Abstract
Landscape features shape patterns of gene flow among populations, ultimately determining where taxa lay along the continuum between panmixia to complete reproductive isolation. Gene flow can be restricted, leading to population differentiation in two non-exclusive ways: “physical isolation”, in which geographic distance in combination with the landscape features restricts movement of individuals promoting genetic drift, and “ecological isolation”, in which adaptive mechanisms constrain gene flow between different environments via divergent natural selection. In central Iberia, two fire salamander subspecies occur in parapatry across elevation gradients along the Iberian Central System mountains, while in the adjacent Montes de Toledo Region only one of them occurs. By integrating population and landscape genetic analyses, we show a ubiquitous role of physical isolation between and within mountain ranges, with unsuitable landscapes increasing differentiation between populations. However, across the Iberian Central System, we found strong support for a significant contribution of ecological isolation, with low genetic differentiation in environmentally homogeneous areas, but high differentiation across sharp transitions in precipitation seasonality. These patterns are consistent with a significant contribution of ecological isolation in restricting gene flow among subspecies. Overall, our results suggest that ecological divergence contributes to reduce genetic admixture, creating an opportunity for lineages to follow distinct evolutionary trajectories.
Similar content being viewed by others
Introduction
Understanding how landscapes shape biodiversity patterns is a long-standing goal in ecology and evolution (Wright 1943; Nosil 2012; Balkenhol et al. 2016). Landscapes can influence gene flow and, consequently, patterns of genetic variation in two inherently distinct ways. First, through “physical isolation”, in which neutral genetic divergence increases with geographic distance (isolation by distance [IBD]; Wright 1943), alone or in combination with landscape features (e.g., topography, climate or land cover) that create general resistance to the movement of individuals (isolation by resistance [IBR]; McRae 2006). Second, through “ecological isolation”, in which adaptive mechanisms constrain gene flow between different environments via divergent natural selection (isolation by environment [IBE]; Wang and Bradburd 2014). Importantly, although physical and ecological isolation are associated with distinct evolutionary forces, they are not mutually exclusive. Physical isolation facilitates the fixation of locally adaptive alleles (Nosil 2012), and transitions between ecologically divergent taxa tend to be coincident with areas of low habitat suitability and population density (Barton and Gale 1993). Understanding the evolutionary processes that shape biodiversity requires disentangling their relative contribution in generating and maintaining barriers to gene flow (Wang et al. 2013; Balkenhol et al. 2016).
Ecological isolation can result from various factors, including (i) selection against migrants, in which individuals dispersing among contrasting environments show lower fitness relative to residents, (ii) selection against hybrids, when hybrids are less fit than parentals in their environments, (iii) habitat preference, if individuals show preferential associations with natal habitats relative to alternative ones, and (iv) temporal (allochronic) isolation, occurring when distinct environments are associated with distinct activity and breeding periods (Wang and Bradburd 2014; Hendry 2017). Processes driving ecological isolation are expected to predominate in heterogeneous environments that facilitate ecological contrasts (Hendry 2017). Studies inferring the relative role of physical and ecological isolation in restricting gene flow have been conducted across numerous taxa and environments (e.g., Sexton et al. 2014; Wang and Bradburd 2014; Hendry 2017), but rarely among well-defined intraspecific phylogeographic groups (Velo‐Antón et al. 2013; Gray et al. 2019). Also, the relative contribution of these factors at different stages of divergence, from panmixia to complete reproductive isolation, remains unclear.
The Iberian Peninsula is a topographically and climatically heterogeneous region, offering multiple opportunities for physical and ecological divergence. Its role as a center of diversification is supported by high levels of endemism and intraspecific variability across multiple taxa (Gomez and Lunt 2007; Abellán and Svenning 2014). Patterns of genetic variation in Iberia have been investigated in depth in multiple taxa (from insects, e.g., Noguerales et al. 2017; freshwater fishes, e.g., Méndez et al. 2019; to large carnivores, e.g., Silva et al. 2018), but especially in ectotherms (e.g., Miraldo et al. 2011; Gutiérrez-Rodríguez et al. 2017a, b; Maia-Carvalho et al. 2018; Sánchez-Montes et al. 2019; Dufresnes et al. 2020; Martínez-Freiría et al. 2020), revealing pronounced patterns of genetic structure that have been attributed to processes of physical and/or ecological divergence. The fire salamander, Salamandra salamandra (Linnaeus 1758), is a remarkable example on both fronts. Broadly distributed across Europe, it is in Iberia where it shows its highest levels of genetic and phenotypic diversity (Burgon et al. 2021), occurring across a wide range of habitats and environmental conditions (Velo-Antón and Buckley 2015; Fig. 1). Across Europe, several genetically differentiated lineages of S. salamandra hybridize in contact zones of varying widths, usually associated with strong mito-nuclear discordances (e.g., García-París et al. 2003; Pereira et al. 2016; Bisconti et al. 2018), and sometimes coincident with sharp ecological transitions. In such cases, parental populations have evolved differences in color pattern (Beukema et al. 2016; Burgon et al. 2020), morphology (Alarcón-Ríos et al. 2020), and reproductive modes (García-París et al. 2003; Velo-Antón et al. 2007; Lourenço et al. 2019) that are suggestive of local adaptation and are taxonomically recognized as subspecies. The integration of population and landscape analyses with the study of hybrid zones in Salamandra can help to infer the relative role of landscape features and ecological factors in reproductive isolation.
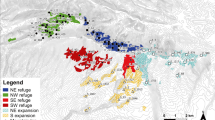
When mtDNA is available, localities are marked with colors from the adjacent mitochondrial tree, adapted from Pereira et al. (2016) (white dots represent newly sampled localities for which mtDNA data was not available). The mtDNA based distribution range of S. s. bejarae and S. s. almanzoris is displayed in transparent blue and red respectively. Inset map shows ranges from other S. salamandra subspecies present in Iberia (S.s bejarae – blue, S. s. almanzoris – red, S. s. gallaica – pink, S. s. longirostris – dark gray, S. s. morenica – green, S. s. crespoi – yellow, S. s. bernardezi – dark red, S. s. fastuosa – orange, S. s. terrestris – brown). ICS stands for Iberian Central System and MTR for Montes de Toledo Region. Sampling localities used for population and landscape genetic analyses identified with numbers (1–18, see details in Methods).
Across the Iberian Central System (ICS), populations from two distinct subspecies, S. s. bejarae and S. s. almanzoris, occur in parapatry across multiple regions associated to elevation gradients (Martínez-Solano et al. 2005; Pereira et al. 2016). The high-elevation subspecies S. s. almanzoris shows a restricted distribution in three isolated mountain tops, while the subspecies S. s. bejarae inhabits the surrounding lower elevation habitats (Fig. 1). From an ecological viewpoint, S. s. almanzoris is described as a distinct S. salamandra “ecotype” (defined as an independent evolutionary unit showing adaptations to specific environments; e.g., Hendrix et al. 2017; Czypionka et al. 2018; Burgon et al. 2020), mainly because of its longer larval period, small body size and reduced size of the yellow spots (but see Bosch and López-Bueis 1994), in comparison with the parapatric S. s. bejarae (Velo-Antón and Buckley 2015). Yet, because larval traits and adult body size are known to be subject to some degree of plasticity within S. salamandra subspecies (e.g., Alcobendas and Castanet 2000; Alcobendas et al. 2004; Alarcón-Ríos et al. 2020), it remains unclear whether such ecological divergence is genetically based or the result of developmental phenotypic plasticity. A previous study on this system revealed that the two subspecies are highly divergent at the mtDNA level (Fig. 1), whereas nuclear DNA shows low or no differentiation across parapatric boundaries and high divergence across allopatric boundaries (Pereira et al. 2016). The highly divergent mtDNA lineages are fixed within populations and have been used to delineate the two subspecies, although they co-occur in regions of secondary contact (Martínez-Solano et al. 2005). The strong mito-nuclear discordances may result from a scenario of vicariant evolution leading to the differentiation of both subspecies followed by nuclear-mediated gene flow across contact zones, potentially following range expansions from western refugia along the ICS (Pereira et al. 2016). South of the ICS, populations of S. s. bejarae in the Montes de Toledo Region (MTR) have probably evolved in isolation from S. s. almanzoris during the most recent glacial cycles (Pereira et al. 2016). Whether physical and ecological isolation is maintaining genetic differentiation between subspecies and between populations of the same subspecies remains an open question, which can be answered by integrating population and landscape genetic analyses.
Here, we infer the relative contribution of physical and ecological isolation in patterns of population differentiation in two mountain systems (ICS and MTR) characterized by distinct evolutionary and environmental contexts. The relative role of landscape features and ecology is expected to depend highly on the spatial extent of analyses (Hendry 2017; Fletcher and Fortin 2018). Specifically, a correlation between genetic differentiation and geographic distance (IBD) or landscape resistance (IBR) across the landscape is consistent with a major role of “physical isolation” restricting gene flow, whereas an additional correlation between genetic differentiation and local environmental conditions would be consistent with a role of “ecological isolation” constraining gene flow (IBE). Considering this, we conducted population and landscape genetic analyses across three distinct spatial extents (Total, ICS and MTR) and assessed the relative role of physical isolation (IBD/R) and ecological isolation (IBE) in explaining patterns of population connectivity and differentiation.
Material and methods
Specimen sampling
Sampling was designed to cover the known high-elevation isolates of the subspecies S. s. almanzoris and their potential contact zones with S. s. bejarae along the ICS (Fig. 1). We also collected samples from the allopatric populations of S. s. bejarae across the MTR (see Fig. 1). Samples already available from Pereira et al. (2016) were combined with new collections, resulting in a total of 385 georeferenced tissue samples (mostly tail clips from larvae, but also some toe clips from metamorphosed individuals) (Fig. 1; Table S1). Importantly, to avoid potential biases arising from uncertainties related with the distribution of each subspecies, our analyses do not rely on a priori assumptions regarding their classification. Instead, we identify evolutionary lineages based on our nuclear data and assign them to subspecies based on diagnostic mitochondrial data (Martínez-Solano et al. 2005) only a posteriori, to aid interpretation of results (detailed below).
Laboratory procedures
For the new samples, DNA was extracted using NucleoSpin Tissue-Kits (Macherey-Nagel). A total of 14 microsatellites (Steinfartz et al. 2004; Hendrix et al. 2010), distributed in four optimized multiplex reactions (see Table S2), were amplified by polymerase chain reaction (PCR). Each PCR was performed in a total volume of 10 µl: 5 µl of Multiplex PCR Kit Master Mix (QIAGEN), 3 µl of distilled water, 1 µl of primer multiplex mix and 1 µl of DNA extract (~50 ng/µl). To identify possible contaminations, a negative control was used in all reactions. PCR touchdown cycling conditions were equal in all multiplexes: starting with an initial step at 95 °C for 15 min, 19 cycles at 95 °C for 30 s, 90 s of annealing at 65 °C (decreasing 0.5 °C each cycle), 72 °C for 40 s, followed by 25 cycles of 95 °C for 30 s, 56 °C for 60 s, 72 °C for 40 s, and ended with a final extension of 30 min at 60 °C. PCR products were sent to Secugen S. L. for genotyping and alleles were scored in GENEMAPPER 4.0 (Applied Biosystems). Genotyped samples or markers presenting more than 40% of missing data were discarded.
Microsatellite genetic data pre-treatment
Individual samples were grouped into populations using 1 km radius buffer intersections (distance below the maximum dispersal distance of ~2 km recorded in Salamandra salamandra; Hendrix et al. 2017). Locations with a sample size ≥8 were considered populations and used for subsequent population and landscape genetic analyses. This threshold was defined to maximize the number of populations and spatial coverage for posterior landscape genetic analyses. A threshold of 10 was used in preliminary analyses and revealed comparable patterns of differentiation and diversity. Localities with less than 8 individuals were incorporated for analyses of genetic structure. Deviations from Hardy–Weinberg equilibrium (HWE) and linkage disequilibrium (LD) were examined in each population (locations with ≥8 samples), that is 254 samples (distributed across 18 populations), corresponding to ~66% of our complete dataset. This was done by performing exact tests implemented in GENEPOP 4.5.1 (Rousset 2008; dememorization = 10,000, batch length = 10,000, batch number = 1000). We applied the false discovery rate (Benjamini and Hochberg 1995) to correct p values (p < 0.05) from HWE and LD multiple exact tests. The presence of null alleles was investigated for each population in INEST 2.0 (Chybicki and Burczyk 2009) with a total of 200,000 iterations thinned every 50 iterations and a burn-in of 10% for the individual inbreeding model.
Genetic diversity and structure
To characterize patterns of genetic diversity, we used populations as previously defined (i.e., locations with a sample size ≥8 and with 1 km radius buffer intersections, see Fig. 1). We used GenAlEx 6.502 (Peakall and Smouse 2006) to estimate observed heterozygosity (HO), expected heterozygosity (HE), and mean number of alleles (NA). Additionally, unbiased allelic richness (AR) and private allelic richness (P-AR) were calculated in HP-RARE 1.0 (Kalinowski 2005). Allelic richness was spatially interpolated to visualize the geographic distribution of genetic diversity. Genetic differentiation was estimated using the pairwise FST (Weir and Cockerham 1984) calculated in R (R Development Core Team 2018) package diveRsity version 1.9.89 (Keenan et al. 2013). Respective 95% confidence intervals (CIs) were estimated using 1000 permutations, and pairwise estimates were considered significant when 95% CIs did not overlap zero.
Genetic structure was assessed using the Bayesian algorithm implemented in STRUCTURE 2.3.4 (Pritchard et al. 2000) in a hierarchical manner (i.e., a first assessment of the total dataset, followed by independent assessments of the major genetic groups recovered – corresponding to ICS and MTR mountains, see Results). Ten independent runs were performed for a number of clusters (K) ranging between 1 and 10. Runs consisted of a burn-in period of 1 × 10−5 iterations, followed by 1 × 10−6 iterations, with a correlated allele frequencies admixture model and no prior information regarding population of origin. The K showing the highest ΔK statistic value (Evanno et al. 2005) was identified using STRUCTURE HARVESTER 0.6.94 (Earl and VonHoldt 2012) and deemed as the best K describing the observed genetic data. For the optimal K, runs were summarized in Pophelper 1.0.10 (Francis 2016) and graphically displayed in R. Additionally, a discriminant analysis of principal components (DAPC; Jombart et al. 2010) was used as an alternative, complementary description of population structure, because this method does not assume HWE nor linkage equilibrium within clusters. DAPC summarizes the data to minimize genetic differentiation within previously defined groups “(i.e., locations with a sample size ≥8) while maximizing it between groups, potentially unraveling population substructure linked to complex isolation patterns (e.g., IBD, IBR and/or IBE; Jombart et al. 2010). For this analysis, we assumed six genetic clusters based on the Bayesian Information Criterion and ΔK (see Results), while the number of retained principal components and discriminant functions were selected following the package’s manual guidelines to avoid overfitting.
Landscape resistance data for physical isolation analyses
Pairwise landscape resistances among populations, representing how gene flow is constrained across the landscape, were obtained for: geographic distance alone (IBD), climatic resistance (IBR-Climate), topographic resistance (IBR-Slope), land cover resistance (IBR-Landcover) and a complex model incorporating all the previous information (IBR-All). It should be noted that all IBR models constructed incorporate IBD in order to test if IBR models incorporating environmental variables explain the observed data better. The workflow used to obtain IBD/IBR matrices can be summarized as follows. First, ecological niche models (ENMs) were built independently for each set of variables (Climate, Slope, Landcover and All) and used to obtain landscape suitability layers. Those were then converted to resistance layers and used as input in CIRCUITSCAPE 4.0 (McRae 2006) to obtain pairwise matrices of resistance among populations. Below we address each of these steps in detail.
Ecological niche models (ENMs) were built using as input occurrence records encompassing the distribution of both subspecies obtained from our own field work, collaborators, and the Spanish Herpetological Atlas (Table S3). To avoid statistical biases arising from spatial autocorrelation (Fourcade et al. 2014), a maximum of one presence record per 10 × 10 km grid cell and a minimum distance of 2 km (maximum dispersal distance per generation recorded for the species) was allowed. The Nearest Neighbour index function of ArcGIS (ESRIR) was used to assess the degree of data clustering (occurrence data was not significantly different than random, z-score = −0.81). As previously mentioned, occurrence records of S. salamandra were used in an undifferentiated way, allowing us to avoid assumptions related to the distribution or the ecological differentiation of the different subspecies. Nevertheless, it should be kept in mind that, if the described distributions of both subspecies (Fig. 1) are correctly defined, most occurrence records belong to S. s. bejarae (28 out of 38), which could bias patterns of suitability. Spatial data representing the different landscape characteristics (geographical distance, climate, slope, and landcover; see Table S4) were obtained from the following sources. First, we downloaded data from nineteen climatic variables with a 30 arc-seconds resolution (~1 × 1 km) from WorldClim Version 1.4 (1960–1990). Studies disentangling the effect of geographic and ecological isolation in a reptile have used the same cell resolution with success (Wang et al. 2013). Also, studies in other salamanders (Velo‐Antón et al. 2013) and S. salamandra (Antunes et al. 2018) were able to extract meaningful conclusions regarding the role of landcover, climate, and topography in shaping patterns of genetic structure using the same baseline information for spatial analyses. In order to represent climatic conditions and avoid the possible statistical effects of collinearity among predictor variables on niche modeling, we retained five variables with pairwise Pearson’s r ≤ 0.70: Isothermality (ISO), Temperature Annual Range (TAR), Mean Temperature of Wettest Quarter (MTWQ), Precipitation of Wettest Quarter (PWQ) and Precipitation of Driest Quarter (PDQ). Slope was derived from an elevation layer (~1 × 1 km; SRTM), using the Slope function of ArcGis. Finally, categorical land cover data with 100 m resolution was downloaded from https://land.copernicus.eu/pan-european/corine-land-cover/clc-2012. The distribution area for model construction, validation and projections was created using a 200 km radius buffer around occurrence records. Bootstrap subsampling was used to build 30 model replicates, retaining 30% of the data records to test each replicate. Ecological niche models built for broadly similar Salamandra datasets (Antunes et al. 2018; Dinis et al. 2018) used the same percentage of training data and performed well, based on high area under the receiver operating characteristic curve (AUC) scores. Default values were used for all other settings. Model performance was evaluated using the AUC in MAXENT, which ranges from 0.5 (complete randomness) to 1 (perfect discrimination) (Phillips et al. 2006).
Habitat suitability layers depicting cell suitability (S) obtained from the ENMs were converted into resistance layers (i.e., resistance to movement) representing cell resistance (R = (1–S)*100). These layers were subsequently used as input for CIRCUITSCAPE 4.0 (McRae 2006) analyses to obtain the pairwise landscape resistance matrices among populations for each resistance model. CIRCUITSCAPE calculates averaged pairwise resistance to dispersal among populations based on all possible paths (unlike the least-cost path), thus better explaining the movement of individuals among widely separated regions (McRae et al. 2008). For CIRCUITSCAPE analyses, the pairwise mode and a cell connection scheme of eight neighbors were set, while remaining parameters were set as default.
Environmental dissimilarity data for ecological isolation analyses
The role of environmental dissimilarity in shaping patterns of genetic connectivity among populations (IBE) was assessed using climate-based dissimilarity measures between population locations, summarizing and capturing the environmental variation using a Principal Component Analysis (PCA). This was done explicitly by calculating the mean values of 19 bioclimatic variables (WorldClim Version 1.4.) and elevation (www.diva-gis.org/gdata) within a 2 km radius buffer around each locality. Obtained values were centered to zero and scaled to a standard deviation of one and data complexity was reduced with PCA (see Fig. S1). This was done independently for the Total, ICS and MTR ranges. The first three Principal Components (PCs) of each analysis (Total, ICS and MTR) were selected for downstream analyses, as they represented most of the total environmental variation (Fig. S1). Then, Euclidean distances between populations were obtained for each PC, representing environmental dissimilarities associated with distinct environmental characteristics. We also ran hierarchical clustering analyses for visualization purposes. This resulted in trees representing environmental dissimilarity among populations, which help identify populations driving connectivity patterns (i.e., pairs of populations with higher or lower environmental dissimilarity than most pairwise comparisons). Furthermore, to interpret the PCs, we noted the relative weight of all climatic variable loadings in the PCs explaining most of the observed variability (Fig. S2). All analyses were implemented using R (R Core Team 2018) package raster (Hijmans and van Etten 2016).
Analyses of physical and ecological isolation – landscape genetics
Models inferring the relative role of physical and ecological isolation in genetic differentiation were obtained independently for the three spatial extents considered (Total, ICS, and MTR). Models used matrices of genetic differentiation (pairwise FST) as the response variable and matrices of physical isolation (i.e., landscape resistance [IBD, IBR-Climate, IBR-Slope, IBR-Landcover, and IBR-All]) and ecological isolation (i.e., environmental dissimilarity [IBE-PC1, -PC2 and -PC3]) for -Total, -ICS, and -MTR, as predictors. Model building and selection was done using two independent regression-based methods [multiple matrix regression with randomization approach (MMRR, Wang 2013) and a maximum likelihood population effects model (MLPE, Clarke et al. 2002)] in two steps. First, because matrices of physical isolation are highly correlated among themselves (r > 0.90, see Fig. S3) we independently regressed them against the matrices of pairwise genetic differentiation (pairwise FST). Results from these univariate regressions allowed us to find the best model of physical isolation for each region (based on the highest r2 and lowest conditional Akaike information criterion [AICc], for MMRR and MLPE, respectively, see Table S5). Then, we proceeded to models incorporating both physical and ecological isolation matrices. Model selection for MMRR was done using a backward elimination procedure, starting with a full model that included all predictor variables (the best physical isolation matrix [higher r2 from univariate regressions, for the respective range, Table S5], and all ecological isolation matrices) and then iteratively removing the variable with the highest (and not significant, p > 0.05) p value, while checking the overall increase in model fit. MLPE model selection was done based on the lowest AICc among all combinations of predictors (i.e., the best physical isolation matrix [lowest AICc from univariate regressions, for the respective range, Table S5], and all ecological isolation matrices). Only the final models selected for each region are displayed and discussed.
Results
Microsatellite genetic data pre-treatment
A total of 352 samples were successfully genotyped. One out of the 14 loci was not successfully amplified. Analyses assessing deviations from HWE and identification of null alleles revealed two loci (SST-A6-I and SalE2) as potentially containing null alleles in some populations. As this was only true for MTR populations (localities 1–8 in Fig. 1), we maintained these loci for downstream analyses. Nonetheless, we replicated some key analyses for the Total and MTR ranges removing potentially problematic loci (SM – Dataset 11 loci, for replicated analyses and results). No pairs of loci showed signs of linkage disequilibrium.
Patterns of genetic diversity and structure
Genetic diversity varied across the study area, with lower values in the easternmost populations in both ICS and MTR, where the species is at its range limit (Pop. 1 - NA = 3.31, HE = 0.40 and AR = 2.47; Pop. 18 - NA = 3.46, HE = 0.45 and AR = 2.54; Table 1 and Fig. 2). In contrast, the highest diversity values were found in western and central populations (Pop. 8 - NA = 5.54, HE = 0.72 and AR = 4.45; Pop. 12 - NA = 6.62, HE = 0.70 and AR = 4.24; Table 1 and Fig. 2). Both clustering analyses (i.e., STRUCTURE and DAPC) recovered concordant results, separating ICS and MTR at the highest hierarchical level of genetic structure (STRUCTURE K = 2, see Fig. 2 and S4; DAPC with PC1 separating both groups, see Fig. 3). The ICS and MTR groups were further substructured in four and two groups, respectively (Figs. 2 and 3). Population pairwise genetic differentiation (FST) ranged from 0.001 to 0.511 across the Total range, from 0.003 to 0.411 across the ICS range, and from 0.012 to 0.344 across the MTR range (Table 2).
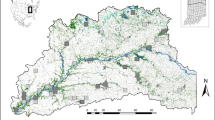
Pie charts, with size representing sample size, display population averages of ancestry proportions according to STRUCTURE results (K2 for MTR and K4 for ICS range). STRUCTURE barplots are shown below the map (Total, K2; MTR, K2; ICS, K4). On the bottom, another barplot displays assignment of individuals to mtDNA lineages (when available; gray color indicates lack of mtDNA data). A spatial interpolation of allelic richness (AR) is displayed in a green (high) to red (low) transparent gradient.
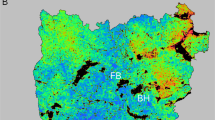
A Map showing gradient of precipitation seasonality (coefficient of variation) and genetic clusters from DAPC. B Tree representing environmental dissimilarity among populations from the ICS based on IBE-PC2-ICS (mostly related with precipitation seasonality, see Results). Colors match genetic clusters from DAPC analyses. C Relative densities of genotyped individuals plotted against the discriminant function 1 from DAPC analyses (DAPC-DF1), representing genetic divergence among groups. Arrows indicate clusters where genetic differentiation is strongly associated with “ecological isolation” (IBE). Below, hypothesized gradient bar showing subspecies hybridization (see Discussion).
Patterns of physical and ecological isolation
Under the MMRR approach, univariate analyses inferring the best model of physical isolation were all significant (p < 0.05) across the three ranges (see Table S5 for details in every model). For the Total and ICS ranges, the physical isolation model with the best fit incorporates all variables (IBR-All, r2 = 0.681 and r2 = 0.661, respectively). For the MTR range, a simpler model incorporating only climate resistance provided the best fit (IBR-Climate, r2 = 0.633). Model selection under the MLPE approach (based in AICc), recovered exactly the same top physical only models (Table S5).
Model selection using MMRR models, incorporating both physical and ecological isolation as predictors, produced distinct results among the three ranges. At the scale of the Total range, a model incorporating both physical (IBR-All) and ecological isolation (IBE-PC2 and IBE-PC3) was the best model (r2 = 0.747, Table 3). In this model, the regression coefficient for physical isolation was about four times greater (βIBR-All = 0.76; Table 3) than the regression coefficient for each ecological isolation component (βIBE-PC2-Total = 0.23, βIBE-PC3-Total = 0.21; Table 3). This supports a stronger role of physical isolation (IBR) across the Total range, but also reveals a significant role of ecological isolation (IBE) in driving patterns of genetic differentiation. At the scale of the MTR range, all ecological isolation predictors were excluded by the backward elimination procedure, and thus a model only considering physical isolation was selected as the best (IBR-Climate, r2 = 0.633; Table 3). At the scale of the ICS range, the best model included both physical (IBR-All) and ecological isolation (IBE-PC2) as predictors (r2 = 0.817, Table 3). In this model, the regression coefficient for ecological isolation (βIBE-PC2-ICS = 0.51) was similar to the regression coefficient for physical isolation (βIBR-All = 0.46; Table 3), consistent with the similar importance of both physical and ecological isolation predicting genetic differentiation in this area. Models selected under the MLPE approach (based in AICc) retrieved equal results for ICS and MTR. Across the Total range, the best model was different from the MMRR approach, with IBR-All + IBE-PC1 + IBE-PC2 showing the lowest AICc, and the IBR-All regression coefficient (βIBR-All-Total = 0.70, Table 3) about three times higher than IBE-PC2 coefficients (similar to MMRR; βIBE-PC2-Total = 0.27, Table 3) and five times higher than IBE-PC1 coefficients (βIBE-PC1-Total = 0.14, Table 3).
The hierarchical clustering trees representing environmental dissimilarity among populations of the Total range revealed stronger environmental dissimilarity between two main groups for PC2-Total (~Temperature Mean Diurnal Range, Precipitation Seasonality among other correlated variables; see Figs. S2 and S5): one including all MTR populations (bejarae) plus populations 10 (bejarae) and 11 (almanzoris) from the ICS, and another one with the remaining ICS populations (almanzoris, Fig. S5). For PC3-Total (~Isothermality mostly but also Precipitation Seasonality and others; Fig. S5), population 18 (almanzoris) was identified as being most environmentally distinct from all others (Fig. S5). The PC1-Total segregated populations at higher elevation across the Total (populations 10, 11 and 18, Fig. S5). The hierarchical clustering trees representing environmental dissimilarity for the ICS range segregated populations mostly by their levels of precipitation seasonality (Fig. 3). Specifically, population 18 occurred at lower levels of precipitation seasonality (~27%, average from localities in its genetic cluster, in red in Fig. 3), followed by populations 16 and 17 (almanzoris), located in an environmental transition zone (~32%, average from localities in their genetic cluster, in yellow in Fig. 3), populations 11 to 15 (almanzoris and bejarae) at similar levels (~34%, average from localities in their genetic cluster, in orange in Fig. 3), and finally population 10 at the other extreme (~39%, average from localities in their genetic cluster, in blue in Fig. 3). Hierarchical clustering trees representing environmental dissimilarity for the MTR range are not interpreted because model selection excluded all ecological isolation proxies as predictors for patterns of genetic differentiation.
Discussion
The role of “physical isolation” and genetic drift determining patterns of genetic differentiation between populations, subspecies and species is well established in the literature. Yet, such physical landscape elements may also interact with ecological factors, shaping patterns of genetic differentiation. When divergent natural selection contributes to patterns of genetic differentiation, gene flow within ecologically homogeneous habitats is expected to be higher relative to that between distinct habitats. In this study, we show that although “physical isolation” is a ubiquitous phenomenon predicting genetic differentiation between populations and subspecies of fire salamanders, “ecological isolation” plays an equally important role between populations from distinct subspecies.
The relative role of physical and ecological isolation in genetic differentiation
In central Iberia, S. salamandra is distributed across spatially heterogeneous and temporally dynamic environments, with populations in close proximity often experiencing independent evolutionary forces. This is the case in our study area, where the relative role of physical and ecological processes in population divergence differed between and within two mountain systems (ICS and MTR) characterized by contrasting levels of topographic and climatic complexity and by differences in historical factors associated with their colonization history.
Our results show that patterns of genetic differentiation between and within the ICS and MTR were largely explained by physical isolation (IBD and IBR) (Table S5). This is not surprising considering the low dispersal capacity of salamanders (2 km per generation; Hendrix et al. 2017). Furthermore, models explicitly accounting for other features of the landscape (i.e., IBR models) sometimes outperformed IBD models (Table S5). However, the ranking of IBR models was different among the three tested spatial ranges (Total, ICS and MTR, Table S5), with particularly contrasting results between mountain ranges (ICS vs MTR, Table S5). These results can be attributed to the differences in the evolutionary forces and environmental contexts across study areas. Climate-based models of physical isolation, for example, outperformed all models in MTR, while across the Total range and ICS regions they provided particularly poor predictions (worse than IBD, Table S5). Slope-based models produced poor predictions across all areas, also performing worse than IBD models. The low performance of these models could be partially explained by subspecies having different ecological niches. Our ENMs were built from records not distinguishing subspecies, and thus patterns of suitability may deviate from reality (and by extension resistance) if subspecies respond in distinct ways to environmental factors. Landcover-based models, on the other hand, always behaved similarly between regions, improving, although slightly, genetic differentiation predictions from IBD models in the ICS and MTR (Table S5). If we consider this together with the fact that forest cover had the highest weight in landcover ENMs (Table S6), our results add to the cumulative evidence that forested areas play a key role in genetic connectivity in salamander populations (Cushman 2006; Antunes et al. 2018; Emel et al. 2019; Lourenço et al. 2019; Arntzen and van Belkom 2020).
One landscape feature posing strong resistance to gene flow was the Tagus river basin, as shown by clustering analyses, which recovered two main genetic groups north (ICS) and south (MTR) of the river (Fig. 2 and S4). This is in agreement with other studies showing that large river basins can act as major barriers to gene flow in salamanders, an effect that is likely enhanced by the reduced forest cover of these flat, open areas (García-París et al. 1998; Antunes et al. 2018; Dinis et al. 2018). In addition to this major north-south break, we also observed a clear pattern of decreasing genetic diversity from west to east in both mountain systems (Fig. 2). This is consistent with the hypothesis of current populations resulting from range expansions originating in western refugia and proceeding eastwards along the ICS and the MTR (Pereira et al. 2016). This pattern is also consistent with the expected pattern of genetic variation in core-peripheral populations, with western populations being contiguous and maintaining high genetic diversity through migration, while the populations at the eastern margin of the species distribution are less connected and more subject to drift and thus have reduced diversity.
Results from multiple regression models accounting for both physical and ecological isolation varied between mountain ranges. Across the Total range, models incorporating physical and ecological isolation improved predictions of patterns of genetic differentiation, albeit the contribution of the ecological component was about three to five times weaker than that of the physical component. When multivariate landscape genetic analyses were conducted separately for each mountain range (ICS and MTR), results differed strongly, showing that physical and ecological isolation play an equally important role across the ICS, while across the MTR, physical isolation alone predicted patterns of genetic differentiation (Table 3).
Disentangling the relative contribution of physical and ecological isolation in predicting patterns of genetic differentiation between populations can provide insights onto the evolutionary processes determining gene flow. Although in the landscape genetics literature patterns of genetic differentiation are generally interpreted as reflective of gene flow, this is only true when populations are at a migration-drift equilibrium (sensu Wright 1943). Such equilibrium is hard to assess in natural populations, particularly in those affected by range shifts during the current interglacial period (Hewitt 2000), like fire salamanders (García-París et al. 2003). Yet, the fact that sampling localities at lower elevation have not been previously glaciated, and those at higher elevation have become deglaciated some 25,000–30,000 years ago (Domínguez-Villar et al. 2013), suggests that these populations should be close to an idealized migration-drift equilibrium. Further, we did not detect populations departing from HW or linkage equilibrium, which is consistent with demographic stability. In addition to gene flow, differences in effective population size can influence patterns of genetic differentiation because smaller populations experience stronger genetic drift. In our study, most localities show similarly high to intermedium levels of genetic diversity across multiple summary statistics (Table 1), suggestive of overall similar effective population size. Yet, the lower diversity observed in the easternmost populations (Pops. 18 and 1; Fig. 2 and Table 1) are indeed suggestive of smaller effective population sizes in areas colonized more recently. However, the species is locally abundant in this region (Martínez-Solano 2006), and other populations with similar low diversity (Pops. 16, 17 and 14; Fig. 2 and Table 1), are not as strongly differentiated (Table 2), suggesting that potential differences in effective population size are small and perhaps have a negligible contribution to patterns of genetic differentiation. Thus, we regard inferred patterns of genetic differentiation in our study area as most likely reflecting actual restrictions to gene flow.
Other environmental features, and the scale (spatial or temporal) at which they act, can also help to explain observed patterns of genetic differentiation (see for details Epps and Keyghobadi 2015; Winiarski et al. 2020). Some examples include physical isolation driven by the barrier effect of mountain ridges (Sánchez‐Montes et al. 2018), exposure to wind, which is known to significantly constrain the movement of S. salamandra even in less exposed landscapes (Lourenço et al. 2019), or less obvious factors such as the orientation of mountain slopes, that affects overall patterns of humidity and temperature (Mulder et al. 2019). Also, ecological isolation driven by landscape factors rather than the climatic factors (and elevation) explored in this study could play an important role, including the type of water bodies available for breeding and development (e.g., lotic vs. lentic, persistent vs ephemeral) or vegetation cover (e.g., deciduous, coniferous, mixed and shrublands). Considering this, future studies in this system should further explore patterns of IBR and IBE, not only by including new landscape and environmental variables, but also using alternative methodological and conceptual approaches (e.g., genetic based optimization of IBR models, Peterman 2018). At any rate, our results support the ubiquitous role of physical isolation in shaping genetic differentiation patterns among S. salamandra populations in central Iberia, and of ecological isolation playing an important role across the ICS.
The role of ecological isolation in “ecotype” divergence
The fire salamander subspecies S. s. almanzoris has been treated as a distinct ecotype associated with higher altitudinal ranges. Some characteristic features include longer aquatic larval stages, smaller size and reduced number and size of yellow spots, in comparison to parapatric subspecies S. s. bejarae (Velo-Antón and Buckley 2015, and references therein). The delimitation of these subspecies was later supported by high differentiation at the mtDNA level (Martínez-Solano et al. 2005). However, further analyses using five nuclear genes revealed low or no differentiation among subspecies at parapatric boundaries (Pereira et al. 2016), which led to question whether S. s. almanzoris represents an independent evolutionary unit, with distinct ecological features, despite gene flow.
Our population and landscape genetic analyses support a role of ecological divergence in shaping patterns of genetic structure across the ICS. Considering both patterns of gene flow and the distribution of mtDNA lineages (Fig. 2) we propose a scenario where MTR and West-ICS populations (genetic clusters in blue tones; Fig. 3) represent pure S. s. bejarae, at higher levels of precipitation seasonality, while the East-ICS populations (red genetic cluster) represent pure S. s. almanzoris, at lower levels of precipitation seasonality (Fig. 3). In this scenario, gene flow between pure subspecies is seemingly restricted by ecological isolation, associated with sharp transitions in precipitation seasonality (arrows indicating IBE; Fig. 3). In contrast, in areas characterized by intermediate levels of precipitation seasonality, gene flow between Western S. s. almanzoris and Eastern S. s. bejarae is less restricted (clusters in orange and yellow; Fig. 3). Although such patterns of gene flow match geographic distance, they were significantly better predicted when accounting for both physical and ecological isolation, with both processes having almost equal contributions (Table 3 and Fig. 3). We note, however, that correlation between matrices of physical (IBR-All) and ecological isolation (IBE-PC2-ICS) in the best model for the ICS range (Table 3) was considered not severe (r2 = 0.60, VIF = 2.5 < 10; Prunier et al. 2015). MMRR models were not tested with such levels of correlation and accuracy of relative weights can be questioned (Wang et al. 2013). Nonetheless, subspecies differentiation seems to be maintained in the Eastern ICS by the combined effect of physical and ecological isolation. Examples from S. salamandra ecotypes in which gene flow is restricted by processes of ecological divergence have already been documented in parapatric pond- and stream-breeding fire salamanders across the Kottenforst (Hendrix et al. 2017; but see Arntzen and van Belkom 2020).
Observed patterns of ecological divergence can underline either adaptive or plastic processes, or a combination of both (Wogan et al. 2020). Characters like the length of the larval stage and adult body size are known to be subject to developmental plasticity within S. salamandra subspecies (e.g., Alcobendas and Castanet 2000; Alcobendas et al. 2004). Nonetheless, some of those traits are also known to have a genetic basis, like color patterns (Burgon et al. 2020), growth rate, or body size (Alcobendas and Castanet 2000). Interestingly, one study exploring the evolutionary responses of ectothermic vertebrates to climatic variation identified variation in precipitation seasonality as the best predictor for variation in the number of thoracic vertebrae (a trait linked to body size) in S. salamandra (Ficetola et al. 2016).
While we cannot infer the relative importance of each process in ongoing ecological divergence, our results are consistent with a scenario of environmental adaption, where genetic divergence occurs via physical and ecological isolation and maintains S. s. almanzoris phenotypic traits in high-elevation environments despite extensive gene flow in neutral parts of the genome (Pereira et al. 2016). Contemporary patterns of neutral genetic variation (Fig. 2) show that previously described mito-nuclear discordances probably result from post-glacial range expansions of S. s. bejarae into the range of S. s. almanzoris, with both subspecies merging across most of the ICS, possibly favored by shallower ecologic transitions. The coincidence of sharp genetic and ecological transitions in the easternmost ICS suggests that this region may be the only remnant of the original (unmixed) S. s. almanzoris. Their importance as an independent evolutionary unit with a distinct ecology should be considered in conservation strategies.
Data availability
Individual genotypes have been deposited in Dryad (https://doi.org/10.5061/dryad.rjdfn2z97).
References
Abellán P, Svenning JC (2014) Refugia within refugia–patterns in endemism and genetic divergence are linked to Late Quaternary climate stability in the Iberian Peninsula. Biol J Linn Soc 113:13–28
Alarcón-Ríos L, Nicieza AG, Kaliontzopoulou A, Buckley D, Velo-Antón G (2020) Evolutionary history and not heterochronic modifications associated with viviparity drive head shape differentiation in a reproductive polymorphic species, Salamandra salamandra. Evol Biol 47:43–55
Alcobendas M, Castanet J (2000) Bone growth plasticity among populations of Salamandra salamandra: interactions between internal and external factors. Herpetologica 56:14–26
Alcobendas M, Buckley D, Tejedo M (2004) Variability in survival, growth and metamorphosis in the larval fire salamander (Salamandra salamandra): effects of larval birth size, sibship and environment. Herpetologica 60:232–245
Antunes B, Lourenço A, Caeiro-Dias G, Dinis M, Gonçalves H, Martínez-Solano I et al. (2018) Combining phylogeography and landscape genetics to infer the evolutionary history of a short-range Mediterranean relict, Salamandra salamandra longirostris. Conserv Genet 19:1411–1424
Arntzen JW, van Belkom J (2020) ‘Mainland-island’ population structure of a terrestrial salamander in a forest-bocage landscape with little evidence for in situ ecological speciation. Sci Rep 10:1–15
Balkenhol N, Cushman SA, Waits LP, Storfer A (2016) Current status, future opportunities, and remaining challenges in landscape genetics. In: Balkenhol N, Cushman SA, Storfer AT, Waits LP (eds) Landscape genetics: concepts, methods, applications, John Wiley and Sons Ltd, Chichester, pp 247–255.
Barton NH, Gale KS (1993) Genetic analysis of hybrid zones. Hybrid zones and the evolutionary process. In: Harrison RG (ed) Hybrid zones and the evolutionary process. Oxford University Press, Oxford, p 13–45
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Stat Methodol 57:289–300
Beukema W, Nicieza AG, Lourenço A, Velo‐Antón G (2016) Colour polymorphism in Salamandra salamandra (Amphibia: Urodela), revealed by a lack of genetic and environmental differentiation between distinct phenotypes. J Zool Syst Evol 54:127–136
Bisconti R, Porretta D, Arduino P, Nascetti G, Canestrelli D (2018) Hybridization and extensive mitochondrial introgression among fire salamanders in peninsular Italy. Sci Rep 8:13187
Bosch J, López-Bueis I (1994) Comparative study of the dorsal pattern in Salamandra salamandra bejarae (Wolterstorff, 1934) and S. s. almanzoris (Müller & Hellmich, 1935). Herpetol J 4:46–48
Burgon JD, Vieites DR, Jacobs A, Weidt SK, Gunter HM, Steinfartz S et al. (2020) Functional colour genes and signals of selection in colour polymorphic salamanders. Mol Ecol 29:1284–1299
Burgon JD, Vences M, Steinfartz S, Bogaerts S, Bonato L, Donaire-Barroso D, Martínez-Solano I, Velo-Antón G, Vieites DR, Mable BK, Elmer KR (2021) Phylogenomic inference of species and subspecies diversity in the Pal earctic salamander genus Salamandra. Molecular Phylogenetics and Evolution 157:107063
Chybicki IJ, Burczyk J (2009) Simultaneous estimation of null alleles and inbreeding coefficients. J Hered 100:106–113
Clarke RT, Rothery P, Raybould AF (2002) Confidence limits for regression relationships between distance matrices: estimating gene flow with distance. JABES 7:361
Cushman SA (2006) Effects of habitat loss and fragmentation on amphibians: a review and prospectus. Biol Conserv 128:231–240
Czypionka T, Goedbloed DJ, Steinfartz S, Nolte AW (2018) Plasticity and evolutionary divergence in gene expression associated with alternative habitat use in larvae of the European Fire Salamander. Mol Ecol 27:2698–2713
Dinis M, Joger U, Slimani T, Martínez-Freiría F, Merabet K, Donaire D et al. (2018) Allopatric diversification and evolutionary melting pot in a North African Palearctic relict: the biogeographic history of Salamandra algira. Mol Phylogenet Evol 130:81–91
Domínguez-Villar D, Carrasco RM, Pedraza J, Cheng H, Edwards R, Willenbring JK (2013) Early maximum extent of paleoglaciers from Mediterranean mountains during the last glaciation. Sci Rep. 3:2034
Dufresnes C, Pribille M, Alard B, Dubey S, Perrin N, Gonçalves H et al. (2020) Integrating hybrid zone analyses in species delimitation: lessons from two anuran radiations of the Western Mediterranean. Heredity 124:423–438
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Emel SL, Olson DH, Knowles LL, Storfer A (2019) Comparative landscape genetics of two endemic torrent salamander species, Rhyacotriton kezeri and R. variegatus: implications for forest management and species conservation. Conserv Genet 20:801–815
Epps CW, Keyghobadi N (2015) Landscape genetics in a changing world: disentangling historical and contemporary influences and inferring change. Mol Ecol 24:6021–6040
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Ficetola GF, Colleoni E, Renaud J, Scali S, Padoa‐Schioppa E, Thuiller W (2016) Morphological variation in salamanders and their potential response to climate change. Glob Chang Biol 22:2013–2024
Fletcher R, Fortin M (2018) Spatial ecology and conservation modeling. Springer International Publishing, Cham
Fourcade Y, Engler JO, Rödder D, Secondi J (2014) Mapping species distributions with MAXENT using a geographically biased sample of presence data: a performance assessment of methods for correcting sampling bias. PLoS ONE 9:e97122
Francis RM (2016) pophelper: An r package and web app to analyse and visualize population structure. Mol Ecol Resour 17:27–32
García-París M, Alcobendas M, Alberch P (1998) Influence of the Guadalquivir river basin on mitochondrial DNA evolution of Salamandra salamandra (Caudata: Salamandridae) from southern Spain. Copeia 1998:173–176
García-París M, Alcobendas M, Buckley D, Wake DB (2003) Dispersal of viviparity across contact zones in iberian populations of fire salamanders (Salamandra) inferred from discordance of genetic and morphological traits. Evolution 57:129–143
Gomez A, Lunt DH (2007) Refugia within refugia: patterns of phylogeographic concordance in the Iberian Peninsula. In: Weiss S, Ferrand N (eds). Phylogeography of Southern European Refugia. Springer: Dordrecht. pp 155–188
Gray LN, Barley AJ, Poe S, Thomson RC, Nieto‐Montes de Oca A, Wang IJ (2019) Phylogeography of a widespread lizard complex reflects patterns of both geographic and ecological isolation. Mol Ecol 28:644–657
Gutiérrez-Rodríguez J, Barbosa AM, Martínez-Solano I (2017a) Present and past climatic effects on the current distribution and genetic diversity of the Iberian spadefoot toad (Pelobates cultripes): an integrative approach. J Biogeogr 44:245–258
Gutiérrez-Rodríguez J, Barbosa AM, Martínez-Solano I (2017b) Integrative inference of population history in the Ibero-Maghrebian endemic Pleurodeles waltl (Salamandridae). Mol Phylogenet Evol 112:122–137
Hendrix R, Schmidt BR, Schaub M, Krause ET, Steinfartz S (2017) Differentiation of movement behaviour in an adaptively diverging salamander population. Mol Ecol 26:6400–6413
Hendrix R, Susanne Hauswaldt J, Veith M, Steinfartz S (2010) Strong correlation between cross-amplification success and genetic distance across all members of “True Salamanders” (Amphibia: Salamandridae) revealed by Salamandra salamandra-specific microsatellite loci. Mol Ecol Resour 10:1038–1047
Hendry AP (2017) Eco-evolutionary dynamics. Princeton University Press, Princeton
Hewitt G (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hijmans RJ, Van Etten J (2016) raster: Geographic Data Analysis and Modeling. R package version 2.5-8. Available from: http://CRAN.R-project.org/package=raster
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:1–94
Kalinowski ST (2005) HP-RARE 10: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189
Keenan K, Mcginnity P, Cross TF, Crozier WW, Prodöhl PA (2013) DiveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788
Linnaeus C (1758) Systema Naturae per Regna Tria Naturae, Secundum Classes, Ordines, Genera, Species, cum Characteribus, Differentiis, Synonymis, Locis. 10th Edition. Volume 1. Stockholm, Sweden: L. Salvii
Lourenço A, Gonçalves J, Carvalho F, Wang IJ, Velo-Antón G (2019) Comparative landscape genetics reveals the evolution of viviparity reduces genetic connectivity in fire salamanders. Mol Ecol 28:4573–4591
Maia-Carvalho B, Vale CG, Sequeira F, Ferrand N, Martínez-Solano I, Gonçalves H (2018) The roles of allopatric fragmentation and niche divergence in intraspecific lineage diversification in the common midwife toad (Alytes obstetricans). J Biogeogr 45:2146–2158
Martínez-Freiría F, Freitas I, Zuffi MAL, Golay P, Ursenbacher S, Velo-Antón G (2020) Climatic refugia boosted allopatric diversification in Western Mediterranean vipers. J Biogeogr 47:1698–1713
Martínez-Solano I (2006) Atlas de distribución y estado de conservación de los anfibios de la Comunidad de Madrid. Graellsia 62:253–291
Martínez-Solano I, Alcobendas M, Buckley D, García-París M (2005) Molecular characterisation of the endangered Salamandra salamandra almanzoris (Caudata, Salamandridae). Ann Zool Fenn 42:57–68
McRae BH (2006) Isolation by resistance. Evolution 60:1551–1561
McRae BH, Dickson BG, Keitt TH, Shah VB (2008) Concepts and synthesis emphasizing new ideas to stimulate research in ecology using circuit theory to model connectivity in ecology, evolution, and conservation. Ecology 89:2712–2724
Méndez L, Perdices A, Machordom A (2019) Genetic structure and diversity of the Iberian populations of the freshwater blenny Salaria fluviatilis (Asso, 1801) and its conservation implications. Conserv Genet 20:1223–1236
Miraldo A, Hewitt GM, Paulo OS, Emerson BC (2011) Phylogeography and demographic history of Lacerta lepida in the Iberian Peninsula: multiple refugia, range expansions and secondary contact zones. BMC Evol Biol 11:170
Mulder KP, Rodriguez NC, Grant EHC, Brand A, Fleischer RC (2019) North ‐ facing slopes and elevation shape asymmetric genetic structure in the range ‐ restricted salamander Plethodon shenandoah. Ecol Evol 9:5094–5105
Noguerales V, Cordero PJ, Ortego J (2017) Testing the role of ancient and contemporary landscapes on structuring genetic variation in a specialist grasshopper. Ecol Evol 7:3110–3122
Nosil P (2012) Ecological speciation. Oxford University Press, New York
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pereira RJ, Martínez-Solano I, Buckley D (2016) Hybridization during altitudinal range shifts: nuclear introgression leads to extensive cyto-nuclear discordance in the fire salamander. Mol Ecol 25:1551–1565
Peterman WE (2018) ResistanceGA: an R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol Evol 9:1638–1647
Phillips SB, Aneja VP, Kang D, Arya SP (2006) Modelling and analysis of the atmospheric nitrogen deposition in North Carolina. IJGEI 6:231–252
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Prunier JG, Colyn M, Legendre X, Nimon KF, Flamand MC (2015) Multicollinearity in spatial genetics: Separating the wheat from the chaff using commonality analyses. Mol Ecol 24:263–283
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, https://wwwR-project.org/
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106
Sánchez‐Montes G, Wang J, Ariño AH, Martínez‐Solano I (2018) Mountains as barriers to gene flow in amphibians: quantifying the differential effect of a major mountain ridge on the genetic structure of four sympatric species with different life history traits. J Biogeogr 45:318–331
Sánchez-Montes G, Recuero E, Barbosa AM, Martínez-Solano I (2019) Complementing the Pleistocene biogeography of European amphibians: testimony from a southern Atlantic species. J Biogeogr 46:568–583
Sexton JP, Hangartner SB, Hoffmann AA (2014) Genetic isolation by environment or distance: which pattern of gene flow is most common? Evolution 68:1–15
Silva P, López-Bao JV, Llaneza L, Álvares F, Lopes S, Blanco JC et al. (2018) Cryptic population structure reveals low dispersal in Iberian wolves. Sci Rep 8:14108
Steinfartz S, Küsters D, Tautz D (2004) Isolation and characterization of polymorphic tetranucleotide microsatellite loci in the fire salamander Salamandra salamandra (Amphibia: Caudata). Mol Ecol Notes 4:626–628
Velo-Antón G, García-París M, Galán P, Cordero Rivera A (2007) The evolution of viviparity in Holocene islands: ecological adaptation versus phylogenetic descent along the transition from aquatic to terrestrial environments. J Zool Syst Evol 45:345–352
Velo‐Antón G, Parra JL, Parra‐Olea G, Zamudio KR (2013) Tracking climate change in a dispersal‐limited species: reduced spatial and genetic connectivity in a montane salamander. Mol Ecol 22:3261–3278
Velo-Antón G, Buckley D (2015) Salamandra común—Salamandra salamandra. In: Salvador A, Martínez-Solano I (eds), Enciclopedia Virtual de los Vertebrados Españoles, Museo Nacional de Ciencias Naturales, CSIC www.vertebradosibericos.org
Wang IJ (2013) Examining the full effects of landscape heterogeneity on spatial genetic variation: a multiple matrix regression approach for quantifying geographic and ecological isolation. Evolution 67:3403–3411
Wang IJ, Bradburd GS (2014) Isolation by environment. Mol Ecol 23:5649–5662
Wang IJ, Glor RE, Losos JB (2013) Quantifying the roles of ecology and geography in spatial genetic divergence. Ecol Lett 16:175–182
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Winiarski KJ, Peterman WE, Whiteley AR, McGarigal K (2020) Multiscale resistant kernel surfaces derived from inferred gene flow: an application with vernal pool breeding salamanders. Mol Ecol Resour 20:97–113
Wogan GOU, Yuan ML, Mahler DL, Wang IJ (2020) Genome-wide epigenetic isolation by environment in a widespread Anolis lizard. Mol Ecol 29:40–55
Wright S (1943) Isolation by distance. Genetics 28:114–138
Acknowledgements
We thank A. Álvarez, M.J. Fernández Benéitez, D. Fernández Ortín, R. Finat, C. Grande, J. Gutiérrez Rodríguez, P. Hernández Sastre, E. Jockusch, J.R. Mayor, M. Modrell, E. Recuero, G. Sánchez Montes, A. Sánchez Vialas, J.A. Saz, and F. Smith for help in collecting samples, and A. Lourenço and J. Gutiérrez Rodríguez for help with lab work. We appreciate the help of I. Rey and B. Álvarez (Tissue and DNA collection, MNCN-CSIC) for access to samples under their care. The Associate Editor R. Faria, W. Babik and three anonymous reviewers provided constructive comments that contributed to improve the manuscript. GV-A was supported by FCT – Foundation for Science and Technology (CEECIND/00937/2018). This research was supported by the European Science Foundation (Frontiers of Speciation Research, Exchange grant 3318), and by the European Commission (Synthesys grant ES-TAF-1486), granted to RJP. Partial funds were additionally provided by grants EVOVIV: PTDC/BIA-EVF/3036/2012 (FCT, Portugal) to GV-A, and CGL2017-83131-P (FEDER/Ministerio de Ciencia, Innovación y Universidades–Agencia Estatal de Investigación, Spain) to IMS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Rui Faria
Supplementary information
Rights and permissions
About this article
Cite this article
Antunes, B., Velo-Antón, G., Buckley, D. et al. Physical and ecological isolation contribute to maintain genetic differentiation between fire salamander subspecies. Heredity 126, 776–789 (2021). https://doi.org/10.1038/s41437-021-00405-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-021-00405-0
This article is cited by
-
Ancient allopatry and ecological divergence act together to promote plant diversity in mountainous regions: evidence from comparative phylogeography of two genera in the Sino-Himalayan region
BMC Plant Biology (2023)
-
Modelling the effects of climate and land-cover changes on the potential distribution and landscape connectivity of three earth snakes (Genus Conopsis, Günther 1858) in central Mexico
The Science of Nature (2023)
-
Landscape resistance constrains hybridization across contact zones in a reproductively and morphologically polymorphic salamander
Scientific Reports (2021)