Abstract
Recently, altered PLP1 splicing was confirmed as a genetic cause of hypomyelination of early myelinating structures (HEMS). A novel deep intronic deletion in intron 3 of PLP1 (NM_000533.5: c.453+59_+259del) was identified, and an in vitro minigene assay detected abnormal splicing patterns. However, the clinical and radiological findings of the patient were compatible with a severe phenotype of Pelizaeus-Merzbacher disease rather than HEMS, which may be due to undetected abnormal PLP1 splicing.
Similar content being viewed by others
Pelizaeus-Merzbacher disease (PMD: MIM #312080) is one of the major types of hypomyelinating leukodystrophy1. The gene responsible for this disease is the proteolipid protein 1 gene (PLP1), which is located on Xq222. Thus, PMD is recognized as an X-linked recessive disorder. Approximately two-thirds of PMD patients show microduplications, including in PLP1, and the other patients show nucleotide alterations in PLP13. Previously, many variants have been reported as pathogenic. Although missense variants are the major type of variants, a number of splicing variants have also been reported4,5,6,7. In particular, variants in intron 3 (IVS3) have been detected to cause unique radiological findings recognized as hypomyelination of early myelinating structures (HEMS)8,9. Here, we report a novel intronic variant identified in a patient with typical PMD features rather than HEMS.
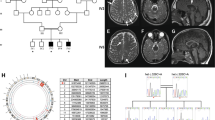
At present, the male patient is 4 years and 6 months old. The boy was born at full term with a birth weight of 2886 g, length of 51.0 cm, and occipitofrontal circumference (OFC) of 32.0 cm. Although there was no remarkable episode during early infancy, pendular nystagmus was noted at 3 months of age. He has never achieved head control and is still bed ridden. There is no verbal communication. There is no stridor, and he can swallow paste food. Owing to the acceleration of the deep tendon reflex, spastic paraplegia was diagnosed. Brain magnetic resonance imaging (MRI) performed at 2 years and 9 months showed diffuse high intensity of the white matter in T2-weighted images in association with the maturation of myelination in the corpus callosum, indicating the typical hypomyelination pattern observed in patients with PMD (Fig. 1A and Supplemental Fig. S1). There were no other abnormalities, including no dilatation of the lateral ventricles. From these findings, PMD was suspected. At present, his height is 93 cm (−2.7 SD), his weight is 11.4 kg (−2.5 SD), and his OFC is 48.5 cm (−1.2 SD), indicating growth deficiency but no microcephaly. Conventional FISH analysis targeting PLP1 did not demonstrate PLP1 duplication. Thus, a comprehensive analysis was planned.
T1- (A) and T2- (B) weighted brain magnetic resonance imaging examined at 2 years of age shows T2-high intensity in the deep white matter. A normal myelination pattern is shown in the corpus callosum. There is no structural abnormality. C Genomic PCR for exons 3–4 of PLP1. In comparison with the normal control (N), an abnormal short band is shown in the patient (P). The mother (M) exhibits both bands, indicating that she is a heterozygous carrier of this variant. L; Gene Ladder 100 (Nippon Gene, Tokyo, Japan).
This study was performed in accordance with the Declaration of Helsinki. We obtained permission from the ethics committee of the institution. The family of this patient provided written informed consent after genetic counseling. Blood samples were obtained from the patient and his parents. Subsequently, genomic DNA was extracted using the standard method, and whole-exome sequencing using trio samples, including parental samples, was performed as described previously10. However, no possible candidate variant related to leukodystrophy was identified. The diagnostic strategy was then reconsidered. Chromosomal microarray testing was performed, and no pathogenic copy number aberration was identified. Further, all coding exons of PLP1 were planned to be reanalyzed by standard PCR-Sanger sequencing. In this process, we could not obtain a PCR amplicon of exon 3, although the other six exons were successfully amplified. Next, we used primers for neighboring exons, including exon 3 forward and exon 4 reverse primers. As a result, a shorter PCR band was confirmed by agarose gel electrophoresis (Fig. 1B). His mother showed both normal and short bands, suggesting that she was a carrier of this PLP1 variant. Sanger sequencing using a PCR amplicon of the short band confirmed a 201-bp deletion in IVS3 59 bp from the exon/intron boundary (NM_000533.5: c.453+59_+259del). Retrospective analysis of the data obtained through whole-exome sequencing showed that there was no mapped read in the identified deletion region, indicating the nullisomy of this region (Supplemental Fig. S2).
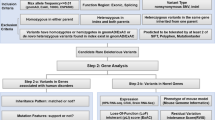
To confirm whether this deletion may cause splicing abnormalities, RNA expression from EB virus-transformed leukocytes was analyzed by nested RT-PCR according to a previously reported method11. However, we could not obtain a PCR amplicon due to the insufficient expression level of PLP1 in the cells. Thus, an in vitro assay was tried as an alternative method. For DNA cloning of the genomic region from IVS3 (−110) to IVS4 (+88) of PLP1, a set of primers (PLP1_insF; TAACTCGAGTAGCCTTGTTAAGGTGCTCGCT, Xba1_PLP1_insR; ATATCTAGACACCACCCTCCTTACACTAAGA) including the restricted enzyme sites Xho1 and Xba1 (indicated by underbars) was designed. Samples from a normal control with the wild-type allele and the genomic DNA of the present patient were used as templates. Then, two PCR amplicons of 1700 bp and 1499 bp were obtained. These amplicons underwent Xho1 and Xba1 restriction digestion and were cloned into the pET 01 Exontrap vector (MoBiTec GmbH, Gottingen, Germany) (Fig. 2A).
A Physical map of the vector used in this study. A partial sequence of PLP1 was cloned into multiple cloning sites. B In comparison with the control, QIAxcel semiquantitative RT-PCR shows 67.0% reduced expression of PLP1 in addition to aberrant large products. C Sanger sequencing confirms intron 3 (IVS3) retention in mRNA.
The successfully constructed vectors were transfected into COS7 cells by lipofection. After 24 h of incubation, total RNA was extracted, and RNA expression from mini-PLP1 was examined by RT-PCR. Two different isoforms, PLP1 and DM20, are expressed from PLP1. DM20 mRNA does not include exon 3b, which is included in PLP1 mRNA. Through the use of the primers designed in the vector region, only DM20 was detected, and PLP1 expression could not be detected as previously reported12. Thus, we focused only on PLP1. For the analysis of PLP1 splicing, a 5ʹ RT-PCR primer (PLP_isoF; GAGCGGGTGTGTCATTGTTT) was designed in an exon 3b-specific region. On the other hand, a 3ʹ RT-PCR primer (PLP_isoR; CTCCACCCAGCTCCAGTTGT) was used in the 3ʹ exon region in the vector. RT-PCR amplicons were analyzed by capillary electrophoresis using a QIAxcel Advanced System (Qiagen, Venlo, The Netherlands). The normal splicing pattern was confirmed in the wild type. On the other hand, the deletion mutant showed a large RT-PCR product in addition to the wild-type product and multiple faint bands (Fig. 2B). The expression level of wild-type RNA was reduced by 67.0%. Sanger sequencing of the large RT-PCR product confirmed that the full sequence of IVS3 was included between exon 3b and exon 4, suggesting that this intron was not normally spliced out (Fig. 2C). Because wild-type IVS3 is 1071 bp in length, the retained intron in this patient is considered to be 870 bp in length (201 bp shorter than wild type) and would insert nine amino acids and a premature termination codon [p.(Lys151_Phe152insValIleIleLeuArgLieLeuTrpGlinTer)]. Thus, the expression of abnormal mRNA related to the large RT-PCR product, which includes this retained intron, would be prevented by nonsense-mediated decay.
Previously, many splicing mutations have been reported4,5,6,7. In the splicing donor site of IVS3, three variants were reported in 200013. Two of them affected the consensus donor site (IVS3+2T>C and IVS3+4A>G). The remaining deletion was a 19-bp deletion in the enhancer region (IVS3+28_+46del), which reduced the expression of PLP1 but not DM20 (ref. 14). This unbalanced expression of PLP1 and DM20 caused a milder phenotype of PMD. In 2015, mutations in exon 3B and IVS3 were identified in patients with HEMS9, which is characterized by unique hypomyelination patterns specifically pronounced in early myelinating structures, including alternating T2 hyperintense–hypointense–hyperintense stripes in the posterior limb of the internal capsule8. The identified mutations in HEMS are predicated to cause abnormal splicing in PLP1 but not in DM20. Thus, unbalanced expression of PLP1 and DM20 is also considered the pathogenesis of HEMS. Because all patients with HEMS are able to achieve unsupported walking8, the clinical features are milder than those with typical PMD.
In this study, a 201-bp deletion (IVS3+59_+259del) was identified in the region neighboring the abovementioned 19-bp deletion (IVS3+28_+46del). Furthermore, this 201-bp deletion removed a 5ʹ-long-distance interaction (LDI) structure15, in which two noncoding mutations have been identified in patients with HEMS9. In this study, an in vitro mini-PLP1 gene assay detected abnormal splicing patterns similar to HEMS. However, the clinical features of the present patient were more severe than those of the patients with HEMS, and the radiological findings were also different from those of patients with HEMS. On the basis of these points, we believe that the previously undetected underlying abnormal PLP1 splicing is the cause of the severe clinical features in this patient. The limitation of this study is that the mini-PLP1 gene cloned into the vector contained only exons 3 and 4. Thus, some other splicing abnormalities may have been missed.
HGV database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at https://doi.org/10.6084/m9.figshare.hgv.2984.
References
Inoue, K. PLP1-related inherited dysmyelinating disorders: Pelizaeus-Merzbacher disease and spastic paraplegia type 2. Neurogenetics 6, 1–16 (2005).
Shimojima, K. et al. Comprehensive genetic analyses of PLP1 in patients with Pelizaeus-Merzbacher disease applied by array-CGH and fiber-FISH analyses identified new mutations and variable sizes of duplications. Brain Dev. 32, 171–179 (2010).
Yamamoto, T. & Shimojima, K. Pelizaeus-Merzbacher disease as a chromosomal disorder. Congenit. Anom. (Kyoto) 53, 3–8 (2013).
Bonnet-Dupeyron, M. N. et al. PLP1 splicing abnormalities identified in Pelizaeus-Merzbacher disease and SPG2 fibroblasts are associated with different types of mutations. Hum. Mutat. 29, 1028–1036 (2008).
Wang, E. et al. Deletion of a splicing enhancer disrupts PLP1/DM20 ratio and myelin stability. Exp. Neurol. 214, 322–330 (2008).
Omata, T. et al. A splicing mutation of proteolipid protein 1 in Pelizaeus-Merzbacher disease. Brain Dev. 38, 581–584 (2016).
Kubota, K. et al. Brain magnetic resonance imaging findings and auditory brainstem response in a child with spastic paraplegia 2 due to a PLP1 splice site mutation. Brain Dev. 37, 158–162 (2015).
Steenweg, M. E. et al. Novel hypomyelinating leukoencephalopathy affecting early myelinating structures. Arch. Neurol. 69, 125–128 (2012).
Kevelam, S. H. et al. Altered PLP1 splicing causes hypomyelination of early myelinating structures. Ann. Clin. Transl. Neurol. 2, 648–661 (2015).
Sasaki, H. et al. Definitive diagnosis of mandibular hypoplasia, deafness, progeroid features and lipodystrophy (MDPL) syndrome caused by a recurrent de novo mutation in the POLD1 gene. Endocr. J. 65, 227–238 (2018).
Yamamoto-Shimojima, K. et al. Elucidation of the pathogenic mechanism and potential treatment strategy for a female patient with spastic paraplegia derived from a single-nucleotide deletion in PLP1. J. Hum. Genet. 64, 665–671 (2019).
Regis, S. et al. Restoration of the normal splicing pattern of the PLP1 gene by means of an antisense oligonucleotide directed against an exonic mutation. PLoS ONE 8, e73633 (2013).
Hobson, G. M. et al. Mutations in noncoding regions of the proteolipid protein gene in Pelizaeus-Merzbacher disease. Neurology 55, 1089–1096 (2000).
Hobson, G. M. et al. A PLP splicing abnormality is associated with an unusual presentation of PMD. Ann. Neurol. 52, 477–488 (2002).
Taube, J. R. et al. PMD patient mutations reveal a long-distance intronic interaction that regulates PLP1/DM20 alternative splicing. Hum. Mol. Genet. 23, 5464–5478 (2014).
Vaz-Drago, R., Custódio, N. & Carmo-Fonseca, M. Deep intronic mutations and human disease. Hum. Genet. 136, 1093–1111 (2017).
Dufner-Almeida, L. G., do Carmo, R. T., Masotti, C. & Haddad, L. A. Understanding human DNA variants affecting pre-mRNA splicing in the NGS era. Adv. Genet. 103, 39–90 (2019).
Acknowledgements
We appreciate the cooperation of the patient and his parents in this study. This work was supported by the Initiative on Rare and Undiagnosed Diseases (Grant Number 20ek0109301) from the Japan Agency for Medical Research and Development (AMED). This work was partially supported by Grants-in-Aid for Scientific Research from Health Labor Sciences Research Grants from the Ministry of Health, Labor and Welfare, Japan, and JSPS KAKENHI (Grant Number 18K07803).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamamoto-Shimojima, K., Akagawa, H., Yanagi, K. et al. Deep intronic deletion in intron 3 of PLP1 is associated with a severe phenotype of Pelizaeus-Merzbacher disease. Hum Genome Var 8, 14 (2021). https://doi.org/10.1038/s41439-021-00144-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-021-00144-y