Abstract
Culture-independent molecular surveys of plant root microbiomes indicate that soil type generally has a stronger influence on microbial communities than host phylogeny. However, these studies have mostly focussed on model plants and crops. Here, we examine the root microbiomes of multiple plant phyla including lycopods, ferns, gymnosperms, and angiosperms across a soil chronosequence using 16S rRNA gene amplicon profiling. We confirm that soil type is the primary determinant of root-associated bacterial community composition, but also observe a significant correlation with plant phylogeny. A total of 47 bacterial genera are associated with roots relative to bulk soil microbial communities, including well-recognized plant-associated genera such as Bradyrhizobium, Rhizobium, and Burkholderia, and major uncharacterized lineages such as WPS-2, Ellin329, and FW68. We suggest that these taxa collectively constitute an evolutionarily conserved core root microbiome at this site. This lends support to the inference that a core root microbiome has evolved with terrestrial plants over their 400 million year history.
Similar content being viewed by others
Introduction
Plant roots harbor limited microbial diversity relative to soil that surrounds them, and are usually dominated by a small number of bacterial lineages. Surveys of root microbiomes associated with angiosperms such as Arabidopsis thaliana 1, 2, maize3, oak4, barley5, rice6, lettuce7, and sugarcane8 typically reveal Actinobacteria and Proteobacteria as the dominant phyla, suggesting that certain members of these lineages may be consistently enriched in the plant root environment. However, most existing root microbiome studies are of domesticated plants that may not be representative of native plants9. Since plants recruit root microbial communities primarily from the soils they inhabit, soil type is considered one of the key determinants of root microbial community composition10. Plant host phylogeny is a secondary factor influencing root microbial community composition, and the effect size appears considerably smaller than soil type as was demonstrated with Arabidopsis ecotypes1, 2. Surveys comparing root-associated communities in maize, sorghum, and wheat11 (all monocots), as well as Arabidopsis and the related species Cardamine hirsuta 12 (eudicots), report greater variation in root community composition between more distantly related plants. These studies hint at a broader influence of host phylogeny on root microbiome composition than is currently appreciated.
Since host effects are subtle in shaping the root microbial community relative to soil type, we predicted that a stronger host effect would be detected when comparing more distantly related plant taxa. To date, small-subunit ribosomal RNA gene (16S) sequencing-based root community surveys have been conducted mostly on angiosperms with a focus on model plants (e.g., Arabidopsis, poplar) and crops (e.g., wheat, maize, rice, barley, sugarcane, lettuce, grapevine, oat, and pea). By contrast, the root communities of non-angiosperms are poorly characterized; to our knowledge, only a few studies describe the root community composition of non-seed and non-flowering plants13,14,15.
Here, we extend the scope of plant host lineages to non-seed (lycopods and ferns) and seed plant phyla (gymnosperms and angiosperms) through investigation of the root microbiomes of 31 plant species in 25 families and 19 orders. The plant species in our study grow in close proximity to one another along a coastal tropical soil chronosequence that spans ~460,000 years. Sites along this chronosequence span 10 km from the youngest to oldest site and experience negligible differences in climate. The chosen chronosequence has a phylogenetically diverse flora composed of ancient and modern plant lineages, and a considerable overlap of plant species between communities facilitating the goal of distinguishing host and soil determinants of root microbial communities. Our study shows that root bacterial community composition is significantly correlated with host phylogeny despite the stronger effect of soils on these communities. Moreover, a core root microbiome was identified at the study site that comprises both well-known plant-associated taxa and poorly characterized and as yet uncultured taxa.
Results
Cooloola study site
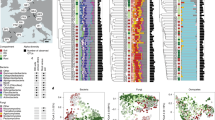
Samples were obtained from a well-characterized chronosequence of coastal sand dunes in the Great Sandy National Park at Cooloola, Queensland, Australia16, 17. The chronosequence consists of six dune systems across an ~ 10 km transect (Fig. 1), each of which harbors phylogenetically diverse flora consisting of lineages that were present on the Australian landmass during Gondwanan times and those whose ancestors dispersed into Australia since it became isolated18, 19. The older dunes have developed into giant podzols over a period of 460,000 years. We chose six plant communities across four dune systems (Fig. 1, Supplementary Fig. 1, Supplementary Table 1) that share component plant species in early succession sclerophyll woodlands, mid-succession forests, and late-succession retrogressive woodland and shrubland17, 19. This setting facilitates comparison of root microbial community similarity and plant phylogenetic distance while controlling for soil type. In addition, the proximity of the chosen sites to one another ensured that any effects of soil type and plant phylogeny on the root microbiome were not confounded by differences attributable to climate. With evidence that plant age can affect root microbial communities20, all sampled plants were mature individuals of perennial or biannual species.
Overview of Cooloola study site. Geographical location of the six plant communities sampled (sites a–f) in the Great Sandy National Park at Cooloola, Queensland, Australia, in relation to dune systems and vegetation types. Site a, open sclerophyll Eucalyptus racemosa early successional woodland; site b, Eucalyptus pilularis tall open moist sclerophyll forest; site c, rainforest (complex notophyll vine forest) with Agathis robusta, Ficus, and Archontophoenix cunninghamiana in fire-sheltered parabolic high dunes; site d, mixed eucalypt conifer open sclerophyll forest with Eucalyptus racemosa, Angophora leiocarpa, and Callitris rhomboidea; site e, retrogression sclerophyll shrubby woodland of Eucalyptus racemosa, Banksia aemula, and Leptospermum species; site f, retrogression Wallum shrubland with Banksia aemula, Xanthorrhoea johnsonii, and a high diversity of heath shrubs. Scale bar represents 1 km
We collected 470 samples (235 root and 235 associated bulk soil) from 31 plant species across six plant communities (Fig. 2). We successfully extracted DNA and amplified 16S rRNA gene amplicon sequences to produce bacterial community profiles for 183 and 225 root and soil samples, respectively. Chimeric sequences were removed and the remaining data were error-corrected, leaving 3,598,535 sequences. These sequences were clustered into 177,758 operational taxonomic units (OTUs) each comprising ≥10 sequences (i.e., no OTUs were represented by less than 10 sequences) at a sequence similarity threshold of 99%, which corresponds approximately to species-level units21. Following taxonomic assignment, chloroplast, mitochondrial, and unassigned sequences were removed. Two approaches were then used to normalize for sequencing depth—the first was a centered log ratio normalization with total sum scaling and the other was sequence rarefication to 1000 reads per sample followed by correction of read counts to adjust for variation in bacterial lineage-specific 16S rRNA gene copy numbers22. Low-abundance OTUs were filtered out from the rarefaction-based OTU table by removing those with less than 0.1% relative abundance in any sample.
A range of soil physicochemical characteristics as well as microbial biomass and activity were determined for five replicate bulk soil samples per plant community. Each bulk soil replicate represented a pool of three to ten soil samples. As expected, chemical characteristics of the upper soil horizon varied between sites; however, pH was relatively uniform only ranging between 4.1 and 4.6, which is in the normal range for ~36% of tropical land area worldwide23. Higher concentrations of metals (aluminum, chromium, iron, potassium, magnesium, manganese, sodium, nickel, strontium, and zinc) were detected in younger soils as these elements coat the silica sand grains that form the original dune substrate but are lost from the upper soil horizons over time (Fig. 3 and Supplementary Table 2). Higher levels of carbon, nitrogen, calcium, phosphorus, and sulphur were detected in rainforest soils, with carbon and nitrogen enrichment as a result of biological processes, and enrichment of other elements the consequence of plants extracting these nutrients from the deeper soil24. The most ancient soils are the most nutrient-depauperate as net nutrient losses occur with repeated wildfires and rainfall leaching nutrients into the deep soil out of reach of roots17, 19 (Fig. 3). Soil microbial biomass and total enzyme activity peaked in the mid succession rainforest soils (sites c and d), but phosphatase activity remained high in the ancient soils, indicating an increased relative microbial investment in phosphorus acquisition (Fig. 3).
Soil chemical characteristics, microbial biomass, total enzyme, and phosphatase activity measured in bulk soils. Values shown are the average of five soil samples from each plant community sampling site, each sample a pool of three to ten individual soils (see Supplementary Table 2 for standard error of the mean and statistical analyses). Color shading represents concentration, biomass, or enzyme activity, and is based on standardized z-scores. Values above the respective means are colored red, values below the mean are colored blue
Bulk soil microbiomes
In total, 17,429 bacterial OTUs were detected in soil samples across the chronosequence at greater than 0.1% relative abundance in at least one sample, of which only 20.5% were shared between all soils, but these accounted for 76.1% of the average relative abundance. The most abundant soil taxa were members of the Alphaproteobacteria, Actinobacteria, and Acidobacteria consistent with previous studies25, 26. Collectively, they represented 79.6% of taxa in each soil based on relative abundance (Supplementary Fig. 2).
To determine the relative contributions of soil chemical characteristics and plant phylogeny to changes in soil microbial community composition, we used permutational multivariate analysis of variance (PERMANOVA). Firstly, we used principal component analysis (PCA) to summarize the variation in elemental composition (C, N, P, K, S, Mg, Mn, Fe, Al, Zn, Na, Cu, Ni, Ba, Ca, Cr, and Sr) between soils (Supplementary Fig. 3a). This analysis captured a combined 76.3% of variation in soil chemical characteristics between soils in the first two axis scores, which were then used as predictor variables in the PERMANOVA model. Secondly, we used PCA to summarize variation in plant phylogenetic relatedness (Supplementary Fig. 3b) as represented by a distance matrix generated from a multiple sequence alignment of plant ribulose-1,5-bisphosphate carboxylase gene (rbcL) sequences (Supplementary Data 1). This analysis captured a combined 83.1% of variation in phylogenetic relatedness between plants in the first two axis scores, which in combination with those representing variation in soil chemical characteristics, were used as predictor variables to explain turnover in soil microbial community composition. Our PERMANOVA model revealed that variation in soil microbial community composition was significantly associated with soil chemical characteristics but not with plant phylogeny (Table 1a). Hierarchical clustering, PCA, and redundancy analysis (RDA) ordination of the same data gave consistent results in that bulk soil communities predominantly clustered by soil type (Supplementary Figs. 4–6), with the composition of rainforest soil communities being distinct from other soils. Alpha diversity metrics indicated that rainforest soil communities were the most phylogenetically diverse, whereas observed species richness and estimated richness (Chao1) were comparable across all soils (Fig. 4).
Alpha diversity metrics of Cooloola root and bulk soil microbial communities across the six sampling sites. See Fig. 1 for number of samples in each group. Box and whisker plots showing a observed species richness, b estimated species richness (chao1), and c Faith’s phylogenetic diversity. White rectangles represent bulk soil communities and grey rectangles represent root communities. The centre line within rectangles represents the median values, and the two ends of the rectangles represent upper and lower quartiles. The upper whisker extends to the highest value within 1.5× the interquartile range above the upper quartile, whereas the lower whisker extends to the lowest value within 1.5× the interquartile range below the lower quartile. Values outside this range are represented by black dots. Total number of root samples is 183 and soil samples is 225
Root-associated microbiomes
Since the diversity of root microbial communities varies along the longitudinal root axis27, 28, we isolated DNA only from root apices (five centimeters to the root tip) as this is the primary site of root exudation29. A total of 15,991 bacterial OTUs were detected in root samples at greater than 0.1% relative abundance in at least one sample. When compared with bulk soil, root communities had consistently lower species richness and diversity1, 2, 5 (observed species richness, Chao1, and Faith’s phylogenetic diversity; p < 0.001, Mann–Whitney U-test). This trend included root bacterial communities in the rainforest despite greater phylogenetic diversity of microbial communities in rainforest soil than the other sites (Fig. 4). Diversity metrics of root communities were also largely comparable between plant orders, except for the eudicot Dilleniales (Hibbertia scandens) with consistently lower scores relative to other eudicots (Supplementary Fig. 7). Between plant phyla, root communities of the basal lycopod lineage scored, albeit not statistically significant (Kruskal–Wallis test), higher estimated species richness and phylogenetic diversity compared to other phyla (Supplementary Fig. 7). It is possible that these higher values reflect a less selected root microbiome in lycopods before the evolution of root communities with more recent plants. However, additional lycopod root samples collected from different sites are required to verify this hypothesis as we identified lycopods only in one of the Cooloola plant communities (Fig. 2).
At high taxonomic ranks (phylum and class), root bacterial communities were similar to each other and to bulk soils, with communities being dominated by Alphaproteobacteria (average 41.7% relative abundance of root-associated community), Actinobacteria (19.1%), and Acidobacteria (17.3%; Supplementary Fig. 2). Despite a lower relative abundance compared to these more dominant lineages, Betaproteobacteria were enriched approximately fourfold in roots (average 5.7% relative abundance in root communities) relative to soil (1.4%), possibly indicating selective enrichment in the root environment1, 2, 6. This gross similarity between roots and their respective bulk soil communities broadly reflects that root-associated communities are enriched subsets of populations predominantly acquired from the surrounding soil microbiome1, 2, 30.
At the OTU level, the effect of soil type and host phylogeny on the root bacterial community composition was discernible. For example, hierarchical clustering (Supplementary Fig. 4) and ordination (Supplementary Figs. 5 and 8) of root bacterial communities showed localized groupings by both factors. To determine the relative contributions of soil chemical characteristics and plant phylogeny to changes in root bacterial community composition, we used PERMANOVA as described above for bulk soils. Root bacterial community composition was significantly associated with both soil chemical characteristics and host phylogeny (Table 1b). In contrast, the composition of bulk soil bacterial communities was strongly associated with soil chemical characteristics only (Table 1a). Further support for an association between host phylogeny and root bacterial community composition was provided by Procrustes analysis (correlation = 0.20, p = 0.02, number of permutations = 3000) and Mantel test (Spearman r = 0.11, p = 0.018) that revealed a small but significant correlation between ordinations summarizing variation in root bacterial community composition and plant phylogenetic distance. These findings suggest that within the Cooloola chronosequence, root bacterial communities have evolved in concert with their hosts. Similar studies comparing root communities between the monocot grasses maize, sorghum, and wheat11, and Arabidopsis species and the closely related Cardamine hirsuta 12 have also implicated host phylogeny as a contributing factor to root community diversification, albeit secondary to other influences, which may include agricultural management, host–microbe, and microbe–microbe interactions31. The significant correlation found in the present study between root-associated bacterial communities and host phylogeny across a wide range of plant species suggests that root microbiomes have evolved with their plant hosts at least since the divergence of lycopods ~400 million years ago. Nonetheless, as soil type has a stronger influence on root community composition, root microbiome surveys across multiple plant phyla should be replicated in separate locations to determine whether the influence of host phylogeny on root community composition is consistent across different geographic conditions.
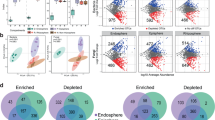
Cooloola core root microbiome
The number of Cooloola core root OTUs varies depending on analysis—369 with indicator species analysis32 (Table 2 and Supplementary Table 3), 302 using a univariate Welch’s t-test implemented in STAMP33 (Supplementary Table 4), and 30 with sparse partial least squares discriminant analysis (sPLSDA) implemented in mixOmics34 (Supplementary Table 5). Nevertheless, core root OTUs from the three analyses completely overlap phylogenetically except for the bacterial genus Methylovirgula indicated in STAMP (Supplementary Fig. 9 and Supplementary Table 6), and represent 47 and 40 classifiable and unclassifiable bacterial genera, respectively. The core root OTUs comprise up to 33.2% of root communities based on relative abundance depending on analysis (Supplementary Tables 3–5), some of which are well-known plant–root-associated bacteria, notably Bradyrhizobium 35, Rhizobium 36, Burkholderia 37, and Azospirillum 38 . Bradyrhizobium and Rhizobium are best known as root-nodulating bacteria of legumes, and supply their hosts with biologically fixed nitrogen35. The high relative abundance of Bradyrhizobium across multiple plant phyla in the present study suggests that their association with non-legumes may be more widespread than previously appreciated39. Non-leguminous plants including Arabidopsis, corn, and tomato respond to Bradyrhizobium nodulation factors using a common molecular mechanism40, suggesting that this association predates the evolution of legumes within the eudicots.
The genera Burkholderia and Azospirillum also contain multiple species of recognized root-associated bacteria41, 42 that have been detected in roots of crops such as lupin43, maize44, 45, and sugarcane46, 47, and are thought to contribute to plant fitness primarily through biological nitrogen fixation41 and phytohormone production42. Other relatively abundant lineages in the core Cooloola root microbiome include Mycobacterium and Rhodoplanes (Table 2), which have been detected in plant roots4, 48, 49, but their ecology and function are unknown. Core genera with cultured representatives that are not well recognized in the context of rhizosphere microbiology include Actinospica, Asticcacaulis, and Salinispora, which have been isolated from roots50, soils51,52,53, or marine sediments54, 55. The core set also contains taxa belonging to as yet unnamed lineages with no or few isolates, including candidate phylum WPS-2, the alphaproteobacterial order Ellin329, and order FW68 in the phylum Armatimonadetes (Supplementary Table 3). Members of these lineages have been detected in soil habitats56, 57, but not specifically associated with plant roots. Soil isolates belonging to the order Ellin329 metabolize xylan, arabinose, rhamnose, and starch58, 59, and are speculated to play a role in plant litter decomposition59. Whether these unfamiliar root-associated taxa are actively recruited into the root microbiome, are root-proximal opportunists feeding on rhizodeposits, or interact only with other root-associated bacteria remains to be determined.
Biogeographical considerations
Biogeography is often an important factor in shaping the composition of root microbiomes1,2,3, 6, 8; thus, it is possible that the Cooloola core root microbiome identified in this study (Table 2 and Supplementary Table 3) is specific to the region, or to the continent of Australia. To assess potential biogeographic variation, we cross-referenced the core Cooloola root taxa with root and associated bulk soil microbiome surveys of plants grown in Australia8 and other countries1, 3, 5, 60. We reanalyzed 16S rRNA gene amplicon data from these studies to predict core OTUs via the indicator species method used in the present study for consistency. Several core taxa were shared across multiple studies including Streptomyces, Mesorhizobium, Agrobacterium, Rhizobium, Sphingomonas, and Rubrivivax (Table 2), suggesting that these taxa may be globally important root-associated bacteria. However, other Cooloola core taxa were not identified in the cross-referenced studies, which may indicate regional differences (localized evolution), although methodological variations between the studies (e.g., DNA extraction method) cannot be ruled out as an important contributing factor to compositional differences. It was also noted that the Cooloola data set shared the greatest number of core root taxa (32 of 60, Table 2 and Supplementary Table 3) with Australian sugarcanes8, possibly reflecting a continental biogeographical signal. From these comparisons we predict that a global core set of plant root microbiota will be considerably more restricted than the list provided in the present study (Table 2 and Supplementary Table 3).
Conclusion
We identified significant correlation between root community composition and host phylogeny in a survey encompassing plant species from multiple plant phyla growing in close proximity. A core root microbiome dominated by a small number of bacterial taxa was identified. These findings suggest that a core root bacterial community was established before the evolution of modern plant lineages, and root-associated bacterial communities have evolved with their plant hosts. By extension, it is likely that core functionality of the root microbiome is also conserved. Independent root and endosphere metagenome studies have reported a shared functionality relating to traits such as bacterial motility, nitrogen metabolism, iron acquisition, and metabolism, and protein secretion systems in the rhizospheres of rice, cucumber, and wheat61, 62. In light of these findings, this study provides a list of bacterial lineages for investigation into their specific plant–microbe interactions including recruitment into the rhizosphere, persistence, function, and turnover, knowledge of which could be used to enhance agricultural crop productivity.
Methods
Study site
An ~10 km transect across a well-characterized coastal dune chronosequence in the Great Sandy National Park (S 25.964, E 153.077) located in Cooloola, south-east Queensland, Australia, was selected as the study site. This location features at least six distinct soil types representing a chronosequence in soil development spanning from young soils several thousand years old to ancient soils ~460,000 years old16. The chronosequence exhibits progressive and retrogressive vegetation succession from which we selected multiple plant species representing diverse lineages of the plant kingdom. Climate was identical across the study site thereby minimizing other environmental influences between samples. The rainforest plant community receives similar rainfall but differs in rarity or absence of fire compared to the other fire-prone sclerophyll plant communities.
Sample collection
Approval for sample collection at the Great Sandy National Park was obtained from the Queensland Government Department of Environment and Heritage Protection (Permit number: WITK09457411). We sampled the chronosequence in March 2013 after summer rains to obtain a snapshot of the root microbial community composition of 31 plant species. Plants were identified morphologically. Smaller plants (~10–30 cm) were uprooted to access the root system for sampling while larger plants were partially excavated to access roots. Corresponding soil samples were collected from soil (top 10 cm) adjacent to the sampled plant. Where possible, at least three replicate root and soil samples for each plant species were collected. Leaf vouchers were also obtained from each plant sampled. Samples were stored on dry ice in the field and then at −20 °C in the laboratory until further processing.
Bulk soil nutrient analyses
Bulk soil samples were pooled into replicates of five according to study site for microbial biomass, activity, and soil chemical compositions. Microbial biomass was measured using chloroform fumigation/extraction followed by a ninhydrin assay for nitrogen content. Microbial activity was assayed by measuring fluorescein diacetate hydrolysis63. Soil moisture content was determined gravimetrically (drying at 105 °C for 48 h). Elemental carbon and nitrogen concentrations were measured by combustion in a Dumas apparatus followed by analysis using a LECO TruSpec analyser. Concentrations of other elements were measured by analyzing microwave-digested samples using a Varian Vista Pro inductively coupled plasma optical emission spectrometer.
Plant phylogeny construction
Leaf samples approximately 2 × 2 cm were cleaned by dipping in 80% ethanol solution for 1 min followed by washing in sterile water. DNA was extracted from cleaned leaf samples using the PowerSoil® DNA Isolation Kit following manufacturer’s instructions. The rbcL gene sequence was PCR amplified using primers rbcLa-F 5′-ATGTCACCACAAACAGAGACTAAAGC-3′ and rbcLa-R 5′-GTAAAATCAAGTCCACCRCG-3′. Thermocycling conditions were: 95 °C for 3 min followed by 32 cycles of 95 °C for 30 s, 53 °C for 30 s, 74 °C for 1 min, and finally 74 °C for 10 min. PCR amplicons were cleaned using Agencourt AMPure XP beads (Beckman Coulter Inc.) and capillary sequenced with both forward and reverse primers to obtain a complete amplicon sequence by alignment in Geneious R664.
Microbial DNA extraction and sequencing
Root tissue up to 3 cm from the root tip was first separated from root samples using a sterile scalpel. Separated root tissues were rinsed with sterile phosphate buffered saline with 0.02% Silwet L-77 surfactant to remove adhering bulk soil particles. DNA was extracted directly from these processed root tissue and soil samples using PowerSoil® DNA Isolation Kits (MO BIO Laboratories, Carlsbad, CA) following manufacturer’s instructions. Extracted DNA was quantified using a Qubit fluorometer with Quant-it dsDNA BR assays (InvitrogenTM) and then normalized to 4 ng/µl using sterile water. Normalized DNA samples were PCR amplified and sequenced using the 454 GS FLX Titanium pyrosequencing platform. Briefly, 16S rRNA genes were PCR-amplified in 50 µl volumes containing 20 ng DNA, 1X PCR buffer, 0.2 mM of each dNTPs, 1.5 mM MgCl2, 0.3 mg bovine serum albumin, 0.02 U Taq DNA polymerase and 0.2 µM each of primers 27F 5′-AGAGTTTGATCMTGGCTCAG-3′ and 519R 5′-GWATTACCGCGGCKGCTG-3′ modified to contain the 454 FLX Titanium Lib L adapters B and A, respectively. The 519R primer contained a barcode sequence between the primer sequence and adapter. A unique barcode was used to amplify DNA from each sample to facilitate sample identification and demultiplexing after sequencing. Thermocycling conditions were: 95 °C for 5 min followed by 30 cycles of 95 °C for 30 s, 55 °C for 45 s, 72 °C for 90 s; and finally 72 °C for 10 min.
Sequence data processing for community composition
Sequence reads were demultiplexed based on their barcode sequences. Adapter, primer and barcode sequences were subsequently removed, and reads were filtered for chimeras using usearch v6.1.544 and corrected for homopolymer errors using Acacia v1.5265. Error-corrected sequences were clustered at 99% sequence identity roughly corresponding to species-level units21 using UCLUST v1.2.22 and cluster representative sequences were assigned taxonomy by BLAST alignment to the Greengenes 16S database66 (August 2013 release). Chloroplast, mitochondria and low abundance OTUs represented by 10 or fewer sequences in all samples were removed. Sampling depth was rarefied to 1000 reads per sample to calculate alpha diversity metrics. The sequence processing procedures described above were performed using QIIME v1.8.067 except for homopolymer error correction using Acacia. Scripts related to the procedures described in this section are provided as Supplementary Software.
Community diversity and indicator species analyses
Alpha diversity metrics including observed species richness, Chao1 and Faith’s phylogenetic diversity were calculated for all samples using QIIME v1.8.0 based on a rarefied sequence depth of 1000 sequences per sample. For beta diversity analyses, a centered log ratio normalization was first applied to non-normalized OTU sequence counts. Differences in microbial community composition were then visualized using PCA and RDA ordination methods implemented in the R statistical software68 vegan package69. The relative effects of soil type and host phylogeny on root and soil bacterial community composition were assessed using PERMANOVA, available in the vegan package, and principal component scores representing soil chemical characteristics and host phylogeny. Briefly, two distance matrices were constructed, one based on soil chemical characteristics and the other based on plant rbcL gene sequence alignments. Soil chemical measurements were first standardized using z-scores, and then principal component scores extracted from PCA performed on the standardized values. Similarly, a distance matrix representing host phylogeny was constructed using EMBOSS70 Distmat v6.6.0 (Jin-Nei gamma distance), and principal component scores extracted from PCA performed on this matrix. The latter matrix was constructed using rbcL gene sequences amplified from leaf tissue collected during sampling or from public databases if amplification was unsuccessful. Correlation between root/soil community composition and host phylogeny was assessed using the Procrustes and Mantel tests available in the vegan package. Cooloola core root OTUs were determined using indicator species analyses32 implemented in the R labdsv package71 on total sum-scaled OTU relative abundances (relative abundance >0.5% in at least one sample) to discriminate between root-associated and soil-associated OTUs. Indicator species analysis was also performed on a rarefaction-normalized OTU table (1000 reads, Supplementary Table 7). The core root community was also assessed using Welch’s t-test in STAMP v2.1.333 and sPLSDA implemented in mixOmics v6.1.135 on centered log ratio-transformed OTU counts. R commands are provided as Supplementary Software.
Root microbiome core taxa comparison with published studies
External root microbiome survey data sets with associated bulk soil profiles were downloaded from public repositories and processed identically to the Cooloola data set. Indicator OTUs were determined using indicator species analysis in the labdsv package comparing total sum-scaled relative abundances of OTUs in rhizosphere and/or root to soil samples to determine core root taxa.
Data availability
The sequence data have been deposited in the NCBI Sequence Read Archive under BioProject accession code PRJNA328519. The authors declare that all other relevant data supporting the findings of the study are available in this article and its Supplementary Information files, or from the corresponding author upon request.
References
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012).
Peiffer, J. A. et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl Acad. Sci. USA 110, 6548–6553 (2013).
Uroz, S., Buée, M., Murat, C., Frey-Klett, P. & Martin, F. Pyrosequencing reveals a contrasted bacterial diversity between oak rhizosphere and surrounding soil. Environ. Microbiol. Rep. 2, 281–288 (2010).
Bulgarelli, D. et al. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17, 392–403 (2015).
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl Acad. Sci. USA 112, E911–E920 (2015).
Schreiter, S. et al. Effect of the soil type on the microbiome in the rhizosphere of field-grown lettuce. Front. Microbiol. 5, 144 (2014).
Yeoh, Y. K. et al. The core root microbiome of sugarcanes cultivated under varying nitrogen fertilizer application. Environ. Microbiol. 18, 1338–1351 (2016).
Pérez-Jaramillo, J. E., Mendes, R. & Raaijmakers, J. M. Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol. Biol. 90, 635–644 (2016).
Bulgarelli, D., Schlaeppi, K., Spaepen, S., van Themaat, E. V. L. & Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 64, 807–838 (2013).
Bouffaud, M. L., Poirier, M. A., Muller, D. & Moënne‐Loccoz, Y. Root microbiome relates to plant host evolution in maize and other Poaceae. Environ. Microbiol. 16, 2804–2814 (2014).
Schlaeppi, K., Dombrowski, N., Oter, R. G., van Themaat, E. V. L. & Schulze-Lefert, P. Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc. Natl Acad. Sci. USA 111, 585–592 (2014).
Valverde, A. et al. Specific microbial communities associate with the rhizosphere of Welwitschia mirabilis, a living fossil. PLoS ONE 11, e0153353 (2016).
Uroz, S. et al. Specific impacts of beech and Norway spruce on the structure and diversity of the rhizosphere and soil microbial communities. Sci. Rep. 6, 27756 (2016).
Rigg, J. L. et al. Variation in soil microbial communities associated with critically endangered Wollemi pine affects fungal, but not bacterial, assembly within seedling roots. Pedobiologia 59, 61–71 (2016).
Tejan-Kella, M. S. et al. Thermoluminescence dating of coastal sand dunes at Cooloola and North Stradbroke Island, Australia. Soil Res. 28, 465–481 (1990).
Jones, A. R., Sanderman, J., Allen, D., Dalal, R. & Schmidt, S. Subtropical giant podzol chronosequence reveals that soil carbon stabilisation is not governed by litter quality. Biogeochemistry 124, 205–217 (2015).
Wardle, D. A., Walker, L. R. & Bardgett, R. D. Ecosystem properties and forest decline in contrasting long-term chronosequences. Science 305, 509–513 (2004).
Peltzer, D. A. et al. Understanding ecosystem retrogression. Ecol. Monograph. 80, 509–529 (2010).
Chaparro, J. M., Badri, D. V. & Vivanco, J. M. Rhizosphere microbiome assemblage is affected by plant development. ISME J. 8, 790–803 (2014).
Stackebrandt, E. & Goebel, B. M. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Evol. Microbiol. 44, 846–849 (1994).
Angly, F. E. et al. CopyRighter: a rapid tool for improving the accuracy of microbial community profiles through lineage-specific gene copy number correction. Microbiome 2, 1 (2014).
Sanchez, P. A. & Logan, T. J. Myths and science about the chemistry and fertility of soils in the tropics. SSSA Spec. Publ. 29, 35–46 (1992).
Jobbágy, E. G. & Jackson, R. B. The uplift of soil nutrients by plants: biogeochemical consequences across scales. Ecology 85, 2380–2389 (2004).
Lauber, C. L., Hamady, M., Knight, R. & Fierer, N. Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl. Environ. Microbiol. 75, 5111–5120 (2009).
Bissett, A. et al. Introducing BASE: the biomes of Australian soil environments soil microbial diversity database. GigaScience 5, 1 (2016).
Marschner, P., Neumann, G., Kania, A., Weiskopf, L. & Lieberei, R. Spatial and temporal dynamics of the microbial community structure in the rhizosphere of cluster roots of white lupin (Lupinus albus L.). Plant Soil. 246, 167–174 (2002).
Dennis, P. G. et al. A novel method for sampling bacteria on plant root and soil surfaces at the microhabitat scale. J. Microbiol. Method 75, 12–18 (2008).
Dennis, P. G., Miller, A. J. & Hirsch, P. R. Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbiol. Ecol. 72, 313–327 (2010).
Hacquard, S. et al. Microbiota and host nutrition across plant and animal kingdoms. Cell Host Microbe 17, 603–616 (2015).
Lareen, A., Burton, F. & Schäfer, P. Plant root-microbe communication in shaping root microbiomes. Plant Mol. Biol. 90, 575–587 (2016).
Dufrêne, M. & Legendre, P. Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol. Monograph. 67, 345–366 (1997).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Rohart, F., Gautier, B., Singh, A. & Lê Cao, K. A. mixOmics: an R package for ‘omics feature selection and multiple data integration. doi: https://doi.org/10.1101/108597. Preprint at http://www.biorxiv.org/content/early/2017/05/05/108597 (2017).
Long, S. R. Rhizobium-legume nodulation: life together in the underground. Cell 56, 203–214 (1989).
van Rhijn, P. & Vanderleyden, J. The Rhizobium-plant symbiosis. Microbiol. Rev. 59, 124–142 (1995).
Compant, S., Nowak, J., Coenye, T., Clément, C. & Barka, E. A. Diversity and occurrence of Burkholderia spp. in the natural environment. FEMS. Microbiol. Rev. 32, 607–626 (2008).
Michiels, K., Vanderleyden, J. & Van Gool, A. Azospirillum—plant root associations: a review. Biol. Fertil. Soils 8, 356–368 (1989).
Lafay, B., Bullier, E. & Burdon, J. J. Bradyrhizobia isolated from root nodules of Parasponia (Ulmaceae) do not constitute a separate coherent lineage. Int. J. Syst. Evol. Microbiol. 56, 1013–1018 (2006).
Liang, Y. et al. Nonlegumes respond to rhizobial Nod factors by suppressing the innate immune response. Science 341, 1384–1387 (2013).
Estrada-De Los Santos, P., Bustillos-Cristales, R. & Caballero-Mellado, J. Burkholderia, a genus rich in plant-associated nitrogen fixers with wide environmental and geographic distribution. Appl. Environ. Microbiol. 67, 2790–2798 (2001).
Bashan, Y., Holguin, G. & de-Bashan, L. E. Azospirillum-plant relationships: physiological, molecular, agricultural, and environmental advances (1997–2003). Can. J. Microbiol. 50, 521–577 (2004).
Weisskopf, L., Heller, S. & Eberl, L. Burkholderia species are major inhabitants of white lupin cluster roots. Appl. Environ. Microbiol. 77, 7715–7720 (2011).
de Salamone, I. G., Döbereiner, J., Urquiaga, S. & Boddey, R. M. Biological nitrogen fixation in Azospirillum strain-maize genotype associations as evaluated by the 15N isotope dilution technique. Biol. Fertil. Soils 23, 249–256 (1996).
Dalmastri, C., Chiarini, L., Cantale, C., Bevivino, A. & Tabacchioni, S. Soil type and maize cultivar affect the genetic diversity of maize root–associated Burkholderia cepacia populations. Microbiol. Ecol. 38, 273–284 (1999).
Tejera, N., Lluch, C., Martinez-Toledo, M. V. & Gonzalez-Lopez, J. Isolation and characterization of Azotobacter and Azospirillum strains from the sugarcane rhizosphere. Plant Soil 270, 223–232 (2005).
Paungfoo‐Lonhienne, C. et al. A new species of Burkholderia isolated from sugarcane roots promotes plant growth. Microb. Biotechnol. 7, 142–154 (2014).
Child, R. et al. Polycyclic aromatic hydrocarbon-degrading Mycobacterium isolates: their association with plant roots. Appl. Microbiol. Biotechnol. 75, 655–663 (2007).
Srinivas, A., Sasikala, C. & Ramana, C. Rhodoplanes oryzae sp. nov., a phototrophic alphaproteobacterium isolated from the rhizosphere soil of paddy. Int. J. Syst. Evol. Microbiol. 64, 2198–2203 (2014).
Zhu, L. et al. Asticcacaulis endophyticus sp. nov., a prosthecate bacterium isolated from the root of Geum aleppicum. Int. J. Syst. Evol. Microbiol. 64, 3964–3969 (2014).
Golinska, P., Zucchi, T. D., Silva, L., Dahm, H. & Goodfellow, M. Actinospica durhamensis sp. nov., isolated from a spruce forest soil. Antonie Van Leeuwenhoek 108, 435–442 (2015).
Kim, S., Gong, G., Park, T. H. & Um, Y. Asticcacaulis solisilvae sp. nov., isolated from forest soil. Int. J. Syst. Evol. Microbiol. 63, 3829–3834 (2013).
Cavaletti, L. et al. Actinospica robiniae gen. nov., sp. nov. and Actinospica acidiphila sp. nov.: proposal for Actinospicaceae fam. nov. and Catenulisporinae subord. nov. in the order Actinomycetales. Int. J. Syst. Evol. Microbiol. 56, 1747–1753 (2006).
Ahmed, L. et al. Salinispora pacifica sp. nov., an actinomycete from marine sediments. Antonie Van Leeuwenhoek 103, 1069–1078 (2013).
Maldonado, L. A. et al. Salinispora arenicola gen. nov., sp. nov. and Salinispora tropica sp. nov., obligate marine actinomycetes belonging to the family Micromonosporaceae. Int. J. Syst. Evol. Microbiol. 55, 1759–1766 (2005).
Nogales, B. et al. Combined use of 16S ribosomal DNA and 16S rRNA to study the bacterial community of polychlorinated biphenyl-polluted soil. Appl. Environ. Microbiol. 67, 1874–1884 (2001).
Nemergut, D. R. et al. The effects of chronic nitrogen fertilization on alpine tundra soil microbial communities: implications for carbon and nitrogen cycling. Environ. Microbiol. 10, 3093–3105 (2008).
Ueki, A. et al. Rhizomicrobium palustre gen. nov., sp. nov., a facultatively anaerobic, fermentative stalked bacterium in the class Alphaproteobacteria isolated from rice plant roots. J. Gen. Appl. Microbiol. 56, 193–203 (2010).
Harbison, A. B., Carson, M. A., Lamit, L. J., Basiliko, N. & Bräuer, S. L. A novel isolate and widespread abundance of the candidate alphaproteobacterial order (Ellin 329), in southern Appalachian peatlands. FEMS Microbiol. Lett. 363, fnw151 (2016).
Zarraonaindia, I. et al. The soil microbiome influences grapevine-associated microbiota. mBio 6, e02527–14 (2015).
Sessitsch, A. et al. Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol. Plant Microbe Interact. 25, 28–36 (2012).
Ofek-Lalzar, M. et al. Niche and host-associated functional signatures of the root surface microbiome. Nat. Commun. 5, 4950 (2014).
Schnürer, J. & Rosswall, T. Fluorescein diacetate hydrolysis as a measure of total microbial activity in soil and litter. Appl. Environ. Microbiol. 43, 1256–1261 (1982).
Kearse, M. et al. Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Bragg, L., Stone, G., Imelfort, M., Hugenholtz, P. & Tyson, G. W. Fast, accurate error-correction of amplicon pyrosequences using Acacia. Nat. Methods 9, 425–426 (2012).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2016).
Oksanen, J. et al. Vegan: community ecology package. https://cran.r-project.org/web/packages/vegan/index.html (2017).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Roberts, D. W. Labdsv: ordination and multivariate analysis for ecology. https://cran.r-project.org/web/packages/labdsv/index.html (2016).
Acknowledgements
We thank Andrew R. Jones for assistance with landscape and dune system classification, Lyn Cook for advice on plant phylogeny and terminology, Maren Westermann, Harshi Gamage, Jitka Kochanek, Scott Buckley, and Deng Yan for processing root samples in the laboratory, Deng Yan also for assistance with sample collection, and David Wood for assistance with statistical analyses. This research was supported through access to facilities managed by Bioplatforms Australia and funded by the Australian Government National Collaborative Research Infrastructure Strategy and Education Investment Fund Super Science Initiative. It was also supported by a Queensland Smart State Co-investment Fund awarded to M.A.R., P.H., and S.S. Y.K.Y. was supported by a University of Queensland International PhD Scholarship.
Author information
Authors and Affiliations
Contributions
Y.K.Y., P.G.D., C.P.-L., S.S., and P.H. designed the experiment. L.W. identified vegetation types and plants at the study site. Y.K.Y., L.W., S.S., and P.H. collected samples, R.B. performed soil microbial and nutrient measurements, and Y.K.Y. and C.P.L. prepared samples for DNA sequencing. Y.K.Y., P.G.D., and P.H. analyzed the data. Y.K.Y., S.S., and P.H. wrote the manuscript, and all authors reviewed drafts of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yeoh, Y., Dennis, P.G., Paungfoo-Lonhienne, C. et al. Evolutionary conservation of a core root microbiome across plant phyla along a tropical soil chronosequence. Nat Commun 8, 215 (2017). https://doi.org/10.1038/s41467-017-00262-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-017-00262-8
This article is cited by
-
Unraveling the interplay between root exudates, microbiota, and rhizosheath formation in pearl millet
Microbiome (2024)
-
Plant effects on microbiome composition are constrained by environmental conditions in a successional grassland
Environmental Microbiome (2024)
-
Effects of core soil microbial taxa on soil carbon source utilization under different long-term fertilization treatments in Ultisol
Soil Ecology Letters (2024)
-
Influences of plant organ, genotype, and cultivation site on the endophytic bacteriome of maize (Zea mays L.) in the semi-arid region of Pernambuco, Brazil
Brazilian Journal of Microbiology (2024)
-
Metagenomic analysis of soybean endosphere microbiome to reveal signatures of microbes for health and disease
Journal of Genetic Engineering and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.