Abstract
Population genomics offers innovative approaches to test hypotheses related to the source and timing of introduction of invasive species. These approaches are particularly appropriate to study colonization of island ecosystems. The brown rat is a cold-hardy global invasive that has reached most of the world’s island ecosystems, including even highly isolated archipelagoes such as the Faroe Islands in the North Atlantic Ocean. Historic records tell of rats rafting to the southern island of Suðuroy in 1768 following a shipwreck off the coast of Scotland, then expanding across the archipelago. We investigated the demographic history of brown rats in the Faroes using 50,174 SNPs. We inferred three independent introductions of rats, including to Suðuroy, the islands of Borðoy and Viðoy, and onto Streymoy from which they expanded to Eysturoy and Vágar. All Faroese populations showed signs of strong bottlenecks and declining effective population size. We inferred that these founder events removed low frequency alleles, the exact data needed to estimate recent demographic histories. Therefore, we were unable to accurately estimate the timing of each invasion. The difficulties with demographic inference may be applicable to other invasive species, particularly those with extreme and recent bottlenecks. We identified three invasions of brown rats to the Faroe Islands that resulted in highly differentiated populations that will be useful for future studies of life history variation and genomic adaptation.
Similar content being viewed by others
Introduction
Invasive alien species are one of the major causes of species extinctions and threaten ecosystems worldwide (Doherty et al. 2016; Russell and Blackburn 2017). Islands are especially vulnerable to invasive species because many insular taxa evolved in relative isolation, and the effects of invasions interact with other threats such as climate change (Russell et al. 2017). Rats (Rattus norvegicus Berkenhout, 1769 and R. rattus L., 1758) and house mice (Mus musculus L., 1758) are the most widespread and destructive invasive vertebrates around the world (Genovesi et al. 2012; Pimentel et al. 2000), and have particularly harmed island bird species (Doherty et al. 2016). Despite the destructiveness of invasive rodents, only recently has the history of rodent invasions been examined using population genetic approaches (Johnson and Munshi-South 2017).
Island invasions by rodents present several questions of broad interest to ecology and evolution that largely remain unanswered. For one, are islands typically invaded a single time followed by population expansion, or are invasive populations established through multiple introductions? Invasive species such as rats often exhibit explosive population growth, and thus single invasions could populate entire islands. Once established, these founding populations may exhibit a priority effect whereby they exclude subsequent invaders entirely, or swamp out any genetic admixture from other source populations or lineages (Combs et al. 2018; Puckett et al. 2016). Older invasions may have been poorly documented, and thus we are forced to rely on incomplete or apocryphal historical narratives about the origins of invasive populations. Even without a clear historical record or zooarchaeological remains, invasion histories may be recoverable using population genetic analysis of contemporary populations (Searle et al. 2009). Success at such reconstruction requires adequate sampling of both the invasive and source populations, or at least lineages related to the original source. The genetic markers used for reconstruction must also have sufficient variation to identify relationships between invasive and the true source population(s). Recently, the advent of genome-wide SNPs (single nucleotide polymorphisms), whole exome, and/or whole genome resequencing combined with coalescent-based modeling approaches have provided unprecedented power for the study of invasive populations (Puckett and Munshi-South 2019; Puckett et al. 2016). However, the severe founder effects and/or population bottlenecks that often characterize island invasions may complicate our ability to reconstruct historical invasions even with whole-genome data.
Early work on invasive rodents relied heavily on mitochondrial phylogeography and is most developed for house mice. This invader accompanied Western peoples for centuries and serves as a proxy for the history of human movements (Jones et al. 2013). Phylogeographic analyses of black rats (R. rattus) have also revealed a long history of transport by humans and multiple evolutionary origins of commensalism (Aplin et al. 2011; Lack et al. 2012). Studies of brown rats (R. norvegicus) have identified the greatest mtDNA haplotype diversity in the putative center of origin in east Asia (Song et al. 2014), and multiple invasions of North America (Lack et al. 2013). While these studies of mtDNA diversity identified broad patterns of relatedness between invasive populations, they were limited in their ability to infer complex histories of recent invasion from multiple source populations. The first studies to use genome-wide data found that R. norvegicus moved out of China into southeast Asia from which it became a truly global menace a few hundred years ago when brown rats were moved all over the world by European colonial powers (Puckett et al. 2016). Molecular data used in coalescent models estimated European rats diverged approximately 1227 AD (90% Highest Density Probability (HDP): 589-1781 AD) (Puckett and Munshi-South 2019); this time frame was congruent with brown rat archeological remains found in Tarquinia, Italy where the site was dated from 1390-1436 (Clark et al. 1989).
Population genomic approaches are needed to understand complex histories of invasion and reinvasion, particularly on islands that may have received founders from multiple lineages (Robins et al. 2016). In this study, we analyzed thousands of genome-wide SNPs from brown rats sampled throughout the world to understand the routes of invasion onto the Faroe Islands. We also examined the historical demography and population structure of brown rats within the Faroes. The Faroe Islands are a highly isolated archipelago in the north Atlantic (Fig. 1). Topographically the western and northern shorelines of the islands are characterized by precipitous cliffs, while inland the terrain consists of valleys, mountain ridges, and upland hills. The climate is oceanic with mean temperatures of 4 and 11 °C during the coldest (January - February) and warmest (July) months, respectively. Mean annual precipitation varies locally (823–3261 mm) and fog or strong winds frequently prevail (Cappelen and Laursen 1998). Human settlements consist of small densely built-up areas with adjoining cultivated land (mostly for haymaking and potatoes), which is usually separated from the surrounding outfield by stonewalls or other kinds of fences. There is no natural tree vegetation on the islands, but small plantations (mostly coniferous) occur in some of the settlements (Rutherford 1982). Given the range of the brown rat extends from the sub-tropics to the northern Holarctic, the climate of the Faroe Islands is within the range for the species. Seabirds and marine animals dominate the native fauna with no native land mammals on the islands following the Pleistocene; however, house mouse, brown rat, and mountain hare (Lepus timidus L., 1758) were introduced by humans and are now found on many islands. Additionally, black rats have also lived and reproduced on the islands in the past (Svabo 1783). Although black rats are considered extirpated, they have been occasionally and unintentionally reintroduced mainly from Russian fishing boats, but they do not breed and thus die off (Bloch and Fuglø 1999). Faroe house mice currently occupy six of the islands and exhibit strikingly different morphology than source populations on the European mainland (Berry et al. 1978; Davis 1983). Genetic analyses established that they mainly arrived from south-western Norway with the Vikings around 1,200 years ago (~800 AD), although mice on Sanðoy were introduced from Great Britain (Jones et al. 2011). Mountain hares were intentionally introduced from Norway in 1855 for hunting and now occur on 15 of the islands (Bloch and Fuglø 1999).
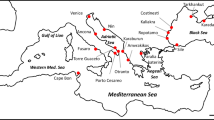
Map of the Faroe Islands with island names in capitals and cities with documented brown rat invasion histories denoted with asterisks (*). Islands with continued rat presence are shown in grey. Cities that were first colonized by rats for the three independent introductions are denoted with a circle around the asterisk; dashed arrows indicate the expected dispersal patterns. Beside each city name is the year of the earliest record of brown rats and associated reference including: (1) Svabo (1783), (2) Kyrjarheyg (2014), and (3) Skarði (1956). Black dots denote the 23 sampling locations
Brown rats were first detected in the Faroes in 1768. It is thought that the first individuals arrived on the southernmost island, Suðuroy, via the wreck of a Norwegian ship that stranded on the Scottish Isle of Lewis. The wreckage drifted northwards and landed in the village of Hvalba, Suðuroy (Fig. 1) (Jensen and Magnussen 2015). Within 10 years, brown rats expanded their range to Streymoy, Eysturoy, and Vágar northwest of the original invasion site either through movement from the first invaded island, additional introductions, or a combination of the two (Svabo 1783). According to the historic record, rats were introduced to the northern islands after Norwegians built and operated a whaling station in the village Norðdepil, Borðoy from 1898 to 1920 (Kyrjarheyg 2014; Skarði 1956). From there, the brown rat spread to the neighbouring islands Viðoy and Kunoy (Fig. 1). Today the brown rat is found on seven of the 18 Faroese islands (Fig. 1), and is common in and around human habitations as well as in the wild.
The objective of this study was to test if the demographic history inferred through population genomics was consistent with the historic record of the introduction of brown rats to the Faroe Islands. Specifically, we wanted to test three hypotheses: first, that rats were introduced to Suðuroy from Norway; second, that rats from Suðuroy expanded to the northwestern islands; and third, that a second independent introduction from Norway resulted in the invasion of Borðoy and Viðoy. Thus, we genotyped rats across the Faroe Islands and combined the data from a previous study of 326 individuals sampled from 41 cities around the globe (Puckett et al. 2018; Puckett et al. 2016). We used genome-wide SNP genotypes to examine population structure, identify source population(s) to test our hypotheses, and model the demographic history, particularly estimating changes in effective population size (Ne) of rats across the Faroe Islands. Our study provides a foundation for future work to understand life history variation and genomic adaptations to harsh environments, particularly in a comparative framework across climatic and resource gradients.
Materials and methods
We collected rat tissue samples in the Faroe Islands (n = 85), western Russia (n = 10), and Sweden (n = 34) to add to our existing global collection of samples from China, southeast Asia, the Aleutian archipelago, Europe, North America, and South America (Puckett et al. 2018; Puckett et al. 2016). The additional sampling in Russia and Sweden increased the spatial resolution in eastern and northern Europe, thus allowing additional power to test the source of introduction to the Faroe Islands (see below). The Faroese rats were collected between 2011 and 2017 from 24 locations (Fig. 1) where one to 14 specimens were sampled in each location (Jensen and Magnussen 2015). Russian rats were collected from five locations (four in Moscow and one near Tver), where ears were taken postmortem. We used lethal snap-traps outside of areas with known rat activity after obtaining permission from land owners.
DNA Extraction, RAD sequencing, and SNP calling
We extracted DNA following the manufacturer’s protocols using Qiagen DNeasy kits (Valencia, CA). We prepared double digest restriction-site associated DNA sequencing (ddRAD-Seq) (Peterson et al. 2012) libraries with 500–1000 ng of genomic DNA from each sample. Briefly, samples were digested with SphI and MluCI before ligation of unique barcoded adapters. We pooled 48 barcoded samples each in three libraries at equimolar concentrations. We then selected fragments from 340 to 412 bp (target = 376 bp) using a Pippin Prep (Sage Science, Beverly, MA). The size-selected pools were PCR-amplified for 12 cycles using Phusion PCR reagents (New England Biolabs, Ipswich, MA) and primers that added an Illumina multiplexing read index. Final libraries were checked for concentration and fragment size on a BioAnalyzer (Agilent Technologies, Santa Clara, CA), then sequenced (2 × 125 bp paired-end) at the New York Genome Center in two lanes of an Illumina HiSeq 2500.
We demultiplexed the raw reads using the process_radtags script in STACKS v2.3 (Catchen et al. 2013), then aligned reads for each individual to the Rattus norvegicus reference genome (Rnor_6.0) (Gibbs et al. 2004) using Bowtie v2.2.6 (Langmead and Salzberg 2012) with default parameters. We exported four datasets. The first dataset included 32,127 previously identified SNPs with a global sample of brown rats (n = 424; henceforth the “global” dataset) (Puckett et al. 2016). We used a position list in SAMTOOLS v1.3 (Li et al. 2009) to extract genotypes at each position, then BCFTOOLS v1.3 to call SNPs. For the second dataset (i.e. “European” dataset), we reprocessed the data from Puckett et al. (2016) with samples from Europe and the data generated for this paper from the Faroe Islands, Sweden, and Russia to better capture genetic diversity and differentiation within the region. We used the ref_map.pl script provided within STACKS to build a European specific catalog with gstacks, then filtered the data to have a minor allele frequency (MAF) of 0.05, with a single SNP per RAD-tag (--write_single_snp flag). Preliminary analyses with this dataset showed that few private alleles from the Faroe Islands were included; thus, we output another dataset ascertained only on Faroese samples, again with a MAF of 0.05 and a single SNP per RAD-tag. The datasets overlapped at 29,883 loci (Fig. S1); we removed 1,803 loci to maintain one SNP per RAD-tag. We further filtered European and Faroese samples in this dataset using PLINK v1.9 (Purcell et al. 2007) to remove loci with greater than 30% missing data, as well as sites on the Y-chromosome or mitochondrial genome. We removed related individuals within sampling locations by assessing relatedness in KING v1.4 (Manichaikul et al. 2010). For each pair of individuals with relatedness estimators greater than 0.2, one individual was removed from the analysis (n = 13). Thus, our final European dataset included 50,174 bi-allelic SNPs genotyped in 208 samples, of which 73 were from Faroe Islands (Table S1). We estimated observed heterozygosity (HO) in Arlequin v3.5.2.2 (Excoffier and Lischer 2010) and private alleles using the hierfstat package for R (Goudet 2005) for each sampling location and broader geographic grouping. To make the third dataset (henceforth “TreeMix”), we used the whitelist function in STACKS with these 50k SNPs and output all genotypes across European, Faroese, southeast Asian (n = 25), and black rat (n = 33) samples for use in TREEMIX (see below).
The fourth dataset (henceforth “SFS”) included one-dimensional site frequency spectra (SFS) for each of eight evolutionary clusters. We estimated the SFS using ANGSD v0.9.2.0 (Korneliussen et al. 2014). We limited the estimation of the SFS by first running ANGSD with the European and Faroese samples where bases had a minimum mapping quality (-minmapq) of 30, minimum quality score (-minQ) of 20, and greater than 80% of samples genotyped at a locus using the genotype likelihood function from SAMtools (-GL 1). This procedure resulted in 15,379,536 basepairs. We then limited estimation of the folded SFS (-fold 1) to these sites for each population using the realSFS program within ANGSD.
Population genomic analyses
We performed three principal component analyses (PCA) in PLINK (--pca) to place samples from the Faroe Islands within the context of global brown rat diversity. The first PCA utilized the global dataset with R. rattus samples included, and confirmed that all samples genotyped for this study were R. norvegicus. We ran a second PCA of the global dataset with R. rattus excluded. The third PCA used the European dataset with 50k SNPs and European samples only.
Using the European dataset, we estimated the number of evolutionary clusters by running 20 independent iterations of ADMIXTURE v1.23 (Alexander et al. 2009) at each cluster (K) from 1 to 20. We used the cross-validation error to determine the best supported admixture model. We also estimated fine-scale spatial structure using FINESTRUCTURE v4.0.1 (Lawson et al. 2012) on data from the 20 autosomes (49,241 SNPs) of the European dataset. SNPs on each chromosome were phased and imputed using fastPHASE v1.2 (Scheet and Stephens 2006). In FINESTRUCTURE, we used default settings except for the following parameters: 25% of the data were used for initial EM estimation; 750,000 iterations of the MCMC were run (375,000 of which were burnin) with 1000 samples retained, 20,000 tree comparisons, and 500,000 steps of the tree maximization were run. We viewed MCMC trace files to confirm stability of all parameters.
Estimation of effective population size
Given that the Faroe Islands were likely founded by a limited number of individuals, we tested for a bottleneck. Based upon the clustering results, we separated the data into nine clusters: 1- Faroe Islands- Suðuroy (n = 6); 2- Faroe Islands- Borðoy and Viðoy (n = 7); 3- Faroe Islands- Eysturoy (n = 18); 4- Faroe Islands- Streymoy; 5- Faroe Islands- Vágar (n = 8); 6- Great Britain (n = 12); 7- France and Spain (n = 29); 8- central Europe (Netherlands and Germany; n = 21); and 9- northern Sweden (villages of Örsundsbro, Uppsala, and Lövsta, Sweden; n = 15). We calculated the extent of linkage disequilibrium (LD) as a function of inter-SNP distance for both continental and Faroese populations using the European dataset. Specifically, we used PLINK to calculate r2 in windows of 1000 kb for variants with a minor allele frequency (MAF) equal to or greater than 0.15 within populations (PLINK commands --r2 --ld-window 99999 --ld-window-r2 0 --maf 0.15) similar to vonHoldt et al. (2011). We binned the inter-SNP distances into the following classes: 2.5, 5, 7.5, 10, 15, 20, 30, 40, 60, 80, 115, 150, 213, 275, 388, 500, 738, 975, 1000 kb in program R (R Core Team 2013) and plotted the mean r2 of each class using ggplot2 (Wickham 2016). We also calculated MAF for each locus within each of the eight clusters using PLINK and plotted the density using ggplot2.
To quantify the magnitude of the bottleneck, we estimated Ne through time using Stairway Plot v2 (Liu and Fu 2015). Stairway Plot models multiple epochs similar to skyline plots (Drummond et al. 2005) given the input of a one-dimensional SFS. Using the SFS dataset, we ran four iterations of Stairway Plot each with a different number of random break points that corresponded to the sample size of the analysis as suggested in the manual, and accepted the best model of the four based on the composite likelihood. To convert the estimated θ to Ne, we used an estimated rat mutation rate of 1.97 × 10−9 (Deinum et al. 2015) and a generation time of four months (i.e. three generations per year).
Population Tree Analyses
To estimate the population tree topology, we ran TREEMIX v1.12 (Pickrell and Pritchard 2012) with the TreeMix dataset. We included R. rattus as the outgroup, and southeast Asian brown rat populations due to shared ancestry with the sources that expanded into Europe (Puckett and Munshi-South 2019). We ran this analysis at the spatial scale of cities (n = 29), including locations with four or more samples, except for Norðdepil, Borðoy and Sandavágur, Vágar which each had three samples. We ran TREEMIX without migration edges, then added up to six edges allowing the tree topology to be estimated independently each run (versus fixing it to the tree estimated without migration edges); we used blocks of 500 SNPs and the sample size correction. We assessed models by considering both the proportion of variance that the tree explained and the residuals.
Results
Population structure
Our PCA of global brown rat samples showed differentiation of 55 of the 67 Faroe Islands samples along the first PC axis (Fig. 2a), indicating that the Faroe Islands rats are highly differentiated. The remaining 12 samples clustered near Western Europe. The European dataset included 50,157 SNPs where 13,628 overlapped with the global dataset (Fig. S1). In the PCA using only European samples, the first PC axis described 26.1% of the variation and distinguished the highly differentiated Faroe Islands samples (Fig. 2b) identified in the global PCA (Fig. 2a). The second PC axis described 8.1% of the variation and differentiated continental European samples from the remaining Faroe Islands individuals. This axis also suggested clinal geographic variation from the Iberian Peninsula to northern Fennoscandia across continental Europe.
Principal components analyses of Rattus norvegicus variation using either a 32k nuclear SNPs with global and Faroe Islands samples (green), or b 50k SNPs comparing only continental European and the Faroe Islands. Geographic locations of individuals (triangles- Faroe Islands; circles- global) are indicated by colors in the respective legends, where panel B corresponds to K = 13 in Fig. S3
Our analysis of hierarchical clustering identified the lowest cross-validation error at K = 6 clusters (Fig. S2). At K = 2, we observed a Faroe Islands cluster composed of the islands of Streymoy, Eysturoy, and Vágar (Fig. 3a, S3). At the best supported K = 6, the previously observed Faroese cluster split in two (Streymoy, and Eysturoy and Vágar), and two additional Faroese clusters were observed on Suðuroy, and Borðoy and Viðoy (Fig. S3). The last two clusters were western and northern Europe which had a continuum of ancestry values, as previously reported (Puckett et al. 2016). We also visualized K = 13 due to observations of localized clustering patterns, including differentiation across the island of Streymoy, differentiation of Vágar, and clustering of individual cities (i.e. Bergen, Malmo, and Moscow; Fig. S3). Fine-scale clustering of brown rats within and between sampling sites was observed in our FINESTRUCTURE analysis where we estimated 87 populations within the 208 samples from 53 cities (Fig. 3c).
a Inset of the Faroe Islands and b full map of continental Europe Rattus norvegicus sampling locations with average proportion of ancestry per site inferred using 50k SNPs. Ancestry proportions were based on ADMIXTURE estimates at six clusters (see Fig. S3C). c Coancestry heat map analyzed in FINESTRUCTURE, where yellow and black respectively denote lower and higher coancestry
Our FINESTRUCTURE analysis suggested that the western Faroes (Streymoy, Eysturoy, and Vágar) are genetically differentiated from continental Europe, Suðuroy, and Borðoy and Viðoy (Fig. 3c), in concordance with our PCA results (Fig. 2). The western Faroes show overall low coancestry with Europe, but an elevated amount with Great Britain (Fig. 3c). Suðuroy had highest coancestry with Great Britain, France, and Spain; the coancestry tree suggests Suðuroy and Bergen, Norway share genetic similarity where the lower coancestry between these populations may be due to drift following divergence (Fig. 3c). Borðoy and Viðoy had highest coancestry with Great Britain and Suðuroy (Fig. 3c).
Observed heterozygosity in the European dataset was similar between Faroe Islands and continental European regions (Table S1). Our dataset had more private alleles in the Faroe Islands than continental Europe. Both of these results may be related to our genotyping strategy that aimed to identify variation in the Faroe Islands. Specifically, calling SNPs only on the Faroese samples meant the number of alleles that had to be observed for a MAF > 0.05 was reduced when compared to using Faroese and European samples, thus private or rare variation within the focal populations was included in the European dataset. Even given the genotyping strategy, we observed that the Faroe Islands had larger proportions of MAF = 0 than continental populations (Fig. S4), suggesting that low to moderate frequency variants were fixed during colonization.
Bottleneck and effective population size estimations
We observed increased signatures of LD in six of the eight clusters we tested (Fig. 4), matching expectations for bottlenecked populations. The Western Europe (Great Britain, France, and Spain) and Central Europe (Netherlands and Germany) clusters showed the fastest decay of LD and the lowest r2, whereas northern Sweden (villages of Örsundsbro, Uppsala, and Lövsta) had higher LD despite their location on continental Europe. The Faroe Islands Streymoy and Eysturoy had elevated LD although the geographically proximate island of Vágar had significantly higher LD (Fig. 4). This pattern may reflect the colonization history (Fig. 1), differences in census size on the larger islands, and/or sample size as the smaller island Vágar may have experienced a second bottleneck after the one that brought rats to Streymoy. Suðuroy, and Borðoy and Viðoy had similarly high LD (Fig. 4). Thus, all of the Faroese Islands exhibited signatures of elevated LD; when combined with our clustering and coancestry results, we interpret the LD patterns as indicative of bottlenecks.
To test the strength and identify if the inferred bottlenecks were concordant with founder events on each island, we estimated the change in Ne through time (Fig. 5). Here, we expected to observe sharp declines in Ne at times associated with founder events due to range expansion. Across populations we observed a bottleneck between 10 and 22 kya which likely corresponds to the Last Glacial Maximum (LGM: 18-22 kya) yet with variation within populations resulting from independent sampling of the coalescent process. Patterns differed between Central and Western Europe, where a steady decline was observed in Central Europe and Great Britain but a recovery was observed in France and Spain (Fig. 5a, b). We observed a stairstep pattern of declining Ne in northern Sweden (Fig. 5b); the model was likely overfit to the data, although the overall decreasing population size and temporal estimation of putative bottlenecks was consistent with other European populations. Populations with more samples had smoother declines. We observed sharp population declines in all populations ~1–3 kya and interpret this as the initial range expansion out of the ancestral habitat; the estimates from the stairway plot analyses were near the upper bound of the 90% highest density probability from a global analysis of brown rat demography (Puckett and Munshi-South 2019). Within the four European populations, we estimated the maximum Ne between 273,000 and 372,000 individuals (Fig. 5a, b). These Ne estimates reflect variation in the estimate of the ancestral population size prior to both the LGM and global range expansion. We estimated the contemporary population sizes in Central Europe, Great Britain, France and Spain, and northern Sweden at 1900, 836, 64,000, and 486 individuals, respectively.
Estimates of effective population size (Ne) in Rattus norvegicus populations over time (kya) with 95% highest density probability confidence intervals shown in dotted lines. Populations included: a Great Britain (light blue) and France and Spain (dark teal); b central (dark purple) and northern Europe (medium purple); c Streymoy (gold), Eysturoy (medium green), and Vágar (lime green); d Suðuroy (medium blue) and Borðoy and Viðoy (blue)
Effective population size declined through time on Streymoy and Vágar but appeared to remain steady in Eysturoy (Fig. 5c). Low sample sizes for Vágar contributed to the coarseness of the estimate over time; population size decreased on Vágar around 0.25 and 0.15 kya with a contemporary population size of 1490 individuals. In Eysturoy, population size leveled then increased to the current size of 25,000 effective individuals; whereas in Streymoy there was a continual decline in Ne including several sharp declines that may be indicative of founder events during the global range expansion (Fig. 5c). We estimated the contemporary population size in Streymoy at 123 individuals despite sampling across multiple cities.
The extent of the temporal data for Suðuroy, and Borðoy and Viðoy ends ~250 years ago, where the other populations were estimated closer to the present (Fig. 5d). This is a feature of all flavors of coalescent estimators for Ne, particularly where the last two- or three-time steps are estimated with increased error as there has been insufficient time for coalescence of recent mutations. Thus, although we see declining population sizes around 11 kya and 1.9 kya, as observed in other populations, the temporal extent of the data does not have additional resolution. Therefore, we do not observe a bottleneck during the founding times hypothesized from the historic records of Suðuroy or Borðoy and Viðoy despite the suggestion from the LD analysis (Fig. 4). Further, we do not present an estimate of contemporary population size given this data limitation.
Population tree analysis
We inferred the relationships between sampling sites by estimating the population tree. The population tree with no migration edges explained 98.9% of the variation in the data; we report the tree with two edges based upon the error structure and increase of the variation explained to 99.2% (Fig. S5). Within the tree Suðuroy diverged separately from the other five islands (Fig. 6). The trees also show Borðoy, Streymoy, Eysturoy, and Vágar sharing a population ancestor, where Borðoy shared an ancestor with the western Faroes (Fig. 6). The divergence of Suðuroy was in a part of the tree where other western European sites diverged; similarly, the other islands shared common ancestry with cities in Great Britain (Fig. 6). One migration edge suggested black rat ancestry in the population in Uppsala, Sweden; the second edge estimated movement of brown rats from Sweden to Borðoy (Fig. 6).
Discussion
We identified four evolutionary clusters of brown rats in the Faroe Islands that largely coincided with separate islands (Fig. 3 and S3). We interpret our coancestry and clustering results as evidence for three independent colonizations of brown rats onto the islands of Streymoy, Suðuroy, and Borðoy (Fig. 3 and S3). We utilized the historic record of years of first occurrence to interpret that rats founded Streymoy, expanded across the island, then founded Eysturoy and Vágar seven to ten years later (Fig. 1). This explains the two genetic clusters observed (Fig. S3) as structure established following the founder event, particularly if a northwestern Streymoy population was the source onto both Eysturoy and Vágar which were more closely related than to Streymoy across analyses (Figs. 2b and 3c). Both our coancestry and population tree analyses suggested Great Britain as the source population for this introduction (Figs. 3 and 6). For the introduction to Suðuroy, our coancestry analysis suggested either Great Britain or Norway as the source of the invasion. Our results were least conclusive for Borðoy and Viðoy as coancestry values were low with all samples and suggest the source was not captured within our geographic sampling.
Our results provided mixed support for the three historical hypotheses regarding the origin of brown rats on the Faroe Islands. The first hypothesis, that rats rafted to Suðuroy following a 1768 shipwreck, was partially supported by our coancestry analysis (Fig. 3c). Specifically, we observed elevated coancestry between Suðuroy, and both Norway and Great Britain, where the latter was greater. This result contrasted with the TREEMIX analysis where Suðuroy diverged in the same region of the population tree as both western and central European locations (including Bergen, Norway) but did not share a most recent common ancestor with Great Britain (Fig. 6). Given the historic narrative that the Norwegian ship wrecked off the coast of Scotland, it is possible that rats from Norway and/or Scotland (with presumably shared ancestry with southern populations in Great Britain) founded Suðuroy. Thus, our results do not exclude that the 1768 shipwreck served as the source of rats, although our study would be strengthened by additional sampling along the Norwegian coastline and Scotland. Unfortunately, we were unable to test the timing of this founding (see below), which may provide additional support for the shipwreck hypothesis.
The second hypothesis, that Suðuroy was the source population that expanded to Streymoy, Eysturoy, and Vágar, was not supported by any of our analyses (Figs. 2, 3, 6, and S3). Our population tree inferred an independent introduction of rats to these islands that shared ancestry with populations in Great Britain (Fig. 6). Our clustering analyses suggested that rats first invaded Streymoy and then subsequently spread to Eysturoy and Vágar; this pattern was consistent with the historic record of years of first occurrence of rats on each island (Fig. 1).
Our third hypothesis, that Norwegian whalers introduced rats to Borðoy and Viðoy, was inconclusive. Our coancestry analysis identified that Borðoy and Viðoy were distinct from other Faroe Islands, and showed low coancestry with all continental European populations (Fig. 3c). Coancestry was highest with Great Britain and Suðuroy, and our population tree suggested a shared common ancestor with Great Britain (Fig. 6). Similar to our inference about the founding of Suðuroy, additional sampling in Norway would likely add resolution and aid in inference of the source population which was likely unsampled in this analysis. However, given the elevated coancestry with Suðuroy and the similar demographic histories (Fig. 5d), an alternative hypothesis may be that rats moved from Suðuroy to Borðoy and Viðoy. This movement could have been facilitated by whaling ship traffic between the ports of Nordepli, Borðoy and Lopra, Suðuroy which were founded in 1898 and 1901, respectively. We view some type of human assisted movement as more likely than rats swimming between the islands. Brown rats may swim up to 800 m in water with moderate temperatures; swimming trials in cold water, such as the Faroes where summer ocean temperatures reach 11 °C, estimated they can swim up to 15 min (references within Russell et al. 2008). Given distances between islands, rats could swim from Streymoy to Eysturoy or Vágar, or from Borðoy to Viðoy or Kunoy especially if narrow crossings were used (Fig. 1). However, swimming would be unlikely to explain movement from Suðuroy to Borðoy, as we would expect island hopping to have resulted in colonization of additional islands that have remained rat free.
While each of our hypotheses were explicitly spatial in nature, they also had a temporal component given information on the year of first occurrence of brown rats on each island. We were unable to estimate the timing of founder events onto the Faroe Islands. If populations had gone through strong founding bottlenecks as suggested by the LD analysis (Fig. 4), we may be able to see that signature within the temporal Ne analysis. For the introductions into Suðuroy and Borðoy and Viðoy, there was no inference of Ne more recent than 250 years ago (Fig. 5d). Given the high LD in these populations (Fig. 4), we hypothesized that the founding bottlenecks were so severe that many low frequency variants were removed, thus coalescence only occurred for older events. Our observation of elevated MAF = 0 (i.e. fixation of the major allele) on each of the Faroe Islands sampled supported this hypothesis (Fig. S4). The short time period since founding has not produced sufficient new mutations to estimate more recent demographic events, or at least not enough to be genotyped using a reduced representation method such as ddRAD-Seq. Notably, the SFS dataset for the temporal Ne analysis did not apply a filter on MAF, thus the result of limited temporal inference was not due to filtering out sites containing recent signal (Linck and Battey 2019). We interpret the concordance in Ne between Suðuroy and Borðoy and Viðoy as consistent with both introductions coming from western Europe along with limited sample sizes which produced the strong stairstep pattern. In contrast, temporal patterns of Ne on Streymoy and Vágar showed sharp declines approximately 50 and 150 years ago (Fig. 5c). A 150-year Ne decline would occur around 1865 AD given our assumption of three generations per year, which if associated with a founding bottleneck would be later than expected given the historic record, yet we note wide confidence intervals around the estimates spanning 0 - 1000 years (especially given that rat generation time may be shorter or longer than we assumed, and may vary over time). Interestingly, the 50-year decline in Ne was also observed in Great Britain and coincides with the deployment of warfarin as a rodenticide starting in the 1950s.
There appears to be little to no gene flow either between the Faroe Islands themselves or between the archipelago and continental Europe (Fig. S3). This result was concordant with global and local findings that hypothesized that resident rats excluded potential immigrants from resources and/or breeding opportunities (Calhoun 1962; Combs et al. 2018; Puckett et al. 2016). While this hypothesis needs both observational and experimental field testing, if true it has important implications for rat eradication programs, particularly on isolated islands. First, should rats reappear following an eradication program, monitoring would be aided by genotyping individuals on islands then comparing co-ancestry patterns before and after eradication efforts to determine if eradication was incomplete or new colonization occurred. Second, the low gene flow inferred between islands indicates that eradication efforts could be successful within the Faroe Islands. Streymoy, which is the largest of the Faroe Islands, stands out for having greater genetic differentiation across the island, which may indicate more connectivity between towns on each island than between the islands. This pattern was also observed between towns in northern Streymoy and Eysturoy (Fig. S3).
History of continental Europe
Although our study focused on the history of the Faroe Islands, the data from Sweden and western Russia coupled with the population tree allowed us to make additional inferences across Europe. Our PCA identified a clinal pattern of differentiation from the Mediterranean coast to northern Fennoscandia (Fig. 2b), supporting previous results suggesting rats entered central Europe before expanding bi-directionally across the continent (Puckett and Munshi-South 2019). Our genetic results support Donaldson’s repudiation (Donaldson 1924) of Pallas (Pallas 1778), who suggested that brown rats invaded continental Europe from Eurasia following a massive crossing of the Volga River in 1727. The Volga River is east of Moscow, and our population tree does not support that eastern European cities served as the source for central and western Europe, and our previous work does not support the timing (Puckett and Munshi-South 2019). Notably, our results do not refute the observation of rats crossing the Volga, only the resultant inference. Our samples from Moscow and the Tver region in Russia cluster with other central European locations (Figs. 3 and 6). We interpret these results as part of an eastward expansion following invasion of central Europe (Puckett and Munshi-South 2019), followed by isolation which resulted in the elevated coancestry and unique clustering (Fig. 3 and S3).
Limitations and future directions
Our genotyping strategy and the demographic history of invasive brown rats limited the inferences that we could make concerning the origins of rats on the Faroe Islands. These limitations may apply to many invasive species introductions. Specifically, we observed that the most recent temporal information in the data for Suðuroy and Borðoy and Viðoy was approximately 2 kya (Fig. 5d), thus we could not infer the demographic history of those populations since that time. We believe that the founder events onto these islands removed many low frequency variants (Fig. S4) that would coalesce at shallower temporal depths (Linck and Battey 2019). Population-specific mutation rates and genetic drift will eventually shift the allele frequency spectra in these populations towards low frequency variants; however, these populations have had only 250 years (500-750 rat generations) or fewer to accumulate variants. Thus, there is generally a low probability that a reduced representation genotyping method will detect recently derived alleles. Moreover, the application of a standard filter for MAF > 0.05 applied across all samples from source and invaded populations further reduces the chance that recent population specific variants occur in the final dataset. Here, we observed that typical ddRAD-Seq data processing and filtering resulted in few private alleles in either continental or Faroes populations, as well as many fixed variants in the focal populations. Thus, we recalled SNPs on the Faroe Islands samples alone with the aim to include private and low-frequency alleles within the focal geographic region. We suggest that this strategy may be appropriate for other systems with extremely strong bottlenecks.
Brown rats in the Faroe Islands have great potential to contribute to understanding life history variation and genomic adaptation to cold climates. The generation time of rats is an open debate, as well as an important parameter for demographic modeling. Rats living in the Faroe Islands are first, in cold northern latitudes near the edge of the species range (although there are rat habitats in Iceland, Fennoscandia, and Russia that are further north); and second, present fewer resources (particularly human food) than large cities at similar latitudes. Thus, there is a unique opportunity for comparative analyses of life history evolution in relation to climate and resource availability. We also see a unique opportunity to investigate Faroe Island rats for rapid adaptation to harsh climates. Particularly given the three genomic backgrounds that exist across the islands, scans for selection could focus on genomic regions showing both independent and parallel selective sweep signatures. These future directions would greatly expand our knowledge of rat biology and take advantage of the isolation and lack of gene flow both from Europe and within the Faroes.
Data archiving
DNA sequence data for 103 samples using ddRAD-Seq methods has been deposited in the NCBI SRA under BioProject PRJNA344413.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19(9):1655–1664
Aplin KP, Suzuki H, Chinen AA, Chesser RT, ten Have J, Donnellan SC et al. (2011) Multiple geographic origins of commensalism and complex dispersal history of black rats. PLoS ONE 6(11):e26357
Berry RJ, Jakobson ME, Peters J (1978) The House mice of the Faroe Islands: a study in microdifferentiation. J Zool 185(1):73–92
Bloch D, Fuglø E (1999) Villini Súgdjór í Útnorðri. Føroya Skúlabókagrunnur: Tórshavn, Faroe Islands
Calhoun JB (1962) US Department of Health E, and Welfare (ed.). Public Health Service: Washington DC, pp 288.
Cappelen J, Laursen EV (1998) The climate of the Faroe Islands- with climatological standard normals, 1961–1990. Dan Meteorol Inst 98-14:62. Technical Report
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) STACKS: an analysis tool set for population genomics. Mol Ecol 22(11):3124–3140
Clark G, Costantini L, Finetti A, Giorgi J, Jones A, Reese D et al. (1989) The food refuse of an affluent urban household in the late fourteenth century: Faunal and botanical remains from the Palazzo Vitelleschi, Tarquinia (Viterbo). Pap Br Sch Rome 57:200–321
Combs M, Puckett EE, Richardson J, Mims D, Munshi-South J (2018) Spatial population genomics of the brown rat (Rattus norvegicus) in New York City. Mol Ecol 27(1):83–98
Davis SJM (1983) Morphometric variation of populations of House mice Mus domesticus in Britain and Faroe. J Zool 199(4):521–534
Deinum EE, Halligan DL, Ness RW, Zhang Y-H, Cong L, Zhang J-X et al. (2015) Recent evolution in Rattus norvegicus is shaped by declining effective population size. Mol Biol Evol 32(10):2547–2558
Doherty TS, Glen AS, Nimmo DG, Ritchie EG, Dickman CR (2016) Invasive predators and global biodiversity loss. Proc Natl Acad Sci USA 113(40):11261–11265
Donaldson HH (1924) The rat: data and reference tables for the albino rat (Mus norvegius albinus) and the Norway rat (Mus norvegius), 2nd edn. Wistar Institute, Philadelphia
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22(5):1185–1192
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10(3):564–567
Genovesi P, Carnevali L, Alonzi A, Scalera R (2012) Alien mammals in Europe: updated numbers and trends, and assessment of the effects on biodiversity. Integr Zool 7(3):247–253
Gibbs RA, Weinstock GM, Metzker ML, Muzny DM, Sodergren EJ, Scherer S et al. (2004) Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428(6982):493–521
Goudet J (2005) Hierfstat, a package for r to compute and test hierarchical F-statistics. Mol Ecol Notes 5(1):184–186
Jensen J-K, Magnussen E (2015) Occurrence of fleas (Siphonaptera) and lice (Phthiraptera) on brown rats (Rattus norvegicus) on the Faroes Islands. Nor J Entomol 62:154–159
Johnson MTJ, Munshi-South J (2017) Evolution of life in urban environments. Science 358:6363
Jones EP, Eager HM, Gabriel SI, Jóhannesdóttir F, Searle JB (2013) Genetic tracking of mice and other bioproxies to infer human history. Trends Genet 29(5):298–308
Jones EP, Jensen J-K, Magnussen E, Gregersen N, Hansen HS, Searle JB (2011) A molecular characterization of the charismatic Faroe house mouse. Biol J Linn Soc 102(3):471–482
Korneliussen TS, Albrechtsen A, Nielsen R (2014) ANGSD: Analysis of Next Generation Sequencing Data. BMC Bioinforma 15(1):1–13
Kyrjarheyg M (2014). Stóri týnarin: Rottan. In: Bø Lí (ed) Jústa-ættin - og aðrar Viðareiðissøgur, pp 210–212
Lack J, Hamilton M, Braun J, Mares M, Van Den Bussche R (2013) Comparative phylogeography of invasive Rattus rattus and Rattus norvegicus in the U.S. reveals distinct colonization histories and dispersal. Biol Invasions 15(5):1067–1087
Lack JB, Greene DU, Conroy CJ, Hamilton MJ, Braun JK, Mares MA et al. (2012) Invasion facilitates hybridization with introgression in the Rattus rattus species complex. Mol Ecol 21(14):3545–3561
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359
Lawson DJ, Hellenthal G, Myers S, Falush D (2012) Inference of population structure using dense haplotype data. PLoS Genet 8(1):e1002453
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25(16):2078–2079
Linck E, Battey CJ (2019) Minor allele frequency thresholds strongly affect population structure inference with genomic data sets. Mol Ecol Resour 19(3):639–647
Liu X, Fu Y-X (2015) Exploring population size changes using SNP frequency spectra. Nat Genet 47:555
Manichaikul A, Mychaleckyj JC, Rich SS, Daly K, Sale M, Chen WM (2010) Robust relationship inference in genome-wide association studies. Bioinformatics 26(22):2867–2873
Pallas P (1778). Novae species Quadrupedum e Glirium mordine, Vol 1: Erlangen.
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: An inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS ONE 7(5):e37135
Pickrell JK, Pritchard JK (2012) Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 8(11):e1002967
Pimentel D, Lach L, Zuniga R, Morrison D (2000) Environmental and economic costs of nonindigenous species in the United States. BioScience 50(1):53–65
Puckett EE, Micci-Smith O, Munshi-South J (2018) Genomic analyses identify multiple Asian origins and deeply diverged mitochondrial clades in inbred brown rats (Rattus norvegicus). Evolut Appl 11(5):718–726
Puckett EE, Munshi-South J (2019) Brown rat demography reveals pre-commensal structure in eastern Asia prior to expansion into Southeast Asia. Genome Res 29:762–770
Puckett EE, Park J, Combs M, Blum MJ, Bryant JE, Caccone A et al. (2016) Global population divergence and admixture of the brown rat (Rattus norvegicus). Proc R Soc B: Biol Sci 283(1841):1–9
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D et al. (2007) PLINK: A tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575
R Core Team. (2013) R Foundation for Statistical Computing: Vienna, Austria.
Robins JH, Miller SD, Russell JC, Harper GA, Fewster RM (2016) Where did the rats of Big South Cape Island come from? NZ J Ecol 40(2):229–234
Russell JC, Blackburn TM (2017) Invasive alien species: Denialism, disagreement, definitions, and dialogue. Trends Ecol Evol 32(5):312–314
Russell JC, Meyer J-Y, Holmes ND, Pagad S (2017) Invasive alien species on islands: Impacts, distribution, interactions and management. Environ Conserv 44(4):359–370
Russell JC, Towns DR, Clout MN (2008) Review of rat invasion biology: Implications for island biosecurity. Sci Conserv 286:1–54
Rutherford GK (1982) The physical environment of the Faroe Islands. Dr. W. Junk Publishers, Hague
Scheet P, Stephens M (2006) A fast and flexible statistical model for large-scale population genotype data: Applications to inferring missing genotypes and haplotypic phase. Am J Hum Genet 78(4):629–644
Searle JB, Jones CS, Gündüz İ, Scascitelli M, Jones EP, Herman JS et al. (2009) Of mice and (Viking?) men: phylogeography of British and Irish house mice. Proc R Soc Lond B: Biol Sci 276(1655):201–207
Skarði J (1956) Føroyski leypurin. Fróðskaparrit 5:108–152
Song Y, Lan Z, Kohn MH (2014) Mitochondrial DNA phylogeography of the Norway rat. PLoS ONE 9(2):e88425
Svabo J (1783) Om firfødde dyr Indberetninger fra en Reise i Færøe 1781 og 1782. Selskabet til Udgivelse af Færøske Kildeskrifter og Studier: Copenhagen, pp 54-59.
vonHoldt BM, Pollinger JP, Earl DA, Knowles JC, Boyko AR, Parker H et al. (2011) A genome-wide perspective on the evolutionary history of enigmatic wolf-like canids. Genome Res 21(8):1294–1305
Wickham H (2016) ggplot2: Elegant graphics for data analysis. Springer-Verlag, New York
Acknowledgements
We thank three anonymous reviewers for comments which substantially improved the manuscript. This work was funded by the US National Science Foundation grants DEB 1457523 and DBI 1531639 to JM-S, and the Russian Science Foundation project number 16-14-10323 to LAK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Puckett, E.E., Magnussen, E., Khlyap, L.A. et al. Genomic analyses reveal three independent introductions of the invasive brown rat (Rattus norvegicus) to the Faroe Islands. Heredity 124, 15–27 (2020). https://doi.org/10.1038/s41437-019-0255-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-019-0255-6