Abstract
Although the biological importance of post-transcriptional RNA modifications in gene expression is widely appreciated, methods to directly detect their introduction during RNA biosynthesis are rare and do not easily provide information on the temporal nature of events. Here, we introduce the application of NMR spectroscopy to observe the maturation of tRNAs in cell extracts. By following the maturation of yeast tRNAPhe with time-resolved NMR measurements, we show that modifications are introduced in a defined sequential order, and that the chronology is controlled by cross-talk between modification events. In particular, we show that a strong hierarchy controls the introduction of the T54, Ψ55 and m1A58 modifications in the T-arm, and we demonstrate that the modification circuits identified in yeast extract with NMR also impact the tRNA modification process in living cells. The NMR-based methodology presented here could be adapted to investigate different aspects of tRNA maturation and RNA modifications in general.
Similar content being viewed by others
Introduction
Over 170 nucleotide modifications are currently reported in RNAs from the three domains of life, the vast majority being found in tRNAs1,2. This family not only displays the largest variety of post-transcriptional decorations among RNA molecules, but also the highest density of modifications per RNA transcript3,4. The introduction of post-transcriptional chemical modifications is central in the maturation process to generate functional tRNA molecules5,6,7. The biogenesis of tRNAs is a multi-step process that comprises the synthesis of a tRNA precursor (pre-tRNA) by transcription of a tRNA gene, the removal of the 5'-leader and 3'-trailer sequences, the addition of the 3'-CCA amino-acid accepting sequence, the removal of intronic sequences when present, and the incorporation of a large number of post-transcriptional chemical modifications8. Modifications are found in two main regions of the L-shaped structure of tRNAs, the tRNA core, and the anticodon-loop region (ACL)4. All cellular functions of tRNAs, including their alternative functions outside translation, are controlled and modulated by modifications5,9,10. While modified nucleotides in the ACL participate in decoding during protein synthesis11,12, those in the tRNA core are collectively implicated in the folding and stability of tRNAs13.
While a simple model would be that modifications are introduced to tRNA independently, several modification circuits have been identified in which one or more modifications stimulate formation of a subsequent modification7,14. This obviously drives a defined sequential order in the tRNA modification process. In addition, the interdependence of multiple modification events might in certain cases be responsible for the variations of tRNA modifications in response to environmental perturbations2,7,15,16,17. The characterization of modification circuits is therefore central to understand the dynamic regulation of modifications in tRNAs, however their identification remains difficult, since monitoring tRNA maturation at a single nucleotide level in a time-resolved fashion is technically challenging18.
In the past 10 years, NMR spectroscopy became a pre-eminent method to investigate post-translational modifications (PTMs) in proteins19,20. Time-resolved NMR measurements provided the means to monitor the establishment of PTMs in vitro, in cellular extracts and in living cells21,22,23,24,25. In addition, monitoring the introduction of PTMs with NMR has provided mechanistic insights into modification hierarchies, with initial modifications exhibiting stimulatory or inhibitory effects on subsequent modification events26,27.
Here, inspired by the NMR monitoring of PTMs in cellular environments, we report an original methodology to monitor RNA modifications in cellular extracts with NMR. Using yeast tRNAPhe as a model system, we demonstrate that multiple modification events can be monitored in yeast extract with NMR in a time-resolved fashion. Using continuous NMR measurements to measure a series of snapshots of the tRNA along the maturation route, we observe a sequential order in the introduction of several modifications. This suggests that modification circuits could control the tRNAPhe maturation process in yeast. We next adopted a reverse genetic approach and investigated the interplay between the different modifications in yeast tRNAPhe with both NMR and mass spectrometry and show that modification circuits identified in the yeast extract on tRNAPhe also influence the process of tRNA modification in living cells.
Results
Monitoring RNA modifications in cellular extracts with NMR
In order to investigate the different steps along the maturation pathway of post-transcriptionally modified RNAs, we sought to implement NMR-based methods for monitoring the introduction of modifications in RNA substrates. As a general concept, we believed that modification reactions by RNA-modifying enzymes could be probed by NMR in cell extracts containing enzymatic activities responsible for the modification of the RNA substrate of interest. Introducing isotope-labeled RNAs into unlabeled cell extracts combined with the use of isotope-filters in NMR experiments enables the detection of the sole RNA of interest within the complex cell extract environment. The non-disruptive nature of NMR provides the means to directly monitor RNA modification events in a continuous and time-resolved fashion, by measuring successive NMR experiments on a single sample (Fig. 1a). In addition, since NMR spectroscopy provides information at atomic-resolution, multiple RNA modifications introduced on the same substrate, e.g. methylations on nearby nucleotides, can be easily distinguished.
Schematic description of the method. a An isotope-labeled RNA (represented as red dots) is introduced in cell extracts in an NMR tube to yield an in extract NMR sample. Successive NMR measurements on a single sample incubated directly in the NMR spectrometer can then be performed. b RNA modifications are monitored on the RNA imino signals with 2D (1H,15N) NMR spectra in a time-resolved fashion. Modifications cause the progressive disappearance of imino signals from unmodified nucleotides (signal M disappearing in blue) and the correlated appearance of new imino signals from modified nucleotide (signal M appearing in red). Modifications also cause disappearance and correlated appearance of imino signals of nearby nucleotides (signal P in green). Nucleotides far away from modification sites have unperturbed imino signals over time (signal U in black)
Our approach for monitoring of RNA post-transcriptional modifications with NMR relies on the fact that imino signals of RNAs are very sensitive to their chemical environment. Imino groups are carried by uridines and guanosines, and are easily observed in 1H–15N correlation spectra on condition that the imino proton is protected from exchange with the solvent by hydrogen bonding in, e.g. a Watson–Crick-like pairing. Different types of behavior for imino signals can in principle be observed in 1H–15N correlation spectra upon enzymatic modifications (Fig. 1b). The incorporation of a chemical group on a defined nucleotide will affect the chemical environment of the modified nucleotide itself but also of nearby nucleotides. Overall, this will cause the progressive disappearance of signals from the unmodified RNA and the correlated appearance of new signals from the modified RNA. For the sake of clarity, the disappearance and the correlated appearance related to the imino signal of a nucleotide experiencing a chemical modification will be thereafter called ‘a direct effect’ (signal M on Fig. 1b), whereas the disappearance and the correlated appearance associated with the imino signal of a nearby nucleotide will be referred to as ‘an indirect effect’ (signal P on Fig. 1b). The possibility to observe indirect effects enables the detection of modifications on adenosines and cytosines, even though they do not carry imino groups.
Yeast tRNAPhe as a model for monitoring RNA modifications
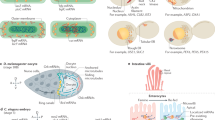
As a proof of concept, we undertook the investigation of the maturation of the yeast tRNAPhe in yeast extract. Matured tRNAPhe contains 14 modified nucleotides (Fig. 2a and Supplementary Table 1), among which: (i) T54, Ψ55, and D16 are found in almost all yeast tRNAs; (ii) m1A58, m22G26, m2G10, Ψ39, m7G46, and m5C49 are frequently found in yeast tRNAs; and (iii) Gm34, yW37, and m5C40 are found uniquely in tRNAPhe in yeast1. To evaluate the ability of yeast extracts to modify yeast tRNAPhe, we first produced an unmodified 15N-labeled tRNAPhe by in vitro transcription (Fig. 2a), which to a certain extent can be regarded as a tRNAPhe precursor (pre-tRNAPhe) with processed 5′- and 3′-termini and spliced intron, but lacking all RNA modifications. This choice of substrate affects the introduction of certain modifications near the anticodon28, but constitutes a simplified system, which has the advantage to spotlight nucleotide modifications over RNA processing steps. The chemical shifts of imino groups involved in secondary and tertiary interactions were assigned using standard NMR procedures for RNAs29. NMR-fingerprints of tRNAs can be efficiently acquired with 1H–15N Band-selective excitation short-transient Transverse relaxation optimized spectroscopy (BEST-TROSY) experiments30. To serve as a reference spectrum of yeast tRNAPhe without any modifications, we measured a 1H–15N BEST-TROSY experiment in vitro (Fig. 2b), in a buffer aiming to approach cellular conditions. We next incubated this tRNA at 30 °C for 12 h in yeast cellular extracts prepared under mild conditions in order to preserve most enzymatic activities of the cell. Cellular extracts were supplemented with enzymatic cofactors, such as S-adenosyl-l-methionine (SAM), the almost universal methyl donor of RNA methylations by methyltransferases31 (MTases), and reduced nicotinamide adenine dinucleotide phosphate (NADPH), a hydride donor implicated in the reduction of uridines to dihydrouridines32. After the 12 h of incubation, we measured a BEST-TROSY experiment (experimental time of 2 h) directly in yeast extracts. The 15N-isotope filter of the experiment enables the exclusive detection of the 15N-labeled tRNAPhe signals, even in this complex environment (Fig. 2b). The comparison of this spectrum with the reference spectrum revealed obvious differences, with apparent additional signals, as well as duplicated or shifted signals (Fig. 2b). Remarkably, tRNA signals are still observed after this long incubation, confirming that tRNA transcripts are not rapidly degraded by RNases down to single nucleotides and are stable for several hours in yeast extracts28. In addition, the line-width of the NMR signals is not particularly broadened in the extracts, meaning that the overall tumbling time of the 15N-labeled tRNAPhe is unchanged compared with the in vitro situation, and therefore that it remains mostly free and does not predominantly associate with proteins or other cellular components within large molecular complexes. These two hurdles, namely substrate degradation and signal line broadening, were identified in a pioneer study as the most prominent difficulties that complicate in depth investigations of nucleic acids with NMR in cellular environments33. We did not face these difficulties in the case of tRNA transcripts in yeast extracts, which opened the way to a thorough investigation of the fate of tRNAs throughout their maturation pathway with NMR.
Modification of yeast tRNAPhe in yeast extract. a Sequence and cloverleaf representation of unmodified yeast tRNAPhe (left) and modified tRNAPhe (right). Main tertiary interactions are represented with thin lines. Modifications are in red. b (left) Imino (1H,15N) correlation spectrum of a 15N-labeled unmodified tRNAPhe measured in vitro in a buffer approaching physiological conditions (see “Methods” section). Assignments of the imino groups are reported on the spectrum. (right) Imino (1H,15N) correlation spectrum of a 15N-labeled tRNAPhe measured after 12 h of incubation at 30 °C in yeast extract. The most obvious changes as compared with the spectrum of the unmodified tRNAPhe are highlighted with red patches
Changes in the NMR-fingerprint of tRNAPhe (Fig. 2b) can be associated with expected chemical modifications. For instance the signal of U55, a residue that gets modified into Ψ55 and that is found near two other modified nucleotides, i.e. T54 and m1A58, gave rise to three individual signals (Fig. 2a, b). Another example comes from the signal of G24, which is split into two (Fig. 2b), and for which the appearance of the second signal could be linked to the m2G10 modification, since the G24:C11 base pair stacks over this modified nucleotide (Fig. 2a). However, the exact assignments of all the changes on the NMR-fingerprint of tRNAPhe, which are potentially generated by direct and indirect effects from all modifications, remains impractical only from the signal assignments of the initial unmodified tRNA transcript. We thus performed further investigations by NMR in order to accurately interpret these changes on the NMR-fingerprint of tRNAPhe upon incubation in yeast extracts.
The NMR signature of individual modifications
To characterize the effect of modifications on the NMR-fingerprint of tRNAPhe, we measured NMR spectra on three distinct tRNAPhe samples differing in their modification content. The first sample is the aforementioned tRNAPhe sample produced by in vitro transcription, which presents none of the modifications (Fig. 3a, left). The second sample corresponds to native tRNAPhe purified from Saccharomyces cerevisiae, and therefore contains all 14 modifications found in fully modified yeast tRNAPhe (Fig. 3a, right). The third sample corresponds to a recombinant yeast tRNAPhe produced in and purified from an Escherichia coli strain overproducing this tRNA with a system previously described34,35. Since E. coli has a less prolific but related tRNA modification machinery than S. cerevisiae, the modification pattern of this sample resembles that of the fully modified yeast tRNAPhe, but overall exhibits fewer modifications (Fig. 3a, center). The modification status of a tRNA expressed in heterologous systems can be inferred from the transposition of the known tRNA modification patterns found in the host organism onto the recombinant tRNA sequence. This approach predicts that yeast tRNAPhe produced in E. coli contains eight modifications, six of which are identical to the ones found in natural tRNAPhe (Fig. 3a). This modification pattern was confirmed by further NMR analysis of the sample.
The comparison of different maturation states gives the NMR signature of individual modifications. a Sequence and cloverleaf representation of unmodified yeast tRNAPhe produced by in vitro transcription (left), yeast tRNAPhe produced and modified in E. coli (middle), and yeast tRNAPhe produced and modified in yeast (right). Main tertiary interactions are represented with thin lines. Modifications are in red. b Imino (1H,15N) correlation spectra and imino group assignments of 15N-labeled tRNAPhe measured in vitro: (left) unmodified yeast tRNAPhe, spectrum in red; (middle) yeast tRNAPhe produced and modified in E. coli, spectrum in cyan; (right) yeast tRNAPhe produced and modified in yeast, spectrum in purple. c Superposition of imino (1H,15N) correlation spectra of various tRNAPhe samples and NMR signature of individual modifications: (left) unmodified (red) and E. coli-modified (cyan); (middle) unmodified (red) and yeast-modified (purple); (right) unmodified (red), E. coli-modified (cyan), and yeast-modified (purple). A selection of imino assignments is reported on the spectra. NMR signatures of modifications are reported with continuous line arrows (direct effects—see main text), or dashed arrows (indirect effects)
We next performed the chemical shift assignments of the imino groups of the two modified tRNAPhe samples, produced in E. coli and yeast, with the same approach used for the unmodified tRNAPhe sample. Imino protons assignments of the fully modified tRNAPhe are consistent with the ones reported by early NMR studies of this tRNA36,37. Chemical shifts assignments of the imino groups of the three samples are reported on 1H–15N BEST-TROSY experiments measured in identical conditions in Fig. 3b. The comparison of the NMR fingerprints of the three samples revealed the NMR signature of individual modifications (Fig. 3c). The identified direct effects and indirect effects are reported in Fig. 3c with solid line arrows and dashed line arrows, respectively. NMR signatures of modifications in the anticodon loop could not be identified on this type of NMR spectra, since their imino protons are not engaged in base pairing and are not protected from exchange with the solvent, and are therefore not detectable. For the identification of the NMR signature of modifications, the 3D structure of yeast tRNAPhe provided a valuable aid, enabling the precise identification of the nearest modifications to an imino group38. The identified NMR signatures (Fig. 3c) might seem puzzling if one looks at the 2D cloverleaf representation of tRNAs, but become more apparent if one has in mind the complicated 3D L-shaped structure of tRNAs (Supplementary Fig. 1). As an example, we describe a few NMR signatures of individual modifications derived from the comparison of the NMR fingerprints. For instance, the comparison of the unmodified with the E. coli-produced tRNAPhe (Fig. 3c, left) reveals the NMR signature of the T54 modification, with a direct effect on the imino group of U54 (solid line) and an indirect effect on U52 (dashed line). Moreover, the comparison of the NMR-fingerprints of the three samples (Fig. 3c, right) reveals an NMR signature for the m1A58 modification, with an indirect effect on the imino group of T54 (dashed line). The indirect effect is very pronounced in this case, most probably because T54 forms a non-canonical base pair with A58, the chemical environment of which is highly perturbed by the positive charged appearing on the A58 base after methylation at position 1. Similarly, the comparison of the unmodified with the E. coli-produced tRNAPhe (Fig. 3c, left) reveals the NMR signature of the Ψ55 modification, with a direct effect on the imino group of U55 (solid line); and the comparison of the NMR-fingerprints of the three samples (Fig. 3c, right) reveals an NMR signature for the m1A58 modification, with an indirect effect on the imino group of Ψ55 (dashed line). Overall, we identified the NMR signatures for nine modifications, most of which are located in the tRNA core, i.e. m2G10, D16, and m22G26 in the D-arm; m5C40 in the anticodon-arm; m7G46 in the variable region; and m5C49, T54, Ψ55, and m1A58 in the T-arm (Fig. 3). This paved the way to a detailed investigation of the introduction of RNA modifications in yeast tRNAPhe by NMR.
Time-resolved NMR monitoring of RNA modifications
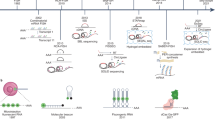
In order to record time-resolved snapshots along the tRNAPhe modification pathway, we incubated the unmodified 15N-labeled tRNAPhe at a concentration of 40 μM in active yeast extract supplemented with the modification enzymes cofactors SAM and NADPH. The incubation was done at 30 °C directly in an NMR tube in the NMR spectrometer, and a series of 1H–15N BEST-TROSY experiments were measured from initial mixing time up to ~24 h after starting the monitoring of tRNA maturation (Fig. 4a). At this concentration of 15N-labeled tRNA, reasonable signal-to-noise ratio (SNR) is achieved with an acquisition time of 2 h, and therefore each NMR fingerprint measurement spreads over a 2 h time period (Fig. 4a). Importantly, all changes observed in the NMR-fingerprints can be rationalized and attributed to specific RNA modifications on the basis of the NMR signatures identified above (Fig. 3). The identifications of specific RNA modification in each snapshot are displayed with solid line and dashed line arrows for direct and indirect effects, respectively (Fig. 4a). We observed different types of behavior for different modifications. First, some modifications appear early in the sequence, such as Ψ55, m7G46, and T54; and some appear late, such as m1A58. Second, some modifications are introduced during a short interval of time, such as Ψ55 and m7G46, whereas others are introduced over a long time period, such as T54 and m2G10, suggesting different levels of intrinsic catalytic activities for the different modification enzymes. In detail, the time-resolved NMR monitoring of RNA modifications in yeast tRNAPhe reveals that the modification Ψ55 is the first one to be introduced and that U55 is completely modified before any other modification can be detected (Fig. 4a, t = 0–2 h), showing that Pus4 is one of the most active modification enzymes in yeast and suggesting that it is highly efficient on a non-modified tRNA transcript and does not require prior modifications. Then, m7G46 and T54 are the two next modifications to be introduced (Fig. 4a, t = 2–4 h), and whereas m7G46 is rapidly fully modified (t = 2–6 h), T54 is introduced over a longer time period (t = 2–18 h), suggesting that Trm2 is a less active enzyme than the Trm8–Trm82 complex. The three next modifications to be observed in the sequence are m2G10, m5C49, and D16 (Fig. 4a, t = 4–6 h). Among these three enzymes, the complex responsible for the introduction of the m2G10 modification (Trm11–Trm112) appears to be the least active one. Even though the T54 modification is observed earlier in the time-course of modification, the three modified nucleotides m2G10, m5C49, and D16 are initially detected at a time point, where U54 is not completely converted into T54, meaning that these modifications could, in principle, be introduced on tRNAs lacking the T54 modification. The strict sequential introduction of m2G10, m5C49, and D16, only after T54 is fully present is also compatible with our data, since ensemble-averaged measurements do not allow us to discriminate between these two possibilities (see below). The last modification to be detected along the tRNA maturation pathway is the m1A58 modification (Fig. 4a, t = 8–10 h). Similarly, the m1A58 modification is initially detected at a time point, where U54 is not completely modified into T54, meaning that it could, in principle, be introduced on tRNAs lacking the T54 modification. However, our data suggest that the m1A58 modification is strictly introduced after T54. Indeed, we detect the m1A58 modification by its indirect effect on the T54 imino group signal (Fig. 3c). If the m1A58 modification is introduced on a tRNA lacking the T54 modification, it would be detected by a large perturbation on the U54 imino group that would then resonate at a different position than that of T54 (δH = 12.27 ppm; δN = 158.2 ppm), since it lacks the T54 modification. Importantly, no unidentified transient signal, which could correspond to tRNAs with m1A58 but still lacking T54, is observed during the incubations. The m1A58 modification of U54-containing tRNAs is therefore not observed in our experiments, suggesting a strict introduction of m1A58 after T54.
Time-resolved NMR monitoring of RNA modifications in yeast tRNAPhe. a Imino (1H,15N) correlation spectra of a 15N-labeled tRNAPhe measured in vitro (first top left spectrum—same as Fig. 2b left) and in a time-resolved fashion during a continuous incubation at 30 °C in yeast extract over 26 h (remaining eight spectra). Each NMR spectrum measurement spreads over a 2 h time period, as indicated. Modifications detected at the different steps are reported with continuous line arrows for direct effects, or dashed arrows for indirect effects (see main text for the distinction between direct and indirect effects). b Schematic view of the sequential order of the introduction of modifications in yeast tRNAPhe as observed with NMR. Each modification is associated with a defined color also used in Figs. 5 and 6
As a summary, the sequence in the introduction of modifications in yeast tRNAPhe as monitored by NMR is schematically represented in Fig. 4b. Two modifications for which NMR signatures have been identified (Fig. 3c) were not observed with these conditions and incubation time, namely m22G26 and m5C40. The lack of the m5C40 modification was expected, since it was reported to strictly depend on the presence of the intron at the level of the anticodon region28. Taken as a whole, we clearly observed a sequential order in the introduction of the modifications found in the heart of the 3D structure of yeast tRNAPhe (Fig. 4b), which could suggest that regulatory circuits are controlling the introduction of these modifications. However, since the modification content observed by NMR is averaged over the total tRNAPhe population, the exact modification status of each tRNA molecule cannot be firmly established. This type of questions that needs to be addressed at a single molecule level, cannot be resolved with currently available methods aimed at quantifying the modification content of RNAs. For instance, in our case it is not possible to know whether or not all m2G10-containing tRNAs also contain T54 and m7G46 modifications. Importantly, with our experimental setup, it is difficult to discriminate between an authentic dependence on the prior introduction of certain modifications and a sequential order caused by different intrinsic catalytic activities of the enzymes. In order to address these important questions concerning tRNA maturation and identify potential interdependencies among RNA modifications, we have adopted a reverse genetic approach and conducted a systematic analysis of tRNAPhe maturation in different yeast strains.
Complex circuits of RNA modifications in yeast tRNAPhe
In order to identify a potential interplay between different modification enzymes, we recorded several time-resolved snapshots of the tRNAPhe modification pathway in identical conditions as in Fig. 4, with the exception that the unmodified 15N-labeled tRNAPhe was incubated in various extracts prepared from yeast strains depleted of one specific modification enzyme at a time (Fig. 5a and Supplementary Figs. 2–8). The modification pattern was then compared with that of the wild-type yeast extract (Fig. 4). First, this provided independent confirmation of the NMR signatures of the individual modifications identified previously, since changes in the NMR spectra associated with a given modification were no longer observed in the corresponding depleted yeast extract. Second, the detailed analysis of the different profiles of the tRNAPhe modification pathways, provided a way to visualize any subtle interdependence between the multiple modification events. To identify these correlated changes, it is best to compare the whole series of snapshots measured in the wild-type and depleted yeast extracts side by side (Fig. 4a and Supplementary Figs. 2–8). However, for ease of visualization, a single time point, intermediate in the time-course of the tRNAPhe modification pathway, and at which most of the differences between the wild-type and depleted strains are apparent, is shown in Fig. 5a.
Complex modification circuits in yeast tRNAPhe. a Imino (1H,15N) correlation spectra of a 15N-labeled tRNAPhe measured after 16 h of incubation at 30 °C in yeast extracts prepared from wild-type cells (top left spectrum) and in yeast extracts depleted of one modification enzyme at a time, namely dus1Δ, pus4Δ, trm1Δ, trm2Δ, trm4Δ, trm8Δ, and trm11Δ. Each NMR spectrum measurement spreads over a 2 h time period (t = 16–18 h). Complete series of time-resolved NMR monitoring of tRNAPhe maturation in the different depleted yeast extracts are given in Supplementary Figs. 2–8. Modifications occurring at this specific step in each extract are reported with continuous line arrows for direct effects, or dashed arrows for indirect effects. Each strain is associated with a defined color also used in panel b. b Schematic view of the modification circuits revealed by the NMR monitoring of tRNAPhe maturation in the different yeast extracts of panel a. Each modification is displayed on the cloverleaf structure with its associated color also used in Figs. 4 and 6. Arrows indicate stimulatory effects and blunted lines inhibitory effects. Thick and thin lines indicate strong and slight effects, respectively
Very strong effects are seen in some cases, as for instance in the case of the pus4∆ strain, where not only the Ψ55 modification is absent after a long incubation in the corresponding yeast extract, as expected, but the m1A58 modification is absent as well. In addition, the incorporation of T54 is substantially hindered in the pus4∆ strain. Likewise, the incorporation of m1A58 is much reduced in the trm2∆ strain (preventing T54 formation), which altogether suggest a regulatory circuit in the T-arm of tRNAPhe. In this circuit of modification, Ψ55 has a positive effect on the introduction of T54 and m1A58 by their respective enzymes (Supplementary Table 1), and T54 has a positive effect on the introduction of m1A58 (Fig. 5b). Another strong effect is observed in the case of the trm11∆ strain (preventing m2G10 formation), where the m22G26 modification is apparent after ~14 h of incubation in the trm11∆ yeast extract, but is undetectable in wild-type and all other depleted strain extracts after ~24 h of incubation. Therefore, m2G10 has a negative effect on the introduction of m22G26 (Fig. 5b). Less drastic effects are also apparent in the different time-courses of the tRNAPhe modification pathway (Fig. 4a and Supplementary Figs. 2–8), and reveal more subtle interplay in the introduction of certain modifications. For instance, dihydrouridines introduced in the D-arm by Dus1 are seen to have a positive effect on the introduction of m1A58 in the T-arm and a negative effect on the introduction of m2G10 in the D-arm (Fig. 4a and Supplementary Fig. 2). Moreover, m22G26 has a positive effect on the introduction of the nearby m7G46 but also on the more distant m1A58 in the T-arm (Fig. 4a and Supplementary Fig. 4). Our analysis has revealed a complex interplay between different modifications in the tRNA core, with the introduction of some modifications appearing to be strongly coupled (Fig. 5b). We next set out to analyze these modification circuits in vivo.
Modification circuits in other yeast tRNAs
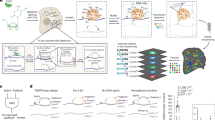
The modification circuits identified here in yeast extracts are very clear, but we wanted to rule out the possibility that these interdependences could be the consequence of monitoring the modification pathways in yeast extracts, in which numerous cellular activities are present but many others are likely missing. In addition, we wondered whether the modification circuits uncovered here on tRNAPhe are specific to this tRNA or whether they are relevant to tRNA modifications in general. To address these two points, we undertook to quantify the modification content of total yeast tRNAs prepared from different yeast strains depleted of one specific modification enzyme at a time. The absolute quantification of the different modified nucleosides was performed by liquid-chromatography coupled with tandem mass spectrometry (LC–MS/MS) and compared with the modification content of wild-type yeast cultured under the same experimental conditions. Modification abundances were quantified by isotope dilution mass spectrometry39 for the following modifications: D, Ψ, m22G, T, m5C, m7G, m2G, and m1A in wild-type and depleted strains as used for the NMR monitoring in yeast extracts, namely dus1Δ, pus4Δ, trm1Δ, trm2Δ, trm4Δ, trm8Δ, and trm11Δ (Fig. 6). Their quantification in total tRNAs (Fig. 6a, b) reflects the sum of the individual modification changes in all tRNA species and therefore alterations specific to a single tRNA are most probably averaged out and masked in these measurements. Substantial changes in the tRNA modification contents of depleted strains therefore reflect changes that must be common to several tRNA species. Apart from the expected loss of modifications corresponding to the depleted gene itself, significant alterations in the modification content of total tRNAs were measured in the pus4Δ and trm2Δ strains (Fig. 6a, b and Supplementary Fig. 9). In the pus4Δ strain, both the levels of T and m1A are much reduced, whereas in the trm2Δ strain, the level of m1A is slightly but significantly reduced (Fig. 6b). This shows that the effects we have observed with NMR in yeast extracts on tRNAPhe (namely that Ψ55 has a positive effect on the introduction of T54 and m1A58, and T54 has a positive effect on the introduction of m1A58) also occur in vivo and must be present in several tRNAs. Our measurements clearly establish the existence of a conserved modification circuit involving T54, Ψ55, and m1A58 in the T-arm of tRNAs in yeast, but the extent to which a given tRNA is responsive to this circuit is likely to be tRNA-species-dependent (see the “Discussion” section). Apart from this modification circuit in the T-arm, other more subtle interdependences detected by NMR on tRNAPhe were not observed on total tRNAs (Figs. 5 and 6a, b), which led us to quantify the modification content of specifically purified tRNAPhe prepared from these same total tRNA samples. The number of modified nucleosides in tRNAPhe from wild-type yeast is shown in Supplementary Fig. 10, while the relative abundance is shown in Fig. 6c, d and Supplementary Fig. 11). Similarly to what was observed for total tRNAs, the levels of T and m1A in the purified tRNAPhe samples confirm interdependence of modifications in the T-arm of tRNAPhe (Fig. 6c, d and Supplementary Fig. 11). It is worth noting that the effects on the T and m1A levels in the pus4Δ strain are even more pronounced for tRNAPhe than in total tRNAs, with almost no m1A detected in this strain in tRNAPhe (Fig. 6b, d). In addition, dihydrouridines introduced by Dus1 in the D-arm and m22G26 introduced by Trm1 have both a positive effect on the introduction of m1A58 in the T-arm (Fig. 6d). These effects were also observed by NMR in yeast extracts (Fig. 5). Finally, our mass spectrometry (MS) measurements suggest other small effects that were not detected by NMR in yeast extracts, such as m22G26 and m5C modifications, which both have a slight but significant positive effect on the introduction of T54 (Supplementary Fig. 11). Overall, the quantification of modifications performed here with MS demonstrates that interdependences between modifications observed by NMR in yeast extracts on tRNAPhe also occur in yeast in vivo and that some of these modification circuits are not limited to tRNAPhe but are common to several yeast tRNAs.
Quantitative analysis of nucleoside modifications in yeast tRNAs with LC–MS/MS. a Heat map depicting the relative comparison of normalized modification levels in total yeast tRNAs prepared from depleted strains (dus1Δ, pus4Δ, trm1Δ, trm2Δ, trm4Δ, trm8Δ, and trm11Δ) using the wild-type levels as reference. The nucleotides quantified by LC–MS/MS are listed on the left side of the map. The scale bar indicates the fold change in modification levels compared with the wild-type strain (increased levels shown as red, no change as white, and decreased levels as blue). b Histograms showing the relative abundance of T and m1A modifications in total yeast tRNAs prepared from the depleted strains using the wild-type levels as reference. c Heat map depicting the relative comparison of normalized modification levels in specifically purified yeast tRNAPhe prepared from the depleted strains using the wild-type levels as reference. The scale bar indicates the fold change as in panel a. d Histograms showing the relative abundance of T and m1A modifications in specifically purified yeast tRNAPhe prepared from the depleted strains using the wild-type levels as reference. In panels b and d, black dots represents individual measurements, data heights represent the mean of the biological replicates. Error bars correspond to the s.d. In all panels, significant changes compared to wild type are reported as *** for p < 0.001, ** for p < 0.01 and * for p < 0.05, n = 3. All other changes are not statistically significant. Statistical analyses of the variations compared to the wild-type strain were performed using a two-sided Student’s t-test. See also Supplementary Figs. 9 and 11 for similar analysis of all modified bases in total tRNA and purified tRNAPhe. Modifications were quantified in three independent biological replicates, except for the modifications Ψ, m7G, and m1A in the specifically purified tRNAPhe of the trm4Δ strain, for which n = 2. Source data are provided as a Source Data file
Discussion
In this study, we developed an original methodology for monitoring the introduction of modified nucleotides in tRNAs using NMR in a time-resolved fashion. Our methodology enabled us to demonstrate that the modifications in yeast tRNAPhe are introduced in a particular order and that there are complex interplays between certain modifications. In particular, modifications T54, Ψ55, and m1A58 in the T-arm are strongly interconnected in a circuit of modifications that drives the modification process along a defined route, namely Ψ55, then T54 and finally m1A58. Our study has important implications for our understanding of the regulation of tRNA modifications in yeast. These aspects are discussed below.
By monitoring the introduction of modifications in tRNAs in a time-resolved fashion with NMR, our methodology enables the identification of sequential events and therefore reveals potential regulatory circuits in the system under study. If the introduction of a certain modification is influenced by the presence of prior modifications, this dependency indeed drives a defined order in the introduction of these modifications. Only a handful of modification circuits have been reported to date, most probably because their study remains difficult, since monitoring the maturation of tRNA in real-time at a single nucleotide level is technically challenging18. Most of the well-documented examples of modification circuits occur in the ACL and involve modifications at the frequently modified positions 34 and 371,4. Although the reasons for which modification circuits exist in tRNAs are not known, it was recently proposed that the first modifications act as additional recognition elements for other modifying enzymes, which provides the mean for adding modifications with considerable variation in the ACL14. This theory is highly convincing for modifications in the ACL, but cannot account for the modification circuit identified here in the T-arm of yeast tRNAs. The three modifications involved are indeed highly conserved, with T54 and Ψ55 being present in all but the initiator tRNA in yeast (the specific case of tRNAiMet is discussed below), and the m1A58 modification being present in about two-thirds of yeast cytosolic tRNAs1. Similarly, other small effects uncovered here in tRNAPhe, for instance the dihydrouridines in the D-arm and the m22G26 modification, which have a positive influence on the introduction of the m1A58 modification in the T-arm (Fig. 5 and Supplementary Fig. 11), cannot be explained by the need for an increased modification variability in the tRNA core, since these modifications are also highly conserved in yeast tRNAs. The modification circuits uncovered here in the tRNA core of yeast tRNAs are reminiscent of the circuits that have been reported in Thermus thermophilus tRNAs40,41,42. Although different in the exact nature of the connections between modifications, both the modification circuits revealed here in yeast and those found in T. thermophilus involve conserved modifications from the tRNA core and link modifications from several tRNA regions such as the T- and D-arms42. In the case of T. thermophilus, the modification circuits influence and regulate the levels of modifications in response to changes in external temperature, a mechanism that has been linked to an adaptation of protein synthesis to temperature change43. Whether the circuits identified here in yeast are part of a regulatory mechanism for adaptation to environmental changes is a stimulating idea that deserves further studies.
It is worth noting the self-coherence of the modification circuits reported here in yeast tRNAs. For instance, the branch D ⊣ m2G10 ⊣ T54 → m1A58 is consistent, regarding the influence of the dihydrouridine modifications on the m1A58, with the direct branch D → m1A58 (Fig. 5b). Similarly, the branch Ψ55 → T54 → m1A58 is consistent with the direct branch Ψ55 → m1A58 (Fig. 5b). It should be remembered that the type of global analysis that we carried out here with NMR or MS, cannot distinguish direct from indirect effects. The effects that appear in the form of a short-branch circuit might therefore be indirect effects arising from accumulated direct effects in a long-branch circuit with identical endpoints. To discriminate direct from indirect effects is an important point that could be addressed in vitro by performing careful enzymology with purified enzymes and tRNA substrates bearing or not pre-existing modifications. Among other techniques, this approach could be implemented with NMR using different types of labeling, e.g. 15N-labeling of the tRNA substrate and a similar experimental setup as presented here, or 13C-labeling of the transferable methyl of the SAM cofactor and a monitoring of the modification process with 1H–13C correlation spectra showing the incorporation of the methyl groups into the tRNA substrate. However, whether some of the effects observed here are direct or indirect does not radically change their fundamental quality: a connection exists between two modifications.
If one compares the effects of modifications observed with NMR on tRNAPhe (Fig. 5) and those measured by MS on total tRNAs (Fig. 6a and Supplementary Fig. 9), it seems reasonable to conclude that the identity of the tRNA has an influence on the conservation and the intensity of a given modification circuit. Apart from the effects of the Ψ55 → T54 → m1A58 modification circuit in the T-arm, the other effects observed on tRNAPhe with NMR are not apparent in the MS analysis of total tRNAs. This means that these particular effects are not found in all the different tRNAs but are averaged out and therefore are not detected in the total tRNA population. The observed differences are not derived from potential discrepancies between NMR and MS analysis, since the MS measurements on purified tRNAPhe are more similar to the NMR measurements than the MS data on total tRNAs (Figs. 5 and 6 and Supplementary Figs. 9 and 11). This conclusion is corroborated by the difference in the absolute quantification of modified nucleosides performed by MS on total tRNA and purified tRNAPhe. As a comment, it is worth mentioning that the differences that exist between the NMR and MS measurements on purified tRNAPhe can have two different origins. First, they can reflect the fact that cell extracts do not fully recapture the enzymatic activities of intact cells, especially regarding protein localization (see below). Second, our approach examines the de novo synthesis of modifications on an unmodified tRNA with NMR and the steady-state levels of modifications in mature tRNAs with MS. Differences can therefore arise from the fact that the relatively weak effects on rates of modifications detected by NMR during de novo synthesis might not give rise to significant changes in the steady-state levels of modified nucleosides measured with MS. Concerning the role of tRNA identity, it has been proposed that some modifications might matter more in certain tRNA species than in others44. Our data are consistent with the extension of this idea to the connections between modifications, meaning that modification circuits may have more pronounced effects in certain tRNA species than in others. From a mechanistic point of view, this means that for a certain tRNA species, which is an intrinsically poor substrate of a given enzyme, decent modification activity is only achieved in the presence of a prior modification. Whereas for another tRNA species, an intrinsically good substrate of the same enzyme, the presence of a prior modification, although beneficial, has only a minor effect on the catalysis. In such a situation, the modification circuits would have a more pronounced effect on the first tRNA species than on the second one. In accordance with this idea, in the Ψ55–T54–m1A58 modification circuit in the T-arm, which is certainly conserved in several tRNAs (Fig. 6b), the strength of the connections most likely depends on the identity of the tRNA. The effects observed in the pus4Δ and trm2Δ strains are indeed more pronounced on tRNAPhe than on total tRNAs (Fig. 6b, d), meaning that the effects of these circuits are necessarily less pronounced on other tRNA species. It is important to mention here that this reasoning can be safely applied in this particular case, since the m1A58 and T54 modifications are the only source of m1A and T in yeast tRNAs1.
Although these extract-based NMR measurements may not accurately reflect the spatially controlled enzyme activities of intact cells, they nevertheless permit comparative assessments of global RNA modification activities and their changes upon depletion of a specific modification enzyme. In other words, even though our NMR approach might be biased by the lack of compartmentalization, by the use of a relatively high concentration of substrate tRNA (40 μM) that stretches the modification process over an artificial time span, by an intrinsic decay of enzymatic activities during incubation that is likely enzyme-dependent, and by the fact that the transcription is here dissociated from the modification process, it enables the identification of connections between modifications. In support of our methodology, most of the modification circuits identified in extracts with NMR have been corroborated by MS data on tRNAs purified from living yeast cells. The most striking difference between the NMR-derived circuits and the MS measurements concerns the strong negative influence of m2G10 on the introduction of m22G26 observed with NMR, a connection which is absent from the MS data (Figs. 5 and 6). We believe that the significant inhibition observed in NMR describes a true reduction of the Trm1 activity in the presence of m2G10, an effect that is perfectly compatible with the three-dimensional structure of yeast tRNAPhe, in which m2G10 and m22G26 stack on one another (Supplementary Fig. 1)38. However, this situation might never arise in cells as a consequence of the subcellular localization of the different enzymes. Indeed, Trm11 (responsible for m2G10) is a cytoplasmic protein whereas Trm1 (producing m22G26) is a nuclear protein, and since the m22G26 formation is insensitive to the presence of the intron, the m22G26 modification most likely occurs before the primary export of tRNAs to the cytoplasm (Supplementary Table 1)45,46,47. The m22G26 modification is thus introduced on tRNAs lacking the m2G10 modification, and in the light of our data it is tempting to speculate that Trm11 is localized in the cytoplasm to avoid it inhibiting the m22G26 modification. In another example, the subcellular location of Trm2 is unknown8, but there is indirect evidence of an m5U54-methylation activity in the nucleus, and Trm2 is therefore thought to be, at least partially, a nuclear protein48,49,50. Our data perfectly agree with a nuclear Trm2 protein. In the trm2Δ strain, the levels of m1A, introduced at position 58 by the Trm6/Trm61 nuclear complex51, are indeed slightly but significantly reduced (Fig. 6b, d). This means that the T54 modification has a slight positive effect on the introduction of m1A58, an effect that is incompatible with a situation, where tRNAs would be modified first with m1A58 in the nucleus by Trm6/Trm61 and modified by Trm2 in the cytoplasm after tRNA export.
In this study, we report a robust modification circuit in the T-arm of yeast tRNAs, with Ψ55 positively influencing the introduction of both T54 and m1A58, and T54 positively influencing the introduction of m1A58 (Figs. 5 and 6). This modification circuit was initially postulated based on the ordered modification process revealed by the time-resolved monitoring of tRNAPhe modifications in yeast extracts (Fig. 4). A survey of the tRNAPhe modification process in yeast extracts using radiolabelled nucleotides and 2D chromatography on thin-layer plates to identify modified nucleotides has been previously reported28. Several aspects of the reported kinetics are in accordance with our observations, with for instance the pseudouridine modifications being the fastest modification introduced, but the ordered modification process in the T-arm was not identified. The T54 modification is indeed introduced with no lag-phase, arguing for T54 being a modification that does not require any prior modification. Unfortunately the m1A58 modification was not detected in their experimental conditions, preventing the examination of a potential ordered modification process involving this nucleotide28. We believe it is worth emphasizing that our methodology has the advantage of providing information on modifications within their sequence context, meaning that one can differentiate, for instance, the m5C40 from the m5C49 modification, or the different pseudouridine modifications, information that is absent when tRNAs are digested down to nucleotides before analysis. Since nucleotides 54, 55, and 58 are almost absolutely conserved as U54, U55, and A58 in tRNA genes, and since these positions are often modified to T54, Ψ55, and m1A58 in the three domains of life4,52, it would be interesting to understand whether the T54–Ψ55–m1A58 modification circuit identified in yeast, is conserved in other organisms. E. coli lacks m1A58 modifications in its tRNAs, but interestingly, the enzyme catalyzing the Ψ55 modification in E. coli (TruB) prefers to bind unmodified tRNA, whereas the enzyme catalyzing the T54 modification (TrmA) binds tRNA containing Ψ55 more strongly (Ute Kothe, personal communication). This suggests that the Ψ55 and T54 modifications are likely introduced in the same order in E. coli and yeast. In T. thermophilus, complex modification circuits exist in the T-arm, including the m1A58 modification, which is stimulated by T5453; the s2T54 modification of T54, which is stimulated by m1A5854; and the s2T54 and m1A58 modifications, which are negatively regulated by Ψ5541. Although the yeast and T. thermophilus circuits have the T54 → m1A58 branch in common, the two circuits are fairly different. First, in T. thermophilus, the m1A58 modification is negatively regulated by Ψ55, whereas it is strongly favored by Ψ55 in yeast (Figs. 5 and 6). And second, the connections in T. thermophilus lead to slight changes in the modification content of the analyzed tRNAs (±10% compared to wild-type), whereas the strong influence of Ψ55 on m1A58 in yeast tRNAPhe produces large changes, with the m1A58 level in the pus4Δ strain decreasing to ~10% of the wild-type level (Fig. 6d). Altogether, even though some connections might be conserved in different species, potential modification circuits in the T-arm of tRNAs are likely organism-dependent.
Finally, it is worth mentioning that the trm6/trm61 genes, coding for the heterodimer catalyzing the m1A58 modification, are among the few genes coding for tRNA modification enzymes that are essential in yeast6. It might seem counterintuitive at first sight that the deletion of the pus4 gene, which causes a dramatic decrease in the level of m1A58 in tRNAs (Fig. 6), results in no detectable phenotype in yeast55. However, the essentiality of the m1A58 modification has been studied in detail and was shown to correspond to an increased and deleterious instability of the initiator tRNAiMet when lacking m1A5856,57. The initiator tRNAiMet is the only yeast cytoplasmic tRNA lacking both the T54 and Ψ55 modifications1. Yeast tRNAiMet has a very peculiar T-loop sequence, and contains unmodified A54 and U55, which together with other features give rise to a particular tRNA elbow structure58. Overall, initiator tRNAiMet has its own pathway of modification in the T-arm, which does not depend on the T54–Ψ55–m1A58 modification circuit identified here, and the level of m1A58 in tRNAiMet is therefore most likely unaffected in the pus4Δ and trm2Δ strains.
Overall, our work establishes NMR spectroscopy as an enlightening technique to analyze tRNA modification pathways. Our innovative methodology indeed reports on the sequential order of modification events and provides valuable information on the regulatory circuits in tRNAs. We expect that our NMR-based methodology will be applicable to investigate several aspects of tRNA maturation and RNA modifications in general, such as the dynamic adaptation of RNA modifications in response to environmental changes.
Methods
Yeast strains
Yeast strains used in this study are listed in Supplementary Table 2. The wild-type S. cerevisiae BY4741 strain and the YKO collection kanMX strains carrying deletions of the genes for modification enzymes (YML080w, YNL292w, YDR120c, YKR056w, YBL024w, YDL201w, and YOL124c) were obtained from Euroscarf and used for tRNA preparations for MS analysis.
The proteinase-deficient S. cerevisiae strain c13-ABYS-8659 was used for the preparation of yeast extracts used in NMR experiments. Chromosomal deletion of genes coding for modification enzymes (YML080w, YNL292w, YDR120c, YKR056w, YBL024w, YDL201w, and YOL124c) was carried out by homologous recombination in the c13-ABYS-86 genetic background. The kanamycin-resistance cassettes kanMX were amplified by high-fidelity PCR (Phusion, ThermoFisher) from the appropriate YKO collection kanMX strains (Euroscarf) followed by transformation of the DNA into the c13-ABYS-86 strain and selection on plates containing G418 at 300 μg/mL. All strain constructions were verified by PCR using appropriate oligonucleotides (listed in Supplementary Table 3).
tRNAPhe samples for NMR
Unmodified yeast tRNAPhe was prepared by standard in vitro transcription with T7 polymerase with unlabeled NTPs (Jena Bioscience) for unlabeled samples or 15N-labeled UTP and GTP (Eurisotop) and unlabeled ATP and CTP (Jena Bioscience) for 15N-[U/G]-labeled samples. The DNA template and T7 promotor primer were purchased from Eurogentec. The transcript was purified by ion exchange chromatography (MonoQ, GE Healthcare) under native conditions, dialyzed extensively against Na-phosphate pH 6.5 1 mM, and refolded by heating at 95 °C for 5 min and cooling down slowly at room temperature. Buffer was added to place the tRNAPhe in the NMR buffer (Na-phosphate pH 6.5 10 mM, MgCl2 10 mM), and the sample was concentrated to ~1.5–2.0 mM using Amicon 10,000 MWCO (Millipore).
The yeast 15N-labeled tRNAPhe sample produced in and purified from E. coli was prepared following previously published procedures60. Briefly, the gene coding yeast tRNAPhe was cloned in the pBSTNAV vector (Addgene ID 45801), expressed in LB for unlabeled sample preparation or 15N-labeled Spectra-9 medium (Eurisotop) for 15N-labeled sample preparation. After standard procedures of phenol extraction of soluble RNAs, yeast tRNAPhe was purified by ion exchange chromatography under native conditions, dialyzed against the NMR buffer and concentrated to ~0.5–1.0 mM.
Unlabeled matured yeast tRNAPhe was purchased from Sigma, resuspended in and dialyzed against the NMR buffer and concentrated to 0.8 mM.
Yeast extract preparation
Yeasts (c13-ABYS-86 strain) were grown in YEPD medium (1% (w/v) yeast extract, 2% (w/v) peptone, 2% (w/v) glucose) for 24 h at 30 °C and harvested by centrifugation. Pellets were stored at −80 °C until further use. Pellets were unfrozen and resuspended in their same weight of lysis buffer (Na2HPO4/KH2PO4 pH 7.0 25 mM, MgCl2 10 mM, EDTA 0.1 mM) complemented with 2 mM dithiothreitol (DTT), 1 mM phenylmethylsulphonyl fluoride (PMSF), 1 mM benzamidine, and 1 μg/mL each of leupeptin, pepstatin, antipain, and chymostatin. Cells were rapidly frozen and lysed in an Eaton pressure chamber61 at 30,000 psi in a hydraulic press. The homogenate was centrifuged at 30,000 × g for 1 h at 8 °C to remove cellular debris. The supernatant was further centrifuged at 100,000 × g for 1 h at 8 °C. The resulting supernatant (15–20 mg/mL of proteins) was quickly aliquoted and frozen in liquid nitrogen. Aliquots were stored at −80 °C until used. Yeast extracts from the depleted strains were prepared following the same procedure.
NMR spectroscopy
All in vitro NMR spectra of yeast tRNAPhe were measured at either 38 or 30 °C on Bruker AVIII-HD 600 MHz and AVIII-HD 700 MHz spectrometers (equipped with TCI 5-mm cryoprobes) with 5-mm Shigemi tubes. Imino resonances of the tRNAPhe samples were assigned using 2D jump-and-return-echo (1H,1H)-NOESY62,63, 2D (1H,15N)-BEST-TROSY30, and standard 2D (1H,15N)-HSQC experiments measured in the NMR buffer (NaH2PO4/Na2HPO4 pH 6.5 10 mM, MgCl2 10 mM) supplemented with D2O 5% (v/v). NMR spectra in yeast extracts were measured at 30 °C on Bruker AVIII-HD 700 MHz spectrometer with 5-mm Shigemi tubes. Unmodified 15N-[U/G]-labeled tRNAPhe were prepared at 40 μM in yeast extracts (final concentration of 11 mg/mL of proteins) supplemented with NaH2PO4/K2HPO4 pH 7.5 150 mM, NH4Cl 5 mM, MgCl2 5 mM, DTT 2 mM, EDTA 0.1 mM, SAM 4 mM, ATP 4 mM, NADPH 4 mM, and D2O 5% (v/v)64. This concentration of tRNAPhe (40 μM) was chosen to achieve sufficient SNR in NMR measurements while seeking to approach cellular tRNA concentrations (as a comparison, the concentration of total tRNAs have been estimated to 100–200 μM in yeast and 200–350 μM in E. coli, with typical concentrations of individual tRNAs of 2–15 μM65,66). For monitoring the maturation of tRNAPhe in yeast extract, each 2D (1H,15N)-BEST-TROSY experiment was measured with a recycling delay of 200 ms, a SW(15N) of 26 ppm, 96 increments for a total experimental time of 120 min. A reference 2D (1H,15N)-BEST-TROSY spectrum of unmodified tRNAPhe was also measured at 303 K in a buffer containing NaH2PO4/K2HPO4 pH 6.75 100 mM, MgCl2 5 mM, ATP 4 mM, NADPH 4 mM and D2O 5% (v/v)64. The data were processed using TOPSPIN 3.5 (Bruker) and analyzed with Sparky (http://www.cgl.ucsf.edu/home/sparky/).
Total tRNA samples from yeast for mass spectrometry
Total tRNA from S. cerevisiae BY4741 wild-type or mutant strains used for mass spectrometry analysis were prepared following previously published procedures67. For each strain, all cultures and tRNA preparations were performed in triplicate for statistical analysis. Briefly, the protocol was adapted as follows. Yeast cells were grown in YEPD medium at 30 °C. Yeasts were collected in logarithmic growth phase and washed with water. Pellets were resuspended in one volume of NaCl 150 mM and mixed with two volumes of water-saturated phenol. After mild shaking at room temperature for 30 min, one volume of chloroform was added and the mixture was vortexed for 15 min. The water and phenol phases were separated by centrifugation for 20 min at 8000 × g at 4 °C. RNAs were precipitated from the aqueous phase for 2 h at −20 °C by adding 2.5 volumes of cold ethanol and 0.1 volume of potassium acetate 2 M. RNAs were pelleted by centrifugation at 10,000 × g for 15 min at 4 °C. Dry pellets were resuspended in 400 μL of Tris–HCl pH 8.0 2 M and tRNAs were incubated for 90 min at 37 °C for aminoacyl-tRNA deacylation. RNAs were precipitated with 2.5 volumes of cold ethanol at −20 °C. Pellets were resuspended in 100 μL of Li/K-acetate buffer (lithium acetate 2 M, potassium acetate 0.1 M pH 5.0). After incubation at 4 °C for 20 min, most of the insoluble ribosomal RNAs were eliminated by centrifugation. Soluble tRNAs in the supernatant were recovered by precipitation with 2.5 volumes of cold ethanol. Finally, tRNAs were resuspended in Tris–HCl pH 7.5 1 mM, magnesium acetate 10 mM, and precipitated with 0.1 volume of ammonium acetate 5 M and three volumes of cold ethanol for 2 h at −20 °C. After centrifugation, tRNAs were dissolved in RNase-free water (Thermofisher). The total tRNA samples prepared with this procedure contains small contaminations of other small RNAs < 200 nts, mainly 5S and 5.8S rRNAs (both the 5S and 5.8S rRNA contain a single pseudouridine in their sequence and no other modifications). Using size exclusion chromatography (SEC), tRNAs were estimated to account for ~80% of the small RNAs in these samples.
Specific isolation of yeast tRNAPhe
Yeast tRNAPhe was isolated from ~1 µg total tRNA samples (RNAs < 200 nts) with a first step of SEC following previously published procedures39,68, and a subsequent purification using T1 Dynabeads (Thermo Fisher Scientific, Product no. 65801D) and a DNA probe specific to tRNAPhe ([Btn]AAATGGTGCGAATTCTGTGGATCGAACACAGGACCTCCAGATCTTC, Sigma-Aldrich, Munich, Germany) as previously reported69.
Digestion of tRNAs to nucleosides
Total tRNA samples (300 ng for each sample) and the purified tRNAPhe samples were digested to single nucleosides for 2 h at 37 °C with alkaline phosphatase (0.2 U, Sigma-Aldrich, St. Louis, MO, USA), Phosphodiesterase I (0.02 U, VWR, Radnor, PA, USA), and Benzonase (0.2 U) in a buffer containing Tris–HCl pH 8.0 5 mM and MgCl2 1 mM. Tetrahydrouridine (THU, 0.5 µg from Merck), butylated hydroxytoluene (BHT, 1 µM), and Pentostatin (0.1 µg) were also added to protect modifications. Afterwards samples were filtered through multi-well plates (Pall Corporation, 10 kDa MWCO) at 4 °C for 30 min at 3000 × g to remove digestive enzymes. Stable isotope-labeled internal standard (SILIS, 0.1 volume of 10X solution) from yeast was added for absolute quantification39.
Mass spectrometry
For quantification, an Agilent 1290 Infinity II equipped with a DAD combined with an Agilent Technologies G6470A Triple Quad system and electro-spray ionization (ESI-MS, Agilent Jetstream) was used. Operating parameters were as follows: positive ion mode, skimmer voltage 15 V, cell accelerator voltage 5 V, N2 gas temperature 230 °C, and N2 gas flow 6 L/min, sheath gas (N2) temperature 400 °C with a flow of 12 L/min, capillary voltage of 2500 V, nozzle voltage of 0 V, and the Nebulizer at 40 psi. The instrument was operated in dynamic MRM mode (individual mass spectrometric parameters for the nucleosides are given in Supplementary Table 4. The mobile phases were: A as 5 mM NH4OAc (≥99%, HiPerSolv CHROMANORM®, VWR) aqueous buffer, brought to pH = 5.6 with glacial acetic acid (≥99%, HiPerSolv CHROMANORM®, VWR) and B as pure acetonitrile (Roth, LC–MS grade, purity ≥ 99.95%). A Synergi Fusion-RP column (Phenomenex®, Torrance, CA, USA; Synergi® 2.5 µm Fusion-RP 100 Å, 150 × 2.0 mm) at 35 °C and a flow rate of 0.35 mL/min was used. The gradient began with 100% A for 1 min, increased to 10% B by 5 min, and to 40% B by 7 min. The column was flushed with 40% B for 1 min and returned to starting conditions to 100% A by 8.5 min followed by re-equilibration at 100% A for 2.5 additional minutes.
For calibration, synthetic nucleosides were weighed and dissolved to a stock concentration of 1–10 mM. Calibration solutions ranging from 0.25 to 100 pmol for each canonical nucleoside and from 0.0125 to 5 pmol for each modified nucleoside were prepared in water (D, Ψ = 0.025–10 pmol). The calibration solutions were mixed with the yeast SILIS and analyzed by LC–MS/MS. The value of each integrated peak area of the nucleoside was divided through the respective SILIS area. The linear regression for each nucleoside’s normalized signal/concentration plot gives the relative response factor for nucleosides (rRFN)39. The data were analyzed by the Quantitative and MassHunter Software from Agilent. Finally, the absolute amounts of the modifications were referenced to the absolute amounts of summed canonical nucleosides. The number of modifications per tRNAPhe was calculated by determining the amount of injected tRNA from the signal of canonicals and the number of canonicals from the tRNAPhe sequence. Statistical analyses of the variations compared to the wild-type strain were performed using a two-sided Student’s t-test.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
A reporting summary for this Article is available as a Supplementary Information file. The source data underlying Fig. 6 and Supplementary Figs. 9–11 are provided as a Source Data file. All data is available from the corresponding author upon reasonable request.
References
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Helm, M. & Alfonzo, J. D. Posttranscriptional RNA Modifications: playing metabolic games in a cell's chemical Legoland. Chem. Biol. 21, 174–185 (2014).
Jackman, J. E. & Alfonzo, J. D. Transfer RNA modifications: nature's combinatorial chemistry playground. Wiley Interdiscip. Rev. RNA 4, 35–48 (2013).
Machnicka, M. A., Olchowik, A., Grosjean, H. & Bujnicki, J. M. Distribution and frequencies of post-transcriptional modifications in tRNAs. RNA Biol. 11, 1619–1629 (2014).
El Yacoubi, B., Bailly, M. & de Crécy-Lagard, V. Biosynthesis and function of posttranscriptional modifications of transfer RNAs. Annu. Rev. Genet. 46, 69–95 (2012).
Phizicky, E. M. & Hopper, A. K. tRNA biology charges to the front. Genes Dev. 24, 1832–1860 (2010).
Barraud, P. & Tisné, C. To be or not to be modified: miscellaneous aspects influencing nucleotide modifications in tRNAs. IUBMB Life https://doi.org/10.1002/iub.2041 (2019).
Hopper, A. K. Transfer RNA post-transcriptional processing, turnover, and subcellular dynamics in the yeast Saccharomyces cerevisiae. Genetics 194, 43–67 (2013).
Raina, M. & Ibba, M. tRNAs as regulators of biological processes. Front. Genet. 5, 171 (2014).
Oberbauer, V. & Schaefer, M. R. tRNA-derived small RNAs: biogenesis, modification, function and potential impact on human disease development. Genes 9, 607 (2018).
Agris, P. F., Vendeix, F. A. P. & Graham, W. D. tRNA's wobble decoding of the genome: 40 years of modification. J. Mol. Biol. 366, 1–13 (2007).
Grosjean, H., de Crécy-Lagard, V. & Marck, C. Deciphering synonymous codons in the three domains of life: co-evolution with specific tRNA modification enzymes. FEBS Lett. 584, 252–264 (2010).
Lorenz, C., Lünse, C. E. & Mörl, M. tRNA modifications: impact on structure and thermal adaptation. Biomolecules 7, 35 (2017).
Han, L. & Phizicky, E. M. A rationale for tRNA modification circuits in the anticodon loop. RNA 24, 1277–1284 (2018).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Chan, C. T. Y. et al. A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet. 6, e1001247 (2010).
Chan, C., Pham, P., Dedon, P. C. & Begley, T. J. Lifestyle modifications: coordinating the tRNA epitranscriptome with codon bias to adapt translation during stress responses. Genome Biol. 19, 228 (2018).
Helm, M. & Motorin, Y. Detecting RNA modifications in the epitranscriptome: predict and validate. Nat. Rev. Genet. 18, 275–291 (2017).
Theillet, F.-X. et al. Cell signaling, post-translational protein modifications and NMR spectroscopy. J. Biomol. Nucl. Magn. Reson. 54, 217–236 (2012).
Smith, M. J. et al. Real-time NMR monitoring of biological activities in complex physiological environments. Curr. Opin. Struct. Biol. 32, 39–47 (2015).
Selenko, P. et al. In situ observation of protein phosphorylation by high-resolution NMR spectroscopy. Nat. Struct. Mol. Biol. 15, 321–329 (2008).
Theillet, F.-X. et al. Site-specific mapping and time-resolved monitoring of lysine methylation by high-resolution NMR spectroscopy. J. Am. Chem. Soc. 134, 7616–7619 (2012).
Theillet, F.-X. et al. Site-specific NMR mapping and time-resolved monitoring of serine and threonine phosphorylation in reconstituted kinase reactions and mammalian cell extracts. Nat. Protoc. 8, 1416–1432 (2013).
Amata, I. et al. Multi-phosphorylation of the intrinsically disordered unique domain of c-Src studied by in-cell and real-time NMR spectroscopy. Chembiochem 14, 1820–1827 (2013).
Mylona, A. et al. Opposing effects of Elk-1 multisite phosphorylation shape its response to ERK activation. Science 354, 233–237 (2016).
Cordier, F. et al. Ordered phosphorylation events in two independent cascades of the PTEN C-tail revealed by NMR. J. Am. Chem. Soc. 134, 20533–20543 (2012).
Liokatis, S. et al. Phosphorylation of histone H3 Ser10 establishes a hierarchy for subsequent intramolecular modification events. Nat. Struct. Mol. Biol. 19, 819–823 (2012).
Jiang, H. Q., Motorin, Y., Jin, Y. X. & Grosjean, H. Pleiotropic effects of intron removal on base modification pattern of yeast tRNAPhe: an in vitro study. Nucleic Acids Res. 25, 2694–2701 (1997).
Fürtig, B., Richter, C., Wöhnert, J. & Schwalbe, H. NMR spectroscopy of RNA. Chembiochem 4, 936–962 (2003).
Farjon, J. et al. Longitudinal-relaxation-enhanced NMR experiments for the study of nucleic acids in solution. J. Am. Chem. Soc. 131, 8571–8577 (2009).
Motorin, Y. & Helm, M. RNA nucleotide methylation. Wiley Interdiscip. Rev. RNA 2, 611–631 (2011).
Lombard, M. & Hamdane, D. Flavin-dependent epitranscriptomic world. Arch. Biochem. Biophys. 632, 28–40 (2017).
Hänsel, R. et al. Evaluation of parameters critical for observing nucleic acids inside living Xenopus laevis oocytes by in-cell NMR spectroscopy. J. Am. Chem. Soc. 131, 15761–15768 (2009).
Meinnel, T., Mechulam, Y. & Fayat, G. Fast purification of a functional elongator tRNAmet expressed from a synthetic gene in vivo. Nucleic Acids Res. 16, 8095–8096 (1988).
Tisné, C., Rigourd, M., Marquet, R., Ehresmann, C. & Dardel, F. NMR and biochemical characterization of recombinant human tRNA(Lys)3 expressed in Escherichia coli: identification of posttranscriptional nucleotide modifications required for efficient initiation of HIV-1 reverse transcription. RNA 6, 1403–1412 (2000).
Heerschap, A. et al. Imino-proton resonances of yeast tRNAPhe studied by two-dimensional nuclear Overhauser enhancement spectroscopy. Eur. J. Biochem. 149, 649–655 (1985).
Roy, S. & Redfield, A. G. Assignment of imino proton spectra of yeast phenylalanine transfer ribonucleic acid. Biochemistry 22, 1386–1390 (1983).
Shi, H. & Moore, P. B. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: a classic structure revisited. RNA 6, 1091–1105 (2000).
Borland, K. et al. Production and application of stable isotope-labeled Internal Standards for RNA modification analysis. Genes 10, 26 (2019).
Tomikawa, C., Yokogawa, T., Kanai, T. & Hori, H. N7-Methylguanine at position 46 (m7G46) in tRNA from Thermus thermophilus is required for cell viability at high temperatures through a tRNA modification network. Nucleic Acids Res. 38, 942–957 (2010).
Ishida, K. et al. Pseudouridine at position 55 in tRNA controls the contents of other modified nucleotides for low-temperature adaptation in the extreme-thermophilic eubacterium Thermus thermophilus. Nucleic Acids Res. 39, 2304–2318 (2011).
Hori, H. Methylated nucleosides in tRNA and tRNA methyltransferases. Front. Genet. 5, 144 (2014).
Hori, H. et al. Transfer RNA modification enzymes from thermophiles and their modified nucleosides in tRNA. Microorganisms 6, 110 (2018).
Phizicky, E. M. & Alfonzo, J. D. Do all modifications benefit all tRNAs? FEBS Lett. 584, 265–271 (2010).
Purushothaman, S. K., Bujnicki, J. M., Grosjean, H. & Lapeyre, B. Trm11p and Trm112p are both required for the formation of 2-methylguanosine at position 10 in yeast tRNA. Mol. Cell Biol. 25, 4359–4370 (2005).
Rose, A. M., Joyce, P. B., Hopper, A. K. & Martin, N. C. Separate information required for nuclear and subnuclear localization: additional complexity in localizing an enzyme shared by mitochondria and nuclei. Mol. Cell Biol. 12, 5652–5658 (1992).
Grosjean, H., Szweykowska-Kulinska, Z., Motorin, Y., Fasiolo, F. & Simos, G. Intron-dependent enzymatic formation of modified nucleosides in eukaryotic tRNAs: a review. Biochimie 79, 293–302 (1997).
Knapp, G., Beckmann, J. S., Johnson, P. F., Fuhrman, S. A. & Abelson, J. Transcription and processing of intervening sequences in yeast tRNA genes. Cell 14, 221–236 (1978).
Hopper, A. K., Furukawa, A. H., Pham, H. D. & Martin, N. C. Defects in modification of cytoplasmic and mitochondrial transfer RNAs are caused by single nuclear mutations. Cell 28, 543–550 (1982).
Nordlund, M. E., Johansson, J. O., von Pawel-Rammingen, U. & Byström, A. S. Identification of the TRM2 gene encoding the tRNA(m5U54)methyltransferase of Saccharomyces cerevisiae. RNA 6, 844–860 (2000).
Macari, F. et al. TRM6/61 connects PKCα with translational control through tRNAi(Met) stabilization: impact on tumorigenesis. Oncogene 35, 1785–1796 (2016).
Jühling, F. et al. tRNAdb 2009: compilation of tRNA sequences and tRNA genes. Nucleic Acids Res. 37, D159–D162 (2009).
Yamagami, R. et al. Folate-/FAD-dependent tRNA methyltransferase from Thermus thermophilus regulates other modifications in tRNA at low temperatures. Genes Cells 21, 740–754 (2016).
Shigi, N. et al. Temperature-dependent biosynthesis of 2-thioribothymidine of Thermus thermophilus tRNA. J. Biol. Chem. 281, 2104–2113 (2006).
Becker, H. F., Motorin, Y., Planta, R. J. & Grosjean, H. The yeast gene YNL292w encodes a pseudouridine synthase (Pus4) catalyzing the formation of psi55 in both mitochondrial and cytoplasmic tRNAs. Nucleic Acids Res. 25, 4493–4499 (1997).
Anderson, J. et al. The essential Gcd10p-Gcd14p nuclear complex is required for 1-methyladenosine modification and maturation of initiator methionyl-tRNA. Genes Dev. 12, 3650–3662 (1998).
Kadaba, S. et al. Nuclear surveillance and degradation of hypomodified initiator tRNAMet in S. cerevisiae. Genes Dev. 18, 1227–1240 (2004).
Basavappa, R. & Sigler, P. B. The 3 A crystal structure of yeast initiator tRNA: functional implications in initiator/elongator discrimination. EMBO J. 10, 3105–3111 (1991).
Heinemeyer, W., Kleinschmidt, J. A., Saidowsky, J., Escher, C. & Wolf, D. H. Proteinase yscE, the yeast proteasome/multicatalytic-multifunctional proteinase: mutants unravel its function in stress induced proteolysis and uncover its necessity for cell survival. EMBO J. 10, 555–562 (1991).
Dégut, C., Monod, A., Brachet, F., Crépin, T. & Tisné, C. In vitro/in vivo production of tRNA for X-ray studies. Methods Mol. Biol. 1320, 37–57 (2016).
Eaton, N. R. New press for disruption of microorganisms. J. Bacteriol. 83, 1359–1360 (1962).
Plateau, P. & Gueron, M. Exchangeable proton NMR without base-line distorsion, using new strong-pulse sequences. J. Am. Chem. Soc. 104, 7310–7311 (1982).
Sklenar, V. & Bax, A. Spin-echo water suppression for the generation of pure-phase two-dimensional NMR spectra. J. Magn. Reson. 74, 469–479 (1987).
Freund, J. & Kalbitzer, H. R. Physiological buffers for NMR spectroscopy. J. Biomol. Nucl. Magn. Reson. 5, 321–322 (1995).
Waldron, C. & Lacroute, F. Effect of growth rate on the amounts of ribosomal and transfer ribonucleic acids in yeast. J. Bacteriol. 122, 855–865 (1975).
Dong, H., Nilsson, L. & Kurland, C. G. Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J. Mol. Biol. 260, 649–663 (1996).
de Crécy-Lagard, V. et al. Biosynthesis of wyosine derivatives in tRNA: an ancient and highly diverse pathway in Archaea. Mol. Biol. Evol. 27, 2062–2077 (2010).
Chionh, Y. H. et al. A multidimensional platform for the purification of non-coding RNA species. Nucleic Acids Res. 41, e168 (2013).
Hauenschild, R. et al. The reverse transcription signature of N-1-methyladenosine in RNA-Seq is sequence dependent. Nucleic Acids Res. 43, 9950–9964 (2015).
Acknowledgements
The authors are grateful to Henri Grosjean for stimulating discussions about RNA modifications, Valérie Heurgué-Hamard (IBPC), Josette Banroques (IBPC), Simon Lebaron (CBI), and Sylvie Auxilien (I2BC) for yeast strains, protocols and helpful discussions about yeast handling, Bruno Sargueil for guidance regarding the Eaton press implementation, and Jacqueline Plumbridge (IBPC) for careful reading of the manuscript. This work was supported by grants from the CNRS, the ANR NMR-VitAmin (ANR-14-CE09-0012), the Labex DYNAMO (ANR-11-LABX-0011), the Equipex Cacsice (ANR-11-EQPX-0008) and the SESAME Île-de-France. P.B. and C.T. also acknowledge access to the NMR infrastructure of the Paris Descartes University that is supported by grants from the Région Île-de-France, the European Union (FEDER) and the Paris Descartes University. M.H. and S.K. are grateful for funding from the Deutsche Forschungsgemeinschaft (KE1943/3-1 and KE1943/4-1).
Author information
Authors and Affiliations
Contributions
C.T. conceived the initial project; P.B. and C.T. refined the project and designed the experiments; P.B. and M.C. prepared tRNA samples for NMR studies; P.B., A.G., and M.C. performed gene deletions in yeast and prepared yeast extracts for NMR studies; P.B. measured and analyzed NMR spectra; A.G. prepared total tRNA samples for MS; M.H. isolated tRNAPhe and digested tRNAs for MS; M.H. and S.K. measured and analyzed MS data; P.B. wrote the manuscript; all authors discussed the results, critically reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barraud, P., Gato, A., Heiss, M. et al. Time-resolved NMR monitoring of tRNA maturation. Nat Commun 10, 3373 (2019). https://doi.org/10.1038/s41467-019-11356-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-11356-w
This article is cited by
-
Quantitative analysis of tRNA abundance and modifications by nanopore RNA sequencing
Nature Biotechnology (2024)
-
Cell culture NAIL-MS allows insight into human tRNA and rRNA modification dynamics in vivo
Nature Communications (2021)
-
Metabolomics and health: from nutritional crops and plant-based pharmaceuticals to profiling of human biofluids
Cellular and Molecular Life Sciences (2021)
-
The occurrence order and cross-talk of different tRNA modifications
Science China Life Sciences (2021)
-
1H, 15N chemical shift assignments of the imino groups of yeast tRNAPhe: influence of the post-transcriptional modifications
Biomolecular NMR Assignments (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.