Abstract
Female somatic X-chromosome inactivation (XCI) balances the X-linked transcriptional dosages between the sexes. Skewed XCI toward one parental X has been observed in several complex human traits, but the extent to which genetics and environment influence skewed XCI is largely unexplored. To address this, we quantify XCI-skew in multiple tissues and immune cell types in a twin cohort. Within an individual, XCI-skew differs between blood, fat and skin tissue, but is shared across immune cell types. XCI skew increases with age in blood, but not other tissues, and is associated with smoking. XCI-skew is increased in twins with Rheumatoid Arthritis compared to unaffected identical co-twins. XCI-skew is heritable in blood of females >55 years old (h2 = 0.34), but not in younger individuals or other tissues. This results in a Gene x Age interaction that shifts the functional dosage of all X-linked heterozygous loci in a tissue-restricted manner.
Similar content being viewed by others
Introduction
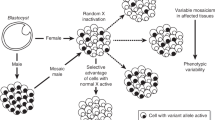
To balance the X-linked transcriptional dosages between the single X chromosome of males and the two X chromosomes of females, one X chromosome is silenced in female placental mammals1. The X-chromosome inactivation (XCI) process starts during preimplantation phases of human embryonic development, presumably at around the eight-cell stage2. XCI is initiated by the transcription of XIST, a 17 kb, alternatively spliced long noncoding RNA mapped to Xq13.2 and exclusively expressed on the inactive X (Xi)3. Once transcribed, XIST molecules spread in cis along the X chromosome4,5 inducing a progressive epigenetic silencing through the recruitment of chromatin remodeling enzymatic complexes, which impose repressive histone and DNA changes on the Xi chromosome6,7. Within each cell, the parental X chromosome selected for inactivation seems to occur at random, and the Xi is mitotically inherited to future somatic daughter cells. This random inactivation results in a mosaic of cells within an individual, where overall, a balanced expression (50:50) of both parental X-linked alleles is expected. Asymmetric selection of the X chromosome to inactivate causes the predominance of one parental Xi in a population of cells, unbalancing the X-linked transcriptional and allelic dosages toward one parental X chromosome. This phenomenon, known as skewed XCI (or nonrandom XCI), occurs when at least 80% of cells within a tissue inactivate the same parental X chromosome. The factors underlying primary skewed XCI are varied and several mechanisms are possible8. Secondary (or acquired) skewed XCI can result from positive selection of cells that after having inactivated a particular parental X, acquire a survival advantage over cells who inactivated the other parental X chromosome. Skewed XCI patterns can also be generated by the stochastic overrepresentation of cell clones in a given tissue, due for instance, to depletion of stem cell populations.
Comprised of 155 MB and containing >800 protein-coding genes, the X chromosome represents approximately the 5% of the haploid human genome. In heterozygous females with skewed XCI, the X-linked transcriptional and allelic dosages of silenced genes are unbalanced and may be functionally homozygous. Skewed XCI is a major cause of discontinuity of dominance and recessiveness, as well as penetrance and expressivity of X-linked traits. How skewed XCI patterns modulate phenotypes in females, and whether they are a cause or a consequence of associated phenotypes is not fully understood. Skewed XCI patterns have been observed in females with X-linked diseases9,10,11, autoimmune disorders12,13, as well as in breast14 and ovarian cancer15. In autoimmune diseases with higher prevalence in females, including rheumatoid arthritis (RA) and systemic lupus erythematosus, XCI is hypothesized to play a role. Chromosome X is enriched for immune-related genes and skewed XCI patterns could cause the breakdown of thymic tolerance induction processes16 conferring a high predisposition to develop autoimmunity (reviewed in17). XCI skewing levels in blood tissues have been associated with ageing, with multiple studies indicating an increase after 50–60 years of age18,19,20,21,22,23. To date, the mechanisms underlying skewed XCI in humans are not fully understood. Several twin studies have reported that genetic factors contribute to XCI skewing in blood-derived cells22,24, while other evidence indicated that most of the XCI skewing levels in human are acquired secondarily25.
Nearly all studies of XCI skewing levels in humans have been carried out in peripheral blood samples or in very small sample sizes26, while XCI patterns in other tissues have not been studied in great detail20,27,28. In this study, we comprehensively assess XCI patterns in a multi-tissue sample of nearly 800 female twins from the TwinsUK cohort29. We quantify the degree of skewing of XCI using a metric based on XIST allele-specific expression (ASE) from paired RNA-seq and DNA-seq data in four tissues (LCLs, whole-blood, fat and skin) and in multiple immune cell types (monocytes, B-cells, T-CD4+, T-CD8+, NK) purified from two identical co-twins. We examine the tissue-specific prevalence of skewed XCI patterns, compare the XCI skewing levels between tissues and across immune cell types within the same individual, and evaluate the association between XCI skewing and age, a complex autoimmune disease and lifestyle (smoking) traits. We show that XCI patterns are highly tissue-specific and shared across immune cell types within an individual, and that XCI skew in haematopoetic tissues increases with age. We investigate the factors underlying the skewed XCI using classical twin models to characterize the extent of the influence of genetic and environmental factors on the tissue-specific skewed XCI. We show that heritability of XCI skew is restricted to blood tissues of females >55 years old (h2 = 0.34), indicating that XCI patterns have both a heritable and environmental (age) basis.
Results
Quantification of degree of XCI skewing in TwinsUK
We assessed XCI patterns in multi-tissue samples from female twin volunteers from the TwinsUK cohort aged 38–85 years old (median age = 60; Supplementary Fig. 1)29,30. We quantified the degree of skewing of XCI using a metric based on XIST ASE from paired RNA-seq and DNA-seq data. XIST is uniquely expressed from the Xi3,28,31, so the relative expression of parental alleles within the XIST transcript are representative of XCI skewing levels in a bulk sample. Skewed XCI patterns can be detected and quantified from the expression levels of XIST-linked heterozygous variants32. Furthermore, transcriptional assays based on single monoallelically expressed X-linked genes, like XIST, have been used as a complement to the HUMARA assay to quantify XCI patterns25,33. We have also calculated the XCI patterns using an alternative method, based on the ASE of all non-pseudo-autosomal region (PAR) heterozygous loci available in a sample34 as done in supplements. We ran a series of benchmarking analyses to compare the non-PARASE calls to the XISTASE-based XCI skew calls (Supplementary Note 1, Supplementary Figs. 2–4) and demonstrated that the XISTASE is the appropriate method to use in our analyses. RNA-seq and genotype data were available for 814 lymphoblastoid cell lines (LCL) samples, 395 whole-blood samples, 766 subcutaneous adipose tissue samples (herein referred as fat) and 716 skin samples. After stringent quality control, we obtained XISTASE calls for 422 LCL samples, 72 whole-blood samples, 378 fat samples and 336 skin samples. The smaller sample size for whole-blood was due to the relatively smaller size of the starting dataset and the relatively lower RNA-seq coverage of this tissue in our dataset. In order to have an absolute measure of the magnitude of the XCI skewing levels in each sample, we calculated the degree of skewing of XCI (DS) from the XISTASE calls. DS is defined as the absolute deviation of the XISTASE from 0.5 (see “Methods”) and it has been similarly used in other investigations to assess XCI patterns21,35,36 and the XCI status of X-linked genes37. In line with previous investigations22,38 we classified samples with DS < 0.3 (corresponding to 0.2 < XISTASE < 0.8) to have random XCI, and samples with DS ≥ 0.3 (corresponding to XISTASE ≤ 0.2 or XISTASE ≥ 0.8) to have skewed XCI. Unless otherwise specified, DS was used in all the analyses performed.
We assessed the robustness of our estimates of the degree of skewing with an alternative DNA-based measure of XCI, the human androgen receptor assay (HUMARA)39. HUMARA was and is still the “gold standard” technique to assess XCI patterns. A previous study has reported good replicability between HUMARA and expression-based quantification of XCI skewing40. We used HUMARA to measure XCI skewing levels in 18 archived whole-blood DNA samples obtained at the same clinical visit as the LCLs samples. Spearman’s correlation between the quantifications was 0.8 (P = 7 × 10−5) revealing a high degree of reproducibility between the both XISTASE and HUMARA methods and the LCLs and whole blood (Fig. 1a).
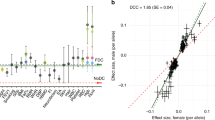
XISTASE is a reproducible and accurate proxy of XCI skewing levels. a Scatter plot showing the comparison between the XCI skewing levels in n = 18 whole-blood and LCLs samples quantified with HUMARA and XISTASE respectively. b Scatter plot of the XISTASE at time point 1 and at time point 2 in n = 16 whole-blood samples. Diagonal line indicates perfect concordance in the data (ρ = 1). HUMARA human androgen receptor assay.
Previous investigations have reported that the XCI skewing levels increase with age in blood tissues, as discussed below. While it would be expected that increases in XCI skewing levels would be observed over relatively large time spans, we would expect minimal variations of XCI skewing levels between two close time points. We therefore reasoned that the sensitivity of our quantifications could also be assessed by comparing the XCI skewing levels in the same individuals at close time points. Briefly, using a publicly-available longitudinal whole-blood RNA-seq dataset from the TwinsUK cohort41, we generated XISTASE calls at two time points (1–2.7 years later) in 16 samples (see “Methods”). The Spearman’s correlation between the XISTASE calls at the first time point and the XISTASE calls at the second time point was 0.94 at P < 2 × 10−15 (Fig. 1b). This indicates that XISTASE is a sensitive proxy when assessing the stability of XCI patterns over short time periods. Overall, these results indicate that XISTASE is a reproducible, accurate and sensitive proxy of XCI skewing levels.
Skewed XCI is tissue-specific with higher prevalence in blood-derived tissues
We observed a wide range of DS values in the four tissues (Fig. 2), with clear differences in the prevalence of skewed individuals between tissues. Blood-derived tissues had the highest incidence of skewed individuals, with skewed XCI observed in 34% of LCLs samples and 28% of whole-blood samples and a lower incidence in the primary tissues, where 12% of fat and 16% of skin samples exhibited skewed XCI (Table 1). In order to examine the extent of similarities of XCI patterns between tissues, we compared the tissue-specific XCI skewing levels in a pairwise manner (Fig. 3). For each tissue–tissue comparison, we included individuals with XISTASE calls in both tissues (Table 2). We found the strongest correlation on XISTASE calls between LCLs and whole blood (n = 59, Spearman’s ρ = 0.78, P = 2 × 10−13), indicating that blood-derived tissues share highly similar XCI skewing levels. We also found a good degree of similarity between the XCI skewing levels in fat and skin tissues (n = 252, Spearman’s ρ = 0.47, P = 2 × 10−15; Fig. 3). However, low concordance was observed between skin and whole blood (n = 47, Spearman’s ρ = 0.3, P = 0.04) and fat and whole-blood (n = 57, Spearman’s ρ = 0.33, P = 0.01). Our data demonstrate that tissue-specific XCI skewing within an individual is common in the population, indicating that XCI patterns are partially controlled by tissue-specific regulatory mechanisms.
Skewed X-inactivation varies across tissues with a higher prevalence in blood than fat and skin tissues. Distribution of the degree of skewing (top row) and XISTASE (bottom row) in LCLs (n = 422 samples), whole-blood (n = 72 samples), fat (n = 378 samples) and skin (n = 336 samples) tissues. XISTASE XIST allele-specific expression.
Tissue–tissue comparison of XISTASE reveals the highest similarity of XCI skew between blood tissues. Each plot shows the comparison of XISTASE between two tissues from an individual. Each dot represents an individual. In each comparison all individuals with available XISTASE calls for both tissues were included. Diagonal line indicates perfect concordance in the data (ρ = 1).
The active or inactive state of each X chromosome in a cell is clonally passed on to daughter cells. In a pool of cells derived from a single clone (or patch), the XCI patterns are expected to be completely skewed. Patch size refers to the amount of cell clones in a pool of cells (e.g. in a tissue biopsy). We considered the possibility that patch size might bias our quantification of XCI patterns in fat and skin samples. This is likely to occur in biopsies that are smaller than the tissue patch size. However, several considerations led us to exclude the possibility that patch sizes in fat and skin biopsies might confound our XISTASE calls. First, the biopsies included skin samples of 8 mm3 in size, which were cut into two skin and three fat samples. As reported in another study, this size is large enough to measure the XCI ratio without being confounded by patch size42. Second, most individuals exhibit random XCI patterns in fat and skin tissues, which is unlikely if patch size was larger than the biopsies. We therefore conclude that the biopsies used in this study are large enough to accurately assess the XCI patterns without being biased by patch size.
LCLs in this study are representative of XCI skewing in-vivo in blood tissues
LCLs generated by Epstein–Barr virus (EBV) mediated transformation of B-lymphocyte cells have been and are widely used in gene expression studies. However, the possibility that the cell lines are monoclonal and/or polyclonal due to selection in the transformation process or clonal expansion in cell culture, and hence not be representative of the in vivo XCI skewing levels, is a potential problem when using LCLs to assess XCI skewing43. As the profiled RNA in this study was extracted from the LCLs very shortly after transformation with limited passaging or time in culture we expected this effect to be minimal, however, to address the possibility we performed the following analyses. First, as described above and shown in Fig. 1a, the degree of skewing in LCLs were highly correlated with the HUMARA-based quantifications of XCI patterns in paired whole-blood samples (Spearman’s ρ = 0.8, P = 7 × 10−5, n = 18). We would not expect such high similarity between the two quantifications if clonal propagation had occurred in LCLs samples after preparation. This was confirmed by the high correlation between LCLs and whole-blood XISTASE values (Spearman’s ρ = 0.78, n = 59; Fig. 3) and overall similarity in the prevalence of skewed XCI in LCLs and whole blood (Table 1). Finally, we assessed the degree of skewing in monocytes, B, T-CD4+, T-CD8+ and natural-killer (NK) cells purified from two monozygotic (MZ) twins exhibiting skewed XCI patterns in LCLs and from one individual exhibiting random XCI patterns in LCLs (Supplementary Fig. 5). We found that in both MZ twins showing skewed XCI in LCLs, the majority of immune cell types exhibited skewed XCI patterns. Conversely, none of the immune cell types purified from the nonskewed individual exhibited skewed XCI patterns (Table 3). These data indicate that within an individual, XCI skewing levels are shared across hematopoietic cells. We conclude that the XCI skewing levels of LCLs in this study are representative of XCI skewing in vivo in blood tissues.
XCI skewing levels are positively associated with age in blood-derived tissues
XCI skewing levels in peripheral blood have been shown to increase with age in multiple studies18,19,20,21,22,24,40,44,45. The age-related increase of XCI skewing levels continues throughout life, since centenarians exhibit higher XCI skewing levels than 95 years old females22. However, there is very limited knowledge on the relationship between XCI patterns and ageing in tissues other than blood. In order to explore this, we investigated the association between age and degree of skewing in LCLs, fat, and skin. Our whole-blood estimates were excluded from analysis due to the low sample size (n = 72). Age was positively associated with XCI skew in LCLs (n = 422, P < 0.01), but we did not detect any association between XCI skew and age in skin (n = 336, P = 0.4) or in fat (n = 378, P = 0.7).
We next explored the dynamics of DS and age progression in each tissue, using the lowess procedure. Lowess curve detected an increase of DS beginning at around 55 years old in LCLs (Supplementary Fig. 6), in agreement with what was found in other studies20,22. Since the increase of DS starts at around 55 years, we divided LCLs samples into a younger group (n = 141, age < 55) and an older group (n = 281, age ≥ 55). We found that the mean DS in LCLs was significantly higher in older than in younger females (DSyounger = 0.2, DSolder = 0.24, T test, P = 0.03; Fig. 4). Accordingly, we found that the frequency of skewed XCI in LCLs was significantly higher in older (38%) than in younger (28%) females (χ2 test, P = 0.04; Fig. 4). In agreement with the lack of association between the DS and age, we did not detect significant differences between the mean DS in young and older females in fat (DSyounger = 0.15, DSolder = 0.15) or in skin tissues (DSyounger = 0.16, DSolder = 0.17). To acquire a more detailed view of the tissue-specific prevalence of skewed XCI in different groups of age, we categorized the samples into four age groups (40–50, 50–60, 60–70, and >70) and calculated the frequency of skewed XCI in each category (Fig. 4). We found that the frequency of skewed XCI increased with age in LCLs, with 41% of individuals >65 years old demonstrating skewed XCI patterns. We did not observe any increase in the skewed XCI frequencies with age in fat and skin tissues. Overall, these data further confirm that XCI skewing levels increase with age in blood-derived tissues, supporting previous investigations. However, we find that there is no increase in XCI in fat and skin tissue from the same individuals, suggesting that acquired XCI skewing with age is a distinctive feature of blood-derived tissues.
Skewed XCI increases with age in blood-derived tissues but not in fat or skin tissues. Left panel, boxplots show the degree of skewing in LCLs samples <55 (n = 141) and ≥ 55 (n = 281). Median (black line in the middle), first and third quartile (boxplot bounds), and whiskers (dashed lines) are shown. Right panel, line plots show the frequencies of skewed XCI at different ages in LCLs (blue), fat (red) and skin (green) tissues. XCI X-chromosome inactivation.
Heritability of skewed XCI is dependent on tissue and age
Twin studies are a powerful strategy to investigate the heritability of complex traits. Previous twin studies have reported that skewed XCI in blood-derived samples is heritable, with h2 estimates of 0.68 in granulocytes of elderly twin pairs and 0.58 in peripheral blood cells22,23,24, however, these studies have not investigated heritability outside of blood. To estimate the influence of additive genetic effects (heritability) and environmental factors on the observed variance in XCI in the three tissues, we implemented the ACE twin model. The ACE statistical model quantifies the contribution of additive genetic effects (A), shared environment (C) and unique environment (E) to the phenotype variance. In order to investigate whether heritability varies with age, we stratified the twin pairs into a younger group (age < 55) and an older group (age ≥ 55; Supplementary Table 1). Age 55 was chosen as it was identified as the inflection point at which XCI skew begins to increase in the lowess analysis (Supplementary Fig. 6), in agreement with other studies20,22. We found that XCI skewing is heritable in LCLs of older females (ACE model, h2 = 0.34, P = 9.6 × 10−6), but not younger females (h2 = 0, P = 1). There was no evidence of heritability of XCI skew in fat or in skin tissues at any age (Table 4). The highest proportion of variance was explained by unique environmental factors in all tissues of both younger and older females (E2LCLs_younger = 0.99, E2LCLs_older = 0.66, E2Fat_younger = 0.73, E2Fat_older = 0.92, E2Skin_younger = 1, E2Skin_older = 1). As a complement to the heritability analysis, we calculated the intraclass Spearman correlation (IC) of XCI skew within MZ and DZ twin pairs of all ages, and within younger and older MZ and DZ twin pairs (Table 5). IC analyses of twin pairs is often used to demonstrate the existence of genetic effect in smaller sample sizes. The IC of XCI skew within MZ twins pairs was positive and statistically significant (ICMZ_allAges = 0.31, P = 0.02). We found significant IC of XCI skew within older MZ twin pairs (ICMZ_older = 0.42, P = 0.005), but not within young MZ twin pairs (ICMZ_younger = 0.06, P = 0.8). We did not detect significant IC within DZ twin pairs at any age, in agreement with previous study in blood22. The higher IC of XCI skew within MZ twin pairs compared with DZ twin pairs indicates the involvement of genetic determinants in the regulation of XCI skew in blood-derived tissues. The increase of IC in older compared to younger MZ twin pairs and the fact that the heritability of XCI skew is observed only in females older than 55, confirm a role for genetic variants as age-dependent regulators of the acquired XCI skew in blood-derived tissues. Presumably, genetically-determined secondary cell selection processes act in haematopoietic cell lineages, with the high mitotic rates contributing to the manifestation of their effects in blood-derived tissues. Results also highlight an age-independent role for environmental factors as regulators of XCI skew in blood, fat, and skin tissues.
Individuals with autoimmunity exhibit more skewed XCI than unaffected co-twins
Chomosome X is enriched for immune-related genes. Most autoimmune disorders have higher prevalence in females than males17. Klinefelter syndrome (47,XXY) males have up to 14-fold higher risk of autoimmunity than 46,XY males46,47. These observations support an X-dosage effect in the pathogenesis of autoimmune diseases. As a mechanism of X-dosage compensation, the XCI process could be involved in the etiology of autoimmune disorders. Unbalanced X-linked transcriptional dosages toward one parental haplotype caused by skewed XCI patterns could influence the functions of the immune system. In particular during development, random XCI patterns in dendritic cells allow balanced expression of both parental X-linked self-antigens, a crucial event for the identification and negative selection of autoreactive T-cells in the thymus16,17. In line with the loss of mosaicism hypothesis48, we postulated that skewed XCI patterns may promote a breakdown of the tolerance induction processes with consequent release of autoreactive immune cells into the circulatory system. Supporting this hypothesis, higher frequencies of skewed XCI patterns have been observed in females affected with autoimmune disorders than in healthy controls12,13, however it is not known if this is a cause or consequence of disease, and these studies have not taken into account the underlying genetic predisposition of the cases and controls.
In order to address the association between autoimmune disease and XCI skewing while controlling for genetics, we investigated our samples for MZ twin pairs discordant for autoimmune disease. We identified eight MZ twins pairs discordant for RA in our study. RA is a chronic autoimmune condition affecting the lining of the synovial joints and associated with progressive disability49. RA is up to three times more frequent in females than males, and the most common age of onset ranges between 50 and 6050. Prevalence of RA ranges from 0.5% to 1%, but it significantly rises with age51. We found that the degree of XCI skewing in LCLs significantly differed between unaffected and affected co-twins (mean_DShealthy = 0.21; mean_DSaffected = 0.35; paired Wilcoxon’s test P < 0.05; Fig. 5). In seven out of eight twin pairs the affected co-twin was more skewed than their unaffected sister. These results are consistent with patterns seen in twins discordant for systemic lupus erythematosus and autoimmune thyroid disease52,53. Only four of the eight RA-discordant twin pairs had available XCI skew calls in fat and skin, and there was no significant difference in the XCI skew between the unaffected and affected co-twins in either tissue. Identical twins share 99% of the genome, age, and multiple environmental traits including in-utero growth, early life, and in most cases, socioeconomic statuses. Differences in the XCI patterns in blood may generate differences in the X-linked transcriptional dosages of immune-related genes and affect identical twins differently. Discordance in RA between identical twins could be well explained by differences in XCI patterns, however, we note that with cross-sectional data it is not possible to determine if changes in XCI drive RA or if RA causes changes in XCI skewing, or if both are independently associated to a third factor. Due to the low sample size, we acknowledge these results are not generalizable at a population level; larger sample sizes will be required to fully test the association of skewed XCI with twin discordance for RA and other autoimmune traits.
XCI patterns are more skewed in blood-derived tissues of individuals affected with rheumatoid arthritis than in their unaffected identical co-twins. Dotplot shows the degree of XCI skewing in LCLs from monozygotic twin pairs that are discordant for rheumatoid arthritis (n = 8 twin pairs). Blue dots on the left represent the twin without rheumatoid arthritis and red dots the co-twins diagnosed with rheumatoid arthritis. Co-twins are connected by a line. XCI X-chromosome inactivation.
XCI skew is associated with smoking status in older females
Tobacco smoking has been reported to induce epigenomic changes including DNA methylation variation (reviewed in ref. 54). Smoking is a well characterized risk factor in cancer55 and, as more recently discovered, in the etiology of autoimmunity56. Although smoking-related X-linked DNA methylation sites have been discovered57, no previous studies, to our knowledge, have investigated the relationship between smoking and XCI patterns. We reasoned that changes of XCI patterns may result from smoking, and affect in turn short-term and long-term health. In order to test our hypothesis, we used the 270 individuals in our dataset for which we had smoking status at the time of sample collection, including 233 never smokers and 37 current smokers58. We found no difference in the frequency of skewed XCI patterns between never and current smokers (36% and 35%, respectively) in LCLs. To take into account the effects of age on the degree of skewing in blood-derived tissues and to examine the relationship between smoking status and degree of skewing at different ages, we split the dataset into a younger (age < 55) and an older group (age ≥ 55; Supplementary Table 2). While the frequencies of skewed XCI were very similar between young smokers and young never smokers (27% and 28%, respectively), we detected a higher prevalence of skewed XCI in older smokers compared with older never smokers (47% and 40%, respectively). Accordingly, we found an overall positive association between XCI skew and smoking status in older (P = 0.02), but not in younger individuals (P = 0.5). The data suggest a role for smoking as a modulator of XCI skew in blood-derived tissues of females older than 55. Presumably, the association between smoking and XCI skew changes is complex, and further investigations are needed to characterize the genetic and molecular mechanisms underlying this phenomenon.
Discussion
In this study, we used multi-tissue transcriptomic data from twins to comprehensively characterize XCI patterns in LCLs, whole-blood, fat, and skin tissues from a healthy twin cohort. We show XCI patterns to be tissue-specific and that blood-derived tissues exhibited the highest prevalence of skewed XCI and share the highest similarity of XCI patterns. These findings indicate that XCI patterns are partially driven by tissue-specific mechanisms, and that the XCI skew measured in blood is not a reliable proxy for the skew in other tissues. Skewed XCI patterns limited to disease-relevant tissues and cells have been observed in multiple conditions9,10,14,15,59,60 but except for several cases of X-linked diseases, their roles in disease etiology and predisposition remain largely unknown. Our results demonstrate that tissue-specific XCI patterns within an individual is common in this healthy population.
We show that XCI skewing levels in blood tissues increase with age, with an inflection point at around 55, in line with previous reports18,19,20,21,22,24,40,44,45. In this study, more than 41% of females >65 years old demonstrate skewed XCI patterns in blood-derived tissues, indicating that acquired skewed XCI is a highly prevalent phenotype in ageing populations. We show age-related increase in XCI skew is a distinctive feature of blood-derived tissues, with no evidence for an age-related increase in fat or skin. Age-related increase in XCI skew partially explains the higher incidence of skewed XCI in blood than fat and skin tissues. The effects of age-related skewing of XCI on healthy ageing remain largely unknown, but may have a broad impact on the immune system. We demonstrate that within an individual, the XCI patterns in blood-derived tissues are shared across multiple immune cell types including monocytes, B-cells, T-CD4+, T-CD8+ and NK cells. Hematopoietic stem cells and the immune system continue to develop throughout life. Presumably, in line with the loss of mosaicism hypothesis48, imbalanced X-linked immune-related gene expression toward one parental haplotype leads to a reduced molecular diversity, which may translate in a decline of immune repertoire as well as poor sustenance of the immunological memory. Thus, by influencing the immune system, age-acquired skewed XCI could influence the predisposition to and manifestation of age-related traits, such as hematopoietic disorders, in women. We support an involvement of skewed XCI in the regulation of the immune system by showing that XCI patterns in LCLs, and consequently in multiple immune cell types, are consistently more skewed in individuals affected with autoimmunity than in healthy identical co-twins. Identical twins share nearly 100% of the genome, including chromosome X. Discordance in an autoimmune phenotype between twins, could partially be attributed to differences in the X-linked allelic and transcriptional dosages of X-linked immune-related genes resulting from difference in the XCI skew.
Previous twin studies have reported that XCI patterns in blood have a genetic component22,24. To our knowledge, this is the first study to investigate heritability of XCI skewing levels in other tissues. We found that the heritability of XCI skewing level is limited to blood-derived tissues of females >55 years old (h2 = 0.34), with no evidence of heritability in fat or skin or younger individuals in any tissue. The restriction of heritability to blood of older individuals is of interest given the link between skewed X-inactivation and clonal haematopoiesis. Positive selection of cells carrying an advantageous somatic mutation will lead to clonal haematopoiesis and skewed XCI patterns as the selected cells will carry the same inactivated parental X. Somatic mutation-driven clonal haematopoiesis is now known to be common in blood of healthy older individuals and is often referred to as clonal haematopoiesis of indeterminate potential (CHIP)36,61,62,63. CHIP is associated with increased risk of both cancer and all-cause mortality, and likewise skewed XCI, the prevalence of CHIP increase with age64,65. The higher skew in the XCI patterns in individuals affected with RA than in unaffected co-twins, is well explained by both the occurrence and the age-related increase in the prevalence of clonal hematopoiesis in RA patients66. The increase in XCI skew in older smokers in our study is also consistent with the increase in clonal haematopoiesis observed in smokers63,67,68. All together, these data converge in suggesting a link between XCI skew and clonal hematopoiesis. It is currently unknown to what extent CHIP accounts for age-acquired XCI skew, however, if it is a major driver this would suggest that like age-related XCI skew, CHIP has a significant germline genetic component. Stochastic selection of cells could also contribute to the variance of XCI skewing levels, but, in agreement with previous works22,24, we reason that their contribution is minimal. If stochastic selection of cells was a dominant mechanism, the correlation of XCI patterns between twin pairs would decrease with age.
Overall, the data presented in this study indicate a gene × age interaction that shifts the functional allelic dosages of X-linked heterozygous loci in a tissue-restricted manner. The high prevalence of skewed XCI and tissue-restricted XCI in a healthy population could complicate discovery of Chromosome X variants associated with a trait and subsequent genetic risk prediction, as an individual’s genotype may not match their functional genotypic dosage in the relevant tissue. Further investigations of the heterogeneity of XCI patterns across tissues and how this is regulated are essential to clarify the biomedical implications of skewed XCI and its role in healthy ageing in women.
Methods
Sample collection
The study included 856 female twins from the TwinsUK registry29,30 who participated in the MuTHER study69 . Study participants included both MZ and dizygotic (DZ) twins, aged 38–85 years old (median age = 60; Supplementary Fig. 1) and were of European ancestry. Volunteers received detailed information regarding all aspects of the research project and gave a prior signed consent to participate in the study. Peripheral blood samples were collected and LCLs were generated via EBV-mediated transformation of the B-lymphocyte fraction. Punch biopsies of subcutaneous adipose tissue were taken from a photo-protected area adjacent and inferior to the umbilicus. Skin samples were obtained by dissection from the punch biopsies. Adipose and skin samples were weighed and frozen in liquid nitrogen. This project was approved by the research ethics committee at St Thomasʼ Hospital London, where all the TwinsUK biopsies were carried out. Volunteers gave informed consent and signed an approved consent form prior to the biopsy procedure. Volunteers were supplied with an appropriate detailed information sheet regarding the research project and biopsy procedure by post prior to attending for the biopsy.
Genotyping and phasing
Whole-Genome Sequence data were generated and genotypes called within the UK10K project70. Briefly, DNA was sheared to 100–1000 bp and used for a paired-end library preparation. DNA libraries were sequenced on a Illumina HiSeq machine and 100 bp paired-end reads were generated. Reads were aligned to the GRCh37 reference genome using the Burrows-Wheeler Aligner71. SAM to BAM format conversion and sorting were performed using either Picard (broadinstitute.github.io/picard/) or samtools72. Variants were called from pooled alignments with samtools and bcftools (www.htslib.org/doc/bcftools.html) and VCF files were generated. For chrX and chrY variant calls, PAR of chrY was masked and male samples were called as diploid in their X-linked PAR and haploid in non-PAR regions. Sites were recalled and recalibrated with GATK UnifiedGenotyper and VariantRecalibrator73. SNV with a VQSLOD (variant quality score odds ratio) corresponding to a truth sensitivity <99.5% and with a Hardy–Weinberg equilibrium P-value < 1e−6 were discarded. Overall sequencing accuracy was assessed by comparing genotype concordance on chromosome 20 in 22 MZ twin pairs. Five hundred and fifty-seven individuals had both X-chromosome sequence data and RNA-seq data for at least one tissue. For individuals with unavailable X-chromosome sequence data, X-linked genotypes data were retrieved from the TwinsUK genotypes previously imputed into the 1000 Genomes Project phase 1 reference panel74,75,76. Haplotypes of X-linked SNPs with a MAF >5% were then phased using shapeit v2.r83777,78, with the–chrX flag to set up all functionalities for the phasing of non-PARs of the X chromosome, along with a phasing window of 2 Mb and 1000 conditional states. Phasing was performed using the genetic map b37 and the 1000 Genomes Project phase three reference panel of non-PAR X-linked haplotypes78. Phased X-linked SNPs were used for further analysis.
RNA-sequencing data
The Illumina TruSeq sample preparation protocol was used to generate the cDNA libraries for sequencing. Samples were sequenced on an Illumina HiSeq 2000 machine and 49 bp paired-end reads were generated. Adapter and polyA/T nucleotide sequences were removed and sequencing reads were aligned to the UCSC GRCh37/hg19 reference genome with the Burrows-Wheeler Aligner v.0.5.971. Samples that failed library preparation (according to the manufacturer’s guidelines) or had less than 10 million reads were discarded. Genes were annotated using the GENCODE v10 reference panel79.
Longitudinal RNA-sequencing data
Peripheral blood samples were collected 1–2.7 years apart from 114 female twins of the TwinsUK registry and were processed with the Illumina TruSeq protocol, sequenced on a HiSeq 2000 machine and 49 bp paired-end reads generated41. Adapter and polyA/T nucleotide sequences were trimmed using trim_galore and PrinSeq tools80, respectively. Reads were aligned to the UCSC GRCh37/hg19 reference genome with the STAR v.2.5.2a aligner81. Alignments containing non-canonical and unannotated splice junctions were discarded. Properly paired and uniquely mapped reads with a MAPQ of 255 were retained for further analysis.
Purified immune cells RNA-sequencing data
Monocytes, B, T-CD4+, T-CD8+ and NK cells were purified using fluorescence activated cell sorting (FACS) from two MZ twins exhibiting skewed XCI patterns in LCLs and from one individual exhibiting random XCI patterns (Supplementary Fig. 5). Total RNA was isolated and cDNA libraries for sequencing were generated using the Sureselect sample preparation protocol. Samples were then sequenced in triplicates on an Illumina HiSeq machine and 126 bp paired-end reads were generated. Adapter and polyA/T nucleotide sequences were trimmed using trim_galore and PrinSeq tools80, respectively. Human and prokaryotic rRNAs were identified using sortmerna v.2.182 and removed. Reads were aligned to the UCSC GRCh37/hg19 reference genome using STAR v.2.5.2a81. Alignments containing noncanonical and unannotated splice junctions were discarded. Properly paired and uniquely mapped reads with a MAPQ of 255 were retained for further analysis.
Correction of RNA-seq mapping biases
To eliminate mapping biases, all RNA-seq data were re-aligned within the WASP pipeline for mappability filtering83. The WASP tool has an algorithm specifically designed to identify and correct mapping biases in RNA-seq data. In each read overlapping a heterozygous SNP, the allele is flipped to the SNP’s other allele (generating all possible allelic combinations) and the read is remapped. Reads that did not remap to the same genomic location indicate mapping bias and were discarded. Reads overlapping insertions and deletions were also discarded. Properly paired and uniquely mapped reads were retained for analysis.
Quantification of allelic read counts and allele-specific expression (ASE)
Allelic read counts at heterozygous SNPs within XIST were quantified from paired RNA-seq and X-linked genotypes data using GATK ASEReadCounter84. Reads flagged by ASEReadCounter as having low base quality were discarded. No retained reads overlapped more than one heterozygous position in an individual. X-linked SNPs with a read depth of least ten reads were used. To increase the confidence that genotypes were truly heterozygous, only SNPs with both alleles detected in RNA-seq data at least once were retained for analysis, as previously done30. Samples with at least 1 heterozygous XIST-linked SNP passing all filters were retained as informative of XCI skewing levels (422 LCLs, 72 whole-blood, 378 fat and 336 skin samples). To quantify the ASE of each heterozygous XIST-linked SNP, the read count at the reference allele was divided by the read depth at the site84. The SNP’s ASE values range from 0 to 1, where 0 and 1 indicate monoallelic expression and 0.5 indicates biallelic expression.
Quantification of XIST ASE and degree of XCI skewing (DS)
In each sample, the XCI skewing levels were quantified by averaging the ASE values of heterozygous SNPs within XIST. All SNPs were phased prior to averaging as detailed above. The measure, called XISTASE is defined as follow:
where XIST_SNPASE are the ASE values of heterozygous SNPs within XIST and n is the number of heterozygous SNPs within XIST in the sample. XIST is uniquely expressed from the inactive X chromosome3,28,31, and thus the relative expression of parental alleles within XIST transcript are representative of XCI skewing levels in a bulk sample. The expression levels of polymorphisms within XIST have been used to infer XCI skewing levels in a sample32,33. We have also calculated XCI skew using the ASE of all non-PAR genes available in a sample, as previously done34. We compared our XISTASE calls to the non-PAR-based calls and concluded that the XISTASE calls are a better proxy for XCI skew in our analyses (Supplementary Note 1, Supplementary Figs. 2–4). Within each sample, the XISTASE values range from 0 to 1; an XISTASE value of 0.5 indicates equal inactivation of the two parental chromosomes (completely random XCI patterns, 50:50 XCI ratio), whereas a value of 0 or 1 indicates complete inactivation of one parental chromosome (completely skewed XCI patterns, 100:0 XCI ratio). To be consistent with previous literature22,38, we classified samples with XISTASE ≤ 0.2 or XISTASE ≥ 0.8 to have skewed XCI patterns, and samples with 0.2 < XISTASE < 0.8 to have random XCI patterns. To have an absolute measure of the magnitude of the XCI skewing levels in each sample, (or effect size of XISTASE), the degree of skewing of XCI (DS) was calculated. DS is the absolute deviation of XISTASE from 0.5. In each sample, DS was calculated as follow:
DS does not take into account the direction of XCI skewing, but the degree of deviation from a 50% XCI patterns (XISTASE = 0.5). DS is then a measure of the magnitude of XCI skewing levels in a sample. DS values range from 0 to 0.5, where 0 means random XCI and 0.5 completely skewed XCI patterns. Samples with DS ≥ 0.3 were classified to have skewed XCI, while samples with DS <0.3 were classified to have random XCI.
Heritability analysis of DS
The relative contributions of additive genetic factors (A), shared (C) and unique environmental factors (E) to the tissue-specific variance of DS, were calculated using the twinlm() function in the mets R package85. For each tissue, samples were split into a young (<55) and an older (≥55) group according to their ages (Supplementary Table 1). Due to the low number of MZ and DZ twin pairs in each group, whole-blood was excluded from heritability analysis. To further assess the contribution of genetic effects, the intraclass Spearman’s correlation (IC) of DS in blood-derived tissues of young and older MZ and DZ twin pairs was also calculated.
Differences in the DS between identical co-twins discordant for RA
We used a subset of eight MZ twin pairs where the co-twins of each pair are discordant for RA. Diagnoses were either confirmed during visits at the rheumatologist clinics at St Thomas’ Hospital in London, or confirmed by phone-interview by a rheumatology clinical fellow to confirm the diagnosis of RA based on the American College of Rheumatology 1987 criteria86. In case of unclear diagnosis of RA, participants were reviewed in clinic or were excluded. Difference in the distribution of the degree of XCI skewing in LCLs between the two groups (twins affected with autoimmunity vs healthy co-twins) was evaluated using paired Wilcoxon test. A P-value ≤ 0.05 was considered to be statistically significant.
Association between degree of skewing and smoking status
Association between the degree of skewing in LCLs and self-reported smoking status was tested in the 270 individuals with reliable smoking status recorded58. Dataset included 270 females classified either as current smokers (n = 37) or never smokers (n = 233; Supplementary Table 2). To examine the association between DS and smoking status, the smoking status was converted into a binary trait (0 = no smoker, 1 = smoker). A linear model of the DS as a function of the smoking status was then implemented for younger (age < 55) and older (age ≥ 55) individuals separately. Age was used as covariate. A P-value ≤ 0.05 was considered to be statistically significant.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
TwinsUK RNA-seq data is available from EGA (Accession number: EGAS00001000805). TwinsUK genotypes and phenotypes are available upon application to TwinsUK Data Access Committee (https://twinsuk.ac.uk/resources-for-researchers/access-our-data/). All other data are contained in the manuscript and its supplementary information.
References
Lyon, M. F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190, 372–373 (1961).
van den Berg, I. M. et al. X chromosome inactivation is initiated in human preimplantation embryos. Am. J. Hum. Genet. 84, 771–779 (2009).
Brown, C. J. et al. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature 349, 38–44 (1991).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
Pinter, S. F. et al. Spreading of X chromosome inactivation via a hierarchy of defined Polycomb stations. Genome Res. 22, 1864–1876 (2012).
Galupa, R. & Heard, E. X-Chromosome Inactivation: a crossroads between chromosome architecture and gene regulation. Annu Rev. Genet. 52, 535–566 (2018).
Gribnau, J., Barakat, T. S. X-chromosome inactivation and its implications for human disease. Preprint at https://www.biorxiv.org/content/10.1101/076950v1 (2017).
Puck, J. M., Nussbaum, R. L. & Conley, M. E. Carrier detection in X-linked severe combined immunodeficiency based on patterns of X chromosome inactivation. J. Clin. Investig 79, 1395–1400 (1987).
Migeon, B. R. et al. Adrenoleukodystrophy: evidence for X linkage, inactivation, and selection favoring the mutant allele in heterozygous cells. Proc. Natl Acad. Sci. USA 78, 5066–5070 (1981).
Plenge, R. M., Stevenson, R. A., Lubs, H. A., Schwartz, C. E. & Willard, H. F. Skewed X-chromosome inactivation is a common feature of X-linked mental retardation disorders. Am. J. Hum. Genet. 71, 168–173 (2002).
Ozbalkan, Z. et al. Skewed X chromosome inactivation in blood cells of women with scleroderma. Arthritis Rheumatol. 52, 1564–1570 (2005).
Chabchoub, G. et al. Analysis of skewed X-chromosome inactivation in females with rheumatoid arthritis and autoimmune thyroid diseases. Arthritis Res. Ther. 11, R106 (2009).
Kristiansen, M. et al. High frequency of skewed X inactivation in young breast cancer patients. J. Med. Genet. 39, 30–33 (2002).
Buller, R. E., Sood, A. K., Lallas, T., Buekers, T. & Skilling, J. S. Association between nonrandom X-chromosome inactivation and BRCA1 mutation in germline DNA of patients with ovarian cancer. J. Natl Cancer Inst. 91, 339–346 (1999).
Chitnis, S. et al. The role of X-chromosome inactivation in female predisposition to autoimmunity. Arthritis Res. 2, 399–406 (2000).
Libert, C., Dejager, L. & Pinheiro, I. The X chromosome in immune functions: when a chromosome makes the difference. Nat. Rev. Immunol. 10, 594–604 (2010).
Busque, L. et al. Nonrandom X-inactivation patterns in normal females: lyonization ratios vary with age. Blood 88, 59–65 (1996).
Hatakeyama, C. et al. The dynamics of X-inactivation skewing as women age. Clin. Genet. 66, 327–332 (2004).
Knudsen, G. P., Pedersen, J., Klingenberg, O., Lygren, I. & Orstavik, K. H. Increased skewing of X chromosome inactivation with age in both blood and buccal cells. Cytogenet Genome Res. 116, 24–28 (2007).
Wong, C. C. et al. A longitudinal twin study of skewed X chromosome-inactivation. PLoS ONE 6, e17873 (2011).
Kristiansen, M. et al. Twin study of genetic and aging effects on X chromosome inactivation. Eur. J. Hum. Genet. 13, 599–606 (2005).
Christensen, K. et al. X-linked genetic factors regulate hematopoietic stem-cell kinetics in females. Blood 95, 2449–2451 (2000).
Vickers, M. A., McLeod, E., Spector, T. D. & Wilson, I. J. Assessment of mechanism of acquired skewed X inactivation by analysis of twins. Blood 97, 1274–1281 (2001).
Bolduc, V. et al. No evidence that skewing of X chromosome inactivation patterns is transmitted to offspring in humans. J. Clin. Investig. 118, 333–341 (2008).
Gale, R. E., Wheadon, H., Boulos, P. & Linch, D. C. Tissue specificity of X-chromosome inactivation patterns. Blood 83, 2899–2905 (1994).
Bittel, D. C. et al. Comparison of X-chromosome inactivation patterns in multiple tissues from human females. J. Med. Genet. 45, 309–313 (2008).
Tukiainen, T. et al. Landscape of X chromosome inactivation across human tissues. Nature 550, 244–248 (2017).
Moayyeri, A., Hammond, C. J., Hart, D. J. & Spector, T. D. The UK Adult Twin Registry (TwinsUK Resource). Twin Res. Hum. Genet. 16, 144–149 (2013).
Buil, A. et al. Gene-gene and gene-environment interactions detected by transcriptome sequence analysis in twins. Nat. Genet. 47, 88–91 (2015).
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527–542 (1992).
Rupert, J. L., Brown, C. J. & Willard, H. F. Direct detection of non-random X chromosome inactivation by use of a transcribed polymorphism in the XIST gene. Eur. J. Hum. Genet. 3, 333–343 (1995).
Amos-Landgraf, J. M. et al. X chromosome-inactivation patterns of 1,005 phenotypically unaffected females. Am. J. Hum. Genet. 79, 493–499 (2006).
Shvetsova, E. et al. Skewed X-inactivation is common in the general female population. Eur. J. Hum. Genet. 27, 455–465 (2019).
Gentilini, D. et al. Age-dependent skewing of X chromosome inactivation appears delayed in centenariansʼ offspring. Is there a role for allelic imbalance in healthy aging and longevity? Aging Cell 11, 277–283 (2012).
Busque, L. et al. Recurrent somatic TET2 mutations in normal elderly individuals with clonal hematopoiesis. Nat. Genet. 44, 1179–1181 (2012).
Cotton, A. M. et al. Analysis of expressed SNPs identifies variable extents of expression from the human inactive X chromosome. Genome Biol. 14, R122 (2013).
Naumova, A. K. et al. Heritability of X chromosome–inactivation phenotype in a large family. Am. J. Hum. Genet. 58, 1111–1119 (1996).
Allen, R. C., Zoghbi, H. Y., Moseley, A. B., Rosenblatt, H. M. & Belmont, J. W. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am. J. Hum. Genet. 51, 1229–1239 (1992).
Mossner, M. et al. Skewed X-inactivation patterns in ageing healthy and myelodysplastic haematopoiesis determined by a pyrosequencing based transcriptional clonality assay. J. Med Genet. 50, 108–117 (2013).
Bryois, J. et al. Time-dependent genetic effects on gene expression implicate aging processes. Genome Res. 27, 545–552 (2017).
de Hoon, B., Monkhorst, K., Riegman, P., Laven, J. S. & Gribnau, J. Buccal swab as a reliable predictor for X inactivation ratio in inaccessible tissues. J. Med. Genet. 52, 784–790 (2015).
Pastinen, T. et al. A survey of genetic and epigenetic variation affecting human gene expression. Physiol. Genom. 16, 184–193 (2004).
Gale, R. E., Fielding, A. K., Harrison, C. N. & Linch, D. C. Acquired skewing of X-chromosome inactivation patterns in myeloid cells of the elderly suggests stochastic clonal loss with age. Br. J. Haematol. 98, 512–519 (1997).
Tonon, L. et al. Unbalanced X-chromosome inactivation in haemopoietic cells from normal women. Br. J. Haematol. 102, 996–1003 (1998).
Scofield, R. H. et al. Klinefelter's syndrome (47,XXY) in male systemic lupus erythematosus patients: support for the notion of a gene-dose effect from the X chromosome. Arthritis Rheumatol. 58, 2511–2517 (2008).
Seminog, O. O., Seminog, A. B., Yeates, D. & Goldacre, M. J. Associations between Klinefelter's syndrome and autoimmune diseases: English national record linkage studies. Autoimmunity 48, 125–128 (2015).
Ozcelik, T. X chromosome inactivation and female predisposition to autoimmunity. Clin. Rev. Allergy Immunol. 34, 348–351 (2008).
Guo, Q. et al. Rheumatoid arthritis: pathological mechanisms and modern pharmacologic therapies. Bone Res. 6, 15 (2018).
Mueller, R. B. et al. Is radiographic progression of late-onset rheumatoid arthritis different from young-onset rheumatoid arthritis? Results from the Swiss prospective observational cohort. Rheumatology 53, 671–677 (2014).
Scott, D. L., Wolfe, F. & Huizinga, T. W. Rheumatoid arthritis. Lancet 376, 1094–1108 (2010).
Brix, T. H. et al. High frequency of skewed X-chromosome inactivation in females with autoimmune thyroid disease: a possible explanation for the female predisposition to thyroid autoimmunity. J. Clin. Endocrinol. Metab. 90, 5949–5953 (2005).
Huang, Q., Parfitt, A., Grennan, D. M. & Manolios, N. X-chromosome inactivation in monozygotic twins with systemic lupus erythematosus. Autoimmunity 26, 85–93 (1997).
Gao, X., Jia, M., Zhang, Y., Breitling, L. P. & Brenner, H. DNA methylation changes of whole blood cells in response to active smoking exposure in adults: a systematic review of DNA methylation studies. Clin. Epigenetics 7, 113 (2015).
Ma, Y. & Li, M. D. Establishment of a strong link between smoking and cancer pathogenesis through DNA methylation analysis. Sci. Rep. 7, 1811 (2017).
Mahdi, H. et al. Specific interaction between genotype, smoking and autoimmunity to citrullinated alpha-enolase in the etiology of rheumatoid arthritis. Nat. Genet. 41, 1319–1324 (2009).
Klebaner, D. et al. X chromosome-wide analysis identifies DNA methylation sites influenced by cigarette smoking. Clin. Epigenetics 8, 20 (2016).
Tsai, P. C. et al. Smoking induces coordinated DNA methylation and gene expression changes in adipose tissue with consequences for metabolic health. Clin. Epigenetics 10, 126 (2018).
Andreu, N. et al. Wiskott-Aldrich syndrome in a female with skewed X-chromosome inactivation. Blood Cells Mol. Dis. 31, 332–337 (2003).
Li, G. et al. Skewed X chromosome inactivation of blood cells is associated with early development of lung cancer in females. Oncol. Rep. 16, 859–864 (2006).
Abelson, S. et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 559, 400–404 (2018).
Loh, P. R. et al. Insights into clonal haematopoiesis from 8,342 mosaic chromosomal alterations. Nature 559, 350–355 (2018).
Zink, F. et al. Clonal hematopoiesis, with and without candidate driver mutations, is common in the elderly. Blood 130, 742–752 (2017).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Steensma, D. P. Clinical consequences of clonal hematopoiesis of indeterminate potential. Blood Adv. 2, 3404–3410 (2018).
Savola, P. et al. Clonal hematopoiesis in patients with rheumatoid arthritis. Blood Cancer J. 8, 69 (2018).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Coombs, C. C. et al. Therapy-related clonal hematopoiesis in patients with non-hematologic cancers is common and associated with adverse clinical outcomes. Cell Stem Cell 21, 374–382 e374 (2017).
Grundberg, E. et al. Multiple tissue human expression resource, C. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat. Genet. 44, 1084–1089 (2012).
Consortium, U. K. et al. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Zheng, H. F. et al. Performance of genotype imputation for low frequency and rare variants from the 1000 genomes. PLoS ONE 10, e0116487 (2015).
Genomes Project, C. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Suhre, K. et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 477, 54–60 (2011).
Delaneau, O., Zagury, J. F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Delaneau, O. & Marchini, J. Genomes Project, C. & Genomes Project, C. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat. Commun. 5, 3934 (2014).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Kopylova, E., Noe, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Castel, S. E., Levy-Moonshine, A., Mohammadi, P., Banks, E. & Lappalainen, T. Tools and best practices for data processing in allelic expression analysis. Genome Biol. 16, 195 (2015).
Scheike, T. H., Holst, K. K. & Hjelmborg, J. B. Estimating heritability for cause specific mortality based on twin studies. Lifetime Data Anal. 20, 210–233 (2014).
Webster, A. P. et al. Increased DNA methylation variability in rheumatoid arthritis-discordant monozygotic twins. Genome Med. 10, 64 (2018).
Acknowledgements
This study was supported by MRC Project Grant (MR/R023131/1) to K.S.S. The TwinsUK study was funded by the Wellcome Trust and European Community’s Seventh Framework Programme (FP7/2007-2013). The TwinsUK study also receives support from the National Institute for Health Research (NIHR)-funded BioResource, Clinical Research Facility and Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust in partnership with King’s College London. This work was also supported by the a joint UK Economic and Social Research Council (ESRC) and Biotechnology and Biological Sciences Research Council (BBSRC) grant (ES/N000277/1) to CCYW. P.C.T. is supported by a Chang Gung Memorial Hospital Research Funding grant (CMRPD1J0081). This project was enabled through access to the MRC eMedLab Medical Bioinformatics infrastructure, supported by the Medical Research Council [grant number MR/L016311/1].
Author information
Authors and Affiliations
Contributions
A.Z. M.D. and K.S.S conceived and designed the project. A.Z. performed analysis. S.N. contributed data discussion. P.C.T and J.T.B. contributed data. R.A.E. performed RNA isolation and FACS experiments. S.R. and C.Y.W performed HUMARA experiments. A.Z. and K.S.S. wrote the paper. All authors read and approved the manuscript. The authors thank Julia El-Sayed Moustafa and Amy Roberts for providing feedback on the manuscript and Alessandro Morea for assistance with HUMARA assays. We also thank Philippa Wells, Flore Zufferey, and Frances Williams for their work identifying cases of rheumatoid arthritis in the TwinsUK cohort.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zito, A., Davies, M.N., Tsai, PC. et al. Heritability of skewed X-inactivation in female twins is tissue-specific and associated with age. Nat Commun 10, 5339 (2019). https://doi.org/10.1038/s41467-019-13340-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-13340-w
This article is cited by
-
Characterization of a novel heterozygous frameshift variant in NDP gene that causes familial exudative vitreoretinopathy in female patients
Molecular Genetics and Genomics (2024)
-
X chromosome inactivation skewing is common in advanced carotid atherosclerotic lesions in females and predicts secondary peripheral artery events
Biology of Sex Differences (2023)
-
A novel quantitative targeted analysis of X-chromosome inactivation (XCI) using nanopore sequencing
Scientific Reports (2023)
-
Investigation of the role of X chromosome inactivation and androgen receptor CAG repeat polymorphisms in patients with recurrent pregnancy loss: a prospective case–control study
BMC Pregnancy and Childbirth (2022)
-
BEXCIS: Bayesian methods for estimating the degree of the skewness of X chromosome inactivation
BMC Bioinformatics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.