Abstract
Nanopore-based sensors are advancing the sensitivity and selectivity of single-molecule detection in molecular medicine and biotechnology. Current electrical sensing devices are based on either membrane protein pores supported in planar lipid bilayers or solid-state (SS) pores fabricated in thin metallic membranes. While both types of nanosensors have been used in a variety of applications, each has inherent disadvantages that limit its use. Hybrid nanopores, consisting of a protein pore supported within a SS membrane, combine the robust nature of SS membranes with the precise and simple engineering of protein nanopores. We demonstrate here a novel lipid-free hybrid nanopore comprising a natural DNA pore from a thermostable virus, electrokinetically inserted into a larger nanopore supported in a silicon nitride membrane. The hybrid pore is stable and easy to fabricate, and, most importantly, exhibits low peripheral leakage allowing sensing and discrimination among different types of biomolecules.
Similar content being viewed by others
Introduction
The advent of single-molecule detection is having an unparalleled impact on the speed with which structural and dynamic aspects of molecules can be probed1. In this regard, nanopores have shown much promise as electrical2,3,4,5,6,7 and combined electro-optical sensors8,9,10 and several nanopore-based systems are now being adopted as primary tools for DNA11,12,13 and RNA14 sequencing. Despite recent progress, identification and quantification of molecular species in solution15,16,17,18,19,20,21,22,23,24,25,26,27,28 requires a reproducible nanopore platform that affords physical stability and structural precision. Sophisticated applications, such as electro-optical sensing, necessitate the pore position be geometrically defined to allow precise alignment of the optical system. While synthetic nanopores fabricated in solid-state (SS) membranes offer physical robustness29,30,31, pore-to-pore variability often limits the reproducibility of experiments, necessitating additional control checks and validation. On the contrary, protein channels embedded in organic thin membrane, e.g., a lipid bilayer, offer the highest reproducibility due to the precise folding and repetitive nature of the constituting multisubunit protein oligomers32,33, but their supporting membrane is typically less chemically and physically robust and, further, the pore position is variable due to in-plane diffusion of the protein channel34. While the use of amphiphilic polymers and polymerizable lipids have improved the lifetime, mechanical stability and voltage tolerance of the biomimetic support membranes for biological nanopores, as evidenced by the devices produced by Oxford Nanopore Technologies, SS membranes are likely to afford additional pressure and temperature resistance. Hybrid nanopore devices, in which channel-containing proteins are embedded in larger pores made in a SS matrix, have been proposed as a strategic solution for combining the benefits, while overcoming the limitations of existing nanopores35. Although initial experiments based on inserting pore-containing proteins with lipophilic regions into a SS pore looked promising35, challenges in inserting such proteins into a SS pore and in controlling the protein orientation have remained major obstacles in the applicability of hybrid nanopores to nanotechnology.
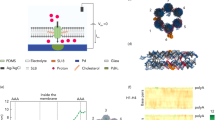
Inspired by natural DNA pores, we designed a novel lipid-free hybrid nanopore based on the hydrophilic portal protein derived from the thermostable virus G20c36. In double-stranded DNA viruses, the portal protein is incorporated into the capsid shell (Fig. 1a), thereby serving as a natural pore through which DNA is moved in and out37. The protein contains a tight tunnel constriction with a repetitive chemical character, being made up by a circle of identical tunnel loops, contributed by 12 subunits38. In previous work, we engineered this protein to reprogram its physico-chemical properties (GG)33 to create a portal with a larger minimum aperture of ~2.3 nm defined by the substitution of two bulky residues at the tip of the tunnel loop with glycines. Additional features of this portal system include substitution of an externally facing residue, located around the outside of the protein, by cysteine (designated C, e.g., CGG or CD/N) which allows chemical labeling and surface immobilization of the portal protein, as we have previously demonstrated for the lipid membrane insertion of the portal protein33. In this work, we have electrostatically engineered a portal protein variant (D/N) by replacing aspartic acid (D) residues at the internal tunnel surface with asparagines (N). This altered the charge of the lower part of the internal tunnel’s surface from negative to positive (Fig. 1b and Supplementary Figure 1), a change that was crucial for electrical sensing of net negatively charged biomolecules. Here, we use this structurally programmable portal protein as a nanoscale adapter by electrokinetically embedding it snugly inside a larger pore made in a freestanding silicon nitride (SiN) membrane to form a lipid-free hybrid nanopore. Using the CD/N portal with engineered internal pore properties (Fig. 1b and Supplementary Figure 1), we characterized the electrical properties of the hybrid pore and applied it to electrically detect different biomolecules. We demonstrate that a folded protein larger than the pore interior does not enter the hybrid portal. In contrast, biopolymers including single-stranded DNA (ssDNA), double-stranded DNA (dsDNA) that contains a single-stranded tail, and a peptide predicted to have a random coil conformation with a 10-amino acid α-helix at the C terminus can all be discriminated based on their distinct signal amplitudes in a way that is commensurate with their molecular cross-section. Our results indicate that the hybrid portal is a versatile sensor of various biopolymer types which may, with further development, find uses in genomic mapping as well as polypeptide and oligonucleotide sequencing.
Design of a bio-inspired lipid-free hybrid nanopore. a Cartoon of the DNA packaging machine of a dsDNA virus. Viral genomic DNA (red) is translocated into the preformed virus capsid by the packaging ATPase (yellow) through the portal protein (aqua) embedded in viral capsid (gray). b Left, electrostatic properties of the tunnel in wild-type and mutant portal proteins. Slice through the middle of molecular surface colored according to charge from red (−1 kT e−1) to blue (+1 kT e−1). b Right (gray box), dimensions of the portal protein (teal, L) and the SS nanopore (pink, R). c Insertion of the purified portal protein into a nanopore fabricated in a thin solid-state (SS) membrane. Portal protein is applied to the trans chamber of a SS nanopore device containing an electrolyte solution of 20 mM Tris pH 7.5, 0.5 M NaCl. The protein electrokinetically inserts into the SS pore during application of a positive voltage. d Cartoon image of the hybrid pore, in which application of voltage results in ion current through the pore (blue arrows), as well as leakage current that is peripheral to the pore (red arrows)
Results
Portal is a nanoscale adapter for SS nanopores
Electrokinetic corking of the G20c portal protein into the SS nanopore occurs when the force on the protein, induced by applied voltage, is sufficient to squeeze the portal into the SS pore (Figs. 1c, 2 and Supplementary Figures 2, 3). We find that stable insertion required specific geometric parameters for the SS nanopore, namely, a diameter between 5.4 to 6 nm and a nominal membrane thickness of 30 nm. Given the dimensions of the portal assembly33 (Fig. 1b), the geometric constraints set by the SS pore restrict the range of possible orientations of the portal pore in it, such that the stem is inserted within the SS nanopore constriction, and the wider cap self-orients towards the top of the trans chamber (Fig. 1d). The larger size of the cap, as compared with the SS pore diameter, prevents the entire protein from moving through the SS nanopore. Remarkably, interactions between the portal protein squeezed into the SS pore and the SS pore surface contribute to a stable, self-inserting and self-aligning hybrid (Fig. 1d) that exhibits tolerable peripheral ion leakage, probed using cyclodextrin as a pore current modulator. Our hybrid pores exhibit lifetimes of hours (see Supplementary Figure 4), and similar ion current noise values to a lipid bilayer-supported portal protein nanopore33 (Fig. 2 and Supplementary Figure 5).
Characterization of the hybrid pore formation. a Typical current profile over time recorded through a 5.5 nm diameter SS pore at +100 mV. After injection of 0.1 nmol of portal protein, short current drops are detected, interpreted as portal collisions with the solid-state nanopore. b A representative current vs time trace recorded for a 5.4 nm SS nanopore at +80 mV, showing stable insertion of a portal protein. c Current as a function of the applied voltage for a 5.7 nm diameter SS pore recorded before (red markers) and after insertion of a portal protein (purple). d Current noise analysis of a 5.5 nm diameter solid-state nanopore before (red) and after insertion of a portal protein (purple). e Conductance of solid-state nanopore vs conductance of hybrid SS-portal pores formed after electrokinetic corking of the portal protein within the SS nanopore (n = 32 for CD/N hybrids and n = 15 for CGG hybrids). Experiments were performed in 0.5 M NaCl, 20 mM Tris, pH 7.5
After confirming the base current of stable SS nanopores of the desired diameter, addition of the portal protein to the trans chamber resulted in reversible partial blockades of the ionic current (Fig. 2a, b, Supplementary Figures 2 and 3). We interpret these short-lived events as portal protein collisions with the SS nanopore without stable insertion, where the ion current is partially blocked as the protein approaches the SS pore, prior to movement away. These short-lived events were usually followed by long-lived events (Fig. 2b, Supplementary Figures 2, 3), of comparable current blockade levels, events that were only observed in SS pores with diameters of 5.4 to 6 nm. We interpret the long-lived events as stable insertion of a portal protein into the SS nanopore to form a hybrid nanopore. The average conductance (mean and s.d.) of these hybrid pores was calculated (Fig. 2e) to be 1.50 ± 0.48 nS and 1.33 ± 0.42 nS for the CD/N (from 32 hybrid nanopores) and the CGG (from 15 hybrid nanopores) variants, respectively. Surprisingly, such hybrid pores remain stable at both positive and negative voltages (Fig. 2c). However, applying significant negative bias generally results in uncorking of the protein from the SS nanopore, and an example of an ejection at −80 mV is shown in Fig. 2c (red markers). These data are consistent with the protein insertion and removal being electrokinetically driven.
Obtaining a sufficient increase in the signal-to-noise ratio is a major challenge for properly identifying transport events by nanopore sensing. Power spectral densities of the current noise for a SS nanopore before, and after, insertion of a portal protein (Fig. 2d and Supplementary Figure 4) showed that the 1/f noise at low frequencies decreased upon formation of the hybrid pore. This 1/f noise reduction is consistent with a reduced pore conductance, as well as an indicator of the reduced surface charge fluctuations that are hallmarks of silicon nitride surfaces39. This, along with the observation that capacitance-dominated noise at high frequencies was comparable for both pores, suggests that no new source of noise was introduced by hybrid pore formation. We deduced that the observed variation in the open pore current for different hybrid nanopores (Supplementary Figures 2, 3) was likely to be caused by differences in SS geometry and the associated leakage currents around the portal protein. We attempted to measure the extent of ion leakage from the pore by measuring β-cyclodextrin interactions with the hybrid pore for the CGG mutant, a mutant that we previously embedded into a lipid membrane (Supplementary Figure 6)33. Our results show that β-cyclodextrin does not translocate the pore, in contrast to the same experiment conducted on the lipid-embedded version of the same portal protein. While this precludes an accurate measurement of the leakage, these results suggest that corking the protein into a snug SS nanopore slightly reduces the innermost pore constriction. The reproducible signals obtained from biomolecules, as well as the steady baselines of the hybrid, allow current blockades as low as ∼20 pA to be accurately measured (Supplementary Figure 6). These data demonstrate that despite a low level of steady peripheral leakage, these hybrid pores are unique lipid-free protein-based pore sensors.
Hybrid nanopores as sensors of biomolecules
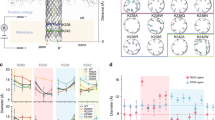
We then investigated the sensing capabilities of these hybrid nanopores by analyzing the transport of a peptide, comprising residues 1–43 of the human TPX2 protein, as a function of applied voltage (Fig. 3). The TPX2 peptide is negatively charged at pH 7.5 (pI = 3.7) and was added to the cis chamber, on the opposite side of the membrane to which the portal protein was introduced (see inset to Fig. 3b) to facilitate electrophoretically driven translocation. Adjusting the applied voltage from +30 to + 60 mV resulted in an increased baseline ion current through the hybrid pore, as well as the frequency of observed current blockades (Fig. 3a). Two kinds of current blockades associated with two different events were detected: bumping events, characterized by brief, low-level current blockades, arising from diffusion of the peptide close to the hybrid pore entrance; and translocation events, characterized by larger current blockades of longer duration. These two types of events are typically seen during translocation of DNA40,41,42 and proteins43,44,45 through protein channels. The inter-event time distribution is well fit by a single exponential equation (Supplementary Figure 7). The entry frequency (Fig. 3b) of the peptide into the hybrid pore is described by Van’t Hoff Arrhenius relationship44,46 f = f0 exp(V/V0), consistent with both translocation of DNA40,41,42, proteins43,47 and peptides46,48,49,50,51 through either α-hemolysin or aerolysin; and a significant entropic barrier for peptide entry into the pore. The dwell time distributions were well fit by a double-exponential equation (Supplementary Figure 7b), which are typically due to two types of processes, normally associated with short bumping and longer translocation events16. We found that the average frequency for both types of events increases exponentially (Fig. 3b), while the average dwell time for the long events decreased exponentially with the applied voltage (Fig. 3c). Based on prior work that employed the α-hemolysin and aerolysin nanopores,43,45 we conclude that the long events represent transport of the peptide through the hybrid pore to the trans chamber.
Dynamics of TPX2 peptide transport. a Current vs time trace recorded through a hybrid pore at +30, +40 and +55 mV in the presence of 10.3 μM TPX2 peptide. b Semi-log plot of the event frequency as a function of the applied voltage. c Semi-log plot of the peptide dwell time as a function of the applied voltage. The lines in (b, c) are exponential fits to the data. Experiments were performed in 0.5 M NaCl, 20 mM Tris, pH 7.5. Data shown in semi-log plots are mean and s.d. of 23,516 events (total for all points) from one hybrid nanopore
In order to further demonstrate the sensing capabilities of this hybrid pore, we investigated the transport of other biopolymers: dsDNA that contains a ssDNA tail, ssDNA, a folded protein as well as the TPX2 peptide (Fig. 4 and Supplementary Figures 7-10). Since all of these polymers are negatively charged at pH 7.5, electrophoresis allows molecular capture into the base of the portal protein following their introduction to the cis chamber (the opposite side of the SS membrane to portal insertion). After addition of each biopolymer: 36.0 μM insulin (Fig. 4a and Supplementary Figure 8), 7.7 μM hairpin-polydT50 (Fig. 4b and Supplementary Figure 9), 10.3 μM TPX2 peptide (Figs. 3, 4c), 6.9 μM 60bp-polydT30 (Supplementary Figure 10) and 16.6 μM ssDNA polydA20dC20dA20 (Fig. 4d and Supplementary Figure 11), reversible partial blockades of the ionic current are observed at +60 mV. Similar short-lived bumping events as well as longer events were observed for each biopolymer, as described above for the TPX2 peptide (Fig. 3). These types of blockades were also observed at several different voltages for DNA molecules (Supplementary Figures 9–11), with voltage-dependent changes in event frequency and duration for ssDNA polydA20dC20dA20 consistent with translocation occurring (Supplementary Figure 11), as noted for the peptide above (Fig. 3). Conversely, the folded, globular molecule of insulin with a smallest dimension of ~3 nm (Protein Data Bank (PDB) code: 1zeh)52,53 is too large for the ~2 nm constriction of the hybrid pore and therefore does not translocate. It is however possible that insulin explores the cavity at the portal tunnel’s entrance (∼5 nm) without being transported to the trans chamber, producing structured events that are long-lived and have a low current blockade level. Such events have been previously observed for nanoreactors, where biomolecules are captured or tethered within ClyA and FraC nanopores20,54.
Sensing different biopolymers using a hybrid nanopore. Current vs time trace recorded through the hybrid pore at +60 mV in the presence of a 36.0 μM insulin, b 7.7 μM DNA hairpin, c 10.3 μM TPX2 peptide and d 16.6 μM ssDNA. The data in a were filtered at 10 kHz (gray) or 0.5 kHz (green). e Scatter plot of ΔI vs dwell time for the DNA hairpin (red, n = 5883 events), the peptide (purple, n = 3368 events) and the ssDNA (orange, n = 18,812 events)). Experiments were performed in 0.5 M NaCl, 20 mM Tris, pH 7.5
Lastly, we compare the event characteristics for different biopolymers at the same applied voltage of +60 mV by overlaying their scatter plots of ΔI vs. dwell time, as shown in Fig. 4e. Crucially, the level of current blockade, ΔI, appeared to be biopolymer dependent. We found current blockades (mean and s.d.) for the dsDNA (Fig. 4e) of ΔI = 34.6 ± 4.2 pA (5883 events), while in contrast, we found ΔI = 18.1 ± 3.2 pA (18,812 events) for ssDNA. This is nearly 2 times less than for the partially dsDNA, and is consistent with values found for dsDNA and ssDNA in SS nanopores55, where the difference in conductance was found to be ~2.75-fold. For the peptide, we found ΔI = 30.1 ± 5.5 pA (3368 events). Since we showed that the peptide is transported through the pore (Fig. 3) and is predicted to contain an α-helix of ∼1.4 nm in diameter, as seen in the structure of the TPX2 peptide bound to its partner kinase, Aurora A (PDB: 1ol5), the data are compatible with translocation through the narrowest constriction of the hybrid pore (∼2 nm diameter). The ΔI value found for the peptide is consistent with the α-helical region being the main cause of the blockade, and with its diameter being intermediate between that of dsDNA and ssDNA. These data suggest that the predicted α-helix is present in the isolated peptide under these experimental conditions. While transport of structured biopolymers has been reported for nucleic acids56,57, only a single report to our knowledge presents transport of an α-helical peptide through a protein nanopore45.
Discussion
Hybrid nanopores, supported by SS membranes, could offer superior properties to both the planar lipid bilayer-based pores (that are sensitive to temperature, osmotic pressure and applied electric field strength and suffer from uncontrollable positional parameters) and SS nanopores (that are prone to edge erosion and are difficult to reproducibly fabricate with diameters <5 nm). However, despite having been the subject of industrial and academic research, development of a device that can be easily fabricated has proven difficult. For example, producing a hybrid pore based on DNA origami or α-hemolysin, a membrane protein, noted substantial leakage current58,59 or relatively short hybrid pore lifetime and required complex protein modifications35. In contrast, the hybrid nanopore described here is based on a soluble, stable and relatively hydrophilic viral portal protein, whose chemical properties, including those inside tunnel, can be easily tuned. This work demonstrates that the lipid-free hybrid nanopore comprising the G20c portal protein inserted into a thin SS SiN membrane is easy to assemble, with the portal protein readily electrokinetically inserting into the SS pores and typically remaining stable for hours of experimental time. We plainly demonstrate the utility of this hybrid pore as a nanosensor, by observing characteristic readout for dsDNA and ssDNA, as well as a peptide and a globular protein.
The stability and tunability of the hybrid nanopore that combines the advantages of both protein and SS nanopores encourages further work to introduce specific sensing properties for this hybrid pore and produce robust and geometrically controlled devices amenable to sophisticated high-throughput array-based or coupled multi-system (e.g., optical, flow or pressure) applications in nanotechnology.
Methods
Protein engineering and purification
Mutant portal proteins CGG33 and CD/N (Supplementary Figure 1) were expressed in Escherichia coli Shuffle cells at 30 °C overnight after induction with 0.5 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) at 0.8 OD600. The cells were lysed by sonication on ice in 50 mM Tris pH 8, 1 M NaCl, 10 mM imidazole, 100 mM AEBSF, 10 mg mL−1 lysozyme and 2 mM dithiothreitol (DTT). After clarification by centrifugation at 15,000 rpm for 15 min, the protein was purified from the lysate by Immobilized Metal Affinity Chromatography (IMAC; 5 mL HiTrap FF Crude, GE Healthcare) and eluted over a gradient of 10–500 mM imidazole over 10 column volumes. Fractions containing the protein were subjected to a buffer exchange step over a desalting column (HiPrep 26/10; GE Healthcare) to improve 3C cleavage of the histidine affinity tag (50 mM Tris pH 8, 0.5 M NaCl, 50 mM K Glutamate, 1 mM DTT) prior to buffer exchange back into low imidazole buffer before the second IMAC step and purification to homogeneity in 20 mM Tris pH 8, 1 M NaCl, 1 mM DTT, before freezing in liquid nitrogen and storage at −80 °C. Protein was exchanged into 20 mM Tris pH 7.5, 0.5 M NaCl buffer (Zeba Spin Columns, Thermofisher) for use in hybrid nanopore formation. CD/N mutant proteins were characterized for stability and assembly state by nano differential scanning fluorimetry (nanoDSF) and negative stained transmission electron microscopy (TEM; Supplementary Figure 1). Human TPX21-43 peptide was produced in E. coli as a his-tagged GB1 fusion (Marko Hyvonen, University of Cambridge). Expression was induced in BL21 pLysS cells at 0.8 OD600 with 0.5 mM IPTG for 4 h at 37 °C. The fusion protein was purified by IMAC in standard nickel affinity chromatography buffers at pH 7.5 containing 1 mM DTT and eluted over a 10–500 mM imidazole gradient, after which peak fractions were pooled, prior to buffer exchange (desalting column, GE Healthcare) into 3C cleavage buffer 50 mM Tris pH 7.5, 0.5 M NaCl, 10 mM imidazole and 1 mM DTT. After cleavage, a second IMAC step removed the his-tagged GB1 fragment and the unbound fraction containing the TPX2 peptide (GPGSMLSYSYDAPSDFINFSSLDDEGDTQNIDWFEEKANLENLKGGGCQ) was concentrated by centrifugal ultrafiltration using a 3 kDa cutoff filter (Amicon) and further purified over a S75 10/300 size exclusion column (GE Healthcare) in 20 mM Tris pH 7.5, 0.5 M NaCl and 1 mM DTT prior to concentration as before, freezing in liquid nitrogen and storage at −80 °C.
Experimental set-up
Nanopores were fabricated in 30 nm thick SiN membranes using previously reported methods60,61. The pore diameters ranged between 5.4 and 6 nm in order to properly orient the portal protein. Nanopores were cleaned with hot piranha (3:1 H2SO4/ H2O2), followed by hot deionized water before each experiment. After being dried under vacuum, nanopore chips were assembled in a custom cell equipped with Ag/AgCl electrodes, and quick-curing silicone elastomer was applied between the chip and the cell to seal the device and thereby reduce the noise by minimizing the chip capacitance. We introduced 0.5 M NaCl, 20 mM Tris, pH 7.5, as an electrolyte solution onto both sides of the chip. Portal protein was always added to the trans chamber and the biopolymers to the cis chamber. All experiments were carried out at ambient temperature. Human insulin was purchased from Alfa Aesar (Thermofisher), and dsDNA hairpin (5′-GCTGTCTGTTGCTCTCTCGCAACAGACAGC T50-3′), ssDNA (5′-dA20dC20dA20-3′), 60 bp-polydT30 ((5′-TCAGGGTTTTTTTACT)4 T30-3’) and its complementary strand ((3′-AGTAAAAAAACCCTGA-5′)4) were synthesized by Integrated DNA Technology.
Electrical detection and data acquisition
The ionic current through SS nanopores and portal hybrid protein was measured using an Axopatch 200B amplifier (Molecular Devices). Data were filtered at 10 kHz and acquired at 250 kHz using a National Instruments DAQ card and custom LabVIEW software. Data were processed and events were detected using Pythion software (https://github.com/rhenley/Pyth-Ion/). The values for the open pore current (I0) and the standard deviation of the noise (σ) were extracted from the analysis. The threshold (Th) applied in Pythion to separate events from the noise is given by Th = I0–4σ. The average duration of blockades was deduced from the distribution of blockade duration, τt. The two blockade time distributions of independent events are adjusted with a double-exponential function, y = A1exp(t/τ1) + A2exp(t/τ2). All statistical analyses were performed using Igor Pro software (WaveMetrics Inc.).
References
Wang, H. et al. Determining the physical properties of molecules with nanometer-scale pores. ACS Sens. 3, 251–263 (2018).
Derrington, I. M. et al. Nanopore DNA sequencing with MspA. Proc. Natl. Acad. Sci. USA 107, 16060–16065 (2010).
Nivala, J., Marks, D. B. & Akeson, M. Unfoldase-mediated protein translocation through an α-hemolysin nanopore. Nat. Biotechnol. 31, 247–250 (2013).
Rodriguez-Larrea, D. & Bayley, H. Multistep protein unfolding during nanopore translocation. Nat. Nanotech. 8, 288–295 (2013).
Gu, L. Q., Braha, O., Conlan, S., Cheley, S. & Bayley, H. Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398, 686–690 (1999).
Butler, T. Z., Pavlenok, M., Derrington, I. M., Niederweis, M. & Gundlach, J. H. Single-molecule DNA detection with an engineered MspA protein nanopore. Proc. Natl. Acad. Sci. USA 105, 20647–20652 (2008).
Kasianowicz, J. J. et al. Analytical applications for pore-forming proteins. Biochim. Biophys. Acta 1858, 593–606 (2016).
Zhang, M. et al. Thermophoresis-controlled size-dependent DNA translocation through an array of nanopores. ACS Nano 12, 4574–4582 (2018).
Larkin, J., Henley, R. Y., Jadhav, V., Korlach, J. & Wanunu, M. Length-independent DNA packing into nanopore zero-mode waveguides for low-input DNA sequencing. Nat. Nanotech. 12, 1169–1175 (2017).
McNally, B. et al. Optical recognition of converted DNA nucleotides for single-molecule DNA sequencing using nanopore arrays. Nano Lett. 10, 2237–2244 (2010).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Jain, M. et al. Improved data analysis for the MinION nanopore sequencer. Nat. Methods 12, 351–356 (2015).
Loman, N. J., Quick, J. & Simpson, J. T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 12, 733–735 (2015).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Howorka, S. & Siwy, Z. Nanopore analytics: sensing of single molecules. Chem. Soc. Rev. 38, 2360 (2009).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl. Acad. Sci. USA 93, 13770–13773 (1996).
Mohammad, M. M. et al. Engineering a rigid protein tunnel for biomolecular detection. J. Am. Chem. Soc. 134, 9521–9531 (2012).
Robertson, J. W. F. et al. Single-molecule mass spectrometry in solution using a solitary nanopore. Proc. Natl. Acad. Sci. USA 104, 8207–8211 (2007).
Merstorf, C. et al. Wild type, mutant protein unfolding and phase transition detected by single-nanopore recording. ACS Chem. Biol. 7, 652–658 (2012).
Huang, G., Willems, K., Soskine, M., Wloka, C. & Maglia, G. Electro-osmotic capture and ionic discrimination of peptide and protein biomarkers with FraC nanopores. Nat. Commun. 8, 935 (2017).
Piguet, F. et al. Identification of single amino acid differences in uniformly charged homopolymeric peptides with aerolysin nanopore. Nat. Commun. 9, 966 (2018).
Luchian, T., Shin, S.-H. & Bayley, H. Kinetics of a three-step reaction observed at the single-molecule level. Angew. Chem. Int. Ed. Engl. 42, 1926–1929 (2003).
Wescoe, Z. L., Schreiber, J. & Akeson, M. Nanopores discriminate among five C5-cytosine variants in DNA. J. Am. Chem. Soc. 136, 16582–16587 (2014).
Baaken, G. et al. High-resolution size-discrimination of single nonionic synthetic polymers with a highly charged biological nanopore. ACS Nano 9, 6443–6449 (2015).
Fennouri, A. A. et al. Single molecule detection of glycosaminoglycan hyaluronic acid oligosaccharides and depolymerization enzyme activity using a protein nanopore. ACS Nano 6, 9672–9678 (2012).
Lee, J. et al. Semisynthetic nanoreactor for reversible single-molecule covalent chemistry. ACS Nano 10, 8843–8850 (2016).
Willems, K., Van Meervelt, V., Wloka, C. & Maglia, G. Single-molecule nanopore enzymology. Philos. Trans. R. Soc. Lond. B Biol. Sci. 372, pii: 20160230 (2017).
Rosen, C. B., Rodriguez-Larrea, D. & Bayley, H. Single-molecule site-specific detection of protein phosphorylation with a nanopore. Nat. Biotechnol. 32, 179–181 (2014).
Verschueren, D. V., Jonsson, M. P. & Dekker, C. Temperature dependence of DNA translocations through solid-state nanopores. Nanotechnology 26, 234004 (2015).
Oukhaled, A. et al. Dynamics of completely unfolded and native proteins through solid-state nanopores as a function of electric driving force. ACS Nano 5, 3628–3638 (2011).
Yamazaki, H. et al. Label-free single-molecule thermoscopy using a laser-heated nanopore. Nano. Lett. 17, 7067–7074 (2017).
Song, L. et al. Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1866 (1996).
Cressiot, B. et al. Porphyrin-assisted docking of a thermophage portal protein into lipid bilayers: nanopore engineering and characterization. ACS Nano 11, 11931–11945 (2017).
Castell, O. K., Berridge, J. & Wallace, M. I. Quantification of membrane protein inhibition by optical ion flux in a droplet interface bilayer array. Angew. Chem. Int. Ed. 51, 3134–3138 (2012).
Hall, A. R. et al. Hybrid pore formation by directed insertion of α-haemolysin into solid-state nanopores. Nat. Nanotech. 5, 874–877 (2010).
Williams, L. S., Levdikov, V. M., Minakhin, L., Severinov, K. & Antson, A. A. 12-Fold symmetry of the putative portal protein from the Thermus thermophilus bacteriophage G20C determined by X-ray analysis. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 69, 1239–1241 (2013).
Casjens, S. R. & Gilcrease, E. B. Determining DNA packaging strategy by analysis of the termini of the chromosomes in tailed-bacteriophage virions. Methods Mol. Biol. 502, 91–111 (2009).
Lebedev, A. A. et al. Structural framework for DNA translocation via the viral portal protein. EMBO J. 26, 1984–1994 (2007).
Hoogerheide, D. P., Garaj, S. & Golovchenko, J. A. Probing surface charge fluctuations with solid-state nanopores. Phys. Rev. Lett. 102, 256804 (2009).
Henrickson, S. E., Misakian, M., Robertson, B. & Kasianowicz, J. J. Driven DNA transport into an asymmetric nanometer-scale pore. Phys. Rev. Lett. 85, 3057–3060 (2000).
Meller, A. & Branton, D. Single molecule measurements of DNA transport through a nanopore. Electrophoresis 23, 2583–2591 (2002).
Japrung, D., Henricus, M., Li, Q., Maglia, G. & Bayley, H. Urea facilitates the translocation of single-stranded DNA and RNA through the α-hemolysin nanopore. Biophys. J. 98, 1856–1863 (2010).
Cressiot, B. et al. Dynamics and energy contributions for transport of unfolded pertactin through a protein nanopore. ACS Nano 9, 9050–9061 (2015).
Pastoriza-Gallego, M. et al. Dynamics of unfolded protein transport through an aerolysin pore. J. Am. Chem. Soc. 133, 2923–2931 (2011).
Oukhaled, A., Bacri, L., Pastoriza-Gallego, M., Betton, J.-M. & Pelta, J. Sensing proteins through nanopores: fundamental to applications. ACS Chem. Biol. 7, 1935–1949 (2012).
Stefureac, R., Long, Y.-T., Kraatz, H.-B., Howard, P. & Lee, J. S. Transport of alpha-helical peptides through alpha-hemolysin and aerolysin pores. Biochemistry 45, 9172–9179 (2006).
Pastoriza-Gallego, M. et al. Evidence of unfolded protein translocation through a protein nanopore. ACS Nano 8, 11350–11360 (2014).
Wang, H.-Y., Ying, Y.-L., Li, Y., Kraatz, H.-B. & Long, Y.-T. Nanopore analysis of β-amyloid peptide aggregation transition induced by small molecules. Anal. Chem. 83, 1746–1752 (2011).
Sutherland, T. C. et al. Structure of peptides investigated by nanopore analysis. Nano Lett. 4, 1273–1277 (2004).
Meng, H. et al. Nanopore analysis of tethered peptides. J. Pept. Sci. 16, 701–708 (2010).
Mereuta, L. et al. Slowing down single-molecule trafficking through a protein nanopore reveals intermediates for peptide translocation. Sci. Rep. 4, 3885–3885 (2014).
Whittingham, J. L., Edwards, D. J., Antson, A. A., Clarkson, J. M. & Dodson, G. G. Interactions of phenol and m-cresol in the insulin hexamer, and their effect on the association properties of B28 pro -->Asp insulin analogues. Biochemistry 37, 11516–11523 (1998).
Kadima, W. et al. The influence of ionic strength and pH on the aggregation properties of zinc-free insulin studied by static and dynamic laser light scattering. Biopolymers 33, 1643–1657 (1993).
Van Meervelt, V. et al. Real-time conformational changes and controlled orientation of native proteins inside a protein nanoreactor. J. Am. Chem. Soc. 139, 18640–18646 (2017).
Skinner, G. M., van den Hout, M., Broekmans, O., Dekker, C. & Dekker, N. H. Distinguishing single- and double-stranded nucleic acid molecules using solid-state nanopores. Nano Lett. 9, 2953–2960 (2009).
Lin, J., Fabian, M., Sonenberg, N. & Meller, A. Nanopore detachment kinetics of poly(A) binding proteins from RNA molecules reveals the critical role of C-terminus interactions. Biophys. J. 102, 1427–1434 (2012).
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E. & Deamer, D. W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid, and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 (1999).
Bell, N. A. W. et al. DNA origami nanopores. Nano Lett. 12, 512–517 (2012).
Wei, R., Martin, T. G., Rant, U. & Dietz, H. DNA origami gatekeepers for solid-state nanopores. Angew. Chem. Int. Ed. 51, 4864–4867 (2012).
Larkin, J. et al. Slow DNA transport through nanopores in hafnium oxide membranes. ACS Nano 7, 10121–10128 (2013).
Wanunu, M. et al. Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nat. Nanotech. 5, 807–814 (2010).
Acknowledgements
This work was supported by a Bilateral United Kingdom Biotechnology and Biological Sciences Research Council (BBSRC)–United States National Science Foundation (NSF) Lead Agency Pilot Program grant (BB/N018729/1 and NSF-1645671 to A.A.A. and M.W., respectively). We would like to thank Marko Hyvonen, Department of Biochemistry, University of Cambridge (UK), for the expression plasmid for the TPX2 peptide. NanoDSF and negative stained TEM of mutant portal proteins were carried out in the Technology Facility, Department of Biology, University of York (UK).
Author information
Authors and Affiliations
Contributions
A.A.A., M.W. and S.J.G. conceived the study. B.C. and S.J.G. designed the engineered portal protein and the electrical sensing experiments. S.J.G. engineered, purified and biophysically characterized the portal protein variants. B.C. performed the electrical sensing experiments with DNA and protein, and analyzed the data with assistance from M.W. M.M. conducted the cyclodextrin electrical sensing experiments and analyzed the data under supervision of M.W. B.C., M.W. and S.J.G co-wrote the paper with assistance from A.A.A. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cressiot, B., Greive, S.J., Mojtabavi, M. et al. Thermostable virus portal proteins as reprogrammable adapters for solid-state nanopore sensors. Nat Commun 9, 4652 (2018). https://doi.org/10.1038/s41467-018-07116-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-07116-x
This article is cited by
-
Focus on using nanopore technology for societal health, environmental, and energy challenges
Nano Research (2022)
-
Solid-state nanopore sensors
Nature Reviews Materials (2020)
-
Nanopore-based sensing interface for single molecule electrochemistry
Science China Chemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.