., K.R. and D.T.W.J. The PCAWG-Structural Variation Working Group provided valuable advice and feedback. Rameen Beroukhim and Peter J. Campbell were working group or project co-leaders, respectively.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sieverling, L., Hong, C., Koser, S.D. et al. Genomic footprints of activated telomere maintenance mechanisms in cancer. Nat Commun 11, 733 (2020). https://doi.org/10.1038/s41467-019-13824-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-13824-9
This article is cited by
-
The ALT pathway generates telomere fusions that can be detected in the blood of cancer patients
Nature Communications (2024)
-
The regulations of telomerase reverse transcriptase (TERT) in cancer
Cell Death & Disease (2024)
-
Most large structural variants in cancer genomes can be detected without long reads
Nature Genetics (2023)
-
A pan-sarcoma landscape of telomeric content shows that alterations in RAD51B and GID4 are associated with higher telomeric content
npj Genomic Medicine (2023)
-
Histone demethylase KDM2A is a selective vulnerability of cancers relying on alternative telomere maintenance
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.