Abstract
Members of the genus Paeonia, which consists of globally renowned ornamentals and traditional medicinal plants with a rich history spanning over 1500 years, are widely distributed throughout the Northern Hemisphere. Since 1900, over 2200 new horticultural Paeonia cultivars have been created by the discovery and breeding of wild species. However, information pertaining to Paeonia breeding is considerably fragmented, with fundamental gaps in knowledge, creating a bottleneck in effective breeding strategies. This review systematically introduces Paeonia germplasm resources, including wild species and cultivars, summarizes the breeding strategy and results of each Paeonia cultivar group, and focuses on recent progress in the isolation and functional characterization of structural and regulatory genes related to important horticultural traits. Perspectives pertaining to the resource protection and utilization, breeding and industrialization of Paeonia in the future are also briefly discussed.
Similar content being viewed by others
Introduction
The genus Paeonia was placed in the Ranunculaceae before the 20th century. According to the characteristic arrangement of stamens, Worsdell1 separated Paeonia from rose in the Ranunculaceae and moved it to the Paeoniaceae. Paeonia species or cultivars are shrubs or perennial herbs. Wild Paeonia species are mainly distributed in temperate regions of the Northern Hemisphere, although individual species extend into cold temperate regions, with latitudes ranging from 24.4 to 66.5°N2,3. Wild woody Paeonia species are distributed only in China, whereas herbaceous species are widely distributed throughout Central and East Asia, the Himalayas, western North America (USA and Canada), and the Mediterranean4. These widely different habitats have led to very rich genetic variation in Paeonia.
Paeoniflorin and paeonol are unique chemical components in the roots of Paeonia, so several species of Paeonia were initially utilized as medicinal plants5. Wild Paeonia species were introduced and domesticated in China approximately 1500 years ago, as well as in medieval Europe6,7. Members of the genus Paeonia were first used as ornamental plants in China, and then this cultural use spilled over into other East Asian countries such as Japan and South Korea8. In the 18th century, Chinese tree and herbaceous peony cultivars were introduced into Europe, and later, into North America9,10. These Paeonia cultivars were very popular, spread widely throughout Europe and North America, and led to considerable cross-breeding work in the 19th century. The subsequent explosion of new Paeonia cultivars laid a foundation for the industrialization of Paeonia. In recent years, herbaceous peony has been used as a new cut flower7,11,12, and several major cut flower production areas have formed around the world, including in China, the Netherlands, North America, and New Zealand13. In 2011, the Ministry of Health of the People’s Republic of China approved tree peony seed oil as a new food resource, so the scale of oil tree peony planting has expanded rapidly in recent years. A review of oil tree peony and Paeonia medicinal plants will be published separately.
With the rapid development of Paeonia industrialization around the world, an increasing number of excellent cultivars are urgently needed. However, there are currently many problems in Paeonia breeding, such as an incomplete understanding of the extent of germplasm resources, rudimentary breeding techniques, fragmentary breeding information, unclear breeding objectives, and low breeding efficiency, among other limitations. This review provides a summary of the germplasm resources and genetic breeding of Paeonia, focusing primarily on the classification and characteristics of wild species and cultivar resources, phenotypic inheritance, molecular marker-assisted breeding, breeding methods and their applications, molecular breeding, and some prospects for breeding in the future. An important purpose of this review is to provide ideas and inspiration for Paeonia breeders. The method used for literature retrieval and selection for coverage is described in Supplementary Data 1.
Germplasm resources
Wild species resources
Linnaeus14 described the genus Paeonia for the first time based on two specimens from Switzerland and named one species Paeonia officinalis L., with two varieties, feminea and mascula. According to a cultivar of Paeonia introduced to Europe from Guangzhou, China, in 1794, Andrews15 introduced the first tree peony plant and named it Paeonia suffruticosa. In 1818, Anderson published the first monograph of Paeonia, which included 13 species divided into two major species groups, Fruticosa and Herbaceae16. The first Paeonia species of the New World (North America) was recorded by Douglas but later published by Hooker17 and named Paeonia brownii. Since then, Paeonia species all over the world have been systematically recognized. In 1890, Lynch reclassified Paeonia into three subgenera, Moutan, Onaepia, and Paeonia18. Following Lynch’s opinion18 but using the lower rank of section instead of subgenus, Stern19 divided the genus into three major groups: sect. Moutan, including all woody peonies; sect. Onaepia, including all herbaceous peonies in the New World; and sect. Paeon (=Paeonia), including all herbaceous peonies in the Old World (Table 1). Stern’s view was widely accepted by later scholars7,20, and taxonomic studies of Paeonia that employed molecular biology also support Stern’s view4,21,22,23,24.
Section Moutan
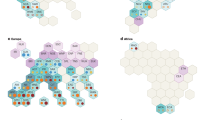
All species of sect. Moutan, which are endemic to China (Fig. 1), are shrubs or subshrubs, with 2n = 10 4. According to morphological differences of the disk, sect. Moutan was divided into two subsections. The species with a thin leathery sheath that at first completely envelops the carpels belong to subsect. Vaginatae, while species with conspicuous fleshy lobes around the base of carpels belong to subsect. Delavayanae24. Species of sect. Moutan have been continuously discovered over the past 100 years2,20,25,26,27,28,29,30,31,32,33,34,35,36. In 1999, Hong and Pan37,38 systematically sorted out the species of sect. Moutan. They reported eight species in this section, three of which included two separate varieties. In the Flora of China, Hong et al.39 revised this group to eight species and two subspecies. Hong40 then elevated the two subspecies of Paeonia decomposita into two separate species, but with no change in the other species (Supplementary Data 2).
Hong’s taxonomic work on sect. Moutan is systematic and comprehensive. Although the vast majority of the taxonomy in sect. Moutan is widely accepted by other scholars, a few different views exist about select species. Hong and Pan41 named a new species, Paeonia cathayana, based on a tree peony found at a farmer’s home in Henan Province, China. The owner claimed that the tree peony was dug from the wild, although no natural populations of this species were found in the wild in a subsequent investigation. Hong et al.36 believed that morphological variation is continuous among Paeonia delavayi, Paeonia lutea and Paeonia potaninii, treating them as single species and without any subspecies. Zhang42 studied the genetic diversity of the P. delavayi complex (including P. delavayi, P. lutea, and P. potaninii) from 14 populations and found large genetic differentiation among populations in different regions. Based on a years-long field investigation and experience in introduction and domestication, Li et al.6 believed that these three species had great differences in morphology and that these differences were stable among offspring. According to their results, Li et al. felt that the types within P. delavayi should be distinguished to allow for their more effective protection and utilization. For simplicity, we treat them as three species in this paper.
Hong and Pan43 claimed that “P. suffruticosa is not a hybrid”. However, many scholars have confirmed that P. suffruticosa is a hybrid formed by repeated hybridization of several species of subsect. Vaginatae, based on morphological and molecular biological evidence5,44,45,46,47. Therefore, we suggest changing the name of this species to Paeonia × suffruticosa.
Section Paeonia
The species of sect. Paeonia are perennial herbs that are widely distributed in East and Central Asia, the Himalayas, and the Mediterranean region (Fig. 1) and have 2n = 10 or 2048. Due to polyploidization and reticulate evolution, many different groups formed in the Mediterranean, making their classification very difficult48,49.
According to the number of leaflets or segments of lower leaves, sect. Paeonia was once divided into two subsections, subsect. Foliolatae and subsect. Dissectifoliae24. Beginning in the 1990s, Hong et al.50,51,52,53,54,55,56,57,58,59 carefully investigated sect. Paeonia distributed in Eurasia and the Mediterranean, studied its type specimens, and then revised the classification of sect. Paeonia comprehensively and systematically. They used the number of flowers per stem and root morphology as the main classification indexes and divided this section into three subsections: subsect. Albiflorae (usually several flowers per stem, root more or less carrot-shaped), subsect. Foliolatae (flower always solitary and terminal, roots carrot-shaped) and subsect. Paeonia (flowers solitary and terminal, lateral roots fusiform or tuberous), including 22 species and 22 subspecies (Supplementary Data 2)24.
In the Flora of China, Paeonia veitchii and Paeonia anomala are listed as separate species39. Hong and Pan55 deemed these species to have similar external morphologies but a difference in the number of flowers per stem, treating them as two subspecies of P. anomala. However, molecular evidence (nuclear and chloroplast genetic markers) indicated that the genetic distance between the two species was far, while P. veitchii and Paeonia sternadia were more closely related48. In the classification of Paeonia, the characteristics of seeds are hardly considered, even though there are great differences between species. We compared the seeds of P. veitchii and P. anomala and found that the testa of P. veitchii was dark blue and fleshy, while that of P. anomala was black and nonfleshy. Based on our assessment and on work conducted by Xia48, it is suggested that the two species should be independent (Yang et al., 2020, unpublished data). Hong4,40 made a breakthrough in the classification of sect. Paeonia. However, the classification of the three subsections was not completely consistent with a phylogenetic tree constructed by other scholars using nuclear and chloroplast genetic markers. Paeonia molecular phylogenetic analyses4,21,23,48,60 revealed that even the parents of different subspecies within the same species were completely different. All these results indicate that the relationships within sect. Paeonia are complex and that its classification requires additional studies.
Section Onaepia
There are only two species (P. brownii and Paeonia californica) in sect. Onaepia, with 2n = 10 4. They are perennial herbs with lateral roots that are carrot-shaped, fusiform or tuberous. The flowers are solitary and terminal or several per stem. The short petals are slightly longer or shorter than the sepals, and the disk is annular and fleshy, enveloping only the base of the carpels4. The distribution areas of the two species mostly overlap: P. californica is limited to the south of California, while P. brownii is distributed from northern California to Washington (Fig. 1). The flowering time of P. californica is from February to April, while that of P. brownii is from June to July61. These two species have very similar morphologies. They were regarded as a single species until a series of detailed morphological, ecological, and cytogenetic studies showed that they are different62. DNA sequencing also supports the above conclusion, lending evidence that the molecular evolution in this section was significantly faster than its morphological evolution22.
Cultivar resources
Paeonia was first domesticated and cultivated in China, including P. × suffruticosa and Paeonia lactiflora cultivars5,63. In the 18th century, Chinese tree peony and herbaceous peony cultivars were introduced into Europe and then spread to many temperate regions around the world9. With the spread of Chinese peony cultivars, different cultivated groups of Paeonia have gradually formed worldwide.
Tree peony cultivars
Wild tree peony may have been introduced and domesticated in China during the Wei and Jin Dynasties. After nearly a thousand years of development, it formed four large cultivar groups (Zhongyuan, northwest, southwest, and Jiangnan) and several smaller cultivar groups, such as the Yan’an and Baokang groups5,6,63. Zhou et al.47 used 14 chloroplast gene fragments and 25 single-copy nuclear genes to study the origin of P. × suffruticosa, concluding that Paeonia rockii, Paeonia qiui, Paeonia ostii, Paeonia jishanensis and P. cathayana participated in the formation of traditional P. × suffruticosa, while P. decomposita and subsect. Delavayanae did not. This conclusion was mostly consistent with those of other studies46,63,64,65. At present, there are more than 1000 tree peony cultivars in China, producing white, pink, red, purple, black, violet, and multicolored flowers with single, half, double and other flower types (Fig. 2)6,11.
1–3, single type; 4–18, semi-double type; 19–22, Japanese type; 23–26, bomb type; and 27–30, full double type. The taxonomy and name of cultivars (species) are shown in Supplementary Data 3
Chinese tree peony cultivars were introduced to Japan starting in the 8th century and largely in the 16th century. In the following two centuries, tree peony was bred independently to meet Japanese esthetic preferences and local climate characteristics on the basis of Chinese tree peony. Japanese tree peony cultivars have distinct characteristics. Most of them produce single or semidouble flowers with bright red or purple colors, beautiful lines, and an upright flower that is higher than the leaf surface. Approximately 200 tree peony cultivars have been bred in Japan8,66.
Most of the early tree peony cultivars in Europe were domesticated from Chinese tree peony8,9. At the end of the 19th century, P. delavayi and P. lutea were found in Yunnan, China25, and then introduced to Europe and North America. A series of cultivars were bred by crossing the two wild species with P. × suffruticosa in France and the USA9. The flower colors of this group of tree peonies include yellow, champagne, orange, and scarlet, which do not exist in P. × suffruticosa. The flower stems are not strong, and some cultivars have drooping flowers. They flower towards the end of spring, a week or two after Chinese or Japanese tree peonies. These hybrid cultivars, which were formed by crossing P. delavayi and P. lutea with P. × suffruticosa or their offspring, interbreed and are classified into the Lutea hybrid group. There are more than 1000 cultivars, mainly in the USA, Europe and Australasia9,11,67,68.
Herbaceous peony cultivars
The earliest record of cultivation of ornamental herbaceous peonies in China was also during the Wei and Jin Dynasties69,70. P. lactiflora is widely distributed in China and has strong adaptability. Chinese herbaceous peony cultivars were bred by domesticating this single species71. Therefore, the cultivars of this group are called Lactiflora or Chinese peony9. There are ~300 cultivars of Lactiflora peony in China, including cultivars producing white, pink, red, purple, purple-black, multicolored, and other-colored flowers (Fig. 2). The flower types include the single, semidouble, double, Japanese and bomb types5,12.
Lactiflora peony was introduced into Japan in the 10th century. Japanese breeders began breeding this peony in the 17th century, creating ~100 cultivars to date66. In the 19th century, Lactiflora peony cultivars became popular in Europe and North America, when much breeding work was carried out. After more than 100 years of development, Lactiflora peony cultivars have developed into the largest group of Paeonia in Europe and North America, with more than 4000 cultivars registered with the American Peony Society (APS)9,68. At the end of the 19th century, European and American breeders attempted to use Lactiflora peonies in crosses with European wild peony species10. These kinds of hybrid cultivars, which are produced by crossing two or more species of sect. Paeonia, are called herbaceous hybrid peonies. The flowering time of herbaceous hybrid peonies is usually earlier than that of Lactiflora peonies, which greatly prolongs their entire flowering period. They also contribute new flower colors, such as yellow, coral, and bright red, which are not available in Lactiflora peonies (Fig. 2)7. At present, there are more than 1000 herbaceous hybrid peonies registered with the APS, and the number is increasing68.
Intersectional hybrid cultivars
The success of intersectional Paeonia hybrids is a tremendous breakthrough in the contemporary history of Paeonia breeding. Japanese breeders successfully hybridized herbaceous peony with tree peony, which was widely developed by American breeders. Thus far, only sect. Paeonia and sect. Moutan have been successfully hybridized among the three Paeonia sections72. At present, most cultivars are obtained by using Lactiflora peonies as the female parent and Lutea hybrid peonies as the male parent. To commemorate the pioneering contribution of the Japanese breeder Toichi Itoh in peony intersectional breeding, the APS refers to these cultivars, which were obtained from the Lactiflora peony (♀) × Lutea Hybrid peony (♂) cross (♀ = female parent, ♂ = male parent), as Itoh hybrid peonies. The flower and leaf types of intersectional hybrid peonies are mostly similar to those of tree peonies, and their ecological habits and flowering times are similar to those of herbaceous peonies. Most intersectional hybrid peonies have strong adaptability, few diseases, and abundant flowers. Their flower colors are rich, including white, yellow, pink, orange, purple, and scarlet (Fig. 2)73. More than 150 cultivars have been registered68.
Horticultural utilization of Paeonia
The application of peony cultivars mainly focuses on three aspects: landscape architecture, potted plants and cut flowers. Special peony gardens are most famous in China and Japan. In Luoyang, Heze and Beijing, China, and Shimane, Japan, there are more than 30 large special peony gardens, with the largest number in Luoyang6. Each year, more than 20 million tourists visit Luoyang during the flowering period, generating total tourism-related revenue that exceeds 17.8 billion yuan (approximately 2.52 billion US$)74. In Europe, North America and other countries, peony cultivars are mainly used for landscape gardening.
Peony cultivars are widely sold as potted plants in Japan, Europe and North America, greatly prolonging the sales period of peony plants. In China, potted tree peonies blooming during the Spring Festival are the most common. In Heze, the annual production of potted tree peonies exceeds 290 thousand pots during the Spring Festival, and the output value exceeds 30 million yuan (Heze Daily, 25 December 2019).
On the international flower market, herbaceous peony is becoming popular as a new cut flower. More than 25 countries are engaged in the production of herbaceous peony cut flowers, with the Netherlands being the largest global producer, annually producing more than 48 million herbaceous peony cut flowers13. According to data from Royal Flora Holland, the global distribution center of cut flowers, in 2019, the sales volume of herbaceous peony cut flowers exceeded 72.4 million stems in the Netherlands and >35 million euros, with a year-on-year growth of 20.39% in sales volume and 26.53% in production value75. Therefore, there is rapid momentum in the development of the herbaceous peony cut flower market.
Genetic breeding
Genetic research
Uncovering genetic mechanisms can serve as an important guide for breeding. Poor self-compatibility76,77,78,79,80, high chromosome heterozygosity, and long juvenility have been the greatest limiting factors in Paeonia genetic research81. In recent years, some scientific institutions have begun to pay attention to Paeonia, and some genetic mechanisms are gradually being studied.
Trait inheritance
Systematic breeding of Paeonia was carried out mainly after the middle of the 18th century. Paeonia breeders tend to pay more attention to the compatibility of different cross combinations but pay less attention to the genetic mechanisms underlying those crosses. Combined with the phenotypic characters of the offspring and the parents of previous hybrids, the blotch at the base of the petal and carpel coat covering the tomentum is a dominant inherent character in Paeonia breeding6,67. In wild peony species, chalconaringenin 2′-O-glucoside, which has been detected only in the petals of subsect. Delavayanae, can be stably inherited by offspring, resulting in yellow or orange flowers in Lutea and Itoh hybrid peonies82. The flower fragrance of subsect. Delavayanae species is pleasant, and the main components are linalool compounds, which can be passed on stably to offspring through hybridization83,84,85. Cheng and Chen analyzed the genetic mechanisms of flower color and flower type in hybrid combinations of Northwest tree peonies and found that these traits may be greatly influenced by the female parent and that the inheritance of white flowers and the single flower type is strong (about half of the offspring had white or single flowers)81. Zhang et al. analyzed 20 phenotypic characters in an F1 genetic population, including 120 individuals, that was obtained by P. ostii ‘Fengdan’ (♀) × (P. × suffruticosa ‘Shin Jitsugets-nishiki’) (♂) and found that the variable coefficients of F1 individuals’ phenotypic traits ranged from 11.03% to 63.49% and heterosis and transgressive segregation for 20 phenotypic traits86,87.
It is necessary to consciously expand the number of hybrid offspring and comprehensively record their phenotypic characters.
Genetic map
Given the limitations stated above, namely, high heterozygosity, poor self-compatibility and a long juvenile period, it is difficult to construct an ideal mapping population of Paeonia. Thus far, only three genetic populations of Paeonia have been cultivated and used to construct genetic maps. Cai et al.88 used P. ostii ‘FengDanBai’ as the female parent and P. × suffruticosa ‘HongQiao’, ‘Kaō’ and ‘Kokuryū-nishiki’ as the male parent and constructed three F1 segregating populations by artificial pollination. Simple sequence repeat (SSR) markers, which were used to check the three F1 populations, revealed that the F1 population in a P. ostii ‘FengDanBai’ (♀) × (P. × suffruticosa ‘HongQiao’) (♂) cross was a better population for constructing a genetic map of tree peony than the other two populations. Using this F1 population, Cai et al.89 performed specific-locus amplified fragment (SLAF) sequencing of the F1 population to identify markers that could be used to construct a genetic linkage map. Finally, a high-density genetic map of tree peony was constructed, including 1189 SLAF markers in five linkage groups and spanning 920.669 centimorgans (cM). Peng et al.90 developed 74 SSR markers with polymorphisms based on the transcriptome data of P. × suffruticosa ‘LuoYang Hong’. These markers were used to construct a genetic map for the same F1 population developed by Cai et al.89. A total of 68 SSR loci were mapped to five linkage groups, 48 of which showed Mendelian inheritance (segregation ratio of 1:1, 1:1:1:1 or 1:2:1). By using P. ostii ‘Fengdan Bai’ (♀) × (P. × suffruticosa ‘Xin Riyuejin’) (‘Xin Riyuejin’ = ‘Shin Jitsugetsu-nishiki’) (♂), an F1 genetic population including 120 individuals was obtained. Zhang et al. used genotyping-by-sequencing (GBS) to develop markers for the Cai et al.91 F1 population. Finally, a genetic map was constructed containing 3868 markers in five linkage groups with a genetic distance of 13,175.5 cM and 322-1224 markers in each linkage group. Guo et al.92 used the same F1 population as Zhang et al.91 to construct a genetic map by using SSR markers. Finally, 35 SSR markers were mapped, covering five linkage groups, each with 3-14 markers. Li et al.93 performed restriction site-associated DNA sequencing (RADseq) of an F1 genetic population with 120 individuals, which were derived from (P. × suffruticosa ‘Qing Long Wo Mo Chi’) (♀) × (P. × suffruticosa ‘Mo Zi Lian’) (♂) (the two cultivars are from the Zhongyuan tree peony group, with purple-black flowers)93. Genetic maps of the parents constructed with RADseq markers showed 1471 markers in the female parent genetic map, in seven linkage groups, and 793 markers in the male parent genetic map, in five linkage groups, covering 965.69 and 870.21 cM, respectively.
The above three genetic populations89,91,93 had different parents, but the findings have not been replicated to date. The number of offspring in the three populations was <200, and the phenotypic characters could not be fully separated, which increased the frequency of partial marker separation. The markers mapped to the genetic map are limited, and their distribution is uneven, even though seven linkage groups were included in the female parent genetic map constructed by Li et al.93. Zhang et al.91 included more molecular markers in their genetic map than in other studies, but the genetic distance that they observed was more than ten times greater than that observed by the other groups, suggesting that additional analyses are needed. Thus far, a limited number of genetic maps have been constructed only for P. × suffruticosa, with no genetic maps in sect. Paeonia. Maps of more and different genetic populations need to be constructed according to their genetic traits.
Molecular markers
The origin and history of peony cultivars are incomplete and complicated. Therefore, it is difficult to meet the requirements of modern breeding by the exclusive use of traditional morphological index markers. Compared with traditional morphological index markers, DNA molecular markers have many advantages, such as a high abundance, a large amount of information, and insensitivity to the environment. At present, molecular marker-assisted breeding of Paeonia mainly involves two processes: (1) early identification of the authenticity of hybrid seedlings and (2) quantitative trait locus (QTL) mapping of phenotypic traits.
There is a certain amount of self-compatibility in Paeonia, and the self-pollinated seed setting rate is <5%78,79,80. During the process of hybridization, improper handling may lead to self-pollination and false hybrids (pollen from unwanted cultivars). Peony seedlings have a long juvenile period (more than 3 years before flowering). To reduce maintenance costs, it is necessary to identify the authenticity of hybrid seedlings early. Several molecular markers, including intersimple sequence repeat (ISSR), amplified fragment length polymorphism (AFLP), sequence-related amplified polymorphism (SRAP), and SSR markers, were used in the early identification of Paeonia hybrids (Supplementary Data 4). Different molecular markers have achieved good results in validating the authenticity of and identifying Paeonia hybrids94,95,96,97,98,99,100,101,102,103. Relatively speaking, SSR markers have the advantages of simple operation, high accuracy, good stability, less demand for DNA, and codominance, among others, making them more suitable than other markers for early authenticity validation and identification of Paeonia hybrids97,103.
There are few genetic populations of Paeonia, and QTL research on phenotypic characters in Paeonia is very limited. Cai used the F1 segregating population and genetic map from P. ostii ‘FengDan Bai’ (♀) × (P. × suffruticosa ‘HongQiao’) (♂) to analyze 27 quantitative characters of branches, leaves, flowers and fruits using composite interval mapping104. Among them, 20 traits were successfully detected for associated QTLs, and QTL-pn-2, which controls the number of petals, explained up to 71.9% of the phenotypic variation. In addition, three QTLs associated with flower color traits were detected, explaining 11.4–12.8% of the phenotypic variation.
Wu et al.101 used 11 pairs of polymorphic SSR markers to analyze 32 phenotypic characters in a natural population of Northwest tree peonies, showing that it was composed of 99 representative individuals without any direct relationships. They also found that five markers were significantly associated with six traits, each explaining 30.4–55.8% of the phenotypic variation. Wu et al.102 used 138 SSR markers to carry out trait association analysis of 462 natural populations and the F1 segregating population of P. ostii ‘FengDan Bai’ (♀) × (P. × suffruticosa ‘HongQiao’) (♂). In natural populations, SSR markers were significantly correlated with flower, leaf and fruit characteristics. Five associations were confirmed in the F1 segregating population, involving four markers and four flower characters: petal width (PS029 and PS296), petal shape (PS029), petal color (PS134), and petal number (PS309). The performance of traits in the above studies101,102,104 was assessed at only one location per year, which did not exclude the interference of environmental factors. Therefore, the reliability of these QTLs would be affected, and they still need to be verified.
Traditional breeding
Based on 30 Paeonia species, more than 8000 cultivars with a rich assortment of flower colors and types have been cultivated through breeding, and the number is growing continuously. Currently, selective breeding and cross-breeding are the most common breeding methods for Paeonia (Fig. 3). In addition to these two methods, other techniques have also been used to obtain new Paeonia cultivars, such as chemical mutation breeding, radiation mutation breeding, space mutation breeding, and others. The seeds of Paeonia are extremely large, and the testa is thick, which limits the penetration of chemical mutagens and rays. Tissue culture is still a challenge in Paeonia, and it is difficult to obtain a large number of seedlings through in vitro culture. All these limitations restrict the development of the above breeding techniques. Therefore, their effectiveness has been very limited11,12,105,106.
Wild species and cultivars are distinguished by different background colors, with wild species indicated above and cultivars indicated below. Parents are shown above the arrows while progenies are shown below the arrows. Brackets combine parts of cultivars groups or types into a large cultivar group
Selective breeding
Selective breeding is a simple method in Paeonia breeding that focuses mainly on selective breeding of seedlings and bud mutations5.
The selective breeding of seedlings is a way to select new cultivars by collecting naturally pollinated seeds that are then sown, and excellent individuals are selected for propagation from the offspring. In China, most of the new Paeonia cultivars were selected using this method before 1960. The largest selective breeding program for tree peony seedlings was carried out under the leadership of Professor Hen Yu in Heze from 1956 to 1975, ultimately generating 252 new cultivars107. In the 19th and 20th centuries, most of the new Lactiflora peony cultivars bred in Europe and the USA were also obtained by this method, such as ‘Pietertje Vriend Wagenaar’, the 2018 APS Golden Award-winning cultivar68. Although this method is simple, sowing seeds and transplanting seedlings require much land, and the workload required to select excellent individuals is very heavy. With the shortage of land and increasing labor cost, this breeding method is gradually being replaced.
Selective breeding via bud mutations involves the discovery of branches with obvious variation and maintaining that variation through asexual propagation. Some Paeonia cultivars were obtained in this way, such as P. × suffruticosa ‘Shima-nishiki’, ‘Hua Er Qiao’ and ‘Ice Age’, the Itoh hybrid peony cultivar ‘White Emperor’, and the Lactiflora peony cultivars ‘Alpine Aire’, and ‘Ann Styer’, among others6,12,68.
Intrasectional cross-breeding
Tree peony
There are two main types of cross-breeding in tree peony: intrasubsectional and intersubsectional.
There is no reproductive isolation between different species (cultivars) in the same subsection of sect. Moutan. A large number of viable seeds can be produced by crossing parents in the same subsection, with fertile offspring. In recent years, to enrich the ornamental traits and enhance the resistance of tree peony cultivars, breeders have carried out different types of cross-breeding within subsect. Vaginatae, mainly the following three types: crossing different cultivar groups of P. × suffruticosa, crossing the wild species in subsect. Vaginatae with P. × suffruticosa and crossing different wild species in the same subsection. In this way, a large number of excellent new cultivars have been cultivated, such as P. × suffruticosa ‘Angel Emily’, ‘Hei Fu Ren’, ‘Wen Hai’, ‘Zi Yu Zui Xue’, and ‘Fei Hua Si Meng’11,67,68 (Fig. 3).
Due to obvious reproductive isolation, it is difficult to obtain viable seeds by crossing subsect. Vaginatae with subsect. Delavayanae6. The species of subsect. Delavayanae were discovered in the late 19th century, and P. delavayi and P. lutea were first used for cross-breeding by European breeders. Louis Henry, a French breeder, announced the first intersubsectional-cross cultivar ‘Souvenir de Maxime Cornu’, which was hybridized by M. Maxime Cornu using P. lutea (♀) × (P. × suffruticosa) (♂). Victor Lemoine carried out similar work in the same period, and some cultivars had bloomed at ~1900 during World War I. These cultivars quickly caused a sensation after they were displayed in the APS. Most of these cultivars have very heavy flowers, which tend to hang down among the foliage. Inspired by Lemoine, American breeder Arthur Saunders began to introduce his own hybrid tree peonies by using P. lutea and P. delavayi as the female parents and Japanese P. × suffruticosa as the male parent. By 1986, a total of 78 cultivars had been registered with the APS. The flower types of these cultivars are mostly single or semidouble, and the flower colors are very rich, from crimson, scarlet, apricot yellow and amber to golden yellow and lemon yellow, while the blooms are held on strong stems well above the leaves7,9.
All the F1 Lutea hybrid peonies from Europe and Saunders’ collections appeared to be sterile. Accidentally, Saunders obtained two seeds from an F1 Lutea hybrid peony (Saunders F2a and Saunders F2b) and managed to develop them into seedlings. The two seedlings were eventually transferred to Nassos Daphnis. Daphnis used these two F2 seedlings as important parents to carry out a large amount of backcross breeding and produced a batch of fertile cultivars. Since then, the sterility bottleneck of Lutea hybrid peonies has been broken9. In the last 30 years, breeders such as Bill Seidl (USA), Bernard Chow (Australia), and Nate Bremer (USA) have crossed fertile Lutea hybrid peonies (most from Daphnis’ cultivars) with each other, and the fertility of an advanced generation of Lutea hybrid peonies has been greatly improved as a result. The breeding objectives have also shifted, with the focus now on cultivars with fortified vigor, larger flowers born on stronger stems and with good form, such as ‘Alicia Nicole’, ‘Aquarius’, ‘Beach Comber’, and ‘Coral Nebula’68.
To improve the adaptability and stem strength of Lutea hybrid peonies, Japanese breeders also attempted to backcross Japanese P. × suffruticosa with F1 Lutea hybrid peonies. Hao et al.99 used the F1 Lutea hybrid peony ‘High Noon’ as the female parent and some Japanese P. × suffruticosa cultivars as the male parent in crosses. Finally, five new cultivars were screened out with strong stems and considerable vigor. In China, intersubsectional cross-breeding work in tree peony began at the end of the 20th century and mainly involved crosses of P. lutea and P. delavayi with P. × suffruticosa, and new cultivars gradually emerged after 2004, such as ‘Cai Hong’, ‘Xiao Xiang Fei’, and ‘Yan Huang Jin Meng’11,67,108 (Fig. 3).
In addition to the cross-breeding combinations mentioned above, breeders have also tried P. × suffruticosa as the pollen parent, P. lutea or P. delavayi as the seed parent, or P. lutea or P. delavayi crosses with wild species of subsect. Vaginatae and obtained some cultivars, such as ‘Nong Yuan Xiang Yue’ and ‘Nong Yan Xiang Yu’9,11,68. Paeonia ludlowii was also used by some Chinese breeders in intersubsectional cross-breeding, but no new cultivars have yet been reported. In April 2019, one of the authors (Yong Yang) visited Heze, Shandong Province, and observed five seedlings of (P. × suffruticosa ‘Xiang Yu’) (♀) × P. ludlowii (♂), which were bred by Xiaojing Zhao, a peony breeder in Heze. The leaf types of these seedlings were intermediate to those of the parents but more inclined towards P. ludlowii. Among them, four plants could not form apical buds in winter, and only one flowered, forming single, yellow flowers. This may be the first excellent individual to be obtained by a cross between P. ludlowii and a member of subsect. Vaginatae.
Herbaceous peony
Herbaceous peony breeding mainly includes intra- and interspecific cross-breeding, with the former mainly focusing on P. lactiflora. In recent years, herbaceous peony, as a new cut flower, has become popular all over the world, and the mainstream cultivars used as cut flowers are Lactiflora peonies13. To meet the demands of the cut flower market, cross-breeding within Lactiflora peony cultivars is still an important direction of herbaceous peony breeding. For example, the excellent P. lactiflora cut flower cultivars ‘Shirley Temple’, ‘Mother’s Choice’, and ‘Shirayuki-hime’ are the products of intraspecific cross-breeding of Lactiflora peonies. In addition to P. lactiflora, P. officinalis and Paeonia teniflora have also produced some intraspecific hybrids used for landscaping68 (Fig. 3).
To expand the diversity of herbaceous peony, interspecific cross-breeding of sect. Paeonia has become a research hot spot in recent years. At the end of the 19th century, European breeders began to cross Lactiflora peonies with local wild herbaceous peony species. The first hybridization was carried out by using P. lactiflora (♀) ×Paeonia wittmanniana (♂), and four varieties were bred. As the most famous herbaceous hybrid peony breeder, Saunders made a pioneering attempt to cross ~28 hybridization groups. Among these groups, Lactiflora peonies generally played a major role in the work of hybridists, serving as parents for over 80% of hybrids9. Finally, more than 180 new herbaceous hybrid peony cultivars were screened out68. Saunders’ work laid a solid foundation for the development of herbaceous peony hybrids, and cultivars such as ‘Nosegay’, ‘Early Windflower’, ‘Moonrise’, ‘Nova’, and ‘Blushing Princess’ were important parents in later herbaceous peony hybrid breeding. To restore fertility and improve the ornamental value of herbaceous peony hybrids, breeders carried out a large number of cross-breeding trials among their offspring based on Saunders’ work10,109. In recent years, a large number of excellent new advanced-generation herbaceous peony hybrid cultivars have been cultivated, such as ‘Salmon Dream’, ‘Lemon Chiffon’, ‘Pastelegance’, and ‘Kim’68 (Fig. 3). The seedlings of the hybrids whose fertility has been restored are all tetraploid10, but the reason is not clear.
Intersectional cross-breeding
The three Paeonia sections have great differences in phenotype and ecological habit, and it was once thought that intersectional cross-breeding was impossible. The first intersectional hybrid cultivars of Paeonia were created by Taichi Itoh, a Japanese breeder, with the Lutea hybrid peony ‘Alice Harding’ (♀) × P. lactiflora ‘Kakohden’ (♂). In 1974, four of the seedlings were registered with the APS by Louis Smirnow and caused an immediate sensation110,111,112. Since then, American breeders have conducted many similar crosses, the three most famous breeders being Don Hollingsworth, Roger Anderson, and Donald Smith7. On the basis of their breeding experience, some excellent hybrid parents were also screened out113,114. Based on US breeders’ experience, Chinese breeders also cultivated some new Itoh hybrid peony cultivars, such as ‘Jing Gui Mei’, ‘Jing Hua Zhao Xia’, and ‘Huang Die’11,12,115 (Fig. 3).
The P. lactiflora (♀) × Lutea hybrid peony (♂) combination is the most successful combination in Paeonia intersectional cross-breeding. To obtain novel varieties, breeders also carried out other types of intersectional cross-breeding. After many failures, some new cultivars were obtained from some new hybrid combinations. The successful cross combinations are as follows: P. lactiflora (♀) × (P. × suffruticosa) (♂)116,117, P. lactiflora (♀) ×P. delavayi (♂), (P. × suffruticosa) or Lutea hybrid peony (♀) ×P. lactiflora (♂)68,116,117,118, and P. delavayi (♀) ×P. lactiflora (♂)68 (Fig. 3). In addition to these cross-breeding combinations, breeders have also tried a variety of intersectional cross-breeding combinations, including those involving species of section Onaepia. However, only select combinations produced a few seeds, and no new cultivars were reported119,120,121,122,123.
Itoh hybrid peonies are generally triploid124,125 with poor fertility and produce few effective pollen grains or full seeds. Smith tried to restore the fertility of Itoh hybrid peonies by crossing some of his own Itoh hybrid peony cultivars with Lactiflora peony or an advanced generation of herbaceous hybrid peony cultivars126. In the past five years, he made many attempts. Finally, two seedlings were retained (private communication with Don Smith). This attempt may change the status of infertility of Itoh hybrid peonies.
Molecular breeding
Molecular breeding is a highly efficient way to compensate for the shortcomings of traditional breeding methods and shorten the breeding cycle. At present, the tissue culture of Paeonia mainly uses zygotic embryos or buds to produce secondary buds directly127,128,129,130. It is difficult for Paeonia plants to dedifferentiate differentiated tissue to form calli and then form seedlings. This limitation has led to the inability to develop a stable genetic transformation system that is the basis of genetic transformation or gene editing in Paeonia. Although there are still many challenges to achieving the molecular breeding of Paeonia, the molecular mechanisms of several Paeonia characters that are linked with production and practical applications have been researched (Table 2). These molecular studies have laid a foundation for the future molecular breeding of Paeonia.
Flower and leaf color
The main factors affecting flower color are the composition and levels of pigments in petals131. The pigments in Paeonia petals are flavonoids, including anthocyanins, flavones, and flavonols82,132,133,134,135,136,137,138. The diversity of flower color is caused by the differential expression of flavonoid biosynthesis-related genes. The metabolic pathway of flavonoids in Paeonia is relatively well resolved139. The structural genes DFR and ANS (the full names and details of all genes are provided in Supplementary Data 5 and 6), which play a key role in the synthesis of anthocyanins, determine the formation of pink and red flowers140,141,142, and the high expression of AOMT promotes the change in flower color from red to purple143. The expression levels of other structural genes, such as CHS, CHI, FLS, PAL, F3H, F3’H, F3'5′H, FLS, ANR, 3GT, 5GT, UF3GT, and UF5GT, also play vital roles in yellow, white and other flower colors144,145,146,147,148.
In addition to structural genes, transcription factors also play important roles in regulating flower coloration in Paeonia. The transcription factors PsMYB and PsWD40 may play a key regulatory role in controlling anthocyanin concentration in red and white petals, and their differential expression mediates the formation of multicolored flowers142. PsMYB12 can activate the PsCHS promoter and play a direct role in the formation of tree peony petal blotch, which is when the color at the base of the petal is significantly darker than that of other parts, forming a blotch148. MYB-1 and MYB-5 may also regulate the formation of tree peony petal blotch143. The transcription factor bHLH3 is highly expressed in white petals of tree peony, while MYB22 is highly expressed in purple petals. Therefore, these two transcription factors may play a role in regulating the shade of tree peony petals147.
The leaf color of most peony cultivars is red when they sprout in spring, giving them ornamental value, but the red gradually fades as the leaves expand. The mechanism of red coloration in Paeonia leaves is consistent with that in petals and is also due to the presence of flavonoids. High expression of the structural genes CHS, DFR, and ANS and low expression of LAR and ANR lead to the accumulation of anthocyanins in red leaves. The anthocyanin repressor MYB2 is activated in spring, replacing MYB1. Regulation of the MYB2 + bHLH1 + WD40-1 complex reduces the expression levels of CHS, DFR, and ANS, leading to a change in leaf color from red to green149,150. In the red and nonred leaf stages of P. × suffruticosa ‘Man Yuan Chun Guang’, the expression of the structural genes PsDFR and PsANS was the same as that of anthocyanin genes, and the expression levels of PsFLS, PsANR and flavonol were consistent; moreover, the transcription factors PsMYB113, PsMYB4 and PsMYBF1 may be important in regulating the expression of these structural genes151.
Flower type
The flowers of all wild Paeonia species are single. After thousands of years of domestication and selection, a rich assortment of flower types has formed, including semidouble, double, Japanese, and bomb. A change in flower type in Paeonia mainly depends on a natural increase in petals combined with centripetal or centrifugal petalization of stamens6. Research on Paeonia flower development revealed that the development of Paeonia floral organs is mainly controlled by the MADS-box gene family, and a large number of functional genes have been cloned152,153,154,155,156. Shu et al.157 cloned PsTM6 in the euAP3 lineage of tree peony. By analyzing the gDNA sequence of this gene in 23 tree peony cultivars, it was concluded that PsTM6 might affect the petalization of stamens. Ge et al.155 cloned the genes of the MADS-box family related to flower organ development in P. lactiflora and analyzed their differences in expression. Their results showed that as the degree of petalization increased, the expression of PlAP1, PlAP2, and PlSEP3 also increased, while the expression of PlAP3-1, PlAP3-2, and PlPI decreased. PlAP1 and PlSEP3 mainly determine sepals and petals, and PlAP3-1, PlAP3-2, and PlPI mainly determine stamens and petals. PlAP2 not only determines sepals and petals but also participates in carpel formation. Wu et al. also found a correlation between PlAP2 and petal formation153. Ren cloned and analyzed the expression differences of genes related to flower type in P. × suffruticosa ‘Zhao Fen’158. The results showed that PsAP1 belonged to the AP1/SQUA subfamily of the MADS-box family, which was mainly expressed in petals, sepals, and carpels, with the highest expression at an early stage of flower bud differentiation. PsPI, PsMADS1, and PsMADS9 were mainly expressed in petals and stamens, regulating the formation of petals and stamens. Moreover, PsAG transformed into Arabidopsis thaliana and Nicotiana tabacum also showed that PsAG was involved in the regulation of floral organ development158. Tang cloned PsMADS5, PsMADS7, and PsMADS12 from the petals of P. × suffruticosa ‘Luo Yang Hong’ and analyzed differences in their expression150. Tang’s results suggested that PsMADS5 was involved in the regulation of calyx development and the petalization of stamens, PsMADS7 initiated the petalization of stamens and cooperated with multiple members of the MADS-box family to jointly regulate the formation of multiple flower organs, and PsMADS12 was significantly expressed in the petals of the ‘Lou Zi’ (crown-proliferation) flower type. These three genes may be key genes for stamen petalization.
Flowering time
Several pathways related to flowering time have been reported in A. thaliana, including the photoperiod, vernalization, gibberellic acid (GA3), and autonomic pathways159. The transcription factor PsCOL4, which is related to flowering time, was cloned from P. × suffruticosa ‘Zi Luo Lan’. The expression of PsCOL4 in the stem and leaves was highest at an early stage, and its expression decreased as the flower bud developed, inducing the expression of downstream genes FT and SOC1 to ensure the flowering process160. Wang et al.161 cloned the MADS-box gene PsSVP, which is mainly regulated by GA3, into P. × suffruticosa ‘Luoyang Hong’. The gene was mainly expressed during vegetative growth, with the highest expression in leaves and stems, but its suppressed expression in aborted flower buds was much greater than that in normal flower buds, inhibiting flower formation. The expression level of AP1 was different among developmental stages in P. lactiflora ‘Jinhui’. The expression levels of AP1 in the inner and outer petals were significantly different, indicating that its expression level might be related to P. lactiflora floral organ development162. Zhou et al.163 isolated PsFT from reblooming (Paeonia×lemoinei ‘High Noon’) and nonreblooming (P. suffruticosa ‘Luo Yang Hong’) cultivars of tree peony. According to differential gene expression analysis and verification of transgenic function, PsFT may be closely related to secondary flowering. Wang et al.164 analyzed the transcriptome of flower buds of seven different flowering types of tree peony and screened the genes related to secondary flowering, namely, PsAP1, PsCOL1, PsCRY1, PsCRY2, PsFT, PsLFY, PsLHY, PsGI, PsSOC1, and PsVIN3. Wei et al.165 found that the expression level of PsFUL1 was highest in flower buds and petals. PsFUL1 expression in different peony varieties was analyzed, and it was found that expression peaked quickly in early blossoms of P. × suffruticosa ‘Luoyanghong’, which promoted the initiation of downstream genes and ensured the occurrence of early blossoms. Therefore, PsFUL1 may be an important transcription factor in floral transition and flowering regulation.
Some research has also focused on how genes artificially regulate the flowering response. PdFT, which was isolated from P. delavayi, had the highest level of expression in flower buds, and its expression could be increased by the exogenous application of 1000 mg/L GA3 and defoliation and decreased under short sunshine (8-h photoperiod) and low temperature (4 °C). These results indicate that PdFT is a key gene in the photoperiodic pathway that determines the switch to flowering166. Exogenous GA3 induced and inhibited the expression of the gibberellin-related genes PsCPS and PsGA2ox in tree peony, and defoliation and exogenous GA3 inhibited the expression of PsNCED and PSbZIP, whose expression coincided with the ethylene release curve167. PsGA2ox is an inhibitory gene for flowering in P. × suffruticosa ‘Luo Yang Hong’168.
Abiotic stress resistance
Plants adapt differently to the environment, and their improved resistance to environmental stresses can expand their adaptive area. Research on the abiotic stress resistance of Paeonia has mainly focused on plant responses to low versus high temperature, drought, salt, and heavy metals. Under high-temperature (35.75/27.5 °C, average daily maximum/minimum temperature) stress, heat-resistant P. lactiflora ‘Zifengyu’ showed enhanced activity of antioxidant enzymes (superoxide dismutase, peroxidase, and catalase) and the expression of HSP, which removed reactive oxygen species effectively and prevented plants from dying169,170. HSP70-overexpressing transgenic plants were tolerant to high temperature171,172. In P. lactiflora, microRNAs played an important role at the posttranscriptional level: miR172c-3p, miR395a, miR397a, miR408-5p, and miR827 expression was upregulated under high-temperature stress, indicating that they were associated with posttranscriptional regulation under high-temperature stress (35.75/27.5 °C, average daily maximum/minimum temperature)173. In salinity stress trials, Hao et al.174 cloned PSK1 from P. × suffruticosa into Arabidopsis, and transgenic lines formed larger green cotyledons on medium with 125 mM NaCl than wild-type (WT) Arabidopsis. Using transcriptome and miRNA sequencing and comparing the response of P. ostii in 100 μM CuSO4·5H2O copper stress and control groups, 12 genes and 30 miRNAs were preliminarily identified as participating in the regulation of copper stress175,176. The tryptophan decarboxylase gene of P. lactiflora (PlTDC) in the melatonin biosynthesis pathway enhanced the tolerance of transgenic tobacco to drought, which was induced for 18 days in the absence of water at 22°C. When the NaCl concentration was 200 mM, there was no difference between transgenic and WT leaf disks, but when the NaCl concentration was >200 mM, the green area of leaf disks of WT lines gradually shrank, especially at 800 mM, where the WT leaf disks all turned brown, in contrast to those of transgenic lines, in which only a few turned brown177. When P. lactiflora was treated at a low temperature (4 °C), the expression level of the cold resistance gene PlGPAT remained high for 72 h, indicating its role in cold resistance178.
The expression level in flower buds of two dehydrin genes, PlDHN1 and PlDHN2, which were cloned from P. lactiflora ‘Dafugui’, was highest in December. PlDHN1 was significantly upregulated in response to heat and waterlogging, and the sensitivity of PlDHN1 to abiotic stresses was ranked as waterlogging > heat > abscisic acid (ABA) > cold179,180. The expression of nucleus-located PlWRKY13, which was also cloned from P. lactiflora ‘Da Fugui’, was upregulated by four abiotic stresses: low and high temperatures, waterlogging, and salt stress. In addition, PlWRKY13-silenced plants were more sensitive to fungal infection, indicating that PlWRKY13 was induced by abiotic stresses but involved in the regulation of disease resistance181. PsDREB is a tree peony transcription factor related to stress resistance. Compared with the WT, transgenic lines did not show wilting, the malondialdehyde levels in vivo were lower than those in the WT, and superoxide dismutase and peroxidase were higher than those in the WT, indicating that PsDREB significantly improved the resistance of transgenic tobacco to drought and salt stress182. Liu et al.183 cloned PsDREB2 and its promoter in tree peony, transferred the promoter into A. thaliana, and expressed it in all parts of transgenic plants. Under drought, low-temperature, high-salt, and ABA stresses, the PsDREB2 promoter upregulated the expression of the stress-related genes DREB1A, CBF1, R29A, and RD29B.
Postharvest
Since peony flowers have become increasingly popular cut flowers, research on postharvest aspects has also increased. At the molecular level, postharvest research has mainly focused on endogenous hormones, water metabolism and sugar accumulation. Ethylene level is an important factor that affects the preservation of cut flowers. By controlling ethylene content, the quality of cut Paeonia flowers can be improved. Zhou et al.184 cloned the key genes PsACS and PsACO involved in ethylene biosynthesis from cut flower petals of P. × suffruticosa ‘Luoyang Hong’184. PsACS1 was the key gene involved in ethylene biosynthesis in tree peony cut flowers, and PsACO1 was mainly regulated at the posttranscriptional level during flower opening184,185. In addition, glucose relies on the hexokinase signal transduction pathway to inhibit the transcription of PsACS1, thus delaying the senescence of tree peony cut flowers186. The expression of PsCTR2 and PsCTR3 in tree peony increased under the effect of ethylene, and the application of an ethylene signaling inhibitor, 1-MCP, could reverse the process, indicating that PsCTR2 and PsCTR3 play a negative regulatory role in response to the ethylene signal187. The quality of P. × suffruticosa ‘Luoyang Hong’ cut flowers was improved by exogenous use of 500 mM glucose, which promoted the expression of PsCTR3 and inhibited the accumulation of PsEIL3 transcripts, thereby reducing the sensitivity of cut flowers to ethylene and prolonging the vase life of cut flowers188. Subsequent studies also confirmed that the transcription factors PsEIL3 (triggering transcription of downstream target genes) and PsERF1 have positive effects on the response to ethylene, playing an important role in the opening and senescence of tree peony cut flowers188,189.
Some studies have also been conducted on the genes related to aquaporins and glycometabolism in Paeonia postharvest. Zhao et al.190 isolated three aquaporin genes, PlPIP1;2, PlPIP2;1 and PlNIP, from P. lactiflora ‘Hongyan Zhenghui’ cut flowers. Nanosilver prolonged the vase life of cut flowers, and the expression of PlPIP1;2 was higher than that in the control group, while the expression of PlPIP2;1 and PlNIP was lower than that in the control group, indicating that aquaporins synergistically maintain the water balance of cut flowers. The same effect of aquaporins was also proven by Xue et al.191. When adding sucrose (20 g/L) to the vase solution, the flowering duration of P. lactiflora ‘Yang Fei Chu Yu’ cut flowers increased by ~1.3 days. Xue et al.192 isolated two sucrose transporter genes (PlSUT2 and PlSUT4) and five invertase genes (PlCWIN1, PlVIN1, PlCIN1, PlCIN2, and PlCIN3) from P. lactiflora ‘Yang Fei Chu Yu’: PlSUT2 and PlSUT4 were induced by sucrose and played a role in sucrose transport, while PlCWIN1 and PlVIN1 were involved in the accumulation of glucose and fructose during the flowering of cut flowers. A study on P. lactiflora ‘Yang Fei Chu Yu’ cut flowers showed that the increased expression of PlSPS1 and PlSPS4 (coding for starch hydrolase) and the decreased expression of PlSUS4 (coding for sucrose synthase) led to starch hydrolysis, increased the amount of sucrose in cut flowers and improved the vase quality of cut flowers of P. lactiflora ‘Yang Fei Chu Yu’193.
Bud dormancy
Paeonia flower buds typically have internal dormancy. Studies on the mechanism of dormancy in flower buds provide a theoretical basis for artificially breaking dormancy and regulating flowering time. Huang constructed a subtractive cDNA library of the genes related to dormancy release in flower buds of P. × suffruticosa ‘Lu He Hong’194. Eight genes (PsPII, PsDHN, PsGA20, PsARP, PsMPT, PsCXE, PsSERK1 and PsPOB) were screened and may be related to the release from internal dormancy. Based on differential gene expression and transgenic analyses, changes in the expression of PsARP may have been related to the transformation between conjugated and free auxin in flower buds, regulating flower bud dormancy at the posttranscriptional level, while PsMPT regulated ATP synthesis and promoted the breaking of bud dormancy in tree peony. Zhang et al. obtained the PsMPT promoter, and in transgenic plants containing a 421-bp PSMPT promoter, the expression and activity of the GUS gene increased significantly after freezing treatment. When the fragment from −421 to −408 base pairs (bp) containing an MYC cis-element of the PSMPT promoter was deleted, the chilling response was not observed. This result confirmed that the MYC element was a key motif for the chilling response of the PsMPT promoter195. In other studies, it was also found that PsPII, PsSERK2, GA20ox, SOC1, PsGRAS1, and PsGRAS2 may play roles in relieving Paeonia flower bud dormancy196,197,198,199,200,201,202. However, only an analysis of differential expression of genes from different treatments was carried out, and their functions were not further verified by transgenic studies196,197,198,199,200,201,202.
DNA methylation in flower buds also inhibits the release of dormancy in Paeonia. Under the action of exogenous 5-azacytidine (5-azaC), the expression of the DNA methyltransferase genes PsCMT3, PsMET1, and PsDRM2 in dormant buds was inhibited, and the transcription of the demethylase gene PsROS1 increased, thus reducing the DNA methylation level and activating the cell cycle, DNA replication and glycol metabolism processes, which accelerated the dormancy release of buds203,204.
Seed dormancy
Paeonia seeds are dormant, and breaking dormancy is of great importance for improving the reproductive efficiency of Paeonia. Research on the molecular mechanisms of seed dormancy in Paeonia is scarce, and the main focus has been on the metabolic pathway of hormones. ABA is an important hormone that regulates seed dormancy. During seed development, ABA expression increases first and then decreases205. PoNCED1 and PoZEP1 may jointly inhibit ABA biosynthesis in P. ostii ‘Feng Dan’ seeds. PoSDR1 induces ABA synthesis in the middle and later stages (104 days to 132 days after pollination). The highest expression of PobZIP1 in ‘Fengdan’ seeds occurred at an early stage of development (earlier than 90 days after pollination), and the highest ABA content occurred later than the highest expression of PobZIP1, suggesting that the transcription factor PobZIP1 played a negative role in dormancy release205,206. The expression of PlCYP70A1 and PlCYP70A2, which are ABA 8′-hydroxylase genes, in the seeds of P. lactiflora ‘Fen Yu Nu’ increased during seed germination, promoted ABA degradation, and played a positive role in breaking dormancy207. The protein phosphatase gene PlPP2C may have a similar mechanism during P. lactiflora ‘Fen Yu Nu’ seed germination208.
Differential gene analysis was performed on the transcriptome of two developmental stages (0 and 40 days) of P. lactiflora ‘Fen Yu Nu’ seeds. The expression of PlGAI1, a member of the DELLA protein gene family that regulates gibberellin biosynthesis, was significantly upregulated during seed germination and thus played a positive role in seed germination209. Ren et al.210 found that the expression of PoGAPC associated with glycolysis in P. ostii ‘Feng Dan’ seeds continued to be suppressed at a low temperature (4 °C), indicating that delayed glycolysis played a key role in dormancy at low temperatures. Auxin may also play a role in the germination of P. lactiflora ‘Fen Yu Nu’ seeds: the expression of the auxin response gene PlSAUR1 was highest in seeds in the period after germination of the seed hypocotyl, PlSAUR2 and PlSAUR3 expression was highest in the period after germination of the seed epicotyl, and PlSAUR4 expression was highest after seed imbibition211.
Conclusions and perspectives
There are still some doubts pertaining to the taxonomy of Paeonia, especially in sect. Paeonia. Taxonomists also need to carry out more in-depth and detailed research on Paeonia. Due to anthropogenic disturbance, many natural distribution areas of wild species of Paeonia are sharply shrinking, and some species are endangered212,213. Therefore, conservation efforts are essential. Some wild Paeonia species, such as P. ludowii, P. sterniana, and P. parnassica, have relatively strict habitat requirements, and their areas of distribution are very narrow, so the most effective protection strategy for them is in situ conservation. To restore wild populations as soon as possible, artificial pollination and seeding are encouraged, while the harvesting of wild plants for any purpose should be prohibited. Ex situ conservation is also an effective method for protecting rare and endangered species of Paeonia, and some significant conservation efforts and achievements have been made in China67.
At present, the number of peony cultivars exceeds 8000, but the utilization of wild resources in breeding is very limited. In sect. Moutan, P. decomposita, P. rotundiloba, P. rockii subsp. rockii, and P. ludlowii are rarely used in breeding46,47; in sect. Paeonia, only approximately seven species (P. lactiflora, P. officinalis, P. peregrina, P. daurica subsp. macrophylla, subsp. mlokosevitschii, P. tenuifolia, P. emodi, and P. anomala) were used in breeding68; and in sect. Onaepia, no cultivars have been bred. Wild Paeonia species are widely distributed around the world, and their phenotypic traits are quite different. The utilization of wild germplasm resources is very important for the development of new cultivars. For example, the use of P. lutea and P. delavayi allowed for the creation of a new cultivar group, Lutea hybrid peony. In addition, the use of parents from different sections in cross-breeding is still very limited, and a large number of wild species and cultivars have still not been utilized. Future breeding efforts should focus on diversification to enrich the gene pool of peony hybrids.
There are still many deficiencies in Paeonia cultivars, such as a relatively narrow flowering period, the unpleasant fragrance of some cultivars, or a limited cultivation range. Therefore, it is necessary to strengthen Paeonia breeding by prolonging the flowering period, creating cultivars with a pleasant fragrance and improving the resistance of cultivars to biotic and abiotic stresses.
The propagation of tree peony depends mainly on division and grafting, but propagation efficiency is very low. To improve the propagation efficiency of Paeonia, many teams have employed tissue culture, but multiple problems have been encountered: explant browning, difficulties in rooting, bottlenecks in the induction of shoots from callus, and the lack of somatic embryogenic protocols127,128,129,130. To mass produce Paeonia cultivars, embryo rescue of seeds from progeny of distantly hybridized germplasm is required214,215. In the future, more attention should be paid to the tissue culture of Paeonia, both herbaceous and tree peonies216,217.
Production of the current cultivars of Paeonia depends almost exclusively on traditional breeding methods, but as modern molecular biotechnology develops, it will be necessary to strengthen and modernize molecular breeding research on Paeonia. Many functional genes and transcription factors have been discovered and employed in Paeonia research. It is necessary to establish a stable genetic transformation system for Paeonia as soon as possible to introduce the functional genes of target characters into homologous plants and to carry out directional breeding. The number of genetic populations of Paeonia is very small. Parents with specific traits should be selected to create more abundant genetic populations. QTL mapping or GWASs should be carried out in genetic populations by using multicomponent and performance data to locate the genes and linkage markers associated with target traits. The application of validated markers for specific traits will expedite the breeding process. Many breeding techniques have been applied to model plants and crops, such as magnetic nanoparticle-mediated transgenic technology218,219,220 or carbon nanotube-mediated DNA delivery technology221, which can be applied to peony breeding without the need to establish a stable genetic transformation system. The draft genome of P. × suffruticosa ‘Luo shen xiao chun’ has been published222. The first genomic data of Paeonia will play a positive role in promoting the modern molecular breeding of members of this genus.
References
Worsdell, W. C. A study of the vascular system in certain orders of the Ranales. Ann. Bot.22, 663 (1908).
Pan, K. Y. Paeonia, In Flora of China (ed. Wu, Z. Y.) Vol. 27, pp. 37–59 (Science Press, Beijing, 1979).
Pan, K. Y. The analysis of distribution pattern in the Paeoniaceae and its formation. Acta Phytotaxon. Sin.33, 340–349 (1995).
Hong, D. Y. Peonies of the world: taxonomy and phytogeography. (Royal Botanical Garden, Kew, London, 2010).
Li, J. J. Chinese tree peony and herbaceous peony. (China Forestry Press, Beijing, 1999).
Li, J. J., Zhang, X. F. & Zhao, X. Q. Tree peony of China. (Encyclopedia of China Publishing House, Beijing, 2011).
Page, M. The gardener’s peony: Herbaceous and tree peonies. (Timber Press, Portland, 2005).
Cheng, F. Y. & Li, J. J. Exportation of Chinese tree peonies (Mudan) and their development in other countries I: cultivated. J. Northwest Nor. Univ.34, 109–116 (1998).
Bigger, M. D. et al. The Peonies. (The American Horticultural Society, Washington, 1962).
Kessenich, G. M. & Hollingsworth, D. The American hybrid peony. (The American Peony Society, Minnesota State, 1990).
Wang, L. Y., Yuan, T., Li, Q. D. & Zhao, X. Z. Chinese tree peony (Continuation). (China Forestry Publishing House, Beijing, 2015).
Yu, X. N. Herbaceous Peonies. (China Forestry Publishing House, Beijing, 2019).
Kamenetsky, R. & Dole, J. Herbaceous peony (Paeonia): genetics, physiology and cut flower production. Floric. Ornam. Biotechnol.6, 62–77 (2012).
Linnaeus, C. Species Plantarum. 530. (Impensis Laurentii Salvii, Stockholm, 1753).
Andrews, H. C. Paeonia suffruticosa. Bot Repos. Vol. 6, tab. 373 (1804).
Anderson, G. A monograph of the genus Paeonia. Trans. Linn. Soc. Lond.12, 248–283 (1818).
Hooker, W. J. Flora Boreali-americana. Vol. 1. (Treuttel & Würtz, London, 1829).
Lynch, R. I. A new classification of the genus Paeonia. J. R. Hortic. Soc.12, 428–445 (1890).
Stern, F. C. Genus Paeonia. J. R. Hortic. Soc.68, 124–131 (1943).
Fang, W. P. Notes on Chinese Paeonies. Acta Phytotax. Sin.7, 297–323 (1958).
Sang, T., Crawford, D. J. & Stuessy, T. F. Documentation of reticulate evolution in peonies (Paeonia) using internal transcribed spacer sequences of nuclear ribosomal DNA: implications for biogeography and concerted evolution. Proc. Natl Acad. Sci. USA92, 6813–6817 (1995).
Sang, T., Crawford, D. J. & Stuessy, T. F. Chloroplast DNA phylogeny, reticulate evolution, and biogeography of Paeonia (Paeoniaceae). Am. J. Bot.84, 1120–1136 (1997).
Sang, T., Donoguhe, M. J. & Zhang, D. M. Evolution of alcohol dehydrogenase genes in peonies (Paeonia): phylogenetic relationship of putative nonhybrid species. Mol. Biol. Evol.14, 994–1007 (1997).
Stern, F. C. A study of the genus Paeonia. (The Royal Horticultural Society, London, 1946).
Franchet, A. R. Plantae Yunnanenses. Bull. Soc. Bot. Fr.33, 382–383 (1886).
Brühl, P. J. Some new or critical Ranunculaceae from India and adjacent regions. Ann. R. Bot. Gard.5, 114–115 (1896).
Rehder, A. New species, varieties and combinations from the herbarium and the collections of the Arnold Arboretum: Paeonia suffruticosa var. spontanea. J. Arnold. Arbor.1, 193–194 (1920).
Handel-Mazzetti, H. F. von. Plantae Sinenses (Paeonia). Acta Hortic. Gothob.13, 37–40 (1939).
Stern, F. C. & Taylor, G. Paeonia lutea var. ludlowii. Bot. Mag.169, 209 (1953).
Hong, T., Zhang, J. X., Li, J. J., Zhao, W. Z. & Li, M. R. Study on the Chinese wild woody peonies (1): new taxa of Paeonia L. sect Moutan DC. Bull. Bot. Res.12, 223–234 (1992).
Hong, T. & Osti, G. L. Study on the Chinese wild woody peonies (II): new taxa of Paeonia L. sect Moutan DC. Bull. Bot. Res14, 237–240 (1994).
Pei, Y. L. & Hong, D. Y. Paeonia qiui – a new woody species of Paeonia from Hubei, China. Acta Phytotax. Sin.33, 91–93 (1995).
Hong, T. & Dai, Z. L. Study on the Chinese wild woody Peonies (III): new taxa of Paeonia L. sect Moutan DC. Bull. Bot. Res.17, 1–5 (1997).
Hong, D. Y. Paeonia (Paeoniaceae) in Xizang (Tibet). Novon7, 156–161 (1997).
Hong, D. Y. Notes on Paeonia decomposita Hand.-Mazz. Kew Bull.52, 957–963 (1997).
Hong, D. Y., Pan, K. Y. & Yu, H. Taxonomy of Paeonia delavayi complex (Paeoniaceae). Ann. Missouri Bot. Gard.85, 554–565 (1998).
Hong, D. Y. & Pan, K. Y. A revision of the Paeonia suffruticosa complex (Paeoniaceae). Nord. J. Bot.19, 289–299 (1999).
Hong, D. Y. & Pan, K. Y. Taxonomical history and revision of Paeonia sect. Moutan (Paeoniaceae). Acta Phytotax. Sin.37, 351–368 (1999).
Hong, D. Y., Pan, K. Y. & Turland, N. J. Paeoniaceae, In flora of China (eds Wu, Z., Raven, P. H. & Hong, D. Y.) Vol. 6, p. 127–132. (Science Press and Missouri Botanic Garden Press, Beijing and St. Louis, 2001).
Hong, D. Y. Peonies of the world: polymorphism and diversity. (Royal Botanical Gardens, Kew, London, 2011).
Hong, D. Y. & Pan, K. Y. Paeonia cathayana, a new tree peony, with revision of P. suffruticosa subsp. yinpingmudan. Acta Phytotax. Sin.45, 285–288 (2007).
Zhang, J. M. Population genetics of Paeonia Sect. Moutan Subsect. Delavayanae (Institute of Botany, Chinese Academy of Science, 2010).
Hong, D. Y. & Pan, K. Y. Notes on taxonomy of Paeonia sect. Moutan DC. (Paeoniaceae). Acta Phytotax. Sin.43, 169–177 (2005).
Yuan, T. Studies on genetic relationship of some species and cultivars/ cultivars groups of chinese tree peony (Beijing Forestry University, 1998).
Zhou, Z. Q., Pan, K. Y. & Hong, D. Y. Advances in studies on relationships among wild tree peony species and the origin of cultivated tree peonies. Acta Hortic. Sin.30, 751–757 (2003).
Cheng, F. Y. Advances in the breeding of tree peonies and a cultivar system for the cultivar group. Int. J. Plant Breed.1, 89–104 (2007).
Zhou, S. L. et al. Multiple species of wild tree peonies gave rise to the ‘King of Flowers’, Paeonia suffruticaosa Andrews. Proc. Biol. Sci.281, 1687 (2014).
Xia, T. Molecular phylogeny of Paeoniaceae - With special emphasis on Paeonia Sect. Paeonia (Institute of Botany, Chinese Academy of Science, 2008).
Liu, J. A study on phylogeny and teraploid speciation in Paeonia (Institute of Botany, Chinese Academy of Science, 2011).
Hong, D. Y., Pan, K. Y. & Li, X. Y. Paeonia in Xinjiang, China. Acta Phytotax. Sin.33, 349–355 (1994).
Hong, D. Y. A subspecies of Paeonia mascula (Paeoniaceae) from W. Asia and SE. Europe. Acta Phytotax. Sin.38, 381–385 (2000).
Hong, D. Y., Pan, K. Y. & Rao, G. Y. Cytogeography and taxonomy of the Paeonia obovata polyploid complex. Plant Syst. Evol.227, 123–136 (2001).
Hong, D. Y. & Pan, K. Y. Paeonia anomala subsp. veitchii, a new combination and its biology. Novon11, 315–318 (2001).
Hong, D. Y. & Zhou, S. L. Paeonia (Paeoniaceae) in the Caucasus. Bot. J. Linn. Soc.143, 135–150 (2003).
Hong, D. Y. & Pan, K. Y. A taxonomic revision of the Paeonia anomala complex (Paeoniaceae). Ann. Missouri Bot. Gard.1, 87–98 (2004).
Hong, D. Y., Wang, X. Q. & Zhang, D. M. Paeonia saueri (Paeoniaceae), a new species from the Balkans. Taxon53, 83–90 (2004).
Hong, D. Y. & Castroviejo, S. Proposal to conserve the name Paeonia broteri against P. lusitanica Mill. Taxon54, 211–212 (2005).
Hong, D. Y., Wang, X. Q., Zhang, D. M. & Seleuk, T. K. On the circumscription of Paeonia kesrouanensis, an east Mediterranean peony. Nord. J. Bot.23, 395–400 (2005).
Hong, D. Y. & Wang, X. Q. The identity of Paeonia corsica Sieber ex Tausch (Paeoniaceae), with special reference to its relationship with P. mascula (L.) Mill. Fedd. Repert.17, 65–84 (2006).
Pan, J. Variation and evolution in Paeonia Sect. Paeonia (Paeoniaceae): Evidence from morphology, chromosome and gene sequences (Institute of Botany, Chinese Academy of Sciences, 2006).
Stebbins, G. L. Cytogenetic studies in Paeonia. II. The cytology of the diploid species and hybrids. Genetics23, 83–110 (1938).
Stebbins, G. L. & Ellerton, S. Structural hybridity in Peaonia californica and P. brownii. J. Genet.38, 1–36 (1939).
Wang, L. Y. Chinese tree peony. (China Forestry Publishing House, Beijing, 1997).
Yuan, T. & Wang, L. Y. Discussion on the origination of Chinese tree peony cultivars according to pollen grain morphology. J. Beijing. Univ.24, 5–11 (2002).
Yuan, T. & Wang, L. Y. The morphological analysis of the origin of Chinese cultivated tree peonies. Shangdong. Sci. Tech.6, 1–3 (2004).
Japan Peony Association. Modern Japanese tree peony and herbaceous peony. (Kohansha Co. Ltd., Tokyo, 1995).
Wang, L. Y., Yuan, T., Wang, F. & Li, Q. D. The ex-situ conservation center of chinese Paeoniaceae wild species and cultivation of new cultivars. (China Forestry Publishing House, Beijing, 2013).
America Peony Society (APS). Peony registry. https://americanpeonysociety.org/cultivars/peony-registry/ (Accessed 18 May 2020)
Qin, K. J. Herbaceous peonies. (China Forestry Publishing House, Beijing, 2004).
Yuan, Q. L. The Culture of chinese herbaceous peony (Beijing Forestry University, 2011).
Guo, X. F., Zang, D. K., Yuan, T., Liu, X. K. & Wang, L. Y. Discussion about the origin of cultivated herbaceous peonies native to China - Based on the comparative study of the geographical distribution and its morphological characters. J. Shandong Agric. Univ. (Nat. Sci.)39, 388–392 (2008).
Smith, D. R. Peony intersectional crosses; a brief review of the progress over the first 50 years. Paeonia27, 6–9 (1997).
Sun, J. F. Introduction and propagation of distant hybrids in Paeonia (Beijing Forestry University, 2007).
Henan Government. The 33rd China Luoyang Peony Culture Festival has come to a successful conclusion and more than 21.74 million visitors were received. (HG, Zhengzhou, 2015). https://www.henan.gov.cn/2015/05-06/568101.html (Accessed 18 May 2020)
Royal Flora Holland. Summary of varieties, quantities and prices of peony cut flowers in recent three years. (RFH, Amsterdam, 2020). https://www.royalfloraholland.com/en (Accessed 18 May 2020)
Luo, Y. B., Pei, Y. L., Pan, K. Y. & Hong, D. Y. A study on pollination biology of Paeonia suffruticosa subsp. spontanea (Paeoniaceae). Acta Phytotax. Sin.36, 134–144 (1998).
Hong, Y. & Liu, Q. A study on the pollination biology of Paeonia lactiflora Pall. Guihaia26, 120–124 (2006).
Bernhardt, P., Meier, R. & Vance, N. Pollination ecology and floral function of Brown’s peony (Paeonia brownii) in the Blue Mountains of northeastern Oregon. J. Poll. Ecol.11, 9–20 (2013).
Li, K., Zheng, B. Q., Wang, Y. & Guo, X. Flowering characteristics and breeding system of Paeonia delavayi. J. Northeast For. Univ.41, 63–67 (2013).
Yang, Y. et al. Studies on floral characteristics and breeding system of Paeonia decomposita. J. Plant Resour. Environ.24, 97–104 (2015).
Cheng, F. Y. & Chen, D. Z. Studies on the selection and breeding of new hybrids from blotched tree peony (Paeonia rockii cvs.) and the cultivars classification of tree peony. J. Beijing. Univ.20, 27–32 (1998).
Li, C. et al. Flavonoid composition and antioxidant activity of tree peony (Paeonia section Moutan) yellow flowers. J. Agric. Food Chem.57, 8496–8503 (2009).
Luo, X. N. et al. Variation of floral volatiles and fragrance reveals the phylogenetic relationship among nine wild tree peony species. Flavour Fragr. J.35, 227–241 (2020).
Li, S., Chen, L., Xu, Y., Wang, L. & Wang, L. Identification of floral fragrances in tree peony cultivars by gas chromatography–mass spectrometry. Sci. Hortic.142, 158–165 (2012).
Smith, D. R. Breeding tree peonies for fragrance. Paeonia27, 1–2 (1997).
Zhang, L. Construction of high-density genetic linkage map based on Gbs technique in tree peony (Henan University of Science and Technology, 2016).
Zhang, L., Guo, L. L., Guo, D. L. & Hou, X. G. Separation analysis and mixed genetic analysis of phenotypic traits in F1 progenies of tree peony. J. Nanjing. Univ. (Nat. Sci. Ed.)42, 51–60 (2018).
Cai, C. F. et al. Selecting optimal F1 segregation population for genetic linkage mapping in tree peony. J. Beijing. Univ.37, 139–147 (2015).
Cai, C. F., Cheng, F. Y., Wu, J., Zhong, Y. & Liu, G. X. The first high-density genetic map construction in tree peony (Paeonia sect. Moutan) using genotyping by specific-locus amplified fragment sequencing. PLoS ONE10, e0128584 (2015).
Peng, L. P. et al. Genetic analyses reveal independent domestication origins of the emerging oil crop Paeonia ostii, a tree peony with a long-term cultivation history. Sci. Rep.7, 5340 (2017).
Zhang, L. et al. Construction of a high-density genetic map and QTLs mapping with GBS from the interspecific F1 population of P. ostii ‘Fengdan Bai’ and P. suffruticosa ‘Xin Riyuejin’. Sci. Hortic.246, 190–200 (2019).
Guo, Q. et al. Construction of a genetic linkage map in tree peony (Paeonia Sect. Moutan) using simple sequence repeat (SSR) markers. Sci. Hortic.219, 294–301 (2017).
Li, S. M. et al. Construction of a high-density genetic map of tree peony (Paeonia suffruticosa Andr. Moutan) using restriction site associated DNA sequencing (RADseq) approach. Tree Genet. Genomes15, 63 (2019).
Suo, Z. L., Zhou, S. L., Zhang, H. J. & Zhang, Z. M. DNA molecular evidences of the inter-specific hybrids between Paeonai ostii and P. suffruticosa based on ISSR markers. Res.17, 700–705 (2004).
Zhang, D. Distant cross-breeding in tree peony and identification of some hybrids by AFLP (Beijing Forestry University, 2008).
Hao, Q. et al. Studies on Paeonia cultivars and hybrids identification based on SRAP analysis. Hereditas145, 38–47 (2008).
Ji, L. J., Wang, Q., Teixeira da Silva, J. A. & Yu, X. N. The genetic diversity of Paeonia L. Sci. Hortic.143, 62–74 (2012).
Wu, J., Cheng, F. Y. & Zhang, D. Utilizing ‘High Noon’ in the crossing breeding of tree peonies and early identification of some hybrids by AFLP markers. Acta Bot. Boreal. Occident. Sin.33, 11551–11557 (2013).
Hao, Q. et al. Crossability of American tree peony ‘High Noon’ as seed parent with Japanese cultivars to breed superior cultivars. Euphytica191, 35–44 (2013).
Yuan, J., Cornille, A., Giraud, T., Cheng, F. & Hu, Y. Independent domestications of cultivated tree peonies from different wild peony species. Mol. Ecol.23, 82–95 (2014).
Wu, J., Cheng, F. Y., Pang, L. Z., Zhong, Y. & Cai, C. F. Association analysis of phenotypic traits with SSR markers in Paeonia rockii. J. Beijing For. Univ.38, 80–87 (2016).
Wu, J., Cheng, F. Y., Cai, C. F., Zhong, Y. & Jie, X. Association mapping for floral traits in cultivated Paeonia rockii based on SSR markers. Mol. Genet. Genomics292, 187–200 (2016).
Liu, J. X. Compatibility Of distant hybridization in Paeonia L. and identification of F1 hybrid progenies (Beijing Forestry University, 2016).
Cai, C. F. High-density genetic linkage map construction and Qtls analyses for phenotypic traits in tree peony (Beijing Forestry University, 2015).
Huo, Z. P. et al. Preliminary report of tree peony and herbaceous peony space breeding research, In The Proceedings of International Tree Peony Summit Form (2013 Luoyang China) (eds Wang, L. Y. & Yuan, T.) p. 108–112 (China Forestry Publishing House, Beijing, 2013).
Wei, D., Tang, Z., Teixeira da Silva, J. A. & Yu, X. In vitro induction of polyploidy by colchicine treatment in herbaceous peony (Paeonia lactiflora Pall.). Acta Hortic.1171, 265–278 (2017).
Yu, H. Some problems in systematizing cultivar resources of Chinese tree peony. Acta Hortic. Sin.9, 65–68 (1982).
Cao, X. J. Introduction and utilization of the advanced generation hybrids of tree peonies and Paeonia Delavayi (Beijing Forestry University, 2010).
Wang, Y., Teixeira da Silva, J. A., Liu, J., Lu, Y. & Yu, X. Evaluation of pollen viability and fertility of triploid herbaceous peony cultivars. Acta Hortic.1171, 137–142 (2017).
Smirnow, L. Remarkable tree peony discoveries. In the best of 75 years (ed. Kessenich, G. M.) Vol. 52 (The American Peony Society, Minnesota State, 1979).
James, W. Improbable peonies. Horiculture March/April. p. 44–49 (2003).
Smith, D. The impossible dream. Bull. Am. Peony Soc.330, 79–109 (2004).
Smith, D. (Martha Washington × Golden Era) intersectional hybrids revisited or “what a difference a year makes”. Paeonia31, 3–6 (2001).
Smith, D. Updated hybridizers list of named and registered intersectional hybrids. Paeonia31, 3–4 (2001).
Zhong, Y. et al. Five new peony cultivars bred from inter-sectional hybrids in Paeonia. Sci. Silvae Sin.52, 164 (2016).
Smith, D. Lactiflora × Suffruticosa and the reverse. Paeonia28, 3–6 (1998).
Hao, Q., Liu, Z. A., Shu, Q. Y., Wang, L. S. & Chen, F. F. Identification of intersectional hybrid between section Moutan and section Paeonia found in China for the first time. Acta Hortic. Sin.35, 853–858 (2008).
Smith, D. Intersectional hybrids from the reciprocal cross. Paeonia31, 1–2 (2001).
Entsminger, H. Lady tessera and prairie moon. Paeonia27, 1–3 (1997).
Smith, D. And then there were two. Paeonia27, 3–6 (1997).
Zhang, D. Distant cross-breeding in tree peony and identification of some hybrids by AFLP (Beijing Forestry University, 2008).
Wang, Y. L. Cross-breeding in tree peony and fertility research of intersectional hybrids (Beijing Forestry University, 2009).
Liu, X. Studies on parental selection of tree peonies and early identification of several hybrids (Beijing Forestry University, 2016).
Yang, L. H. et al. Ploidy identification and karyotype analysis of five Itoh hybrid peonies. Bull. Bot. Res.37, 535–541 (2017).
Yang, L., Zhang, J., Teixeira da Silva, J. A. & Yu, X. Variation in ploidy and karyological diversity in different herbaceous peony cultivar groups. J. Am. Soc. Hortic. Sci.142, 272–278 (2017).
Smith, D. Examining the fertility of the intersectional hybrid. Paeonia29, 1–7 (1999).
Beruto, M., Lanteri, L. & Portogallo, C. Micropropagation of tree peony (Paeonia suffruticosa). Plant Cell Tiss. Org. Cult.79, 249–255 (2004).
Beruto, M. & Curir, P. In vitro culture of tree peony through axillary budding. In protocols for micropropagation of woody trees and fruits (eds Jain, S. M. & Haggman, H.) (Springer, Berlin, 2007).
Wen, S. S. et al. Efficient protocols for the micropropagation of tree peony (Paeonia suffruticosa ‘Jin Pao Hong’, P. suffruticosa ‘Wu Long Peng Sheng’, and P. × lemoinei ‘High Noon’) and application of arbuscular mycorrhizal fungi to improve plantlet establishment. Sci. Hortic.201, 10–17 (2016).
Wen, S. S., He, R. R., Zheng, J. K. & Tian, R. N. Research advance in tissue culture of tree peony. Sci. Silvae Sin.54, 143–155 (2018).
Dai, S. L. Landscape plant genetics. (China Forestry Publishing House, Beijing, 2005).
Wang, L. S. et al. Phenetics in tree peony species from China by flower pigment cluster analysis. J. Plant Res.114, 213–221 (2001).
Wang, L. S. et al. Analysis of petal anthocyanins to investigate flower coloration of Zhongyuan (Chinese) and Daiko Island (Japanese) tree peony cultivars. J. Plant Res.114, 33–43 (2001).
Wang, L. S. et al. Chemical taxonomy of the Xibei tree peony from China by floral pigmentation. J. Plant Res.117, 47–55 (2004).
Zhang, J. J., Wang, L. S. & Liu, Z. A. Recent advances in flower color research of tree peony. Acta Hortic. Sin.6, 1383–1388 (2006).
Zhang, J. et al. Comparison of anthocyanins in non-blotches and blotches of the petals of Xibei tree peony. Sci. Hortic.114, 104–111 (2007).
Jia, N. et al. Analysis of petal anthocyanins to investigate coloration mechanism in herbaceous peony cultivars. Sci. Hortic.117, 167–173 (2008).
Jia, N. et al. Identification and characterization of anthocyanins by high-performance liquid chromatography-electrospray ionization-mass spectrometry in herbaceous peony species. J. Am. Soc. Hortic. Sci.133, 418–426 (2008).
Li, C. H. The flavonoid composition in tree peony petals and their effects on the coloration (Institute of Botany, Chinese Academy of Sciences, 2010).
Zhao, D. Q., Wei, M. R., Liu, D. & Tao, J. Anatomical and biochemical analysis reveal the role of anthocyanins in flower coloration of herbaceous peony. Plant Physiol. Biochem.102, 97–102 (2016).
Shi, M. Expression vector construction and transformation of model plants with flavonid synthesis related genes in herbaceous peony (Paeonia lactiflora Pall.) (Yangzhou University, 2017).
Zhang, X. P., Zhao, L. Y., Xu, Z. D. & Yu, X. Y. Transcriptome sequencing of Paeonia suffruticosa ‘Shima Nishiki’ to identify differentially expressed genes mediating double-color formation. Plant Physiol. Biochem.123, 114–124 (2018).
Du, H. et al. Methylation mediated by an anthocyanin, O-methyltransferase, is involved in purple flower coloration in Paeonia. J. Exp. Bot.66, 6563–6577 (2015).
Zhao, D. Q. et al. Transcriptome sequencing of a chimaera reveals coordinated expression of anthocyanin biosynthetic genes mediating yellow formation in herbaceous peony (Paeonia lactiflora Pall). BMC Genom.15, 689 (2014).
Guo, L. P., Wang, Y. J., Teixeira da Silva, J. A., Fan, Y. M. & Yu, X. N. Transcriptome and chemical analysis reveal putative genes involved in flower color change in Paeonia ‘Coral Sunset’. Plant Physiol. Biochem.138, 130–139 (2019).
Li, X. Cloning and functional analysis of MYB genes in variegation formation of tree peony (Paeonia rockii) (Northwest A&F University, 2019).
Gan, L. X. Analysis of the composition of three color peony Anthocyanins and related gene expression (Northwest A&F University, 2019).
Gu, Z. Y. et al. A novel R2R3-MYB transcription factor contributes to petal blotch formation by regulating organ-specific expression of PsCHS in tree peony (Paeonia suffruticosa). Plant Cell Physiol.60, 599–611 (2019).
Luo, J. R., Shi, Q., Niu, L. & Zhang, Y. Transcriptomic analysis of leaf in tree peony reveals differentially expressed pigments genes. Molecules22, 324 (2017).
Luo, J. et al. Transcriptomic analysis reveals transcription factors related to leaf anthocyanin biosynthesis in Paeonia qiui. Molecules22, 2186 (2017).
Duan, J. J., Luo, J. R., Li, X., Zhang, Q. Y. & Zhang, Y. L. Analysis of pigment changes and related gene expression during the red faded of tree peony leaves in spring. Acta Bot. Boreal. Occident. Sin.38, 1885–1894 (2018).
Li, Y., Lu, J. X., Chang, Y. H., Tang, W. L. & Yang, Q. S. Comparative analysis of tree peony petal development by transcriptome sequencing. Acta Physiol. Plant.39, 216 (2017).
Tang, W. L. Bioinformatics analysis of MADS-box gene family and expression analysis of genes involving to floral organs development in tree peony (Henan Agricultural University, 2018).
Zhang, L. et al. Expression analysis and functional identification of PsAP1 in tree peony. Acta Agric. Boreal. Sin.30, 84–89 (2015).
Ge, J. T. et al. Cloning and expression of floral organ development-related genes in herbaceous peony (Paeonia lactiflora Pall.). Mol. Biol. Rep.41, 6493–6503 (2014).
Wu, Y. Q. et al. Combination of transcriptome sequencing and iTRAQ proteome reveals the molecular mechanisms determining petal shape in herbaceous peony (Paeonia lactiflora Pall.). Biosci. Rep.38, BSR20181485 (2018).
Shu, Q. T. et al. Analysis of the formation of flower shapes in wild species and cultivars of tree peony using the MADS-box subfamily gene. Gene493, 113–123 (2011).
Ren, L. Cloning and expression of floral organ development related genes in tree peony (Chinese Academy of Forestry, 2011).
Wang, Y., Ma, Y. L. & Dai, S. L. The molecular mechanism in regulation of flowering in ornamental plants. Chin. Bull. Bot.45, 641–653 (2010).
Wang, S. L. et al. Molecular cloning, expression and evolutionary analysis of the flowering regulating transcription factor gene PsCOL4 in tree peony. Acta Hortic. Sin.41, 1409–1417 (2014).
Wang, S. L. et al. Molecular characterization and expression patterns of PsSVP genes reveal distinct roles in flower bud abortion and flowering in tree peony (Paeonia suffruticosa). Can. J. Plant Sci.94, 1181–1193 (2014).
Wu, Y. Q., Ge, J. T. & Tao., J. cDNA cloning, sequence analysis and tissue expression detection of APETALA1 gene (AP1) in Paeonia lactiflora Pall. petals of different development stages. J. Agric. Biol.23, 1559–1567 (2015).
Zhou, H., C F, Y., W, J. & He, C. Y. Isolation and functional analysis of Flowering Locus T in tree peonies (PsFT). J. Am. Soc. Hortic. Sci.140, 265–271 (2015).
Wang, S. L. et al. De novo sequencing of tree peony (Paeonia suffruticosa) transcriptome to identify critical genes involved in flowering and floral organ development. BMC Genom.20, 572 (2019).
Wei, W. Q. et al. Molecular cloning and expression analysis of flowering-regulating transcription factor gene PsFUL1 in tree peony (Paeonia suffruticosa). Plant Physiol. J.53, 536–544 (2017).
Zhu, F. Y. et al. Isolation of florigen gene PdFT and its effects on flowering of tree peony (Paeonia delavayi Franch.). Sci. Agric. Sin.47, 2613–2624 (2014).
Xue, J. Q. et al. Elucidation of the mechanism of reflowering in tree peony (Paeonia suffruticosa) ‘Zi Luo Lan’ by defoliation and gibberellic acid application. Plant Physiol. Biochem.132, 571–578 (2018).
Xue, J. Q. et al. Defoliation and gibberellin synergistically induce tree peony flowering with non-structural carbohydrates as intermedia. J. Plant Physiol.233, 31–41 (2019).
Hao, Z. J., Wei, M. R., Gong, S. J., Zhao, D. Q. & Tao, J. Transcriptome and digital gene expression analysis of herbaceous peony (Paeonia lactiflora Pall.) to screen thermo-tolerant related differently expressed genes. Genes Genom.38, 1201–1215 (2016).
Wu, Y. Q., Zhao, D. Q., Han, C. X. & Tao, J. Biochemical and molecular responses of herbaceous peony to high temperature stress. Can. J. Plant Sci.96, 474–484 (2016).
Su, J. H. Cloning and functional study of Hsp70 Gene in herbaceous peony (Paeonia lactiflora Pall.). (Yangzhou University, 2018).
Zhao, D. Q. et al. Overexpression of herbaceous peony HSP70 confers high temperature tolerance. BMC Genom.20, 70 (2019).
Hao, Z. J., Liu, D., Gong, S. J., Zhao, D. Q. & Tao, J. High throughput sequencing of herbaceous peony small RNAs to screen thermo-tolerance related microRNAs. Genes Genom.39, 397–408 (2017).
Hao, Q. et al. Overexpression of PSK1, a SKP1-like gene homologue, from Paeonia suffruticosa, confers salinity tolerance in Arabidopsis. Plant Cell Rep.36, 151–162 (2017).
Jin, Q. J. et al. Identification and characterization of microRNAs from tree peony (Paeonia ostii) and their response to copper stress. PLoS ONE10, e0117584 (2015).
Wang, Y. J., Dong, C. L., Xue, Z. Y., Jin, Q. J. & Xu, Y. C. De novo transcriptome sequencing and discovery of genes related to copper tolerance in Paeonia ostii. Gene576, 126–135 (2016).
Zhao, D. Q. et al. Herbaceous peony tryptophan decarboxylase confers drought and salt stresses tolerance. Env. Exp. Bot.162, 345–356 (2019).
Li, X. T., Liu, P., Yang, P., Fan, C. & Sun, X. Characterization of the glycerol-3-phosphate acyltransferase gene and its real-time expression under cold stress in Paeonia lactiflora Pall. PLoS ONE13, e0202168 (2018).
Chen, Y. Cloning and expression analysis of the dehydrin gene PlDHNs in peony (Paeonia lactiflora) (Shandong University, 2018).
Chen, Y., Ma, Y., Guo, J. & Guo, X. F. Cloning and expression analysis of the dehydrin gene PlDHN1 in peony (Paeonia lactiflora). J. Hortic. Sci. Biotechnol.93, 557–565 (2018).
Wang, X. et al. PlWRKY13: a transcription factor involved in abiotic and biotic stress responses in Paeonia lactiflora. Int. J. Mol. Sci.20, 5953 (2019).
Liu, H. C. et al. Functional analysis of stress-related transcription factor gene PsDREB from Paeonia suffruticosa. J. Zhejiang Univ. (Agric. Life Sci.)42, 679–686 (2016).
Liu, H. C. et al. Identification and characterization of PsDREB2 promoter involved in tissue-specific expression and abiotic stress response from Paeonia suffruticosa. PeerJ.7, e7052 (2019).
Zhou, L. Influence of ethylene on flower opening and senescence process and expression of related genes in tree peony cut flower (Beijing Forestry University, 2007).
Zhou, L. et al. Molecular characterization and expression of ethylene biosynthetic genes during cut flower development in tree peony (Paeonia suffruticosa) in response to ethylene and functional analysis of PsACS1 in Arabidopsis thaliana. J. Plant Growth Regul.32, 362–375 (2013).
Wang, Y. J. Effect of glucose on ethylene biosynthesis and signal transduction in tree peony cut flower during postharvest senescence (Beijing Forestry University, 2013).
Gao, J. et al. Effects of ethylene and 1-MCP treatments on the expressions of CTR genes of tree peony. Acta Bot. Boreal. Occident. Sin.31, 19–26 (2011).
Wang, Y. J. et al. Isolation and expression analysis of three EIN3-like genes in tree peony (Paeonia suffruticosa). Plant Cell Tiss. Organ Cult.112, 181–190 (2013).
Wu, F., Zhang, C., Guo, J., Liu, A. Q. & Dong, L. Isolation and expression analysis of ERF transcription factor genes in tree peony cut flowers. Acta Hortic. Sin.43, 109–120 (2016).
Zhao, D. Q. et al. Nano-silver modifies the vase life of cut herbaceous peony (Paeonia lactiflora Pall.) flowers. Protoplasma255, 1001–1013 (2018).
Xue, J. et al. Dry storage improves the vase quality of cut peony by increasing water uptake efficiency through aquaporins regulation. Plant Physiol. Biochem.148, 63–69 (2020).
Xue, J. et al. Assessment of vase quality and transcriptional regulation of sucrose transporter and invertase genes in cut peony (Paeonia lactiflora “Yang Fei Chu Yu”) treated by exogenous sucrose. Postharvest Biol. Technol.143, 92–101 (2018).
Xue, J. et al. Evaluation of dry and wet storage on vase quality of cut peony based on the regulation of starch and sucrose metabolism. Postharvest Biol. Technol.155, 11–19 (2019).
Huang, X. Cloning and analysis of genes associated with the release of dormant floral buds in tree peony (Paeonia suffruticose Andr.) (Beijing Forestry University, 2008).
Zhang, Y. X. et al. MYC cis-elements in PsMPT promoter is involved in chilling response of Paeonia suffruticosa. PLoS ONE11, e0155780 (2016).
Huang, X. et al. Genes associated with the release of dormant buds in tree peonies (Paeonia suffruticosa). Acta Physiol. Plant.30, 797–806 (2008).
Mu, P., Zhang, J. Q. & Zhang, X. L. Cloning and correlation analysis of PsPII gene associated with the release of dormant buds in tree peony (Paeonia suffruticosa). J. Agric. Biol.21, 292–298 (2013).
Gao, X. K., Zhang, Y. X., Liu, C. Y., Dou, B. L. & Gai, S. P. Cloning of somatic embryogenesis receptor-like kinase gene PsSERK2 and cell division rate analysis during dormancy release in tree peony (Paeonia suffruticosa). Acta Hortic. Sin.45, 511–518 (2018).
Han, L. L. Cloning and functional analysis of GA20-oxidase Gene In Paeonia lactiflora (Shandong Agricultural University, 2017).
Zhang, Y. X. et al. The study of PsGA20ox gene participating in endo-dormancy release of flower buds by chilling treatment in tree peony. Acta Agric. Boreal.-Occident. Sin.29, 22–26 (2014).
Zhang, Y. X. et al. Isolation and characterization of a SOC1-like gene from tree peony (Paeonia suffruticosa). Plant Mol. Biol. Rep.33, 855–866 (2015).
Wu, H., Liu, C. Y., Fu, X. M., Gai, S. P. & Zhang, Y. X. Screening, cloning and expression patterns analysis of PsGRASs associated with dormancy release in tree peony (Paeonia suffruticosa). Acta Hortic. Sin.46, 365–374 (2019).
Xin, H. et al. Morphological, anatomical and DNA methylation changes of tree peony buds during chilling induced dormancy release. Plant Physiol. Biochem.144, 64–72 (2019).
Zhang, Y. X. et al. Application of 5-azacytidine induces DNA hypomethylation and accelerates dormancy release in buds of tree peony. Plant Physiol. Biochem.147, 91–100 (2020).
Xue, J. Q. et al. On the role of physiological substances, abscisic acid and its biosynthetic genes in seed maturation and dormancy of tree peony (Paeonia ostii ‘Feng Dan’). Sci. Hortic.182, 92–101 (2015).
Zhang, P. et al. Expression analysis of the PobZIP1 transcription factor gene in developmental seeds from tree peony and its relationship with ABA content. Acta Hortic. Sin.41, 1642–1650 (2014).
Chen, Z. J. Cloning seeds, germination key genes PlCYP707As in Paeoina lactiflora Pall. and constructing plant expression vector (Shenyang Agricultural University, 2017).
Ma, Y. L. Cloning and genetic transformation of PlPP2C gene related to dormancy and germination of Paeonia lactiflora Seeds (Shenyang Agricultural University, 2018).
Cui, J. Q. Cloning and functional analysis of the PlGAI1 gene related to the dormancy and germination of the seeds in Paeonia lactiflora Pall (Shenyang Agricultural University, 2016).
Ren, X. X. et al. Proteomic analysis of tree peony (Paeonia ostii “Feng Dan”) seed germination affected by low temperature. J. Plant. Physiol.224–225, 56–67 (2018).
Fei, R. W. Cloning of the PlSAURs genes from Paeonia lactiflora Pall. and transformation of Arabidopsis thaliana (Shenyang Agricultural University, 2018).
Hong, D. Y. et al. Current status of wild tree peony species with special reference to conservation. Biodiv. Sci.25, 781–793 (2017).
Yang, Y., Zeng, X. L., Zhang, S. S., Zhang, J. J. & Yu, X. N. Geographical distribution and resource characteristics of 5 wild peonies in southwest China. J. Sichuan Agric. Univ.35, 69–74 (2017). 78.
Shen, M. M., Tang, Z. J., Teixeira da Silva, J. A. & Yu, X. N. Induction and proliferation of axillary shoots from in vitro culture of Paeonia lactiflora Pall. mature zygotic embryos. N. Z. J. Crop Hortic. Sci.43, 42–52 (2015).
Xu, L., Cheng, F. Y. & Zhong, Y. Study on rapid seedling-raising technology of tree peony embryo culture. Bull. Bot. Res.37, 690–699 (2017).
Shen, M. M., Wang, Q., Yu, X. N. & Teixeira da Silva, J. A. Micropropagation of herbaceous peony (Paeonia lactiflora Pall.). Sci. Hortic.148, 30–38 (2012).
Teixeira da Silva, J. A., Shen, M. M. & Yu, X. N. Tissue culture and micropropagation of tree peony (Paeonia suffruticosa Andr.). J. Crop Sci. Biotechnol.15, 159–168 (2012).
Sone, T. et al. A novel gene delivery system in plants with calcium alginate micro-beads. J. Biosci. Bioeng.94, 87–91 (2002).
Zhao, X. et al. Pollen magnetofection for genetic modification with magnetic nanoparticles as gene carriers. Nat. Plants3, 956 (2017).
Jia, Y. J., Guo, X., & Cheng, F. Y. Preliminary analysis on transgenic technology of pollen-mediated by magnetic nanoparticles in tree peony. Mol. Plant Breed. http://kns.cnki.net/kcms/detail/46.1068.S.20190527.0943.002.html (2020).
Demirer, G. S., Zhang, H., Goh, N. S., González-Grandío, E. & Landry, M. P. Carbon nanotube-mediated DNA delivery without transgene integration in intact plants. Nat. Protoc.14, 2954–2971 (2019).
Lv, S. et al. Draft genome of the famous ornamental plant Paeonia suffruticosa. Ecol. Evol. https://doi.org/10.1002/ece3.5965 (2020).
Acknowledgements
We thank Mr. Fu Wang and his wife Mrs. Yuling Fan for their help in Paeonia resource identification and cross-breeding. We are also grateful to Prof. Guangli Liu, Prof. Guangtang Pan and Prof. Huixing Song from the School of Landscape Architecture of Sichuan Agricultural University and Prof. Xiuli Zeng and Miss Shanshan Zhang from the Institute of Vegetables, Tibet Academy of Agricultural and Animal Husbandry Sciences, for their help in the field investigations of wild Paeonia species. This study was financially supported by the National Key R&D Program of China (2018YFD1000406) and the National Natural Science Foundation of China (31772350).
Author information
Authors and Affiliations
Contributions
L.W. and X.Y. planned the outline of the review. Y.Y. collected the available literature and completed the first draft of the paper. L.W. and S.L. carefully compiled the first draft of the paper. M.S. compiled the tables and corrected the references. Q.C. compiled one figure. S.L., Q.C., J.A.T., and A.W. critically reviewed and improved the paper. All authors approved the final paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Y., Sun, M., Li, S. et al. Germplasm resources and genetic breeding of Paeonia: a systematic review. Hortic Res 7, 107 (2020). https://doi.org/10.1038/s41438-020-0332-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-020-0332-2
This article is cited by
-
Agromorphological variability of hoary basil germplasm (Ocimum americanum L.) from Burkina Faso
Genetic Resources and Crop Evolution (2024)
-
Phenotypic diversity analysis of Polygala tenuifolia Willd germplasm resources in China
Genetic Resources and Crop Evolution (2024)
-
The plastome reveals new insights into the evolutionary and domestication history of peonies in East Asia
BMC Plant Biology (2023)
-
Reference genes selection of Paeonia ostii ‘Fengdan’ under osmotic stresses and hormone treatments by RT-qPCR
Molecular Biology Reports (2023)
-
Transcriptome analysis of floral bud development and function analysis of a novel CO gene in Paeonia × lemoinei ‘High Noon’
Scientific Reports (2022)