Abstract
Sex chromosomes are predicted to harbour elevated levels of sexually antagonistic variation due to asymmetries in the heritability of recessive traits in the homogametic versus heterogametic sex. This evolutionary dynamic may manifest as high recessive load specifically affecting the homogametic sex, and the retention of haplotype diversity in small populations. We tested the hypothesis that the Z chromosome in the butterfly Bicyclus anynana carries a high inbred load for male fertility and viability. Homozygosity of Z chromosome blocks was produced by daughter–father backcrosses, and inferred from marker loci positioned via a linkage map. Male sterility was, in general, unrelated to homozygosity in any region of the Z, but there was an extreme deficit of homozygous males within a 2 cM interval in all families. In contrast, no corresponding skew in Z genotype was detected in their (hemizygous) sisters. The same pattern was observed in historically inbred lines, indicating a high frequency of recessive lethals in the ancestral population. Allele-frequency changes between 1993 and 2006 (70 generations at Ne ~ 160) show that, despite the loss of many haplotypes, diversity was retained significantly above the neutral expectation. Effective overdominance in the lethal region can account for this effect locally but not in other parts of the chromosome, that are also associated with persistent linkage disequilibrium. These unexpected patterns suggest the operation of other factors, such as epistatic selection, recombination suppression, assortative mating and meiotic drive. Our results highlight the role of balancing selection in maintaining the inbred load and linked genetic diversity.
Similar content being viewed by others
Introduction
Understanding the factors that maintain the alleles responsible for inbreeding depression is relevant to explaining fitness variation in natural populations (Charlesworth 2015), to predicting the risk of extinction of small populations (Saccheri et al. 1998; Bijlsma et al 2000) and to interpreting changes in genetic marker diversity with respect to effective population size and selection (Zhao and Charlesworth 2016). Mutational input of unconditionally deleterious variants, kept at low-equilibrium frequencies by purifying selection, is the primary underlying process. However, it is increasingly acknowledged that a significant component of the inbred load is due to alleles maintained at intermediate frequencies by various forms of balancing selection (Charlesworth and Charlesworth 1999; Charlesworth 2015). The idea that overdominant loci, even if relatively few in number, could make a disproportionate contribution to fitness variation, has been around for a long time (reviewed in Crow 1987), but lacks strong empirical support (Charlesworth and Willis 2009; Liberatore et al. 2013). An alternative and perhaps more widespread balancing mechanism is antagonistic pleiotropy, where a single allele is beneficial through one trait, but deleterious through another trait (Rose 1982; Charlesworth and Hughes 2000).
In species with separate sexes, sexually antagonistic pleiotropy (intralocus sexual conflict; Bonduriansky and Chenoweth 2009) is predicted to affect many components of fitness (Connallon and Clark 2014; Zajitschek and Connallon 2018). This was borne out by the first genome-wide study of sexually antagonistic polymorphisms, which found many deeply conserved clusters of sexually antagonistic single-nucleotide polymorphisms (SNPs) throughout the genome of Drosophila melanogaster, reflecting structural constraints to resolving functional conflicts (Ruzicka et al. 2019). Surprisingly, given some theoretical predictions (Rice 1984), and earlier studies in D. melanogaster (Innocenti and Morrow 2010) and humans (Lucotte et al. 2016), this study did not find a significant enrichment of these genes on the X chromosome, compared with autosomes. The major contribution from autosomes was attributed, in part, to sex-specific dominance effects, where the same allele confers a (partially) dominant benefit to one sex, but a (partially) recessive cost to the opposite sex, which broadens the conditions (sexually antagonistic cost/benefit ratio) for the maintenance of autosomal polymorphisms (Fry 2010).
Notwithstanding the likely importance of dominance modifiers for autosomal polymorphism (Spencer and Priest 2016), Rice’s (1984) single-locus model predicts that, in species with highly diverged heteromorphic sex chromosomes, intralocus sexual conflict at sex-linked loci may lead to asymmetries in the sex-specific inbred load. This is because a sexually antagonistic allele conferring a recessive cost to the homogametic sex (XX females or ZZ males) can be maintained through its beneficial effect to the heterogametic sex (XY males or ZW females), under the assumption that in the hemizygous state, alleles are always expressed (i.e., dominance restrictions do not apply). In this model (Rice 1984; Gibson et al. 2002), a relatively small pleiotropic benefit to the heterogametic sex can sustain low-equilibrium frequencies of strongly deleterious recessives, whose homozygous cost is rarely exposed in the homogametic sex. Conversely, the conditions for maintaining alleles that are deleterious to the heterogametic sex are much more restrictive and rely on large dominant benefits to the homogametic sex, coupled with relatively mild costs to the heterogametic sex. Thus, we might expect a concentration of larger-effect deleterious recessives affecting the homogametic sex on sex chromosomes. Sex chromosomes are also expected to be enriched for genes whose expression is biased towards the homogametic sex (Ellegren 2011; Huylmans et al. 2017).
No clear pattern emerges as to the sex or gamety that is more sensitive to inbreeding (reviewed in Ebel and Phillips 2016). For example, in D. melanogaster, inbreeding depression for fitness is greater for (XY) males than for (XX) females, due to male–male competition (Enders and Nunney 2010; Mallet and Chippindale 2011), whereas in Caenorhabditis remanei, (XX) females are much more sensitive to inbreeding than are (XO) males (Ebel and Phillips 2016). Lack of a consistent sex chromosome effect is also provided by the pedigreed population of Melospiza melodia, where inbreeding depression for lifetime-reproductive success was substantially greater for (ZW) females than (ZZ) males, predominantly through a reduction in the proportion of eggs hatching (Keller 1998). Other factors, such as the genetic architecture of sex-specific traits, the contribution of these traits to total fitness, the strength of sexual selection and inbreeding imposed by life-history and mating system and the mechanism of dosage compensation, will also influence sexual asymmetries in the inbred load (Janicke et al. 2013; Mank et al. 2014).
The physical distribution of the genetic load (sensu lato) within chromosomes, and the distribution of dominance effects, has major consequences for the pattern of linked molecular genetic diversity. In large populations, background selection against deleterious recessive alleles is expected to accelerate the rate of loss of linked neutral genetic variation, compared with the purely neutral case (Charlesworth et al. 1993). However, in small populations, linkage disequilibrium between two selected loci, each segregating deleterious recessives, and a linked (neutral) marker locus, leads to apparent heterozygote advantage at the marker locus (associative overdominance; Ohta 1971), even though the selected loci do not show true overdominance. Under certain conditions (low coefficient of dominance and weak selection), this type of association can retard the loss of genetic diversity due to genetic drift (Wang et al. 1999; Zhao and Charlesworth 2016). This effect may account for some cases, typically in laboratory populations, where lower-than-expected declines in heterozygosity have been observed (e.g. Schou et al. 2017).
A laboratory population of the Afro-tropical butterfly Bicyclus anynana (Satyridae) has been maintained in captivity since 1988, and offers the opportunity to relate temporal changes in molecular genetic diversity to genetic (recessive) load in a male-homogametic (ZZ) species. Previous studies have revealed that B. anynana carries a very high inbred load (>7 lethal equivalents per zygote), predominantly affecting male fertility and embryo viability (Saccheri et al. 1996; Saccheri et al. 2005). For comparison, the egg–adult inbred load in Drosophila spp. is of the order of 1–2 lethal equivalents (Dobzhansky et al. 1963; García et al. 1994). In B. anynana, sterility is strongly affected by the inbreeding status of males (inbreeding coefficient F ~ 0.25), whereas the degree of inbreeding of females appears to have essentially no effect on this trait (Saccheri et al. 2005). The qualitatively male-biased sterility effect and generally high inbred load, coupled with theoretical predictions of intralocus sexual conflict models (Rice 1984; Gibson et al. 2002), suggested the presence of one or more Z-linked loci segregating recessive alleles with large deleterious effects. We therefore set out to test the null hypothesis that homozygosity (in diploid males) or haplotype identity (in hemizygous females) of Z chromosome segments (denoted by ƶ) has no effect on fertility or viability. The W chromosome in this species is unusually small and degenerated (van’t Hof et al. 2008), implying few if any functional regions that are homologous with the Z. (For practical reasons at the time of the experiments, the study was restricted to the Z chromosome, effectively ignoring any influence from the 27 autosomes.) In addition to seeking genotype–phenotype associations in segregating sibships, we used archival DNA from the ancestral base population to link changes in Z-haplotype diversity over 70 generations in captivity to selection and drift. A linkage map of the Z chromosome was constructed and used to identify marker polymorphisms, which were then applied to segregating sibships and population samples.
Materials and methods
Overview of samples
This study combined analysis of experimental crosses (linkage-mapping family and backcrosses) and bottlenecked lines, with population samples taken at two time points (1993 and 2006), all derived from the same initial laboratory population (Fig. 1a).
a Timeline of the Bicyclus anynana Leiden/Liverpool lab population, showing the samples genotyped in this study, their relationship to each other and the order in which they were analysed (1–5). b Design of daughter–father backcross, highlighting the identify (by descent) of Z chromosome segments (denoted by ƶi, where i refers to the identity of the paternal Z in generation P). The position of the recombination breakpoints, indicated by change in colour, is arbitrary, as is which of the daughters to backcross (recombination does not occur in female Lepidoptera). ZF and ZM refer to whether the Z copy came from the father (F) or mother (M). This design is expected to generate F2 brothers that differ with respect to which portion of the Z is either homozygous (hom) or heterozygous (het). Autosomes (not shown) are expected to experience the same degree of inbreeding, regardless of sex (probability of identity by descent ~0.25, averaged across all offspring, assuming zero coancestry between P-generation parents). To assess the fertility of F2 males, they were mated to unrelated outbred females. The F2 generation is referred to as BF1 (backcross F1).
Linkage map and genetic markers
Our objective was to obtain 8–10 evenly spaced markers across the Z chromosome; however, inconsistent gene order between Bombyx mori and B. anynana made it necessary to map a larger number of loci (34) to achieve this. The majority were initially targeted using B. mori Z-chromosome scaffold orthologues (Wang et al. 2005). Ten Z-linked genes were already mapped in B. anynana (van’t Hof et al. 2008; Beldade et al. 2009). Prior to the release of the B. anynana genome assembly (Nowell et al. 2017), gene sequences were obtained using degenerate primers based on conserved regions (Table S1), and expressed sequence tags (Beldade et al. 2006). Segregating polymorphisms were most commonly found by screening the father of the mapping family with intron-spanning PCR followed by Sanger sequencing. For some markers, exonic polymorphisms were used, and apterous A was mapped using a BAC sequence polymorphism outside the gene (genbank AC239117). Genotyping methods consisted of PCR-RFLP, indels producing distinct gel bands, and Sanger sequencing (PCR primers in Table S1). DNA was extracted from the head and thorax of frozen adults, following a standard phenol–chloroform protocol (Saccheri and Bruford 1993). One in a hundred dilution of this DNA extract was used as PCR template. PCR reactions were performed in 12-µl volumes, containing 1X LongAmp buffer, 1.2 units of LongAmp polymerase (NEB), 0.4 µM of each primer and 0.5 µl of diluted DNA extract. A standard three-step 40-cycle amplification was used with an annealing temperature (Ta) of 57 °C for species-specific primers and Ta of 50 °C for degenerate primers. Most marker alleles consisted of multiple linked SNPs defining a ‘microhaplotype’.
The mapping family is the same used for a previously published AFLP-based linkage map (van’t Hof et al. 2008) with an additional 92 sibs included giving a total of 184 offspring. The large number of offspring and markers required a cost-effective mapping approach, allowing detailed mapping around recombination breakpoints combined with crude mapping of non-recombining regions. A preliminary linkage map was constructed using indels or PCR-RFLP with distinct allelic bands on agarose gels (‘cheap markers’). This revealed all recombination events in the mapping family. The next step was to roughly position the loci genotyped by Sanger sequencing (‘expensive markers’) relative to the preliminary map based on a subset of offspring. Subsequently, these loci were fine-tuned to their exact position by only sequencing individuals with recombination breakpoints near the ‘expensive marker’. Missing genotypes inside non-recombinant chromosome sections (undisrupted inherited paternal haplotype sections) can then be reliably inferred using their surrounding ‘cheap markers’. Errors caused by undetected double recombinants on the preliminary map are unexpected due to the restrictions of interference, i.e., the absence of double recombinations within small intervals (Muller 1916). Markers near chromosome ends were sequenced in all offspring because their genotypes cannot be inferred reliably. The map was produced with Joinmap 4.0 (Kyazma) using the Haldane mapping method. The procedures to account for the absence of female recombination in Lepidoptera only apply to autosomes, and were therefore not implemented for the Z-chromosome mapping.
Backcross and egg hatching
Female Lepidoptera are heterogametic (ZW), receiving the Z chromosome from their father only, whereas homogametic (ZZ) males inherit one Z copy from each parent. Furthermore, there is no crossing over in females. A daughter–father backcross (father Z1Z2; daughter either Z1W or Z2W) is therefore expected to produce a 1:1 ratio of Z1Z1 (homozygous): Z1Z2 (heterozygous) sons (for the Z1W daughter case) and a 1:1 ratio of Z1W: Z2W daughters. In reality, recombination breaks down the original Z1 and Z2 haplotypes, such that the above expectation holds, not for the whole chromosome, but for large chromosome segments, denoted by ƶ (Fig. 1b). For autosomal genes, the expected probability of identity by descent is 0.25 for both sexes. Fourteen daughter–father backcrosses were established from a laboratory stock population originally derived from Nkhata Bay, Malawi (our lab population had been in captivity for ~90 generations at the time of the experiments). Husbandry was standard for this species: larvae were reared on potted maize plants at 27 °C; adults were fed on mashed banana. Potential backcross fathers were held at 20 °C to retard senescence while their daughters developed into adults. Eggs were collected from mated daughters individually on cuttings of their wild host plant, Oplismenus compositus. Backcross larvae were reared to adults in sleeve enclosures and 11–45 sons per backcross mated to (relatively) unrelated outbred females (F1 hybrids of inter-line crosses). Egg batches from these females were used to estimate the frequency of sterile sons produced by each backcross (proportion of eggs that did not hatch or show visible signs of being fertilised, such as a head capsule). To keep sequencing costs down, only five of the backcross families were selected for genotyping, on the basis of having intermediate levels of sterility (maximising statistical power), with the exception of one family that was 100% sterile, and the presence of paternal polymorphisms at marker loci. In each of these, a sample of sons (mean n = 26) and daughters (n = 16, in one case 14) was genotyped. Different combinations of five or six Z-linked loci were used for different families because some fathers lacked polymorphisms in some regions.
Population samples
Archival DNA representing a random sample of the ancestral (base) population in 1993 (n = 112 males, 112 females), ~20 generations into captivity, was used to estimate the base population frequencies of Z microhaplotypes at eight marker loci (C2745, Kettin, Tpi, Catalase, C5197, Masc, Shaker, Ldh). The 1993 male sample was used to estimate the observed frequencies of homozygotes at these same loci. As a temporal comparison, a sample of 48 females from the parental (backcross) generation, separated in time from the archival sample by ~70 generations in captivity (assuming an average generation time of 10 weeks), was also genotyped for the full set of eight loci. Archival DNA was additionally available for six ‘bottleneck’ lines, each established from a single pair of randomly mated individuals from the same ancestral base population in 1993 (Saccheri et al. 1996, 1999), consisting of the founders and a small sample of the descendant F3. This material was used to assess genotype-frequency distortion due to inbreeding of the outbred base population after relatively few generations in captivity.
Analysis of homozygosity, haplotype diversity and linkage disequilibrium in the ancestral and descendant populations
The complexity and diversity of the marker sequences in the ancestral or base (Leiden-1993) population made it impractical to fully deconvolute heterozygous chromatographs into the constituent microhaplotypes; therefore, microhaplotype frequencies were estimated from the female sample only. It was, however, possible to identify microhaplotype homozygotes in the male sample (those producing clean chromatographs). For each marker locus, the observed number of homozygotes was compared with its Hardy–Weinberg expectation, \(n{\sum} {q_i^2}\), where qi is the frequency of each allele (microhaplotype) and n is the size of the male sample. Chromosomal haplotypes spanning the eight marker loci were determined for each female sample (1993 and 2006). Microhaplotype identification numbers were assigned in order of their frequency in the 1993 sample.
Haplotype diversity (Hd), equivalent to expected heterozygosity, at each marker locus and time point (1993 and 2006), was estimated from the female allele (microhaplotype) frequency spectra as \(\left( {1 - {\sum} {q_i^2} } \right)\frac{n}{{n - 1}}\), where n is the size of the female sample (Nei 1987). Standard error of Hd was calculated as the square root of the variance, as defined by Nei (1987) equation 8.12, replacing 2n by n for the haploid female sample. Allelic richness (the number of microhaplotypes, Hn) for each locus, as implemented in R package hierfstat (version 0.04, Goudet and Jombart 2015), was calculated to account for differing n in 1993 and 2006. A point estimate of the expected Hd for each locus in 2006 due only to genetic drift was calculated as haplotype diversity in 1993 multiplied by (1−(1/1.5Ne))t, where the effective population size (Ne) was assumed to be 250 for (t) 40 generations (in Leiden), followed by 100 for 30 generations (in Liverpool). These effective population sizes are approximate as population sizes were not accurately recorded throughout the period, but census numbers of adults (N) were generally kept at 400–500 in Leiden and 150–300 in Liverpool. We applied a Ne:N ratio of 0.6, previously estimated for equivalent caged populations of B. anynana (Brakefield et al. 2001).
Simulation, implemented in simuPOP (Peng and Kimmel 2005), was used to generate distributions of diversity and LD statistics under neutrality and in the presence of effective overdominance (high frequency and diversity of recessive lethals) in the C5197-Masc region. Population size was set to 450 and 200 for generations 1–40 and 41–70, respectively. Poisson variance in family size of 5, broadly consistent with empirical estimates (Brakefield et al. 2001), produces neutral behaviour expected for Ne of 250 for 40 generations, followed by Ne of 100 for 30 generations. The initial population was created from the observed 1993 haplotype frequencies. Chromosomal crossovers occurred in males only, at recombination rates between neighbouring pairs of loci as specified in the linkage map (Fig. 1a). Females were hemizygous and non-recombining. For simplicity, a high frequency of recessive lethals in the C5197-Masc region was specified by assigning a fitness of zero to all C5197 homozygotes. The simulation was run 1000 times.
The R package boot (version 1.3) was used to generate distributions of the observed Hd in 1993 and 2006, with 1000 replicates for each sample (Davison and Hinkley 1997; Canty and Ripley 2019). Distributions of the change in heterozygosity at each locus were calculated from pairs of values from the two (1993 and 2006) heterozygosity distributions. All calculations were performed in R (version 3.3.3, R Core Team 2017). The simulations were also used to estimate the joint probability of obtaining the observed, or greater, combination of Hn and Hd in the 2006 (Liverpool) sample.
Entropy-based linkage disequilibrium (eLD), an index of linkage disequilibrium for multiallelic loci (Okada 2018), was estimated among all pairs of loci from the ancestral and descendant female samples. To examine eLD in the simulated populations, we generated genotype files for 100 replicates of the forward simulation and calculated the mean and standard deviation of eLD across these populations. To visualise allelic associations among marker loci, alluvial diagrams displaying the structure and distribution of whole-chromosome haplotypes in the 1993 and 2006 populations were generated with Disentangler (Kumasaka et al. 2011). Missing data were coded as null haplotypes indicated by the label “00”.
Results
Linkage map
All 34 markers mapped together, with strong log of odd values (LOD > 10), to a single linkage group of 65 cM (Fig. 2a). The segregation pattern in the mapping family was consistent with female hemizygosity: daughters inherited either of the paternal alleles but not the maternal allele, whereas sons inherited the (single) maternal allele plus one of the paternal alleles. All of the mapped orthologues of B. mori Z genes are also on the B. anynana Z chromosome. However, the evolutionary conservation of synteny does not apply to the ordering of the genes, which is very different in the two species. The order in B. anynana also differs from that in Biston betularia (van’t Hof et al. 2013). The gene Z-MOF (BaC1211), previously flagged as inconsistent between B. mori and B. anynana (Beldade et al. 2009), was confirmed to be Z-linked in B. mori, which has an autosomal paralogue and a Z-orthologue (not included in an earlier gene annotation).
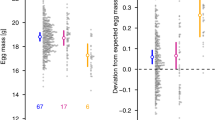
a Linkage map of the Bicyclus anynana Z chromosome, highlighting in blue all the marker loci used in this study (recombination distances in centimorgans). A subset of these was used to genotype population samples. b The distribution of microhaplotype frequencies at eight Z-linked loci in the base (Leiden) laboratory population of Bicyclus anynana in 1993 (black bars) and a descendant (Liverpool) population in 2006 (hatched bars). Haplotype numbers were assigned within each locus according to their frequency in the 1993 sample, and do not indicate direct correspondence between loci. Sample sizes are given in Table 5.
Genotype–phenotype associations in backcrosses
Viability
Five or six marker loci were scored in each of the five backcross families (2BF1, 8BF1, 9BF1, 11BF1 and 13BF1) (Table S3). There was a clear and significant deficit of homozygous adult males (sons) within and close to the interval containing marker loci C5197, Masc and 6PGD (Table 1). In three of the five backcross families (8BF1, 9BF1 and 11BF1), there were no homozygotes in the C5197 to 6PGD interval. In the other two families (2BF1 and 13BF1), there were only one or two homozygous sons at 6PGD and Masc, respectively. The homozygous fraction tends to increase with increasing distance from this region, though it rarely reaches the null expectation of equality with heterozygotes, likely due to only one generation of recombination (in fathers only) to decouple local heterozygosity from heterozygosity in the lethal region. In contrast, both paternal alleles occur at approximately their expected ratio of 1:1 in the daughters of all backcrosses, and there is no striking deviation within the C5197-6PDG interval. The one exception to this overall pattern is for Tpi in 13BF1, where all sons and all daughters carry the same paternal allele, which being the same as the maternal allele, results in no heterozygous sons. The identity of the lethal core haplotype in the C5197-6PGD interval is different in four out of five families (9BF1 and 13BF1 share the same lethal haplotype).
Male fertility
The egg batches produced by the F1 hybrid females mated to unrelated backcross sons showed a wide range of sterility and egg-hatching rates (Table 2), but no association with ƶ homozygosity of fathers at any of the marker loci (Table S4). The average proportion of sterile matings in the 2006 backcross families (0.62, s.d. 0.29) is somewhat higher than it was in similarly inbred parents (inbreeding coefficient = 0.25), produced by full-sib matings from the same population in 1999 and 2002 (0.43, s.d. 0.21, see Table 2 in Saccheri et al. 2005).
Homozygous lethal region in the ancestral population
There was a significant deficit of homozygotes at locus C5197 in the ancestral (Leiden-1993) population, but not at the Masc intronic marker. This pattern contrasts with the close-to-expected number of homozygotes at Tpi, Catalase, Shaker and Ldh (Table 3). Surprisingly, there was a highly significant excess of homozygotes at C2745. This was due to higher-than-expected homozygotes for most of the less common haplotypes, whereas homozygotes for the most common haplotype (H1) were close to expected (15 observed vs. 17 expected). In the partially inbred bottleneck lines (F3 derived from Leiden-1993), there was a similar pattern of homozygote deficit in the C5197-Masc-6PGD interval as observed in the backcross families, and close-to-expected representation of founder alleles in the female sample (Table 4). The allelic identities of the missing homozygotes are diverse. The apparent deficit of homozygotes at Tpi (line 1.1) is in the opposite direction to the observation in 13BF1, and possibly related to long-range linkage disequilibrium with the lethal region, generated through an interaction between drift and selection.
Changes in haplotype diversity and linkage disequilibrium
Comparison of the allele-(microhaplotype) frequency distributions in the base (1993) and descendant (2006) populations (Fig. 2b) reveals that the number of haplotypes declined substantially at every marker locus (45% average loss), but that Hd declined much less than the expected 26% at all marker loci, with the exception of Tpi (Table 5, Fig. 3a, Table S5). At C2745, Catalase and C5197, Hd in 2006 was marginally higher than in 1993; at Masc, Shaker and Ldh it was ~10% lower. The retention of Hd was achieved, despite the loss of many alleles, by balancing the frequencies of the remaining alleles. The big drop at Tpi results from the loss of six out of nine haplotypes, and a disproportionate increase in H1 (from 29 to 71%), which is the same allele that was exclusively inherited in 13BF1 (Table 1); the alternative paternal allele in this family was H7.
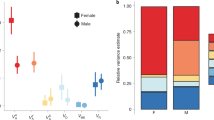
a Distributions of the proportional change between 1993 and 2006 (~70 generations) in haplotype diversity (Hd) at corresponding loci shown on a simplified chromosome b. Bootstrap distribution of the observed data (gold); simulated distribution with overdominance at C5197 (green); neutral expectation (dotted line). c Entropy-based linkage disequilibrium (eLD) among all pairs of loci. Values in the upper-half matrix are for the Leiden population in 1993, and in the lower-half matrix for the descendant Liverpool population in 2006. In parentheses, the mean and standard deviation of eLD in a sample of 100 simulations assuming overdominance at C5197. Colour gradient is proportional to the magnitude of eLD.
The pattern of linkage disequilibrium in the base population (Fig. 3c, upper-half matrix) reveals two blocks of loci with high eLD values: C2745-Tpi (18 cM) and C5197-Ldh (18.3 cM); the former block centred on C2745-Kettin, and the latter block made up of two smaller blocks (C5197-Masc, which are very close to each other, and Shaker-Ldh, which are 7.4 cM apart). Surprisingly, these LD blocks were essentially retained over 70 generations, with some modifications (Fig. 3c, lower-half matrix). The lower block expanded to include Catalase; the upper block weakened locally (C2745-Tpi) but strengthened globally (C2745-Ldh). Some of the LD patterns are visually apparent in the distribution of eight-locus haplotypes (Fig. S2).
Comparison with the simulated changes in Hn and Hd (Fig. S1) confirms that the observed diversity in Liverpool 2006 is significantly greater than predicted under neutrality at six of the eight loci (Table 5). Moreover, the simulation incorporating overdominance at C5197, can explain the excess diversity at C5197 and Masc, but not at C2745, Catalase, Shaker and Ldh, which remain significant outliers (Table 5, Fig. 3a). Similarly, LD in the simulated populations with C5197 overdominance, recovers a closely similar estimate to that observed between C5197 and Masc in 2006, but uniformly low LD everywhere else (Fig. 3c, in parentheses).
Discussion
This experiment was originally motivated to test the prediction that the very high male-specific sterility load in B. anynana (Saccheri et al. 2005) is Z-linked, and maintained through some form of intralocus sexual conflict (Rice 1984). Lack of any consistent association between homozygosity at a sample of loci spanning the entire chromosome and male sterility does not support this hypothesis. The implication, therefore, is that the male-specific sterility load must be due to autosomal loci. In contrast, the Z chromosome does harbour a large proportion of the male-specific viability load, in the form of recessive lethals, with implications for the retention of linked diversity.
The complete, or near-complete, absence of adult males homozygous within the region of the Z chromosome spanning C5197-6PGD in the Liverpool stock (2006) backcrosses indicates a high frequency of recessive lethals in this laboratory population. As this population had been in captivity for some 90 generations, this could be a consequence of genetic drift (the presence of at least four different lethal core haplotypes makes this unlikely). However, the deficit of homozygotes in the Leiden (1993) outbred base population, which will have been subject to minimal genetic drift since being established from the wild, and in the inbred (bottlenecked) lines derived from it, strongly suggests that the high frequency of recessive lethals in this region is a feature of the natural population. Our experiment only measured the adult stage, and therefore does not allow detection of when, during development, the homozygous lethal effect occurs. However, as larval and pupal mortality is very low under benign rearing conditions (Brakefield et al. 2001), sometime during embryo development, prior to hatching, is most likely.
The presence of a few apparent homozygotes within the core region, particularly in the 1993 stock, is likely to reflect the fact that intronic marker microhaplotypes are only loosely linked to extended haplotypes in population samples, so we predict that many of the homozygotes at C5197 and Masc are cryptic haplotype heterozygotes at the functionally lethal region. This interpretation is supported by the complete or near-complete absence of C5197-6PGD homozygotes in the backcross and bottleneck material, where parental haplotypes could be traced. In addition to C5197, Masc and 6PGD, the lethal region includes Z-lethal(3)neo18, C5904, SREBP and 1-Cys peroxiredoxin. The name Z-lethal(3)neo18 is derived from the D. melanogaster orthologue, of which B. anynana has an autosomal and a Z-linked copy. Despite its suggestive name, it remains unclear whether this gene is involved in the observed lethal effect since its function in D. melanogaster is not documented. Based on the B. anynana reference genome (Nowell et al. 2017), several other genes exist in the lethal interval, but none of them are obvious candidates for lethality either.
As demonstrated by our simulation model, heterozygote advantage could account for the observed retention of linked diversity and LD within the lethal-containing region (C5197 and Masc) through ca. 70 generations of genetic drift, despite the complete loss of one-third and one-half of haplotypes, respectively. We cannot at this stage rule out associative overdominance, as opposed to single-locus overdominance (Charlesworth and Willis 2009), which would require different but closely linked lethal-carrying loci in repulsion phase, rather than a single locus with many recessive lethal alleles. A model involving sexually antagonistic pleiotropy is less plausible, given that most, possibly all, alleles appear to be unconditionally deleterious to males, and we did not observe any obvious fitness benefits to females.
The retention of haplotype diversity in other regions of the Z chromosome (C2745, Shaker, Ldh and to lesser extent Kettin and Catalase) is intriguing. There were no obvious deviations from the expected segregation patterns in the backcross families and bottleneck lines, and recombination distances to the male-lethal region are large. Moreover, the family used to create the linkage map, and the backcross families do not suggest unusual recombination or the presence of inversion polymorphisms. However, the very similar pattern in LD between the 1993 and 2006 populations across the lower third of the chromosome (C5197-Ldh) does imply the interaction of reduced recombination and/or epistatic selection (Phillips 2008).
In the specific case of C2745, the excess haplotype diversity is coupled with a significant excess of homozygotes in the ancestral male sample. Assortative mating, with respect to this or nearby loci, could conceivably produce both of these patterns. The restrictive conditions required for stable polymorphism under a one-trait, two-locus model (Hedrick 2017), suggest a more complex architecture, perhaps involving sexual antagonism (Arnqvist 2011) and linkage between (male) attractiveness and (female) preference (Iyengar et al. 2002). In contrast to the C5197-Ldh region, LD in the upper third of the chromosome declined, albeit more slowly than expected. This may relate to the artificially high population-density environment of the caged population, and the associated relaxation of the conditions promoting mate choice behaviour in nature.
One of the marker loci (Tpi) stands out as having lost more haplotype diversity than expected. The highly skewed pattern of inheritance, in both sexes, of the paternal alleles in one of the backcross families, together with the more than doubling in frequency of this specific allele between 1993 and 2006, is suggestive of meiotic drive (Lindholm et al. 2016). The loss of autosomal suppressor alleles, originally present in the ancestral population, due to genetic drift, may explain its spread in captivity. This selective sweep is likely to have contributed to the 1993–2006 increase in LD between Tpi and the neighbouring Catalase, whose most common haplotype (H6) is fully associated with the putative driving haplotype (Tpi-H1, Fig. S2). Interestingly, there is also a suggestion that this haplotype causes male sterility in the homozygous state (all the sons in family 13BF1 were both homozygous for Tpi and sterile).
Our finding of significant retention of heterozygosity on the Z contrasts the near-neutral behaviour over 20 generations of the X in D. melanogaster lines, attributed to low levels of X-linked recessive load (Schou et al. 2017). Greater efficacy of purifying selection is expected to apply to sex-linked genes whose expression is biased towards the heterogametic sex (XY males and ZW females). Conversely, recessive or partially recessive deleterious mutations at sex-linked genes whose expression in biased towards the homogametic sex should experience less-efficient selection. The contrasting pattern in non-synonymous to synonymous nucleotide diversity (πn/πs) expected from this evolutionary dynamic has been confirmed between categories of sex-biased Z-linked genes in two butterfly species (Rousselle et al. 2016). In addition to gamety, differences in sex chromosome-linked load likely reflect the outcome of many other factors, including species demography, life history, phylogeny and intra-/interspecific coevolution.
This exploratory study has raised more questions than it has answered. Future work should be directed at resolving the identity of the male lethals in the C5197-6PGD region, as well as the underlying causes of the joint retention of heterozygosity and LD in the lower third of the Z chromosome, and the homozygote excess at the top end, around C2745. Further experiments are required to confirm the behaviour and identity of the putative meiotic driver in the vicinity of Tpi. Finally, the search for the loci causing inbred male sterility needs to be extended to the autosomes. The overall impression is of a complex set of balancing selection mechanisms, whose net effect is the maintenance of an elevated genetic load resistant to purifying selection, making this species particularly susceptible to inbreeding depression.
Data availability
All the data are provided either as supplementary tables or GenBank accession numbers (listed in Table S1).
References
Arnqvist G (2011) Assortative mating by fitness and sexually antagonistic genetic variation. Evolution 65:2111–2116
Beldade P, Rudd S, Gruber JD, Long AD (2006) A wing expressed sequence tag resource for Bicyclus anynana butterflies, an evo-devo model. BMC Genomics 7:130
Beldade P, Saenko SV, Pul N, Long AD (2009) A gene-based linkage map for Bicyclus anynana butterflies allows for a comprehensive analysis of synteny with the lepidopteran reference genome. PLoS Genet 5:e1000366
Bijlsma R, Bundgaard J, Boerema AC (2000) Does inbreeding affect the extinction risk of small populations? Predictions from. Drosoph J Evol Biol 13:502–514
Bonduriansky R, Chenoweth SF (2009) Intralocus sexual conflict. Trends Ecol Evol 24:280–288
Brakefield PM, El Filali E, Van der Laan R, Breuker CJ, Saccheri IJ, Zwaan B (2001) Effective population size, reproductive success and sperm precedence in the butterfly, Bicyclus anynana, in captivity. J Evol Biol 14:148–156
Canty A, Ripley B (2019) boot: bootstrap R (S-Plus) functions. R package version 1.3-35
Charlesworth B (2015) Causes of natural variation in fitness: evidence from studies of Drosophila populations. Proc Natl Acad Sci USA 112:1662–1669
Charlesworth B, Charlesworth D (1999) The genetic basis of inbreeding depression. Genet Res 74:329–340
Charlesworth B, Hughes KA (2000) The maintenance of genetic variation in life-history traits. In: Singh RS, Krimbas CB (eds) Evolutionary genetics: from molecules to morphology, vol 1. Cambridge University Press, UK, p 369–392
Charlesworth B, Morgan MT, Charlesworth D (1993) The effect of deleterious mutations on neutral molecular variation. Genetics 134:1289–1303
Charlesworth D, Willis JH (2009) The genetics of inbreeding depression. Nat Rev Genet 10:783–796
Connallon T, Clark AG (2014) Evolutionary inevitability of sexual antagonism. Proc R Soc B 281:20132123
Crow JF (1987) Muller, Dobzhansky, and overdominance. J Hist Biol 20:351–380
Davison AC, Hinkley DV (1997) Bootstrap methods and their applications. Cambridge University Press, Cambridge
Dobzhansky T, Spassky B, Tidwell T (1963) Genetics of natural populations. XXXII. Inbreeding and the mutational and balanced genetic loads in natural populations of drosophila pseudoobscura. Genetics 48:361–373
Ebel ER, Phillips PC (2016) Intrinsic differences between males and females determine sex-specific consequences of inbreeding. BMC Evol Biol 16:36
Ellegren H (2011) Emergence of male-biased genes on the chicken Z-chromosome: sex-chromosome contrasts between male and female heterogametic systems. Genome Res 21:2082–2086
Enders LS, Nunney L (2010) Sex-specific effects of inbreeding in wild-caught Drosophila melanogaster under benign and stressful conditions. J Evol Biol 23:2309–2323
Fry JD (2010) The genomic location of sexually antagonistic variation: some cautionary comments. Evolution 64:1510–1516
García N, López-Fanjul C, García-Dorado A (1994) The genetics of viability in Drosophila melanogaster: effects of inbreeding and artificial selection. Evolution 48:1277–1285
Gibson JR, Chippindale AK, Rice WR (2002) The X chromosome is a hot spot for sexually antagonistic fitness variation. Proc R Soc B 269:499–505
Goudet J, Jombart T (2015) hierfstat: estimation and tests of hierarchical F-statistics. R package version 0.04-22
Hedrick PW (2017) Assortative mating and linkage disequilibrium. G3 7:55–62
Huylmans AK, Macon A, Vicoso B (2017) Global dosage compensation is ubiquitous in Lepidoptera, but counteracted by the masculinization of the Z chromosome. Mol Biol Evol 34:2637–2649
Innocenti P, Morrow EH (2010) The sexually antagonistic genes of Drosophila melanogaster. PLoS Biol 8:e1000335
Iyengar VK, Reeve HK, Eisner T (2002) Paternal inheritance of a female moth’s mating preference. Nature 419:830–832
Janicke T, Vellnow N, Sarda V, David P (2013) Sex-specific inbreeding depression depends on the strength of male–male competition. Evolution 67:2861–2875
Keller LF (1998) Inbreeding and its fitness effects in an insular population of song sparrows (Melospiza melodia). Evolution 52:240–250
Kumasaka N, Okada Y, Takahashi A, Kubo M, Nakamura Y, Kamatani N (2011) In: 12th international Congress of human genetics, Montreal
Liberatore KL, Jiang K, Zamir D, Lippman ZB (2013) Heterosis: the case for single-gene overdominance. In: Chen ZJ, Birchler JA (eds) Polyploid and hybrid genomics. John Wiley & Sons, Inc, Ames, Iowa, USA, p 137–152
Lindholm AK, Dyer KA, Firman RC, Fishman L, Forstmeier W, Holman L et al. (2016) The ecology and evolutionary dynamics of meiotic drive. Trends Ecol Evol 31:315–326
Lucotte EA, Laurent R, Heyer E, Ségurel L, Toupance B (2016) Detection of allelic frequency differences between the sexes in humans: a signature of sexually antagonistic selection. Genome Biol Evol 8:1489–1500
Mallet MA, Chippindale AK (2011) Inbreeding reveals stronger net selection on Drosophila melanogaster males: implications for mutation load and the fitness of sexual females. Heredity 106:994–1002
Mank JE, Hosken DJ, Wedell N (2014) Conflict on the sex chromosomes: cause, effect, and complexity. Cold Spring Harbor Persp Biol 6:a017715
Muller HJ (1916) The mechanism of crossing-over. II. IV. The manner of occurrence of crossing-over. Am Nat 50:284–305
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, NY
Nowell RW, Elsworth B, Oostra V, Zwaan BJ, Wheat CW, Saastamoinen M et al. (2017) A high-coverage draft genome of the mycalesine butterfly Bicyclus anynana. Gigascience 6:1–7
Ohta T (1971) Associative overdominance caused by linked detrimental mutations. Genet Res 18:277–286
Okada Y (2018) eLD: entropy-based linkage disequilibrium index between multiallelic sites. Hum Genome Var 5:29
Peng B, Kimmel M (2005) simuPOP: a forward-time population genetics simulation environment. Bioinformatics 21:3686–3687
Phillips PC (2008) Epistasis–the essential role of gene interactions in the structure and evolution of genetic systems. Nat Rev Genet 9:855–867
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Rice WR (1984) Sex chromosomes and the evolution of sexual dimorphism. Evolution 38:735–742
Rose MR (1982) Antagonistic pleiotropy, dominance, and genetic variation. Heredity 48:63–78
Rousselle M, Faivre N, Ballenghien M, Galtier N, Nabholz B (2016) Hemizygosity enhances purifying selection: lack of fast-Z evolution in two satyrine butterflies. Genome Biol Evol 8:3108–3119
Ruzicka F, Hill MS, Pennell TM, Flis I, Ingleby FC, Mott R et al. (2019) Genome-wide sexually antagonistic variants reveal long-standing constraints on sexual dimorphism in fruit flies. PLoS Biol 17:e3000244
Saccheri I, Kuussaari M, Kankare M, Vikman P, Fortelius W, Hanski I (1998) Inbreeding and extinction in a butterfly metapopulation. Nature 392:491–494
Saccheri IJ, Brakefield PM, Nichols RA (1996) Severe inbreeding depression and rapid fitness rebound in the butterfly Bicyclus anynana (Satyridae). Evolution 50:2000–2013
Saccheri IJ, Bruford MW (1993) DNA fingerprinting in a butterfly, Bicyclus anynana (Satyridae). J Hered 84:195–200
Saccheri IJ, Lloyd H, Helyar S, Brakefield PM (2005) Inbreeding uncovers fundamental differences in the genetic load affecting male and female fertility in a butterfly. Proc R Soc B 272:39–46
Saccheri IJ, Wilson IJ, Nichols RA, Bruford MW, Brakefield PM (1999) Inbreeding of bottlenecked butterfly populations: estimation using the likelihood of changes in marker allele frequencies. Genetics 151:1053–1063
Schou MF, Loeschcke V, Bechsgaard J, Schlötterer C, Kristensen TN (2017) Unexpected high genetic diversity in small populations suggests maintenance by associative overdominance. Mol Ecol 26:6510–6523
Spencer HG, Priest NK (2016) The evolution of sex-specific dominance in response to sexually antagonistic selection. Am Nat 187:658–666
van’t Hof AE, Marec F, Saccheri IJ, Brakefield PM, Zwaan BJ (2008) Cytogenetic characterization and AFLP-based genetic linkage mapping for the butterfly Bicyclus anynana, covering all 28 karyotyped chromosomes. PLoS ONE 3:e3882
van’t Hof AE, Nguyen P, Dalíková M, Edmonds N, Marec F, Saccheri IJ (2013) Linkage map of the peppered moth, Biston betularia (Lepidoptera, Geometridae): a model of industrial melanism. Heredity 110:283–293
Wang J, Hill WG, Charlesworth D, Charlesworth B (1999) Dynamics of inbreeding depression due to deleterious mutations in small populations: mutation parameters and inbreeding rate. Genet Res 74:165–178
Wang J, Xia Q, He X, Dai M, Ruan J, Chen J et al. (2005) SilkDB: a knowledgebase for silkworm biology and genomics. Nucleic Acids Res 33:D399–D402
Zajitschek F, Connallon T (2018) Antagonistic pleiotropy in species with separate sexes, and the maintenance of genetic variation in life-history traits and fitness. Evolution 72:1306–1316
Zhao L, Charlesworth B (2016) Resolving the conflict between associative overdominance and background selection. Genetics 203:1315–1334
Acknowledgements
This work was made possible by the enthusiasm of Paul Brakefield, establishing the original lab colony of Bicyclus anynana in 1988 and promoting its use as a model for evolutionary biology. Dave Pye assisted with DNA extractions and sequencing. Comments by three reviewers led to improvements of a previous version of the paper. This work was principally supported by NERC grants (NER/M/S/2001/00040 and NE/N015711/1) and Leverhulme Research Fellowship (RF-2016-654) to IJS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate Editor: Ben Evans
Supplementary information
Rights and permissions
About this article
Cite this article
Saccheri, I.J., Whiteford, S., Yung, C.J. et al. Recessive Z-linked lethals and the retention of haplotype diversity in a captive butterfly population. Heredity 125, 28–39 (2020). https://doi.org/10.1038/s41437-020-0316-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-020-0316-x