Abstract
Isopentenyltransferase (IPT) genes, including those encoding ATP/ADP-IPTs and tRNA-IPTs, control the rate-limiting steps of the biosynthesis of N6-(Δ2-isopentenyl)adenine (iP)-type and trans-zeatin (tZ)-type cytokinins and cis-zeatin (cZ)-type cytokinins, respectively. However, the evolution and roles of these IPTs in angiosperms are not well understood. Here, we report comprehensive analyses of the origins, evolution, expression patterns, and possible roles of ATP/ADP-IPTs and tRNA-IPTs in angiosperms. We found that Class I and II tRNA-IPTs likely coexisted in the last common ancestor of eukaryotes, while ATP/ADP-IPTs likely originated from a Class II tRNA-IPT before the divergence of angiosperms. tRNA-IPTs are conservatively retained as 2–3 copies, but ATP/ADP-IPTs exhibit considerable expansion and diversification. Additionally, tRNA-IPTs are constitutively expressed throughout the plant, whereas the expression of ATP/ADP-IPTs is tissue-specific and rapidly downregulated by abiotic stresses. Furthermore, previous studies and our present study indicate that ATP/ADP-IPTs and their products, iPs/tZs, may regulate responses to environmental stresses and organ development in angiosperms. We therefore hypothesize that tRNA-IPTs and the associated cZs play a housekeeping role, whereas ATP/ADP-IPTs and the associated iP/tZ-type cytokinins play regulatory roles in organ development and stress responses in angiosperms, which echoes the conclusions and hypothesis presented in the accompanying study by Wang, X. et al Evolution and roles of cytokinin genes in angiosperms 2: Do ancient CKXs play housekeeping roles while non-ancient CKXs play regulatory roles? Hortic Res https://doi.org/10.1038/s41438-020-0246-z.
Similar content being viewed by others
Introduction
Cytokinins (CKs) are a class of plant hormones that play essential roles in many aspects of plant development, including the delay of leaf senescence1, root proliferation2,3, apical dominance4, shoot meristem function5,6,7, regulation of reproductive meristem activity8, fruit development9, and nutritional signaling10,11. In addition, CKs play important and complex roles in environmental stress responses12,13. There are two major types of naturally occurring CKs, the cis-zeatin (cZ) type and the iP [N6-(iP]/trans-zeatin (tZ) type14. In many plants, iPs and tZs are the most prevalent CKs in most tissues and stages of the lifespan, while cZ-type CKs are present only in minor quantities15. Recent studies have demonstrated that cZs are the predominant CKs in some plants, such as rice and maize, or in certain developmental stages associated with limited growth16. The presence of iP-type and tZ-type CKs can vary greatly between tissues, developmental stages, and environmental conditions17,18,19.
Isopentenyltransferases (IPTs) catalyze the first and rate-limiting step of CK biosynthesis. IPTs can be classified into the adenylate (ATP/ADP-IPTs and AMP-IPTs) and tRNA types. The IPTs that use ATP, ADP, or AMP as their preferred substrates produce iP-type and tZ-type CKs, while the tRNA-type IPTs, which transfer isopentenyl groups to the N6 atom of adenines in tRNAs, produce cZ-type CKs14,19. In Arabidopsis, nine IPT genes have been identified (AtIPT1–9). Among them, AtIPT1 and AtIPT3–AtIPT8 encode ATP/ADP-IPTs, and the other two AtIPT genes, AtIPT2 and AtIPT9, encode tRNA-IPTs. Mutations in two tRNA-type IPT genes, AtIPT2 and AtIPT9, particularly in combination, result in plant chlorosis and a significant reduction in cZ-type CKs but do not affect the concentrations of iPs and tZs14,20. By comparison, plants with mutant ATP/ADP-IPTs possess scarce amounts of iP-type and tZ-type CKs but contain slightly elevated levels of cZs14,21,22. These mutant plants exhibit increased lateral root primordia and density, fewer leaves, reduced inflorescence, and decreased number of vascular bundles and are unable to form cambium14,21,22.
Genetically mutant plants with altered endogenous CK content display changes in plant tolerance to environmental stresses. A quadruple ATP/ADP-IPT Arabidopsis mutant atipt1;3;5;7 with deficient levels of iPs and tZs but slightly increased levels of cZs displayed significantly enhanced salt and drought tolerance compared to that of wild-type plants23. Plants overexpressing CK degradation genes (CK oxidase/dehydrogenase genes, CKXs), which contain substantially reduced levels of iPs/tZs and moderately reduced levels of cZs, also show elevated tolerance to drought, salt, or heat stress23,24. These studies show that iP and tZ CKs are negative regulators of plant adaptation to environmental stresses. However, it has also been reported that if a stress-associated or senescence-associated promoter is used to drive the expression of an AMP-IPT or ATP/ADP-IPT gene, improved plant tolerance to drought, heat, and other stresses is observed13.
As previously reported19,25, tRNA-IPTs have been found in all major clades of bacteria and eukaryotes but not in archaea. In contrast, adenylate IPTs show a fragmented distribution. The functionally confirmed ATP/ADP-IPTs are found exclusively in flowering plants14,26. Based on the tree topology and taxonomic composition of the IPT gene family, Lindner et al.25 have suggested that eukaryotic IPTs most likely can be traced back to an ancestral α-proteobacterium that was involved in the initial endosymbiotic event that led to extant mitochondria. The ancestral eukaryotic IPT gene was duplicated, which resulted in two classes of tRNA-IPTs (Class I and II). The Class II tRNA-IPTs subsequently lost the capability to bind tRNAs and diversified into the extant ADP/ATP-IPTs found in flowering plants. Recently, Nishii et al.27 performed a broader sampling of IPT genes and hypothesized that Class I tRNA-IPTs represent the direct successors of bacterial miaA genes, while Class II tRNA-IPTs are derived from eukaryotic genes.
In this study, we comprehensively analyzed the differences between ATP/ADP-IPTs and tRNA-IPTs, including their taxonomic distribution, origin, evolutionary history, duplication mechanism, gene/protein structure, and expression patterns in tissues/organs and under various environmental stresses. Our results strongly suggest that tRNA-IPTs in angiosperms can be traced back to the initial endosymbiotic event, while ATP/ADP-IPTs are derived from tRNA-IPTs. In flowering plants, tRNA-IPTs are conservatively retained and constitutively expressed in all tissues and under various environmental conditions. On the other hand, ATP/ADP-IPTs have extensively expanded and functionally diverged, and their expression is tissue-/organ-specific and environmental stress-responsive. Our findings plus previously published results demonstrate that in angiosperms, tRNA-IPT genes and the associated cZ-type CKs most likely play housekeeping roles, while ATP/ADP-IPT genes and the associated iP/tZ-type CKs may be involved in regulating organ development and responses to environmental stresses.
Results
Genome-wide identification of IPT genes among the three domains of life
IPT genes have been identified in various groups of bacteria and eukaryotes but not in archaea19. To verify the absence of IPT homologs in archaea, we used TBLASTN to identify the IPPT domain in archaeal nucleotide databases. Putative IPPT domains were found in two archaeal species, miscellaneous Crenarchaeota group archaeon SMTZ-80 and candidate division MSBL1 archaeon SCGC-AAA382N08, indicating that IPT homologs are also present in archaea. The former archaeal species containing an IPT homolog belongs to the TACK superphylum, which has recently been proposed to be the origin of eukaryotes, and the latter is a member of the unclassified euryarchaeota.
We next performed a broad and balanced sampling of a total of 59 representative species from major lineages of all three domains of life (eukaryotes, bacteria, and archaea, Table S1). IPT homologs in these species were identified. We found that the number of IPT homologs varies considerably among the species in Plantae, ranging from one copy in C. reinhardtii (green algae) and C. merolae (red algae) to nine copies in Arabidopsis thaliana (eudicot) and 11 copies in Z. maize (monocot). By comparison, the number of IPT homologs in the other organisms, i.e., nonplant eukaryotes, bacteria, and archaea, does not vary greatly and ranges from 0 to 3 copies (Table S1). More than 90% (20/22) of the bacteria and nonplant eukaryotes sampled contain putative IPT gene(s) (mostly a single copy), while none of the IPT homologs could be found in the sampled archaea except the two identified by TBLASTN. Taken together, our results show that IPT homologs are distributed sporadically in archaea but widely throughout bacteria and eukaryotes, and show significant expansions in land plants.
Differences in the origin and distribution of tRNA-IPTs and ATP/ADP-IPTs
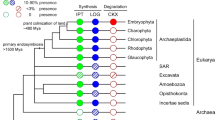
To explore the evolutionary history of the IPT genes, an unrooted ML phylogeny comprised of IPT homologs from 59 representative species was constructed using PhyML (Fig. 1). Based on IPTs with known functions, the entire IPT gene family could be divided into three subfamilies, designated tRNA-IPT, ADT/ATP-IPT, and AMP-IPT19. According to our phylogeny, the AMP-IPT subfamily includes only members of D. discoideum and several bacteria, including Agrobacteria, and the ATP/ADP-IPT subfamily is confined to seed plants (Table 1). In comparison, the tRNA-IPT subfamily has a much wider taxonomic distribution.
Phylogenetic tree of IPT genes from major lineages of bacteria, archaea, and eukaryotes constructed in PhyML. Support values (Shimodaira–Hasegawalike test) >0.5 are indicated at the nodes. Color coding: green lines, ATP/ADP-IPT; orange lines, Class I tRNA-IPT; pink lines, Class II tRNA-IPT; gray lines, AMP-IPT; orange and pink backgrounds, putative Class I and II tRNA-IPT homologs, respectively
The two classes of tRNA-IPT genes (referred to as Class I and II hereafter) categorized by Lindner et al.25 are located in two clades and include members of various species (Fig. 1). The Class I tRNA-IPT homologs can be found in all supergroups of eukaryotes except Excavata (Table 1). In particular, every species containing plastids has at least one Class I tRNA-IPT (Table S1), revealing the universal distribution of this type of IPT gene in photosynthetic organisms. Within the phylogeny, all eukaryotic Class I tRNA-IPTs form a clade that branches with the tRNA-IPT homologs from bacterial species belonging to proteobacteria and four other phyla (Fig. 1 and Table S1). It is worth noting that these five bacterial phyla have relatively close relationships in the recent tree of life28. Therefore, the bacterial homologs of Class I tRNA-IPTs may be traced back to the ancestor of these proteobacteria-related bacterial species.
In contrast, the Class II tRNA-IPT subfamily includes homologs from seed plants and two prasinophyte algae in Plantae and all nonplant eukaryotic supergroups (Table 1). In particular, the Class II tRNA-IPTs and ATP/ADP-IPTs in seed plants form a clade that is a sister to that of the putative Class II tRNA-IPTs in prasinophyte algae and nonplant eukaryotes. This phylogenetic relationship indicates that these two types of IPT genes in seed plants have a common ancestor, and the ATP/ADP-IPTs are likely derived from the Class II tRNA-IPTs. Additionally, the eukaryotic Class II tRNA-IPT clade branches with a cluster of IPTs, including AMP-IPTs and bacterial and archaeal tRNA-IPTs. Hence, we categorized these bacterial and archaeal tRNA-IPTs as putative Class II members. The bacterial species containing these putative Class II tRNA-IPTs include cyanobacteria, actinobacteria, and other species except proteobacteria and those species that are phylogenetically closely related to proteobacteria.
In other words, all the bacteria sampled contain only one type of tRNA-IPT homolog, and those with the same type of tRNA-IPT homolog are phylogenetically more closely related than those without a homolog. In the sampled bacteria, there are four bacterial species with two IPT copies, all of which have one AMP-IPT homolog and one tRNA-IPT homolog of either of the two classes. In contrast, all the nonplant eukaryotes with more than one IPT copy contain both types of tRNA-IPTs, with only one exception. Moreover, these nonplant eukaryotic species belong to different supergroups that diverged at the base of the eukaryote phylogenetic tree, indicating that the coexistence of the two classes of tRNA-IPTs probably originated before the divergence of these supergroups.
Differences in evolutionary patterns between tRNA-IPTs and ATP/ADP-IPTs in land plants
Our identification shows that the IPT genes have undergone considerable expansions, specifically in land plants. To further investigate the evolution of IPT genes in land plants, we identified 171 IPTs from 19 sequenced species belonging to every major lineage of land plants (Table S2) and constructed an unrooted ML tree using PhyML (Fig. 2a). According to the tree topology and that of the known IPTs, the land plant IPT gene family was classified into four groups: tRNA-IPT I, tRNA-IPT II, ATP/ADP-IPT I, and ATP/ADP-IPT II. The members of the tRNA-IPT I group are distributed throughout all the main lineages of land plants, while those of the tRNA-IPT II group are confined to seed plants, which is consistent with the distribution of the two tRNA-IPT classes in the phylogeny shown in Fig. 1. The total number of tRNA-IPT genes in each flowering plant species is either two or three (Table 1), revealing that a conserved number of this type of IPT gene is retained in angiosperm genomes. However, six and five tRNA-IPT genes were found in Physcomitrella patens (moss) and Picea abies (gymnosperm), respectively, indicating that lineage-specific expansions have occurred in these two species/lineages.
a The unrooted ML phylogenetic tree constructed by using PhyML based on the full-length amino acid sequences of the IPT proteins from 19 representative land plants. Bootstrap values >70 are shown. Color coding for the different groups is indicated at the bottom. b Inferred expansion history of the IPT gene family in land plants. Inferred duplications and proposed losses are shown with solid circles and black diamonds, respectively. Duplication events that occurred before the divergence of different lineages are marked by black filled circles, whereas lineage-specific duplication events are marked by circles filled with the lineage-specific colors. Colored lines represent the respective subgroups in Table S2. Dotted lines indicate that the generation of ATP/ADP-IPTs in gymnosperms is undetermined
The number of ATP/ADP-IPT genes varies from two copies in Amborella trichopoda to 21 copies in Medicago truncatula, which accounts for 40% of IPTs in the former and 91.3% of IPTs in the latter (Table 2). The high and increased percentages of ATP/ADP-IPTs indicate that the increased number of IPT genes in flowering plants is mainly due to the expansion of ATP/ADP-IPTs. According to the phylogeny, the two groups ATP/ADP-IPT I and II contain each of the two ATP/ADP-IPT genes in the basal angiosperm A. trichopoda (Fig. 2a), respectively, suggesting that two ancestral ATP/ADP-IPTs were likely present in the last common ancestor of angiosperms. In addition, the ATP/ADP-IPT I group can be further divided into four subgroups, Ia, Ib, Ic, and Id. The ATP/ADP-IPT Ia–Ic subgroups contain only eudicot members, while the ATP/ADP-IPT Id subgroup contains exclusively monocots, reflecting the lineage-specific duplications of angiosperm ATP/ADP-IPTs (Fig. 2b and Table S3). Notably, PaIPT3, the P. abies gene that is located in the ATP/ADP-IPT clade of the phylogeny shown in Fig. 1, branches with the tRNA-IPT II group in the land plant trees, which is supported by the low bootstrap value (Fig. 2a). Therefore, whether gymnosperms contain ATP/ADP-IPT homologs could not be determined from the phylogenetic analyses. Nevertheless, the existence and expansion of ATP/ADP-IPTs in flowering plants could be determined.
We further explored the duplication mechanisms responsible for the expansion of ATP/ADP-IPT genes in the 15 angiosperms using the MCScanX package29. Each ATP/ADP-IPT gene was assigned to one of the five modes, including WGD/segmental duplication, tandem duplication, proximal duplication, dispersed duplication, and singleton (Table 2). Approximately 47.5% of the ATP/ADP-IPT genes from every dicot or monocot were shown to be involved in WGD gene duplications, demonstrating that WGD/segmental duplication is a primary mechanism. In contrast, tandem and proximal duplications accounted for 13.2% and 5.7% of the ATP/ADP-IPT duplications, respectively. In particular, 21 of the 23 tandem/proximal duplications, which belong to the ATP/ADP-IPT Ia subgroup, are derived from only two species, M. truncatula and S. lycopersicum. These results indicate that WGD/segmental duplication has extensively contributed to the expansion of ATP/ADP-IPT genes, while tandem duplication has given rise to lineage-specific expansions of the ATP/ADP-IPT genes in flowering plants.
Differences in the gene and protein structures of tRNA-IPTs and ATP/ADP-IPTs
We next compared the structural differences between tRNA-IPTs and ATP/ADP-IPTs, beginning with the investigation of the exon–intron organization of IPT genes in 19 land plants (Fig. 2a). It is interesting to note that 77.7% of the 121 ATP/ADP-IPT genes contain no introns, and 85.2% of the remaining ATP/ADP-IPTs contain only a single intron. By comparison, 90% of the 50 tRNA-IPT genes have 6–8 introns. These results suggest that ATP/ADP-IPT genes are probably derived from an intron-free ancestral tRNA-IPT gene.
We further explored the motif composition of IPT proteins from four representative flowering plants (Figs. 3 and S1). Fifteen conserved motifs were identified. All three types of IPTs, Class I and Class II tRNA-IPTs and ATP/ADP-IPTs, show conserved motif structures among the members of every type. Motifs 11, 13, and 15 and motif 14 could be specifically detected in the Class I and Class II tRNA-IPTs, respectively, while no particular motif could be identified in the ATP/ADP-IPTs. The Class II tRNA-IPT proteins demonstrate a similar motif composition to the ATP/ADP-IPTs rather than the Class I tRNA-IPTs. In fact, the consensus motif structure of the ATP/ADP-IPTs and the Class II tRNA-IPTs is the same, except for motifs 12 and 14, providing direct evidence to support our above-described hypothesis that ATP/ADP-IPT genes are derived from the Class II tRNA-IPTs.
Differences in the tissue/organ expression patterns of tRNA-IPTs and ATP/ADP-IPTs
To investigate the differences in the expression patterns of tRNA-IPTs and ATP/ADP-IPTs, we analyzed RNA-seq data from the basal angiosperm A. trichopoda and the eudicot woodland strawberry. In A. trichopoda, AmIPT2 and AmIPT4 belong to the ATP/ADP-IPT subfamily. The mRNA level of AmIPT4 was low in vegetative organs and meristems and undetectable in female buds, while AmIPT2 was highly expressed in vegetative organs and weakly expressed in meristems and female buds (Fig. S2). In Fragaria vesca, five genes (FveIPT1, FveIPT3–6) belong to the ATP/ADP-IPT subfamily. FveIPT1, FveIPT3, and FveIPT4 were highly expressed in the various stages in carpel and anther. FveIPT5 was highly expressed in ghost and in some stages of cortex and pith, and the mRNA level of FveIPT6 was high in style and some stages of pith (Fig. 4). These different expression patterns indicate that functional diversification of the ATP/ADP-IPT genes after their expansion occurred in eudicots. By comparison, the three genes (AmIPT1, AmIPT3, and AmIPT5) in A. trichopoda and two genes (FveIPT2 and FveIPT7) in F. vesca, which belong to the tRNA-IPT gene family, were relatively highly and stably expressed in all tissues, suggesting that tRNA-IPTs have a constitutive expression pattern similar to that of housekeeping genes.
The x-axis indicates the different stages of F. vesca flowers and early-stage fruits, and the y-axis represents the mRNA length in kilobases per million mapped reads. Data were retrieved from the SGR database (http://bioinformatics.towson.edu/strawberry/). Expression levels were calculated according to the log2 scale. For a detailed description of the stages, please see http://bioinformatics.towson.edu/strawberry/newpage/Tissue_Description.aspx
We further used qRT-PCR to compare the differences in the expression patterns of the F. vesca ATP/ADP-IPT and tRNA-IPT genes in six stages of fruits (little green, big green, white, preturning, pink, and red) and three vegetative organs (leaves, immature roots, and mature roots, Fig. 5). Similar to the results of the above transcriptomic analyses, ATP/ADP-IPT genes showed quite diversified expression patterns. Although the transcript levels of FveIPT1, FveIPT5, and FveIPT6 gradually increased during fruit ripening, FveIPT1 exhibited higher expression levels in leaves, and FveIPT5 and FveIPT6 were highly expressed in immature roots. The expression of FveIPT3/4 was undetectable in all these samples (<5 × 10−5). Meanwhile, the tRNA-IPT genes were constitutively expressed in different developmental stages of fruits, leaves, and roots. We further compared the variation amplitude of FveIPT gene expression in different organs. The variation amplitude of the expression of the tRNA-IPT genes (~2-fold) was substantially smaller than that of the ATP/ADP-IPT genes (30- to 130-fold). Therefore, the qPCR results showed that the ATP/ADP-IPT genes have quite diversified expression patterns, while the expression of the tRNA-IPT genes is relatively consistently high throughout the plant.
Transcript expression was normalized to the expression of the GAPDH gene. The mean ± s.d. of three biological replicates is presented. The different lowercase letters above the bars indicate significant differences (α = 0.05, LSD) among the different tissues. Three biological replicates and three technical replicates were performed for each data point
Differences in the environmental stress responses of tRNA-IPTs and ATP/ADP-IPTs
CKs produced by IPT enzymes also play important roles in the regulation of plant responses to environmental stresses30. We further investigated the expression profiles of the ATP/ADP-IPT and tRNA-IPT genes under salt, dehydration, hot, and cold stress conditions (Fig. 6). Among the ATP/ADP-IPT genes, the expression of FveIPT1 was significantly decreased (>4-fold and P < 0.001) after salt stress, and the expression of FveIPT5 was significantly decreased after salt (>7-fold) and drought stress at 8 h (>3-fold). The transcription of FveIPT6 was significantly decreased by more than 2–3-fold after salt, dehydration, and cold stress and was increased by 2-fold under heat stress. FveIPT3/4 was undetectable after all four types of stress (<4 × 10−5). For the tRNA-IPT genes, the transcription of FveIPT7 was reduced by ~2-fold after salt and dehydration stress (Fig. 6). The changes in ATP/ADP-IPT gene expression levels were drastic in response to environmental stresses, but much smaller changes were observed in the expression levels of tRNA-IPT genes under the same stress conditions. These results show that ATP/ADP-IPT genes may be involved in plant responses to environmental stresses, whereas tRNA-IPTs are minimally involved in stress responses.
The expression levels relative to that of GAPDH were measured by qRT-PCR. Asterisks indicate that the corresponding gene was significantly upregulated or downregulated under the given condition (*p ≤ 0.05). Three biological replicates and three technical replicates were performed for each data point
Discussion
IPT enzymes catalyze the first step of CK biosynthesis26. ATP/ADP-IPTs and tRNA-IPTs use different substrates and produce distinct biologically active CKs in flowering plants31. In the present study, we performed comprehensive analyses of the differences between these two types of IPT genes in terms of their taxonomic distribution, copy number variation, evolutionary history, duplication mechanism, tissue/organ expression, and environmental stress responses (Table 3). Based on the results here and those previously published by others, we hypothesize that ATP/ADP-IPTs are more important in regulating organ development and environmental stress responses, while tRNA-IPTs mainly play a housekeeping role in angiosperms.
Different origins of ATP/ADP-IPTs and tRNA-IPTs
According to phylogenetic analyses, Lindner et al.25 hypothesized that Class I tRNA-IPTs have a mitochondrial origin and that Class II tRNA-IPTs are derived from a duplication of a Class I tRNA-IPT gene in the plant lineages. Nishii et al.27 later proposed that Class II tRNA-IPTs originated from eukaryotic genes. However, based on a broad and balanced sampling of species from major lineages of all three domains of life (eukaryotes, bacteria, and archaea), our analyses indicate that the coexistence of Class I and II tRNA-IPTs can be traced back to the last common ancestor of eukaryotes. Nonplant eukaryotes, except D. discoideum, were found to contain both Class I and Class II tRNA-IPT homologs for the first time in this study. The reason that Nishii et al.27 did not observe this is likely because of their limited sampling of nonplant eukaryotic species. These nonplant species belong to different supergroups that are considered to have diverged from the base of the eukaryotic tree32; thus, our results show that the two classes of tRNA-IPTs coexisted in the last eukaryotic common ancestor (LECA).
In this study, we identified two tRNA-IPT homologs in archaeal species for the first time. tRNA-IPTs have been previously reported to be widely distributed in other kingdoms but not in archaea19,26,31. Although it is unclear whether the archaeal tRNA-IPTs are functional, their identification reveals that tRNA-IPT sequences are distributed throughout all three kingdoms and suggests the possibility of an archaeal origin for tRNA-IPTs. Our phylogeny indicates that bacterial or archaeal species contain only one type of tRNA-IPT homolog (Fig. 1 and Table S1). The Class I tRNA-IPTs are likely to have a closer relationship with the homologs in proteobacteria and proteobacteria-related bacteria, whereas the Class II tRNA-IPTs are more closely related to other bacterial groups and the two archaeal species. Eukaryotes are believed to have resulted from the initial endosymbiotic event involving α-proteobacterial and archaeal cells28. Therefore, we propose that Class I tRNA-IPTs may be derived from the α-proteobacteria that were involved in the initial endosymbiotic event. The Class II tRNA-IPT gene in the LECA appears to have two alternative origins; it is derived from either the archaeal ancestor of eukaryotes or a nonproteobacteria ancestral homolog that was introduced into the LECA via horizontal gene transfer. Because tRNA-IPTs are present in the latest common ancestor of eukaryotes, these tRNA-IPTs are ‘ancient’ CK biosynthesis genes in angiosperms.
Our phylogenetic and motif analyses indicate that ATP/ADP-IPT genes are derived from Class II tRNA-IPTs. Most ATP/ADP-IPT genes contain no or only one intron, whereas Class II tRNA-IPTs contain many introns. This difference in gene structure suggests that either the original ATP/ADP-IPT gene was generated via a retroposition duplication of a Class II tRNA-IPT gene or that intron loss occurred soon after its emergence. Intronless daughter gene(s) and sequences of short direct repeats are the two main molecular features of retroposition33,34. We searched the ~100 kb regions upstream to downstream of all the ATP/ADP-IPT genes in Arabidopsis but did not find any featured short direct repeat sequences. Consequently, although Class II tRNA-IPTs account for the origin of ATP/ADP-IPT genes, the duplication mechanism is still unclear. Nevertheless, because the evidence from previous studies and the current study clearly demonstrates that the ATP/ADP-IPT genes emerged in flowering plants, we call them ‘non-ancient’ CK biosynthetic genes relative to the tRNA-IPTs, which we call ‘ancient’ CK biosynthetic genes.
In addition, ATP/ADP-IPTs were previously proposed to only be present in flowering plants19. In our phylogeny of IPTs from all three domains of life, four of the five gymnosperm IPTs branch with tRNA-IPTs, while the rest (PaIPT3) phylogenetically clusters with angiosperm ATP/ADP-IPTs (Fig. 1). Although the support for this clustering is not great, this close relationship suggests that PaIPT3 is probably an ATP/ADP-IPT gene. ATP/ADP-IPTs are responsible for the biosynthesis of tZ-type CKs. It has been observed that tZs, rather than cZs, are predominant in vegetative shoots/leaves in most gymnosperms and angiosperms35. By comparison, in seedless plants, whose IPTs all phylogenetically branch with tRNA-IPTs, cZs are the most abundant CKs. The general CK composition of gymnosperms, which is similar to that of angiosperms but distinct from that of seedless plants, suggests that gymnosperms have a similar IPT gene composition to that of angiosperms; that is, gymnosperms may already contain ATP/ADP-IPT genes. Future investigations on the biochemical characteristics, including preferred substrates and products, of gymnosperm IPT proteins are needed to functionally confirm that ATP/ADP-IPT genes originate from gymnosperms.
tRNA-IPTs and cZ-type CKs likely play housekeeping roles, while ATP/ADP-IPTs and iP/tZ-type CKs may be involved in organ development and stress responses
Based on the following lines of evidence, we hypothesize that in angiosperms, tRNA-IPTs and associated cZ-type CKs mainly play housekeeping roles to maintain basic cellular functions, whereas ATP/ADP-IPTs and their products, the iP-type and tZ-type CKs, are more likely to be involved in the regulation of organ development and stress responses.
First, expression profiling indicates the functional differences between ATP/ADP-IPTs and tRNA-IPTs. Our genomic transcriptome and qPCR analyses of F. vesca, for instance, demonstrated that ATP/ADP-IPT genes display drastic changes at the expression level in various tissues/organs and developmental stages (Figs. 4 and 5). In contrast, tRNA-IPTs are constitutively and relatively stably expressed in all tissues and developmental stages throughout the plant. Similar expression patterns of the ATP/ADP-IPT and tRNA-IPT genes have also been observed in the tissues/organs of Arabidopsis36, Z. mays37 (summarized in Figs. S3 and S4, respectively) and other species. Therefore, the expression patterns of the IPT genes suggest that ATP/ADP-IPTs play regulatory roles in organ development in angiosperms, while tRNA-IPTs most likely act as housekeeping genes.
Second, the evolutionary histories of ATP/ADP-IPTs and tRNA-IPTs in angiosperms support their different roles. Our results demonstrate that tRNA-IPTs are conservatively retained as two or three copies in flowering plants, while ATP/ADP-IPTs have undergone considerable expansions and are highly variable in gene number (from 2 to 21) among species. Moreover, we found that the ATP/ADP-IPT genes (AmIPT2 and 4) were rarely expressed in the flower buds of A. trichopoda (Fig. S2), an extant basal flowering plant with primitive flower tissues38. In contrast, most ATP/ADP-IPT genes are highly expressed in different floral tissues or developmental stages in core angiosperms, such as F. vesca, Arabidopsis, and maize. This discrepancy suggests that the expansion and functional diversification of ATP/ADP-IPT genes in eudicots and monocots have contributed to their diverse regulatory roles in floral organs.
Third, the manipulation of the expression of different ATP/ADP-IPT and tRNA-IPT genes gives rise to different phenotypic variations. The tRNA-atipt9 single and tRNA-atipt2/9 double mutant plants of Arabidopsis exhibited an overall small and chlorotic phenotype with no obvious alterations in organ differentiation or development14,20. In contrast, the ATP/ADP-IPT-mutant plants atipt3 or atipt5 had increased numbers of lateral roots and lateral root primordia21. The triple mutant atipt3;5;7 exhibited a decrease in dedifferentiation ability, which resulted in the reduced formation of callus tissues from root explants after wounding22. The quadruple ATP/ADP-IPT mutant atipt1;3;5;7 produced fewer leaves, altered inflorescence14, and was unable to develop cambium in the root and stem22. Additionally, the overexpression of an ATP/ADP-IPT gene resulted in reduced root growth, enhanced shoot differentiation, increased branching, and increased flower number along with abnormal floral organ development39. Altogether, the phenotypic changes of these mutant or transgenic plants indicate that ATP/ADP-IPTs regulate organ development in angiosperms but that tRNA-IPTs do not.
Fourth, the iP/tZ-type and cZ-type CKs produced by ATP/ADP-IPTs and tRNA-IPTs, respectively, have been shown to be differentially involved in developmental processes in flowering plants. Higher levels of iPs and tZs than cZs have been detected in apical meristems, fertilized embryos and other tissues with high dedifferentiation and differentiation activities16,40. Furthermore, tZs were much more potent than cZs in an assay of callus and shoot differentiation15,16,41,42. It has been shown that the endogenous content of tZs was substantially elevated within 12 h after the wounding of root explants, which reflected a similar but earlier increase in comparison with the expression of core cell cycle regulator genes key for callus formation22. A decrease in tZs and iPs in the ATP/ADP-IPT mutant atipt3;5;7 resulted in reduced callus formation, while an increase in tZs and iPs by the overexpression of Agrobacterium IPT (AMP-IPT) dramatically enhanced the production of callus and shoots43,44. Additionally, tZs and iPs have been shown to be important in regulating the development of shoot apical meristem, vascular cambium, lateral roots, inflorescence stems, and ovules14,21,22. On the other hand, few changes in cZ levels were detected during wound-induced callus formation, indicating that endogenous cZs were unlikely to play a significant role. The effects of the overproduction of cZs or prenylated tRNAs remain unknown, as there has been no report on tRNA-IPT overexpression so far. However, the exogenous application of cZs are relatively ineffective in stimulating shoot and other organ development under in vitro conditions15,16,41,42. cZ-deficient tRNA-ipt mutant plants of Arabidopsis are chlorotic and show much reduced plant sizes14,20, suggesting that cZ-type CKs may be essential for maintaining the basic functions of cells. Taken together, the results suggest that ATP/ADP-IPTs and the associated iP-type and tZ-type CKs function as important regulators of angiosperm organ development, whereas tRNA-IPTs and the associated cZ-type CKs play basic housekeeping roles in angiosperm cells.
Fifth, ATP/ADP-IPTs and tRNA-IPTs catalyze different biochemical reactions that produce two distinct types of CKs, the iPs/tZs- and the cZs, respectively14. ATP/ADP-IPTs catalyze the rate-limiting step of the de novo biosynthesis of CKs, directly generating iP and tZ nucleotides or nucleosides that are subsequently converted into their active free-base forms19. By comparison, the tRNA-IPTs in angiosperms primarily catalyze the isopentenylation of adenine in tRNA to produce cZs from the degradation of prenylated tRNAs14,19. Although the function of prenylated tRNA remains unclear in plants, studies in bacteria, yeast, and mammalian cells have shown that tRNA prenylation is important for translational efficiency and fidelity45 by preventing frameshifts and the nonsense suppression of the codon UAA40,46. Moreover, despite the relatively high expression of tRNA-IPTs throughout the plant, the content of endogenous cZs is generally low16. The application of cZ to 35S:AtCKX7 plants that had reduced cZ levels could correct their shorter root phenotype, whereas the application of cZ to tRNA-atipt2/9 plants could not20. These results indicate that the reduced root growth observed in tRNA-atipt2/9 mutants may not be due to the loss of cZ production but more likely results from the loss of tRNA prenylation, suggesting that another housekeeping function of tRNA-IPTs is to maintain translational accuracy, which is essential for normal cellular activities.
Sixth, similar to their roles in organ development, angiosperm ATP/ADP-IPTs have also diversified in response to environmental stresses. Our results show that the expression levels of all three expressed ATP/ADP-IPT genes in F. vesca are substantially reduced under heat, cold, drought, or salt stress (Fig. 6). Studies in other angiosperms have observed similar findings. In rice, salinity suppressed, whereas cold or drought stress enhanced the expression of most ATP/ADP-IPT genes47. In Arabidopsis, the expression patterns of ATP/ADP-IPT genes under stress conditions were also differentially regulated in shoots and roots47. Moreover, the tZ-type CKs that are produced by ATP/ADP-IPT vary upon environmental stress, depending on the type and intensity of the stress treatment40. In contrast, tRNA-IPT genes display relatively little variation in their expression level when plants are subjected to environmental stresses (Fig. 6), as shown in previous studies37,40. Although the elevation of cZ levels has been found under stress conditions, it is generally recognized that cZs are only a byproduct of increased prenylated tRNA turnover but are not produced from the expression of the tRNA-IPT genes40,48. These observations indicate that tRNA-IPTs are more likely to play a housekeeping role, whereas ATP/ADP-IPTs make an important contribution to the regulation of plant responses to environmental stresses.
Additionally, along with a reduction in the expression levels of ATP/ADP-IPTs under abiotic stress conditions, ATP/ADP-IPT mutant plants that contain deficient iP/tZ-CK levels but little changed cZ levels, such as the quadruple atipt1;3;5;7 Arabidopsis mutant, display increased tolerance to drought and salt stress23. There have been few experiments that have shown the effects of altered cZ content on plant responses to abiotic stress. However, the overexpression of AtCKX isoforms, which led to substantial decreases in iP/tZ-CKs but either largely reduced or unchanged cZ levels, resulted in improved levels of tolerance to drought or salt stress23. Therefore, compared with iPs/tZs, variations in cZ content appear to have little effect on plant tolerance to abiotic stress. It has been suggested that cZs produced by tRNA-IPTs are more involved in the maintenance of a basal level of CK activity under growth-limiting conditions16,40. Accordingly, it is more likely that the elevation of cZ levels in stressed conditions is associated with their housekeeping role in plant growth and development, whereas iP/tZ-type CKs produced by ATP/ADP-IPTs play an important role in the regulation of plant adaptation to environmental stresses.
In summary, based on our results that are presented in this manuscript and previously published data, we hypothesize that tRNA-IPTs (ancient CK biosynthesis genes) and the associated cZs play a housekeeping role, whereas ATP/ADP-IPTs (non-ancient CK biosynthetic genes) and the associated iP/tZ-type CKs play regulatory roles in organ development and stress responses in angiosperms. An accompanying paper in this issue provides additional evidence to support this hypothesis49. It is our hope that this hypothesis will stimulate more interest in elucidating the origins, evolutionary significance, and roles of different types of IPTs (tRNA-IPTs vs. ATP/ADP-IPTs) and the associated different types of CKs (iP/tZ vs. cZ types) in regulating organ development and responses/adaptation in angiosperms.
Materials and methods
Homolog identification
A total of 59 whole-genome-sequenced species sampled from major lineages of all three domains of life (eukaryotes, bacteria, and archaea, Table S1) were first examined for the presence of IPT genes. To explore the evolution of IPT genes in land plants, 19 sequenced species belonging to every major lineage of land plants (Table S2) were next sampled and analyzed. The complete genomic sequences and corresponding annotation information for all these species were downloaded from the JGI and NCBI databases.
The hidden Markov model profile of the IPPT domain (PF01715) was downloaded from the Pfam database50, which was used as a query to search for homologous sequences in the proteome data sets. Sequences with an expected value (E-value) < 10−4 were considered candidates. The chromosomal locations of all candidates were verified to remove redundant sequences. Short proteins with lengths <100 aa were removed51. We further confirmed the presence of the IPPT domain in each candidate using the PFAM (http://pfam.xfam.org/search)50 and SMART databases (http://smart.embl-heidelberg.de/)52 with an E-value cut off <1e−10. For archaea, TBLASTN was first used to find potential hits for the IPPT domain under the threshold of 1.0. For the significant hits, the corresponding amino acid sequences of the high-scoring segment pairs (HSPs) were retrieved to assess the presence or absence of the IPPT domain via the PFAM and SMART databases.
Phylogenetic analyses
The full-length amino acid sequences of all identified IPTs were aligned using ClustalX 2.053. The best-fit model for protein evolution was selected using the Model-Generator program54. A phylogenetic tree of the IPT family was constructed via the maximum-likelihood (ML) method using PhyML (version 3.0) software55 with the JTT evolutionary model. The tree topology was reconstructed using the best of nearest-neighbor interchange (NNI) and subtree pruning and regraphing (SPR) methods55. Branch supports were estimated using an approximate likelihood ratio test with a Shimodaira–Hasegawalike procedure55. The phylogenetic trees were visualized in FIGTREE.
Gene structure, protein motif, and synteny analyses
The relative intron positions in the IPT genes were extracted from the annotated whole-genome data sets of 19 land plants. The gene structure was viewed in the online software Evolview (http://www.evolgenius.info/evolview/).
MEME56 was used to discover the conserved motifs in IPT proteins from the four representative flowering plants, A. trichopoda, Zea mays, A. thaliana, and F. vesca. The parameters were set as follows: the maximum number of motifs, 15; minimum motif width, 6 aa; maximum motif width, 50 aa.
Synteny analyses of the 15 flowering plant genomes were conducted locally using a method similar to that developed for the PGDD (http://chibba.agtec.uga.edu/duplication/)57. BLASTP was used to search for potential homologous gene pairs (E < 1e−10, top five matches) across multiple genomes. These homologous pairs were then used as the input for MCScanX to identify the syntenic regions29,58. MCScanX was further used to identify the IPT genes resulting from WGD/segmental, tandem, proximal, and dispersed duplications (http://chibba.agtec.uga.edu/duplication/index/downloads).
Transcriptome data analyses
Transcriptome data from different tissues and developmental stages of F. vesca59,60 and A. trichopoda were downloaded from the SGR (http://bioinformatics.towson.edu/strawberry/) and NCBI (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA212863) databases, respectively. The reads per kilobase per million (RPKM-mapped reads) values were directly retrieved from the website, and the expression levels for the IPT genes were plotted on a log2 scale.
Plant growth conditions, stress treatments, and material collection
All plant materials were collected from a seventh-generation inbred line of F. vesca, Ruegen (kindly provided by Janet Slovin). For the strawberry fruit collection, strawberry plants were grown in 10 cm × 10 cm pots in a greenhouse with a 12 h photoperiod at 25 °C with 65% relative humidity. Strawberry fruits at the little green (2–4 days after anthesis), big green (8–10 days after anthesis), white, preturning, pink (slight pink flesh and red seeds) and red stages (2–3 days after the pink stage) as well as leaves, immature roots, and mature roots were collected prior to immediate submersion in liquid nitrogen. The tissues from at least three samples were combined to form one biological replicate, and each tissue type utilized three biological replicates.
For the environmental stress experiments, sterile strawberry seedlings were grown in magenta boxes for 2 months in a growth chamber with a 16 h photoperiod at 22 °C and 3000 lx. For the heat shock treatment, the seedlings were transferred to a growth chamber at 38 °C and 3000 lx and were collected at 1, 3, 4 (3 h heat shock and 1 h recovery at 22 °C) and 8 h (3 h heat shock and 5 h recovery at 22 °C). Prior to the salinity stress treatment, the seedlings were transferred to 1/2 MS liquid media and cultivated with gentle agitation (100 rpm). After 12 h, the seedlings were transferred to 1/2 MS liquid media supplemented with 150 mM sodium chloride. For drought treatment, the seedlings were placed on filter paper under dim light at 22 °C with 65% relative humidity. For cold treatment, the seedlings were transferred to a growth chamber set at 4 °C (in the dark). Salinity-stressed, cold-stressed, and dehydration-stressed seedlings were collected at 1, 3, and 8 h after the beginning of the treatment. All collected plant materials were immediately submerged in liquid nitrogen prior to RNA processing.
Quantitative real-time PCR (qRT-PCR) analysis
A modified CTAB method was used for RNA isolation from all the samples mentioned above. The isolated RNAs were treated with DNase I and used for cDNA synthesis using the Primerscript RT Reagent Kit with gDNA Eraser (Takara). qRT-PCR was performed using SYBR Premix Ex Tag (Takara) with the cDNA as the template. The IDT website (http://sg.idtdna.com/site) was used to design the qRT-PCR primers for the FveIPT genes (Table S3). The sequence similarity between FveIPT3 and FveIPT4 is >90%; thus, we used a single primer set to investigate the combined expression of both genes. The results were analyzed using the −ΔΔCT method61 with GAPDH62 as the control locus. Three biological and three technical replicates were analyzed.
References
Kim, H. J. et al. Cytokinin-mediated control of leaf longevity by AHK3 through phosphorylation of ARR2 in Arabidopsis. Proc. Natl Acad. Sci. USA 103, 814–819 (2006).
Werner, T. et al. Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15, 2532–2550 (2003).
Werner, T., Motyka, V., Strnad, M. & Schmulling, T. Regulation of plant growth by cytokinin. Proc. Natl Acad. Sci. USA 98, 10487–10492 (2001).
Shimizu-Sato, S., Tanaka, M. & Mori, H. Auxin–cytokinin interactions in the control of shoot branching. Plant Mol. Biol. 69, 429–435 (2009).
Higuchi, M. et al. In planta functions of the Arabidopsis cytokinin receptor family. Proc. Natl Acad. Sci. USA 101, 8821–8826 (2004).
Kurakawa, T. et al. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445, 652–655 (2007).
Nishimura, C. et al. Histidine kinase homologs that act as cytokinin receptors possess overlapping functions in the regulation of shoot and root growth in Arabidopsis. Plant Cell 16, 1365–1377 (2004).
Bartrina, I., Otto, E., Strnad, M., Werner, T. & Schmulling, T. Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. Plant Cell 23, 69–80 (2011).
Nardozza, S. et al. Exogenous cytokinin application to Actinidia chinensis var. deliciosa ‘Hayward’ fruit promotes fruit expansion through water uptake. Hortic. Res. 4, 17043 (2017).
Samuelson, M. E. & Larsson, C. M. Nitrate regulation of zeatin riboside levels in barley roots - effects of inhibitors of N assimilation and comparison with ammonium. Plant Sci. 93, 77–84 (1993).
Takei, K., Sakakibara, H., Taniguchi, M. & Sugiyama, T. Nitrogen-dependent accumulation of cytokinins in root and the translocation to leaf: Implication of cytokinin species that induces gene expression of maize response regulator. Plant Cell Physiol. 42, 85–93 (2001).
Giron, D., Frago, E., Glevarec, G., Pieterse, C. M. J. & Dicke, M. Cytokinins as key regulators in plant-microbe-insect interactions: connecting plant growth and defence. Funct. Ecol. 27, 599–609 (2013).
Ha, S., Vankova, R., Yamaguchi-Shinozaki, K., Shinozaki, K. & Tran, L. S. P. Cytokinins: metabolism and function in plant adaptation to environmental stresses. Trends Plant Sci. 17, 172–179 (2012).
Miyawaki, K. et al. Roles of Arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. Proc. Natl Acad. Sci. USA 103, 16598–16603 (2006).
Schmitz, R. Y. & Skoog, F. Cytokinins: synthesis and biological activity of geometric and position isomers of zeatin. Plant Physiol. 50, 702–705 (1972).
Gajdosova, S. et al. Distribution, biological activities, metabolism, and the conceivable function of cis-zeatin-type cytokinins in plants. J. Exp. Bot. 62, 2827–2840 (2011).
Hirose, N. et al. Regulation of cytokinin biosynthesis, compartmentalization and translocation. J. Exp. Bot. 59, 75–83 (2008).
Stirk, W. & Van Staden, J. Flow of cytokinins through the environment. Plant Growth Regul. 62, 101–116 (2010).
Frebort, I., Kowalska, M., Hluska, T., Frebortova, J. & Galuszka, P. Evolution of cytokinin biosynthesis and degradation. J. Exp. Bot. 62, 2431–2452 (2011).
Köllmer, I. et al. Overexpression of the cytosolic cytokinin oxidase/dehydrogenase (CKX7) from Arabidopsis causes specific changes in root growth and xylem differentiation. Plant J. 78, 359–371 (2014).
Chang, L., Ramireddy, E. & Schmulling, T. Lateral root formation and growth of Arabidopsis is redundantly regulated by cytokinin metabolism and signalling genes. J. Exp. Bot. 64, 5021–5032 (2013).
Matsumoto-Kitano, M. et al. Cytokinins are central regulators of cambial activity. Proc. Natl Acad. Sci. USA 105, 20027–20031 (2008).
Nishiyama, R. et al. Analysis of cytokinin mutants and regulation of cytokinin metabolic genes reveals important regulatory roles of cytokinins in drought, salt and abscisic acid responses, and abscisic acid biosynthesis. Plant Cell 23, 2169–2183 (2011).
Mackova, H. et al. Enhanced drought and heat stress tolerance of tobacco plants with ectopically enhanced cytokinin oxidase/dehydrogenase gene expression. J. Exp. Bot. 64, 2805–2815 (2013).
Lindner, A. C. et al. Isopentenyltransferase-1 (IPT1) knockout in Physcomitrella together with phylogenetic analyses of IPTs provide insights into evolution of plant cytokinin biosynthesis. J. Exp. Bot. 65, 2533–2543 (2014).
Sakakibara, H. Cytokinins: activity, biosynthesis, and translocation. Annu. Rev. Plant Biol. 57, 431–449 (2006).
Nishii, K., Wright, F., Chen, Y. Y. & Moller, M. Tangled history of a multigene family: the evolution of ISOPENTENYLTRANSFERASE genes. PLoS One 13, e0201198 (2018).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Argueso, C. T., Ferreira, F. J. & Kieber, J. J. Environmental perception avenues: the interaction of cytokinin and environmental response pathways. Plant Cell Environ. 32, 1147–1146 (2009).
Kamada-Nobusada, T. & Sakakibara, H. Molecular basis for cytokinin biosynthesis. Phytochemistry 70, 444–449 (2009).
Adl, S. M. et al. The revised classification of eukaryotes. J. Eukaryot. Microbiol. 59, 429–514 (2012).
Magadum, S., Banerjee, U., Murugan, P., Gangapur, D. & Ravikesavan, R. Gene duplication as a major force in evolution. J. Genet. 92, 155–161 (2013).
Brosius, J. Retroposons-seeds of evolution. Science 251, 753 (1991).
Montalban, I. A., Novak, O., Rolcik, J., Strnad, M. & Moncalean, P. Endogenous cytokinin and auxin profiles during in vitro organogenesis from vegetative buds of Pinus radiata adult trees. Physiol. Plant. 148, 214–231 (2013).
Miyawaki, K., Matsumoto-Kitano, M. & Kakimoto, T. Expression of cytokinin biosynthetic isopentenyltransferase genes in Arabidopsis: tissue specificity and regulation by auxin, cytokinin, and nitrate. Plant J. 37, 128–138 (2004).
Vyroubalova, S. et al. Characterization of new maize genes putatively involved in cytokinin metabolism and their expression during osmotic stress in relation to cytokinin levels. Plant Physiol. 151, 433–447 (2009).
Buzgo, M., Soltis, P. S. & Soltis, D. E. Floral developmental morphology of Amborella trichopoda (Amborellaceae). Int. J. Plant Sci. 165, 925–947 (2004).
Li, X. G. et al. Cytokinin overproduction-caused alteration of flower development is partially mediated by CUC2 and CUC3 in Arabidopsis. Gene 450, 109–120 (2010).
Schafer, M. et al. The role of cis-zeatin-type cytokinins in plant growth regulation and mediating responses to environmental interactions. J. Exp. Bot. 66, 4873–4384 (2015).
Pertry, I. Identification of Rhodococcus fascians cytokinins and their modus operandi to reshape the plant. Proc. Natl Acad. Sci. USA 106, 929–934 (2009).
Kamínek, M., Vaněk, T. & Motyka, V. Cytokinin activities of N 6-benzyladenosine derivatives hydroxylated on the side-chain phenyl ring. J. Plant Growth Regul. 6, 113–120 (1987).
Li, Y., Hagen, G. & Guilfoyle, T. J. Altered morphology in transgenic tobacco plants that overproduce cytokinins in specific tissues and organs. Dev. Biol. 153, 386–395 (1992).
Smigocki, A. C. & Owens, L. D. Cytokinin gene fused with a strong promoter enhances shoot organogenesis and zeatin levels in transformed plant cells. Proc. Natl Acad. Sci. USA 85, 5131–5135 (1988).
Persson, B. C., Esberg, B., Olafsen, O. & Bjork, G. R. Synthesis and function of isopentenyl adenosine derivatives in tRNA. Biochimie 76, 1152–1160 (1994).
Chimnaronk, S. et al. Snapshots of dynamics in synthesizing N 6-isopentenyladenosine at the tRNA anticodon. Biochemistry 48, 5057–5065 (2009).
Ghosh, A. et al. Evolutionary variation and expression profiling of Isopentenyl transferase gene family in Arabidopsis thaliana L. and Oryza sativa L. Plant Gene 15, 15–27 (2018).
Endres, L., Dedon, P. C. & Begley, T. J. Codon-biased translation can be regulated by wobble-base tRNA modification systems during cellular stress responses. RNA Biol. 12, 603–614 (2015).
Wang, X. et al. Evolution and roles of cytokinin genes in angiosperms 2: Do ancient CKXs play housekeeping roles while non-ancient CKXs play regulatory roles? Hortic Res. https://doi.org/10.1038/s41438-020-0246-z.
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014).
Bennett, T. et al. Paralogous radiations of PIN proteins with multiple origins of noncanonical PIN structure. Mol. Biol. Evol. 31, 2042–2060 (2014).
Letunic, I., Doerks, T. & Bork, P. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40, D302–D305 (2012).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Keane, T. M., Creevey, C. J., Pentony, M. M., Naughton, T. J. & McLnerney, J. O. Assessment of methods for amino acid matrix selection and their use on empirical data shows that ad hoc assumptions for choice of matrix are not justified. BMC Evol. Biol. 6, 29 (2006).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic Acids Res. 43, W39–W49 (2015).
Lee, T. H., Tang, H., Wang, X. & Paterson, A. H. PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res. 41, D1152–D1158 (2013).
Tang, H. B. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Li, Y., Pi, M., Gao, Q., Liu, Z. & Kang, C. Updated annotation of the wild strawberry Fragaria vesca V4 genome. Hortic. Res. 6, 61 (2019).
Kang, C. et al. Genome-scale transcriptomic insights into early-stage fruit development in woodland strawberry Fragaria vesca. Plant Cell 25, 1960–1978 (2013).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Amil-Ruiz, F. et al. Identification and validation of reference genes for transcript normalization in strawberry (Fragaria × ananassa) defense responses. PLoS ONE 8, e70603 (2013).
Acknowledgements
We thank Dr. Janet Slovin (United States Department of Agriculture, USDA) for providing seeds of Fragaria vesca ‘Ruegen’. We also thank members of the Y.L. Laboratory for discussions and comments on the manuscript. This work was supported by National Natural Science Foundation of China grants 31672124 (to L.G.) and 31471860 (to J.D.).
Authors’ contributions
Y.L. and J.D. designed the experiments; X.W. and S.L. performed the experiments and data analyses; D.L. participated in the data analyses; J.D., X.W. and Y.L. wrote the manuscript; Y.L., L.G. and R.M. edited and revised the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Yi Li holds a no pay visiting professor position at Nanjing Agricultural University.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, X., Lin, S., Liu, D. et al. Evolution and roles of cytokinin genes in angiosperms 1: Do ancient IPTs play housekeeping while non-ancient IPTs play regulatory roles?. Hortic Res 7, 28 (2020). https://doi.org/10.1038/s41438-019-0211-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-019-0211-x
This article is cited by
-
Cytokinin biosynthesis in Hexapoda and Insecta: a bioinformatic analysis
Arthropod-Plant Interactions (2024)
-
The tRNA-degradation pathway impacts the phenotype and metabolome of Arabidopsis thaliana: evidence from atipt2 and atipt9 knockout mutants
Plant Growth Regulation (2024)
-
Identification of IPT9 in Brachiaria brizantha (syn. Urochloa brizantha) and expression analyses during ovule development in sexual and apomictic plants
Molecular Biology Reports (2023)
-
Root predominant overexpression of iaaM and CKX genes promotes root initiation and biomass production in citrus
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
A novel adenylate isopentenyltransferase 5 regulates shoot branching via the ATTTA motif in Camellia sinensis
BMC Plant Biology (2021)