Abstract
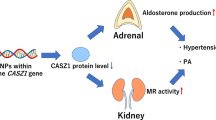
Aldosterone is a biological ligand for mineralocorticoid receptor (MR) that elevates blood pressure by promoting sodium reabsorption in the kidneys. However, the molecular mechanisms of aldosterone-MR-mediated transcription and the role of this transcription in hypertension remain largely unknown. In this study, we aimed to identify novel MR coregulators and elucidate one of the molecular mechanisms of hypertension. We purified MR-interacting factors from HEK293F cells stably expressing FLAG-MR through a biochemical approach and identified the zinc finger protein castor homolog 1 isoform b (CASZ1b) as a candidate novel MR coregulator via liquid chromatography—tandem mass spectrometry analysis. The CASZ1 gene has been implicated in hypertension in genome-wide single-nucleotide polymorphism studies, but its role in the development of hypertension remains unclear. We found that CASZ1b colocalized with MR in the kidneys and interacted with MR in an aldosterone-dependent manner. In luciferase assays using HEK293F cells, overexpression of CASZ1b reduced aldosterone-dependent MR transcriptional activity by ~50%. In contrast, knockdown of CASZ1b via RNA interference increased the expression levels of the aldosterone-induced MR target genes epithelial Na+ channel-α (ENaCα) and serum/glucocorticoid regulated kinase 1 (SGK1) by approximately twofold and 2.3-fold, respectively. Upon aldosterone-MR binding, CASZ1b interacted with MR and formed a protein complex with nucleosome remodeling deacetylase (Mi-2/NuRD), a corepressor complex with chromatin remodeling and histone deacetylation activity, which suppressed ENaCα and SGK1. These findings reveal a critical role of CASZ1b in regulating MR-mediated transcriptional activity and provide new insights into the pathophysiology of hypertension.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Arriza JL, Weinberger C, Cerelli G, Glaser TM, Handelin BL, Housman DE, et al. Cloning of human mineralocorticoid receptor complementary DNA: structural and functional kinship with the glucocorticoid receptor. Science. 1987;237:268–75.
Le Menuet D, Viengchareun S, Muffat-Joly M, Zennaro MC, Lombès M. Expression and function of the human mineralocorticoid receptor: lessons from transgenic mouse models. Mol Cell Endocrinol. 2004;217:127–36.
Valinsky WC, Touyz RM, Shrier A. Aldosterone, SGK1, and ion channels in the kidney. Clin Sci (Lond). 2018;132:173–83.
Riepe FG. Clinical and molecular features of type 1 pseudohypoaldosteronism. Horm Res. 2009;72:1–9.
Mubarik A, Anastasopoulou C, Riahi S, Aeddula NR Liddle Syndrome. StatPearls. StatPearls Publishing Copyright © 2020, StatPearls Publishing LLC.: Treasure Island (FL); 2020.
Pitt B, Zannad F, Remme WJ, Cody R, Castaigne A, Perez A, et al. The effect of spironolactone on morbidity and mortality in patients with severe heart failure. Randomized Aldactone Evaluation Study Investigators. N Engl J Med. 1999;341:709–17.
Pitt B, Remme W, Zannad F, Neaton J, Martinez F, Roniker B, et al. Eplerenone, a selective aldosterone blocker, in patients with left ventricular dysfunction after myocardial infarction. N Engl J Med. 2003;348:1309–21.
Yokota K, Shibata H, Kurihara I, Kobayashi S, Suda N, Murai-Takeda A, et al. Coactivation of the N-terminal transactivation of mineralocorticoid receptor by Ubc9. J Biol Chem. 2007;282:1998–2010.
Zhang W, Xia X, Reisenauer MR, Rieg T, Lang F, Kuhl D, et al. Aldosterone-induced Sgk1 relieves Dot1a-Af9-mediated transcriptional repression of epithelial Na+ channel alpha. J Clin Investig. 2007;117:773–83.
Burrello J, Monticone S, Buffolo F, Tetti M, Veglio F, Williams TA, et al. Is There a Role for Genomics in the Management of Hypertension? Int J Mol Sci. 2017;18:1131.
Levy D, Ehret GB, Rice K, Verwoert GC, Launer LJ, Dehghan A, et al. Genome-wide association study of blood pressure and hypertension. Nat Genet. 2009;41:677–87.
Takeuchi F, Isono M, Katsuya T, Yamamoto K, Yokota M, Sugiyama T et al. Blood pressure and hypertension are associated with 7 loci in the Japanese population. Circulation. 121:2302–9.
Ho JE, Levy D, Rose L, Johnson AD, Ridker PM, Chasman DI. Discovery and replication of novel blood pressure genetic loci in the Women’s Genome Health Study. J Hypertens. 2011;29:62–9.
Kato N, Takeuchi F, Tabara Y, Kelly TN, Go MJ, Sim X, et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat Genet. 2011;43:531–8.
Nakamura T, Kurihara I, Kobayashi S, Yokota K, Murai-Takeda A, Mitsuishi Y, et al. Intestinal mineralocorticoid receptor contributes to epithelial sodium channel-mediated intestinal sodium absorption and blood pressure regulation. J Am Heart Assoc. 2018;7:e008259.
Fujita N, Jaye DL, Kajita M, Geigerman C, Moreno CS, Wade PA. MTA3, a Mi-2/NuRD complex subunit, regulates an invasive growth pathway in breast cancer. Cell. 2003;113:207–19.
Metivier R, Penot G, Hubner MR, Reid G, Brand H, Kos M, et al. Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter. Cell. 2003;115:751–63.
Tong JK, Hassig CA, Schnitzler GR, Kingston RE, Schreiber SL. Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature. 1998;395:917–21.
Xue Y, Wong J, Moreno GT, Young MK, Cote J, Wang WNURD. A novel complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol Cell. 1998;2:851–61.
Liu Z, Lam N, Thiele CJ. Zinc finger transcription factor CASZ1 interacts with histones, DNA repair proteins and recruits NuRD complex to regulate gene transcription. Oncotarget. 2015;6:27628–40.
Liu Z, Lam N, Wang E, Virden RA, Pawel B, Attiyeh EF, et al. Identification of CASZ1 NES reveals potential mechanisms for loss of CASZ1 tumor suppressor activity in neuroblastoma. Oncogene. 2017;36:97–109.
Liu Z, Naranjo A, Thiele CJ. CASZ1b, the short isoform of CASZ1 gene, coexpresses with CASZ1a during neurogenesis and suppresses neuroblastoma cell growth. PLoS ONE. 2011;6:e18557.
Liu Z, Yang X, Li Z, McMahon C, Sizer C, Barenboim-Stapleton L, et al. CASZ1, a candidate tumor-suppressor gene, suppresses neuroblastoma tumor growth through reprogramming gene expression. Cell Death Differ. 2011;18:1174–83.
Wu YY, Chang CL, Chuang YJ, Wu JE, Tung CH, Chen YC, et al. CASZ1 is a novel promoter of metastasis in ovarian cancer. Am J Cancer Res. 2016;6:1253–70.
Amin NM, Gibbs D, Conlon FL. Differential regulation of CASZ1 protein expression during cardiac and skeletal muscle development. Dev Dyn. 2014;243:948–56.
Dorr KM, Amin NM, Kuchenbrod LM, Labiner H, Charpentier MS, Pevny LH, et al. Casz1 is required for cardiomyocyte G1-to-S phase progression during mammalian cardiac development. Development. 2015;142:2037–47.
Guo J, Li Z, Hao C, Guo R, Hu X, Qian S, et al. A novel de novo CASZ1 heterozygous frameshift variant causes dilated cardiomyopathy and left ventricular noncompaction cardiomyopathy. Mol Genet Genom Med. 2019;7:e828.
Huang RT, Xue S, Wang J, Gu JY, Xu JH, Li YJ, et al. CASZ1 loss-of-function mutation associated with congenital heart disease. Gene. 2016;595:62–8.
Liu Z, Li W, Ma X, Ding N, Spallotta F, Southon E, et al. Essential role of the zinc finger transcription factor Casz1 for mammalian cardiac morphogenesis and development. J Biol Chem. 2014;289:29801–16.
Qiu XB, Qu XK, Li RG, Liu H, Xu YJ, Zhang M, et al. CASZ1 loss-of-function mutation contributes to familial dilated cardiomyopathy. Clin Chem Lab Med. 2017;55:1417–25.
Kim B, Jung M, Moon KC. The Prognostic Significance of Protein Expression of CASZ1 in Clear Cell Renal Cell Carcinoma. Dis Markers. 2019;2019:1342161.
Johnsson A, Durand-Dubief M, Xue-Franzen Y, Ronnerblad M, Ekwall K, Wright A. HAT-HDAC interplay modulates global histone H3K14 acetylation in gene-coding regions during stress. EMBO Rep. 2009;10:1009–14.
Kurdistani SK, Robyr D, Tavazoie S, Grunstein M. Genome-wide binding map of the histone deacetylase Rpd3 in yeast. Nat Genet. 2002;31:248–54.
Peserico A, Simone C. Physical and functional HAT/HDAC interplay regulates protein acetylation balance. J Biomed Biotechnol. 2011;2011:371832.
Shahbazian MD, Grunstein M. Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem. 2007;76:75–100.
Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell. 2009;138:1019–31.
Shang Y, Hu X, DiRenzo J, Lazar MA, Brown M. Cofactor dynamics and sufficiency in estrogen receptor-regulated transcription. Cell. 2000;103:843–52.
Acknowledgements
We express our deep appreciation for all the help provided by Dr S Kato. We would like to thank Editage (www.editage.com) for English language editing. This work was supported by JSPS KAKENHI grant number 17K09734 (to KY), the Keio Gijuku Academic Development Funds (to KY and IK), the Japan Foundation for Applied Enzymology (to KY), the Smoking Research Foundation (to KY and IK), Eli Lilly (to KY), the Takeda Science Foundation (to IK), and the Yamaguchi Endocrine Research Foundation (to IK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All animal procedures were approved by the institutional review board of the Animal Care and Use Committee and were conducted in compliance with the animal experimentation guidelines of Keio University School of Medicine.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yokota, K., Shibata, H., Kurihara, I. et al. CASZ1b is a novel transcriptional corepressor of mineralocorticoid receptor. Hypertens Res 44, 407–416 (2021). https://doi.org/10.1038/s41440-020-00562-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41440-020-00562-5
Keywords
This article is cited by
-
Role of the CASZ1 transcription factor in tissue development and disease
European Journal of Medical Research (2023)
-
CASZ1: a promising factor modulating aldosterone biosynthesis and mineralocorticoid receptor activity
Hypertension Research (2023)
-
Mechanisms of mineralocorticoid receptor-associated hypertension in diabetes mellitus: the role of O-GlcNAc modification
Hypertension Research (2023)
-
Lysine-specific demethylase 1 as a corepressor of mineralocorticoid receptor
Hypertension Research (2022)
-
Annual reports on hypertension research 2020
Hypertension Research (2022)