Abstract
Cryo-electron microscopy can determine the structure of biological matter in vitrified liquids. However, structure alone is insufficient to understand the function of native and engineered biomolecules. So far, their mechanical properties have mainly been probed at room temperature using tens of pico-newton forces with a resolution limited by thermal fluctuations. Here we combine force spectroscopy and computer simulations in cryogenic conditions to quantify adhesion and intra-molecular properties of spray-deposited single-strand DNA oligomers on Au(111). Sub-nanometer resolution images reveal folding conformations confirmed by simulations. Lifting shows a decay of the measured stiffness with sharp dips every 0.2–0.3 nm associated with the sequential peeling and detachment of single nucleotides. A stiffness of 30–35 N m−1 per stretched repeat unit is deduced in the nano-newton range. This combined study suggests how to better control cryo-force spectroscopy of adsorbed heterogeneous (bio)polymer and to potentially enable single-base recognition in DNA strands only few nanometers long.
Similar content being viewed by others
Introduction
Nucleic acids (NA)1 are among the most studied biomolecules nowadays due to their biological relevance, but also their nanodevice applications or computing2,3,4. Control over nucleotide sequences as well as knowledge of their folding properties has enabled the rational design of highly elaborate two- and three-dimensional DNA structures, the so-called “DNA origami”5 programmed by Watson–Crick complementarity6. These remarkable advances have been made possible by the accurate determination of nucleotide characteristics beforehand, using noninvasive single-molecule manipulation techniques, such as optical tweezers7,8,9 or magnetic tweezers10,11. Beside these approaches, force spectroscopy based on atomic force microscopy (AFM) also allows direct measurements of mechanical, adhesion12, and tribological properties13, as well as visualizing self-assembly processes. So far, such force spectroscopic experiments on biomolecules have been conducted under ambient conditions in solutions, mostly up to few tens of pico-Newton tensile loads. Mechanical properties are then dominated by thermal fluctuations and folding/unfolding of soft parts14,15,16,17. Only few AFM studies on long polymers strongly bound at both ends reached the nN force level where thermal fluctuations are largely suppressed18. To the best of our knowledge, no features attributable to sub-nanometer structural details have been observed in force versus extension curves recorded under ambient conditions.
Imaging of DNA has also been a long-term challenge. Although numerous groups successfully visualized DNA branches with impressive spatial resolutions in solution under ambient conditions19,20,21,22, the highest accuracy has been reached using scanning tunneling microscopy (STM)/AFM imaging at cryogenic temperatures enabling reduced contamination23,24,25,26. These experiments further required efficient deposition techniques to successfully transfer the macromolecules from solution onto a substrate while maintaining UHV cleanliness standards27,28,29. However, the characterization of adsorbed biomolecules at the sub-nm level, specifically of DNA under such conditions28, still remains rather unexplored. Notably, the recent advances in frequency modulation AFM30 under cryogenic conditions have pushed spatial resolution of adsorbed molecules to the single-bond level31, while force spectroscopy enables complex manipulations of single molecules at surfaces32,33,34,35,36.
Here, we demonstrate that dynamic AFM-based force spectroscopy in cryogenic conditions (5 K) is a promising method for characterizing the mechanics of single-strand DNA (ssDNA) 20-cytosine oligomers down to the sub-nm level. Similar to the advent of cryo-electron microscopy for structure characterization of biosystems37, further investigations along this line could open avenues toward the integration of DNA into solid nanodevices through biomechanical studies at this level of precision.
Results
Real-space imaging of spray-deposited ssDNA
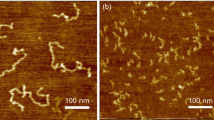
20-Cytosine ssDNA oligomers were electrospray-deposited at room temperature on the precleaned Au(111) kept in ultrahigh vacuum (UHV). The surface was then annealed step by step up to a maximum temperature Tmax of 500 K. After each step, the resulting structures were subsequently imaged at 5 K using constant-current STM. As shown in Fig. 1a, the surface morphology evolves from large aggregates of several nanometers to 4-nm-long isolated oligomers. Clusters of various dimensions are observed by STMsimilar to previous results on ssDNA/Cu(111) deposited using a pulsed-injection technique25,26. These structures are too large (Fig. 1b) compared to the expected size of a single dehydrated ssDNA 20-mer (Fig. 1c). Before annealing, we therefore suppose that a small amount of solvent molecules might also surround single oligomers.
ssDNA morphologies as a function of Au(111) annealing temperature. a Overview of STM images of spray-deposited ssDNA on Au(111) at room temperature, after annealing at 340, 440, and above 500 K, respectively. b STM image of hydrated ssDNA after spray-deposition at room temperature. c Top view of a representative ssDNA structure on Au(111) obtained from MD simulations performed in vacuum at 400 K (see stage 4 in Supplementary Note 3). d STM image of a dehydrated single 20-cytosine ssDNA oligomer after 440-K annealing and the corresponding high- resolution constant-height AFM image, both acquired with a CO-terminated tip. The structural model of c is superimposed on the STM image. e STM image of self-assembled dehydrated ssDNA oligomers after 500-K annealing and the corresponding AFM image. All STM images were recorded at 5 K with I = 2 pA and V = −1.3 V. Properly scaled top views of the representative ssDNA structure are superimposed on b, d, e as a guide for the eye
To promote solvent desorption from the gold surface, we step-by-step annealed the surface that we later imaged by STM after each step (Fig. 1a). This results in a decreased apparent size of the “hydrated ssDNA clusters” (T ≈ 340 K) as water molecules desorb from the surface. At T ≈ 440 K, only “dehydrated ssDNA oligomers” are observed by STM with a length of about 4 nm (Fig. 1d). Their overall size also corresponds to the structure of a folded oligomer adsorbed on Au(111), as systematically predicted by our simulations performed under different adsorption conditions (see Supplementary Note 5) and superimposed top views in Fig. 1b, d, e. When Tmax ≥ 500 K, these dehydrated oligomers coalesce into several nanometer-long structures (Fig. 1e). Sub-nm contrast could be obtained along individual dehydrated oligomers and their assembly not only by STM, but also by using constant-height AFM with CO-terminated tips31 (Fig. 1d, e). In spite of the potential resolution below the molecular level, the determination of the overall ssDNA conformation is rather difficult.
Molecular dynamics simulations of ssDNA adsorption
Instead of simulating the complex electrospray-deposition processes which start and end up in charge-neutral species (see Supplementary Note 1 and Supplementary Fig. 1), in all our simulations, we considered a single ssDNA oligomer together with charge-compensating counterions. For a comparison with Fig. 1b, a 20-cytosine ssDNA oligomer generated from the canonical B-form1 (see Supplementary Note 2 and Supplementary Fig. 2) together with 19 Na+ ions was inserted into a water droplet, equilibrated, and then allowed to adsorb onto an unreconstructed Au(111) surface at room temperature (see Supplementary Note 3). We first used the ssDNA conformation fully embedded in water (Supplementary Note 4 and Supplementary Fig. 4) that we relaxed only considering the first three surrounding hydration layers, i.e., within ~1 nm around the molecule corresponding to 1332 water molecules (see Supplementary Fig. 5). Within the first 10 ns of the simulation, the droplet size decreases due to the surface tension causing a considerable folding of the ssDNA chain into a compact structure of only 3-nm diameter (see Supplementary Fig. 5). The hydration layer and the ssDNA folded structure then remain stable during the rest of the 100-ns-long simulation. Note that much less folding was observed if the chain was completely immersed in water, having a total length of 6.4 nm (see Supplementary Fig. 4).
The droplet so prepared was placed 2 nm above the surface and let free to adsorb (Fig. 2a). At the first stage of the adsorption, a meniscus is formed between the hydration layer and the gold surface, whereas, after the 100-ns-long simulation, the ssDNA adsorbs folded directly on the gold surface, but is elsewhere surrounded by its hydration layer. Compared to the free droplet conformation, the ssDNA structure thus obtained is longer, which is in agreement with the STM images of hydrated oligomers on Au(111) (Fig. 1b). To test the effects of dehydration and thermal annealing on the final adsorption configuration, starting from the adsorbed hydrated ssDNA shown in Fig. 2a, we simulated (i) partial water evaporation by annealing at 450 K (stage 3a in Supplementary Fig. 6), (ii) the effect of removing all water molecules (stage 3b in Supplementary Fig. 7), and (iii) by simulating the adsorption of a single DNA strand at 400 K without any water molecules (stage 4 in Fig. 1c and Supplementary Fig. 7). All adsorption simulations, including stage 3 (hydrated ssDNA at 300 K) led to very similar folded adsorbed conformations ~4 nm in size, in good agreement with the prevalent experimentally observed structures after annealing at 440 K, as shown in Fig. 1.
Molecular dynamics simulations of adsorption and diffusion on Au(111). a Side- and top views of a small water droplet containing one ssDNA oligomer getting adsorbed on the gold surface. Water is represented using a transparent surface. b Five-hundred nanosecond MD simulation of two oligomers assembled by diffusion at 500 K (stage 5 of the simulation protocol—see Supplementary Notes 3 and 6). At 500 K, both oligomers start diffusing, thus favoring self-assembly assisted by intermolecular interactions
The systematic nature of the findings can be traced back to the direct ssDNA/Au(111) contact already established at 300 K before dehydration. Concurrently with this direct contact, the interaction of cytosine bases with gold also induces a systematic flattening of the ssDNA structure. Indeed, most bases lie nearly parallel to the surface, similar to optimum adsorption structures for single-nucleotide bases on Au(111) computed in vacuum using a nonlocal van der Waals density functional38, and very recently by MD simulations in vacuum and in water at 300 K39.
The effect of thermally assisted diffusion of ssDNA on Au(111) has been investigated with 500-ns-long MD simulations of two adsorbed ssDNA at temperatures of 400 and 500 K (see stage 5 in Supplementary Note 3). Diffusion and coalescence of the ssDNA oligomers is observed only at T = 500 K (see Fig. 2b and Supplementary Note 6) which is in agreement with the experimental data. The oligomers preserve their folded adsorption characteristics during diffusion without showing significant flattening of the structure or unfolding. The side-by-side alignment of the molecules is again consistent with the STM/AFM images of Fig. 1c. The relatively high corrugation of the final MD-simulated structures confirms the difficulty to capture the ssDNA conformation from constant-height AFM images with CO-terminated tips (Fig. 1e).
ssDNA cryo-force spectroscopy
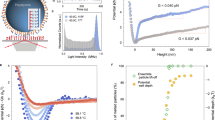
To investigate the mechanical properties of ssDNA adsorbed on Au(111), we have attempted to lift single oligomers from the surface (Fig. 1c, d). We used the protocol introduced in refs. 33,40 to pull off single polyfluorene chains with the AFM tip. Experimentally, the tip was first gently indented into Au(111) to sharpen its apex, and then approached to one end of a selected oligomer until the tunneling current suddenly increased, thus indicating a jump to contact. A retraction curve was then recorded slightly beyond the position where the It and Δf dropped to their noise levels (see Supplementary Fig. 7). The yellow dots shown in the inset of Fig. 3b was the point at which the retraction process was initiated at a constant speed of v = 22 pm s−1. The effective stiffness k ≈ 2k0Δf/f0 (k0 being the deflection sensor stiffness and f0 its resonance frequency) decreases progressively from 23 to about 5 N m−1 as the tip–sample separation Z increases. This variation is interrupted by narrow dips observed every 0.2–0.3 nm followed by an abrupt drop to zero when the tip is approximately at 1.4 nm far from the contact point, well below the length of one ssDNA oligomer in its adsorbed conformation (4 nm in Fig. 2). The premature detachment of the ssDNA from the tip (see Supplementary Notes 7 and 8) is confirmed by comparing STM images before and after the lifting experiment showing the entire oligomer still on the surface (inset Fig. 3b). Experimentally, only partial lifting of single ssDNA oligomers either self-assembled or individually adsorbed (see Supplementary Fig. 9) could be achieved. Thus, it appears that the folded ssDNA conformation and its strong adhesion on gold are the limiting factors of the experimental lifting. Indeed, desorbing a single-folded oligomer with the AFM tip requires not only peeling off the structure from the gold surface, but also unfolding part of the backbone prior to detachment. Each peculiarity of the folding configuration can thus cause an abrupt increase of the required force to lift the molecule and the rupture of the tip–molecule bond.
Mechanical response while lifting a ssDNA oligomer from Au(111). The oligomer is attached at one end to the AFM tip and pulled vertically at constant velocity. a Schematic of the lifting simulations. b Experimental retraction trace, k(Z) ∝ Δf(Z) recorded at 4.8 K. c Force–distance curve F(Z) obtained from MD simulations assuming ktip = 35 N m−1 and d the corresponding computed stiffness k(Z). The blue area shows the Z-range accessed by the experiment. The red curve shows the fit to k maxima according to Eq. (1) with k1 = 32.6 ± 3.9 N m−1, kpin = 18.3 ± 3.3 N m−1, and b cosθ = 0.64 ± 0.05. e Side views of the ssDNA oligomer after lifting 0, 10, and 17 nucleotides revealing the nearly straight configuration of the detached segment
Steered MD simulations41,42 have been run to shed light on the experimental results (see Methods and Supplementary Note 3). A representative theoretical retraction trace is shown in Fig. 3d. To do that, a virtual “tip atom” was connected to the P atom of the backbone between the first two nucleotides by a spring of stiffness ktip representing the tip–molecule bond and was pulled up at a constant speed of 0.1 ms−1 at 5 K (Fig. 3a). From the recorded variations of the noise-averaged normal force 〈F(Z)〉 (Fig. 3c), the effective stiffness was extracted as k = d 〈F(Z)〉/dZ (Fig. 3d), Z being the distance between the tip atom and the pulled P atom at t = 0. In the simulations, the whole ssDNA oligomer detaches from the substrate when the tip has been retracted up to Zoff = 11.8 nm, which is slightly less than its fully stretched length (see Supplementary Fig. 8). In spite of the discrepancy with the experimental value of Zoff, the measured maximum k values exhibit a similar trend in the common Z-range (blue area in Fig. 3d). Not only the stiffness k decreases from comparable initial values of ~15–25 N m−1, but pronounced dips (coinciding with abrupt force drops in the simulations) also appear at repeat distances of about 0.2–0.25 nm. Careful observation of the configurations adopted by the ssDNA atoms during simulated pulling reveals that the repeat distances reflect intermediate stages (peeling, lifting, and detachment) in the successive lifting of cytosine bases. These events are also accompanied by stick-slip-like sliding of the adjacent base over the Au(111) surface (0.28-nm lattice spacing) as well as irregular unfolding of the backbone. Details of such dynamics can be better visualized in Supplementary Movies 1 and 2. Note that the first step of the lifting process involves correlated base detachments and intricate unfolding, which require increased lifting forces. This increase might explain the premature detachment of ssDNA from the tip apex observed experimentally. Similar observations have been reported at room temperature in solution for grafted polymers12 and ssDNA adsorbed on carbon nanotubes17.
It is also remarkable that the computed k variations as a function of Z are very different from those previously reported for polyfluorene chains from Au(111) lifted with the same method33. There, the k maxima (~0.4 N m−1) were constant during retraction. The process ended at a distance corresponding to the number of monomers initially identified by STM on the surface. In that case (and also for graphene nanoribbons34), the much stiffer repeat units weakly adhere to the substrate, allowing nearly frictionless sliding and a complete chain detachment. The oligomer backbone is more flexible and most bases are strongly bonded to the gold surface. As shown in our MD simulation (snapshot of Fig. 3e and Supplementary Movies 1 and 2), the cytosine bases that remain adsorbed do not slide during lifting33,34. As a result, the lifted segment between the tip and the sample becomes essentially straight and inclined as the tip–sample separation increases. This sequential base detachment, similar to peeling off an adhesive tape, also reflects the strong adhesion of adsorbed cytosine bases. Interestingly, “infinite” as opposed to negligible friction was already detected for long ssDNA chains in solution at room temperature when adsorbed on gold13 and graphite15, respectively. Easy sliding of short ssDNA oligomers on graphene under cryogenic conditions was recently predicted by steered MD simulations42.
The gradual reduction of the k maxima arises because the stiffness of the lifted segment decreases as it becomes longer. Focusing on the most pronounced k maxima achieved on the longest nearly linear parts of F(Z), we assume local mechanical equilibrium in the springs-in-series model applied earlier to polyfluorene chains and to unzipped dsDNA hairpins. Including the stiffnesses ktip and kpin of the segment ends anchored to the tip and to the adsorbed part of the oligomer (Fig. 3e and Supplementary Fig. 9), the envelope of k maxima is expected to satisfy
with
k1 being the stiffness per repeat distance b in fully stretched ssDNA and n = int [Z/(b cos θ)] is the number of nucleotides detached from the substrate in the range where the lifted segments are straight and inclined by θ (almost 18° between n = 8 and 18 according to Fig. 3e and the corresponding Supplementary Movies 1 and 2). The resulting fit (red curve) is superimposed on the computed dF/dZ trace in Fig. 3d obtained for ktip = 35 N m−1. The resulting parameters are k1 = 32.6 ± 3.9 and kpin = 18.3 ± 3.3 N m−1. This confirms that ssDNA is strongly adsorbed and that fully stretched ssDNA is much more compliant than polyfluorene33, presumably because the stretched ssDNA backbone stiffness is dominated by bond angle bending. As a consequence, kDNA = k1/19 = 1.7 N m−1 would be the stiffness of the fully stretched 20-cytosine oligomer with one base at each end subject to an average tension of ~2 nN. This load is one to two orders of magnitude larger than the maximum values attained in typical room-temperature investigations of ssDNA12,14,15,16,17. We obtained only slightly different results and fit parameters from independent simulations assuming higher and lower ktip values (see Supplementary Fig. 10). In particular, Zoff, k1, b, and the average tension in the fitting range did not change appreciably.
The present work relies on closely matched scanning tunneling and force measurements, and computer simulations. The first part addresses the adsorption and self-assembly of single-strand DNA cytosine oligomers spray-deposited on Au(111) at the sub-nanometer level. Both theory and experiment showed folded ssDNA conformations arising from the first step of the adsorption. Furthermore, the unfolding of the ssDNA on the surface is not possible upon annealing. In the second part, the mechanical response of single ssDNA oligomers lifted from the gold surface is investigated at the same level using cryogenic force spectroscopy. Multistage detachment is inferred, similar to peeling off an adhesive tape, and reflects the strong adhesion of adsorbed ssDNA bases on gold. We extracted high values for the initial stiffness measured during lifting (~15 N m−1), as well as a comparable pinning stiffness obtained from numerical calculations and the intrinsic stiffness per repeat unit of fully stretched ssDNA (~33 N m−1). This last value corresponds to the stiffness of the fully stretched 20-cytosine oligomer having a maximum length of 12 nm when subject to an average tension of ~2 nN. This load is one to two orders of magnitude larger than those applied at room temperature with single-molecule force spectroscopy on hundreds of nanometer-long ssDNA12,14,15,16,17.
A drawback of the present system for experimental lifting is the complex folding of the ssDNA oligomers in comparison to polymeric systems in similar conditions33,34. This limits the lifting heights to only a fraction of the extended length. Compared to kpin ≈ 0.7 N m−1 obtained for polyfluorene chains33, a complete detachment of ssDNA might be achieved by using end linkers allowing to reinforce the bond between the oligomer and the AFM tip in UHV. Although more difficult than solution chemistry, this task appears feasible in view of the successful detachment of PTDCA from Au(111) following contact to a carboxylic oxygen atom32. In future experiments, we will focus on controlling the adsorption of the ssDNA using, for example, appropriately functionalized tips and patterned surfaces. We believe that such strategy might enable the complete lifting of linearly adsorbed oligomers and permit a meaningful statistical analysis of their detachment. Another alternative is to perform measurements on less adhesive surfaces42. Nevertheless, our results suggest that cryogenic force spectroscopy has the potential to study strongly adsorbed biomolecules or similar nano-sized synthetic systems with sub-nanometer resolution under tensile loads up to a few nano-newtons, i.e., 10–100 times higher than hitherto applied in most single-molecule force spectroscopy studies under ambient conditions. Such studies might better characterize the diffusion properties or enable single-base distinction.
Methods
Sample preparation and electrospray deposition
An Au(111) single crystal purchased from Mateck GmbH was cleaned by several sputtering and annealing cycles in a ultrahigh vacuum (UHV). The single-strand 20-mer DNA molecules were purchased from Microsynth (Switzerland). The solution provided was subjected to high-performance liquid chromatography (HPLC), thus ensuring that only the organic molecules present were 20-cytosine ssDNA oligomers. Subsequently, dialysis was performed in the presence of an excess of NaCl which guaranteed that any other salts generated during synthesis are exchanged by NaCl. The delivered solution contained 14.7 nM of ssDNA and a NaCl concentration sufficient to provide more than twice the required amount of Na+ neutralizing counterions. This solution was further diluted to a concentration of around 1 nmol ml−1 in water and then sprayed in UHV (see Supplementary Note 1). Depending on the applied voltage, the deposition time varies from 1 to 30 min at constant solution flux controlled by a syringe pump with a speed of 2–10 × 10−6 l min−1. Further details on the electrospray-deposition apparatus and characterization can be found in refs. 43,44,45.
STM/AFM microscopy
The STM/AFM experiments were carried out at ~5 K with an Omicron GmbH low-temperature STM/AFM controlled by a Nanonis RC5 electronics. We used commercial tuning fork sensors in the qPlus configuration30, e.g., one prong-fixed, the other with an etched tungsten tip epoxied at the end (unperturbed frequency f0 = 26 kHz, quality factor Q = 10000–25000 in UHV, and nominal spring constant k = 1800 N m−1). These tips were sharpened by slight indentation into the gold surface; some were then terminated by a CO molecule at the apex picked up from the surface. All voltages refer to the sample bias with respect to the tip. The constant-height AFM images were acquired with CO-terminated tips using the noncontact mode by driving the free prong on resonance while maintaining a constant tip oscillation amplitude A = 50 pm. For such small oscillation amplitude, the frequency shift Δf induced by a smoothly varying force acting on the tip is to a good approximation proportional to the gradient k (effective stiffness) of the conservative force along the oscillation direction (perpendicular to the sample surface). The signal-to-noise ratio is then nearly optimal30.
Lifting experiments
Pulling experiments were performed under the same conditions with gold-decorated tips while simultaneously recording the tunneling current at a typical bias of 40 μV. The ssDNA oligomers were picked up by gently pressing the AFM tip to the molecule at one of its extremities. Attachment of the molecule to the apex is revealed by an abrupt jump in the force and current signals, as shown in Supplementary Fig. 7. Force spectroscopic measurements upon retraction were performed at a velocity of 22 pm s−1. In contrast to such measurements on biomolecules in ambient conditions, the gradient of the force along the oscillation direction rather than the pulling force itself is thus measured here.
Atomic-level models and force fields
In our simulations, we considered one Au(111) slab composed of three atomic layers-thick slab, where the positions of the atoms in the lowest layer are fixed during the MD runs using a harmonic restrain of 5 kcal mol−1. Furthermore, we considered surfaces of two different sizes, i.e., 16 × 16 nm2 (Fig. 2 and Supplementary Fig. 5) and 18 × 22 nm2 (see Supplementary Fig. 6). The initial structure of the ssDNA molecule was generated using the software NAB46, thus obtaining a double helix with the canonical B-form, as shown in Supplementary Fig. 2. Then, we removed the complementary sequence and used only one single-stranded 20-cytosine oligomer with the charged phosphate groups in the backbone of the ssDNA and 19 sodium counterions. The ssDNA atoms were described using both the parmbsc047 and the χOL3 refinements48 of the Cornell ff99 force field49. The choice of this force field is motivated by its accuracy to describe the mechanical properties of DNA50 as well as adsorption of biomolecules to surfaces51. The sodium counterions were described using the recently improved Joung/Cheatham parameters52,53. As for the gold atoms, we resorted to the CHARMM-METAL force-field54,55 which simultaneously describes the intrinsic properties of gold, while retaining thermodynamical consistency with all the other force fields used here54,55. This force field has been extensively tested by studying the adsorption of different peptides (charged and uncharged) against both density-functional-theory simulations as well as available experimental results54,55. In the simulations performed in water (Fig. 2 and Suppl. Inf. Fig. S5), the water molecules are explicitly modeled using the TIP3P force field56.
Molecular dynamic (MD) simulation details
MD simulations were carried out using AMBER14 software suite46 with NVIDIA GPU acceleration57,58,59. Periodic boundary conditions and Particle Mesh Ewald (with standard defaults and a real-space cutoff of 2 nm) were used to account for long-range electrostatic interactions. Van der Waals interactions were truncated at the real-space cutoff, and Lorentz–Berthelot mixing rules were used to determine the interaction parameters between different atoms. In all vacuum simulations, the volume of the system was kept fixed and the temperature was adjusted by means of a Langevin thermostat with a damping rate of 1 ps−1 ensuring fast thermalization with a minimal effect on the fast slip dynamics. The SHAKE algorithm was used to constrain bonds containing hydrogen, thus allowing us to use an integration time step of 2 fs. Coordinates were saved every 1000 steps. In all our simulations, we observed that the final configuration was stable as it did not change during the last 40 ns of simulations (which was corroborated by the low, i.e., <0.2 nm, root-mean-square deviation). In the steered MD simulation results shown in Fig. 3 and Supplementary Fig. 10, the conservative force F(Z) was computed as ktip (Z − ZP) and its thermal average <F(Z)> was approximated by a running average over (100 ps), an interval adjusted to obtain a smooth dependence without distortions except close to force drops. The resulting d<F(Z)>/dZ can be safely41 compared to the measured effective stiffness k between negative slips in the intervals where k is positive.
Simulation protocols
In total, we have performed eight different MD simulations, each labeled as a stage and described in Supplementary Note 3 and Supplementary Fig. 3. Here we briefly outline each stage. Stage 1: ssDNA oligomer fully embedded in water at 300 K. Stage 2: ssDNA inside a water droplet in vacuum at 300 K. Stage 3: Adsorption of a water droplet containing one ssDNA onto a Au(111) surface at 300 K. Stage 3a: Water evaporation at 450 K from the same adsorbed droplet. Stage 3b: Evolution of a fully dehydrated ssDNA adsorbed on Au(111) in vacuum at 400 K. Stage 4: Adsorption of a single DNA strand in vacuum onto Au(111) at 400 K. Stage 5: Diffusion and self-assembly of two ssDNA molecules adsorbed on Au(111) in vacuum at 400 and 500 K. Stage 6: Lifting a ssDNA molecule adsorbed on Au(111) in vacuum at 5 K. In all simulations, 19 Na+ counterions per ssDNA molecule were included to ensure overall charge neutrality.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Sinden, R. DNA Structure And Function (Elsevier, New York, 2012) https://www.sciencedirect.com/science/article/pii/B9780080571737500018.
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068–17299 (2017).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol. 2, e424 (2004).
Seelig, G., Soloveichik, D., Zhang, D. Y. & Winfree, E. Enzyme-free nucleic acid logic circuits. Science 314, 1585–1588 (2006).
Han, D. et al. DNA origami with complex curvatures in three-dimensional space. Science 246, 343–346 (2011).
Watson, J. D. & Crick, F. H. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171, 737 (1953).
Bustamante, C., Bryant, Z. & Smith, S. Ten years of tension : single-molecule DNA mechanics. Nature 421, 423–427 (2003).
Neuman, K. C. & Nagy, A. Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy. Nat. Methods 5, 491–505 (2008).
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996).
Strick, T. R., Allemand, J. F., Bensimon, D., Bensimon, A. & Croquette, V. The elasticity of a single supercoiled DNA molecule. Science 271, 1835–1837 (1996).
Strick, T., Allemand, J., Croquette, V. & Bensimon, D. Twisting and stretching single DNA molecules. Prog. Biophys. Mol. Biol. 74, 115–140 (2000).
Sonnenberg, L., Parvole, J., Kühner, F., Billon, L. & Gaub, H. E. Choose sides: differential polymer adhesion. Langmuir 23, 6660–6666 (2007).
Kühner, F. et al. Friction of single polymers at surfaces. Langmuir 22, 11180–11186 (2006).
Lee, G. U., Chrisey, L. A. & Colton, R. J. Direct measurement of the forces between complementary strands of DNA. Science 266, 771–773 (1994).
Manohar, S. et al. Peeling single-stranded DNA from graphite surface to determine oligonucleotide binding energy by force spectroscopy. Nano Lett. 8, 4365–4372 (2008).
Lussis, P. et al. A single synthetic small molecule that generates force against a load. Nat. Nanotechnol. 6, 553–557 (2011).
Iliafar, S., Mittal, J., Vezenov, D. & Jagota, A. Interaction of single-stranded DNA with curved carbon nanotube is much stronger than with flat graphite. J. Am. Chem. Soc. 136, 12947–12957 (2014).
Hugel, T., Rief, M., Seitz, M., Gaub, H. E. & Netz, R. R. Highly stretched single polymers: atomic-force-microscope experiments versus ab-initio theory. Phys. Rev. Lett. 94, 048301 (2005).
Hansma, H. G., Laney, D. E., Bezanilla, M., Sinsheimer, R. L. & Hansma, P. K. Applications for atomic force microscopy of DNA. Biophys. J. 68, 1672–1677 (1995).
Bustamante, C. & Rivetti, C. Visualizing protein–nucleic acid interactions on a large scale with the scanning force microscope. Annu. Rev. Biophys. Biomol. Struct. 25, 395–429 (1996).
Cerreta, A., Vobornik, D. & Dietler, G. Fine DNA structure revealed by constant height frequency modulation AFM imaging. Eur. Polym. J. 49, 1916–1922 (2013).
Ares, P. et al. High resolution atomic force microscopy of double-stranded RNA. Nanoscale 8, 11818–11826 (2016).
Clemmer, C. R. & Beebe, T. P. Jr. Graphite: a mimic for DNA and other biomolecules in scanning tunneling microscope studies. Science 251, 640–642 (1991).
Zhang, Y., Sheng, S. J. & Shao, Z. Imaging biological structures with the cryo atomic force microscope. Biophys. J. 71, 2168–2176 (1996).
Tanaka, H. & Kawai, T. Partial sequencing of a single DNA molecule with a scanning tunneling microscope. Nat. Nanotechnol. 4, 518–522 (2009).
Hamai, C., Tanaka, H. & Kawai, T. Extended structure of DNA oligomer and nucleotide imaging studied by scanning tunneling microscopy. J. Phys. Chem. B 104, 9894–9897 (2000).
Fenn, J. B., Mann, M., Meng, C. K., Wong, S. F. & Whitehouse, C. M. Electrospray ionization for mass spectrometry of large biomolecules. Science 246, 64–71 (1989).
Rauschenbach, S. et al. Electrospray ion beam deposition of clusters and biomolecules. Small 2, 540–547 (2009).
Hinaut, A. et al. Electrospray deposition of structurally complex molecules revealed by atomic force microscopy. Nanoscale 6, 1927–1934 (2018).
Giessibl, F. J. Advances in atomic force microscopy. Rev. Mod. Phys. 75, 949–983 (2003).
Gross, L., Mohn, F., Moll, N., Liljeroth, P. & Meyer, G. The chemical structure of a molecule resolved by atomic force microscopy. Science 325, 1110–1114 (2009).
Wagner, C., Fournier, N., Tautz, F. S. & Temirov, T. Measurement of the binding energies of the organic-metal perylene–teracarboxylic–dianhydride/Au(111) bonds by molecular manipulation using an atomic force microscope. Phys. Rev. Lett. 109, 076102 (2012).
Kawai, S. et al. Quantifying the atomic-level mechanics of single long physisorbed molecular chains. Proc. Natl Acad. Sci. USA 111, 3968–3972 (2014).
Kawai, S. et al. Superlubricity of graphene nanoribbons on gold surfaces. Science 351, 957–961 (2016).
Pawlak, R. et al. Single-molecule tribology: force microscopy manipulation of a porphyrin derivative on a copper surface. ACS Nano 10, 713–722 (2016).
Pawlak, R. et al. Single-molecule manipulation experiments to explore friction and adhesion. J. Phys. D Appl. Phys. 50, 113003 (2017).
Dubochet, J., Adrian, M., Lepault, J. & McDowall, A. W. Emerging techniques: cryo-electron microscopy of vitrified biological specimens. Trends Biochem. Sci. 10, 143–146 (1985).
Rosa, M., Corni, S. & Di Felice, R. Interaction of nucleic acid bases with the Au(111) surface. J. Chem. Theory Comput. 9, 4552–4561 (2013).
Rosa, M., Di Felice, R. & Corni, S. Adsorption mechanisms of nucleobases on the hydrated Au(111) surface. Langmuir 34, 14749–14756 (2018).
Lafferentz, L. et al. Conductance of a single conjugated polymer as a continuous function of its length. Science 323, 1193 (2009).
Park, S. & Khalili-Araghi, F. Free energy calculation from steered molecular dynamics simulations using Jarzynski’s equality. J. Chem. Phys. 119, 3559–3566 (2003).
Vilhena, J. G. et al. Stick-slip motion of ssDNA over graphene. J. Phys. Chem. B 122, 840–846 (2018).
Satterley, C. J. et al. Electrospray deposition of fullerenes in ultra-high vacuum: in situ scanning tunneling microscopy and photoemission spectroscopy. Nanotechnology 18, 455304 (2007).
Kebarle, P. & Verkerk, U. H. Electrospray: from ions in solution to ions in the gas phase, what we know now. Mass. Spectrom. Rev. 28, 898–917 (2003).
Hinaut, A., Pawlak, R., Meyer, E. & Glatzel, T. Electrospray deposition of organic molecules on bulk insulator surfaces. Beilstein J. Nanotech. 6, 1927–1934 (2015).
Case, D. et al. AMBER 14 (University of California: San Francisco, CA, 2014).
Pérez, A. et al. Refinement of the AMBER force field for nucleic acids: improving the description of/conformers. Biophys. J. 92, 3817–3829 (2007).
Zgarbová et al. Refinement of the Cornell et al. nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J. Chem. Theory Comput. 7, 2886–2902 (2011).
Cornell, W. D. et al. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117, 5179–5197 (1995).
Marin-Gonzalez, A., Vilhena, J. G., Perez, R. & Moreno-Herrero, F. Understanding the mechanical response of double-stranded DNA and RNA under constant stretching forces using all-atom molecular dynamics. Proc. Natl Acad. Sci. USA 114, 7049–7054 (2017).
Vilhena, J. G. et al. Adsorption orientations and immunological recognition of antibodies on graphene. Nanoscale 8, 13463–13475 (2016).
Joung, I. S. & Cheatham, T. E. Molecular dynamics simulations of the dynamic and energetic properties of alkali and halide ions using water-model-specific ion parameters. J. Phys. Chem. B 113, 13279–13290 (2009).
Li, P., Roberts, B. P., Chakravorty, D. K. & Merz, K. M. Rational design of particle mesh Ewald compatible Lennard–Jones parameters for +2 metal cations in explicit solvent. J. Chem. Theory Comput. 9, 2733–2748 (2013).
Feng, J. et al. Adsorption mechanism of single amino acid and surfactant molecules to Au111 surfaces in aqueous solution: design rules for metal-binding molecules. Soft Matter 7, 2113 (2011).
Heinz, H. & Ramezani-Dakhel, H. Simulations of inorganic-bio-organic interfaces to discover new materials: insights, comparisons to experiment, challenges, and opportunities. Chem. Soc. Rev. 45, 412–448 (2016).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. Explicit solvent particle mesh Ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Götz et al. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born. J. Chem. Theory Comput. 8, 1542–1555 (2012).
Le Grand, S. L., Götz, A. W. & Walker, R. C. SPFP: speed without compromise A mixed precision model for GPU accelerated molecular dynamics simulations. Comput. Phys. Commun. 184, 374–380 (2013).
Acknowledgments
A.B., A.H., and E.M. and R.P., T.G., and T.M. thank the Swiss National Science Foundation (SNF) and the Swiss Nanoscience Institute (SNI). The COST Action MP1303 is gratefully acknowledged. Ru. P. and J.G.V. thank the financial support of the Spanish MINECO (projects MAT2014-54484-P, MDM-2014-0377, and MAT2017-83273-R) and also the computer resources, technical expertise, and assistance provided by the Red Española de Supercomputación at the Minotauro Supercomputer (BSC, Barcelona). J.G.V. acknowledges funding from a Marie Sklodowska-Curie Fellowship within the Horizons 2020 framework. Dr. Marcin Kisiel is acknowledged for fruitful discussion.
Author information
Authors and Affiliations
Contributions
E.M., Ru.P., R.P. and J.G.V. conceived the experiments. A.H. performed the electrospray deposition. R.P. performed the STM/AFM measurements and lifting experiments. J.G.V. conducted the numerical calculations. R.P. and J.G.V. analyzed the data and co-wrote the manuscript with the help of E.G. and A.B. E.M., Ru.P., R.P., J.G.V., A.H., T.M., T.G., A.B. and E.G. discussed on the results and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Communications thanks Marta Rosa and the other anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pawlak, R., Vilhena, J.G., Hinaut, A. et al. Conformations and cryo-force spectroscopy of spray-deposited single-strand DNA on gold. Nat Commun 10, 685 (2019). https://doi.org/10.1038/s41467-019-08531-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-08531-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.