Abstract
Heritability is a fundamental parameter in genetics. Traditional estimates based on family or twin studies can be biased due to shared environmental or non-additive genetic variance. Alternatively, those based on genotyped or imputed variants typically underestimate narrow-sense heritability contributed by rare or otherwise poorly tagged causal variants. Identical-by-descent (IBD) segments of the genome share all variants between pairs of chromosomes except new mutations that have arisen since the last common ancestor. Therefore, relating phenotypic similarity to degree of IBD sharing among classically unrelated individuals is an appealing approach to estimating the near full additive genetic variance while possibly avoiding biases that can occur when modeling close relatives. We applied an IBD-based approach (GREML-IBD) to estimate heritability in unrelated individuals using phenotypic simulation with thousands of whole-genome sequences across a range of stratification, polygenicity levels, and the minor allele frequencies of causal variants (CVs). In simulations, the IBD-based approach produced unbiased heritability estimates, even when CVs were extremely rare, although precision was low. However, population stratification and non-genetic familial environmental effects shared across generations led to strong biases in IBD-based heritability. We used data on two traits in ~120,000 people from the UK Biobank to demonstrate that, depending on the trait and possible confounding environmental effects, GREML-IBD can be applied to very large genetic datasets to infer the contribution of very rare variants lost using other methods. However, we observed apparent biases in these real data, suggesting that more work may be required to understand and mitigate factors that influence IBD-based heritability estimates.
Similar content being viewed by others
INTRODUCTION
The proportion of phenotypic variance due to additive genetic variation, termed narrow-sense heritability (h2), is perhaps the most fundamental aspect of a trait’s genetic architecture and has both medical and evolutionary significance (Visscher et al. 2008; Tenesa and Haley 2013). Traditionally, h2 has been estimated from family based studies (h2PED), which have suggested that for many complex traits, much of the phenotypic variance is due to additive genetic variance (Polderman et al. 2015). However, h2PED estimates may be biased by factors shared by close relatives, such as non-additive genetic and common environmental effects (Eaves et al. 1978; Coventry and Keller 2005; Yang et al. 2010; Zuk et al. 2012; Tenesa and Haley 2013).
Recently, methods have been developed to estimate the phenotypic variance explained by all genotyped single-nucleotide polymorphisms (SNPs) simultaneously in unrelated individuals, \(\widehat h_{{SNP}}^2\) (Yang et al. 2010; Speed et al. 2012; Bulik-Sullivan et al. 2015). Most of these approaches use a genetic relatedness matrix (GRM) that reflects allele sharing or the average correlation between individuals i and j across genotyped SNPs with entries:
where m is the number of SNPs, xjk is the genotype (coded as 0, 1, or 2) of individual j at the kth locus, and pk is the minor allele frequency (MAF) of the kth locus. The variance–covariance of the phenotype is
where the variance explained by the SNPs (σ2v) and error variance (σ2e) are estimated using restricted maximum likelihood (REML) (Lynch and Walsh 1998). The method, termed GREML, is implemented in packages such as GCTA (Yang et al. 2011). We refer to matrix A (of dimension n × n and with elements Aij) as the “SNP-GRM.” The proportion of the variance explained by all SNPs is an estimate of “SNP-based heritability” (\(\widehat h_{{SNP}}^2\) = σ2v / (σ2v+σ2e)). By using unrelated individuals, these approaches avoid the confounding of non-additive genetic and environmental effects that can occur in family or twin-based studies, and by estimating all marker effects jointly, the contribution from variants with small effect sizes is captured. Using marker-based approaches, \(\widehat h_{{SNP}}^2\) estimates using imputed data have approached h2PED for some complex traits, such as height, suggesting that little of the heritability remains missing (Yang et al. 2015). For other traits, such as BMI, schizophrenia, and neuroticism, \(\widehat h_{{PED}}^2\) estimates remain larger than \(\widehat h_{{SNP}}^2\), and a substantial amount of the heritability remains “still missing” (Lee et al. 2012; Yang et al. 2015).
Advances on the original approach by Yang et al. (2010) have better captured the effects of rare CVs or account for linkage disequilibrium (LD) of markers across the genome, leading to increased \(\widehat h_{{SNP}}^2\) estimates (Yang et al. 2015; Speed et al. 2017). However, even with the best-performing methods such as MAF-stratified and LD-stratified GREML (GREML-LDMS) and large imputation reference panels, downward bias is likely. Imputation quality declines at low MAF, resulting in a downward bias when causal variants are very rare (MAF < 0.0025) and for diverse populations underrepresented in sequencing panels (Yang et al. 2015; Evans et al. 2017). The underestimation of variance due to rare CVs may partly explain why \(\widehat h_{{SNP}}^2\) remains below h2PED for many traits, in addition to factors that may inflate h2PED described above. Thus, developing alternative and better methods to estimate the variation caused by very rare variants while excluding possible confounding factors of close relatives is an important goal.
One such alternative is to leverage information on the proportion of the genome shared identical-by-descent (IBD) between pairs of individuals in a sample (Visscher et al. 2006; Hayes et al. 2009; Zuk et al. 2012; Browning and Browning 2013a), and use a GRM whose elements are the estimated proportions of IBD between all pairs of individuals (IBD-GRM) to drive and estimate of heritability, which we term \(\widehat h_{{IBD}}^2\). This is in some ways similar to classical family based estimates of heritability, which are based on the expected proportion of the genome shared IBD between close relatives (Falconer and Mackay 1996; Lynch and Walsh 1998; Visscher et al. 2006). However, rather than using close relatives, an appealing alternative is to estimate pairwise IBD segments directly between all pairs of unrelated (or technically, distantly related) pairs of individuals in a sample, and then to use these estimated relationship values to estimate the additive genetic variation. Such an IBD-based approach should capture additive genetic variation due to all but the rarest CVs and, so long as close relatives have been removed from the sample, the IBD-based h2 estimate should be uncontaminated by confounding factors shared by close relatives.
Here, we use “IBD” to denote two homologous chromosomal segments that came from the same common ancestor without intervening recombination, such that the sequence identity of the two segments is identical except at sites where new mutations arose since the last common ancestor. The probability that such mutations arose is a function of the number of generations since the last common ancestor and the number of sites, and therefore a function of the length of the shared IBD segment (when its age is unknown) (Wakeley 2009). When two haplotypes match on a sufficiently long stretch of SNPs, the segments are likely to have been inherited intact from a common ancestor. A pair of very long IBD segments is more likely to be found between pairs with a very recent common ancestor, while a pair shorter segments is more likely for two sequences with a more distant common ancestor (Wakeley 2009). Common, and therefore older, alleles are likely to be shared on both long and short IBD segments, but rare variants, which are likely to have arisen more recently, will be captured more frequently on long IBD segments, as those segments are more likely to be shared by individuals with a more recent ancestor (Browning and Thompson 2012). Thus, IBD-GRMs calculated from increasingly long IBD thresholds (i.e., minima) should capture sharing at increasingly rare CVs.
Such IBD-based GRMs have been used in several instances to estimate heritability. Price et al. (2011) and Zaitlen et al. (2013) used IBD segments in an Icelandic data set with close relatives to estimate heritability in quantitative and disease traits, leveraging the known familial relationships within the Icelandic cohort to identify IBD segments. While they demonstrated that IBD could be used for heritability estimation, using close relatives leads to possible confounding of shared environmental or non-additive genetic effects, as noted above. Indeed, Zaitlen et al. (2013) found higher heritability estimates using closer relatives, consistent with confounding from non-additive genetic and/or shared environment effects. Using simulated data, Zuk et al. (2012) demonstrated that the slope estimated from regressing phenotypic similarity (defined as the standardized phenotypic product of individuals i and j, Zi × Zj) on the IBD-GRM elements from long IBD segments—known as Haseman–Elston (H–E) regression—provides an unbiased estimate of the additive genetic variance in isolated founder populations. Browning and Browning 2013a estimated IBD tracts in a Finnish cohort of 5400 individuals, and used the resulting IBD-GRM in both H–E regression and GREML to estimate \(\widehat h_{{IBD}}^2\) for nine quantitative metabolic traits. \(\widehat h_{{IBD}}^2\) was higher than \(\widehat h_{{SNP}}^2\) for only five of the nine traits, and never significantly so. The most notable result of their study was the over two-fold higher standard errors for \(\widehat h_{{IBD}}^2\) (~0.17) compared to \(\widehat h_{{SNP}}^2\) (~0.07), due to the lower variation in the off-diagonal elements of the IBD-GRM compared to the SNP-GRM, suggesting that very large sample sizes are required to obtain meaningful results in non-founder populations.
Several important questions about IBD-based heritability estimation remain in light of these findings. First, how well do IBD-based approaches estimate the heritability due to very rare CVs? Previous studies (e.g., Browning and Browning 2013a; Zaitlen et al. 2013) have simulated CVs from SNPs present on genotyping arrays, which are more common, have generally higher LD, and are more likely to be shared across ancestry groups than whole-genome sequence (WGS) variants. Thus, such simulations do not provide an accurate picture of how h2 estimation methods perform when CVs do not share these same properties, and so it remains unclear whether \(\widehat h_{{IBD}}^2\) estimates are unbiased estimates of h2 due to rare CVs. Second, the studies mentioned above utilized isolated founder populations that were both more homogeneous and more related than non-founder populations. To what extent does stratification within a sample bias \(\widehat h_{{IBD}}^2\), and how feasible are such IBD-based methods in samples from non-founder populations, which are much more readily available? Third, environmental factors can be passed from parents to offspring (called “vertical transmission”), which can increase phenotypic similarity across extended pedigrees (Coventry and Keller 2005), leading to the possibility of confounding with IBD sharing. To what extent do environmental effects shared across distant relatives bias estimates of \(\widehat h_{{IBD}}^2\)?
To address these questions, we used thousands of recently sequenced whole genomes from the Haplotype Reference Consortium (McCarthy et al. 2016) to simulate phenotypes under a range of conditions, including various genetic architectures and levels of stratification, then estimated narrow-sense heritability (\(\widehat h_{{IBD}}^2\)) using an IBD-GRM, either alone or in combination with various SNP-based GRMs. By simulating CVs from whole-genome sequences rather than commercial array SNPs, our study was able to examine the role of all but the rarest frequency classes of CVs in the genome under realistic genomic conditions. We then estimated \(\widehat h_{{IBD}}^2\) for height and BMI in the UK Biobank with over 120,000 individuals.
Materials and methods
Samples and population structure
We tested the \(\widehat h_{{IBD}}^2\) estimation method using simulated phenotypes derived from Haplotype Reference Consortium (HRC) whole-genome sequence data (McCarthy et al. 2016). Briefly, this resource comprises roughly 32,500 individual whole-genome sequences from multiple sequencing studies, with phased genotypes with a minor allele count of at least 5 at all sites. This large sequence dataset allowed us to simulate CVs across all MAF classes down to ~0.0003 with real patterns of LD (within and among chromosomes). It also allowed us to simulate SNP markers available on existing commercial genotyping arrays in order to mimic the process of IBD detection in SNP data. We obtained permission to access the following HRC cohorts (recruitment region & sample size): AMD (Europe & worldwide; 3,189), BIPOLAR (European ancestry; 2,487), GECCO (European ancestry; 1,112), GOT2D (Europe; 2,709), HUNT (Norway; 1,023), SARDINIA (Sardinia; 3,445), TWINS (Minnesota; 1,325), 1000 Genomes (worldwide; 2,495), UK10K (UK; 3,715) (see McCarthy et al. (2016) for additional details of the HRC). This set of cohorts, which included isolated subpopulations of European descent, allowed investigation into the effects of stratification on estimates. The subset totaled 21,500 whole-genome sequences comprising 38,913,048 biallelic SNPs. This is the same set of individuals and simulated phenotypes used in Evans et al. (2017) to compare SNP-based heritability methods. Below, we briefly describe our approach.
Our goal was to assess the accuracy and potential bias of the \(\widehat h_{{IBD}}^2\) estimation method using data similar to those collected for a typical GWAS analysis and \(\widehat h_{{SNP}}^2\)estimation. In order to mimic this kind of data, we first extracted variant positions corresponding to a widely used commercially available genotyping array, the UK Biobank Affymetrix Axiom array. We then identified individuals of primarily European ancestry, using principal components analysis with 133,603 MAF-pruned and LD-pruned markers (plink2 (Chang et al. 2015) command: --maf 0.05 --indep-pairwise 1000 400 0.2) to identify a grouping associated with the 1000 Genomes European individuals in the HRC. This data set comprised 19,478 individuals including Finnish and Sardinian samples (Fig. S1).
From within this European ancestry data set, we identified clusters that contained different levels of genetic heterogeneity within them (Fig. S2). The most structured group contained all samples (N = 19,478). The somewhat structured group excluded Sardinian and Finnish samples (N = 14,424). The low structure group contained northern/western European samples (N = 11,243), and the least structured was a subset of mainly British Isles samples (N = 8,506). We used GCTA (Yang et al. 2011) with LD-pruned and MAF-pruned SNPs to estimate relatedness and remove the minimal number of individuals from pairs with relatedness >0.1 within each of the four samples. In the most homogeneous and smallest sample with no genetic structure, this left 8,201 individuals. In order to eliminate the influence of varying sample size in our comparison across the range of stratification, we randomly chose 8,201 of the unrelated individuals from within each of the other three stratification subsamples. We similarly tested a lower relatedness cutoff of 0.05 within each group (leaving 7,792; 8,115; 8,129; and 8,186 individuals for the four subsamples), and used both subsets later to examine how a 0.1 or 0.05 relatedness cutoff influences \(\widehat h_{{IBD}}^2\) estimates.
Simulated phenotypes using whole-genome sequencing data
We performed two types of simulations to determine how the IBD-based heritability method performed across a range of genetic architectures. First, we used forward-time simulations with the GeneEvolve program, from which we obtained the true IBD segments (Tahmasbi and Keller 2016). As input, we used WGS data from chromosomes 16–22 from 1000 randomly drawn individuals from the “low” stratification subsample described above. We used only these seven chromosomes rather than all autosomes due to computational constraints. We simulated six generations of random mating, with population size increasing by 5000 each generation, and phenotypes derived from 1000 CVs, randomly chosen from all common (MAF > 0.05) or, separately, very rare (MAC > 5 and MAF < 0.0025) sequence SNPs, and a true h2 = 0.5. These simulations allowed us to calculate both true and estimated IBD-GRMs (see Estimating IBD-GRMs below) to determine how inaccuracies in IBD segment calls impacts \(\widehat h_{{IBD}}^2\). These simulations also allowed us to test whether environmental differences between extended families could bias estimates of \(\widehat h_{{IBD}}^2\). To investigate this, we ran simulations using GeneEvolve with h2 = 0.5 and f2 = 0.3, where f2 is the proportion of the phenotypic variance due to vertical transmission—environmental effects passed from parental phenotype to offspring environment—which increases phenotypic similarity within extended pedigrees due to environmental similarity. Thus, this set of simulations served as a test of the robustness of IBD-based heritability estimation to potential confounding by environmental factors that can create similarity within extended pedigrees. We performed 70 replications of each simulation set, using a relatedness cutoff of 0.05 when estimating heritability.
Second, we simulated phenotypes using the 8201 whole-genome sequences within each of the four stratification subsets. This larger sample incorporates complexities of real genomes in a realistically sized sample, which the forward-time simulations did not. We simulated phenotypes from CVs drawn randomly from five MAF ranges: common (MAF > 0.05), uncommon (0.01 < MAF < 0.05), rare (0.0025 < MAF < 0.01), very rare (MAC > 5 and MAF < 0.0025), and all variants randomly drawn with MAC > 5. Phenotypes were generated with 1000 or 10,000 CVs from the model yi = gi + ei, where gi = ∑wikβk, wik is the genotype (coded as 0, 1, or 2) of individual i at the kth CV, and βk is the kth allelic effect size, drawn from (0,1/[2pk(1−pk)]), where pk is the MAF of allele k within each of the four samples, which assumes larger additive effects for rarer variants. The gi’s were standardized (~\({\cal N}\) (0,1)) and residual error was added as ~\({\cal N}\)(0,(1−h2)/h2) for a simulated h2 of 0.5. A total of 400 replications were performed for each CV MAF range and for each of the four stratification subsets.
Mixed models for heritability estimation
We estimated heritability for each simulation using GCTA (Yang and Lee et al. 2011). We tested different models to assess our IBD-based GREML method (GREML-IBD). First, we used the single IBD-GRM with GREML to estimate \(\widehat h_{{IBD}}^2\). Second, to partition the genetic variance into that tagged by common SNPs and that tagged by haplotype sharing, presumably from rarer CVs, we used a two GRM model (GREML-IBD + SNPs) with the IBD-GRM and a common SNP-GRM derived from Axiom array positions with MAF > 0.01. Here, \(\widehat h_{{\rm Total}}^2\) = \(\widehat h_{{IBD}}^2\) + \(\widehat h_{{SNP}}^2\), where \(\widehat h_{{SNP}}^2\) is defined as above and \(\widehat h_{{IBD}}^2{\mathrm{ = }}\frac{{\widehat \sigma _{{IBD}}^2}}{{\widehat \sigma _{{IBD}}^2{\mathrm{ + }}\sigma _e^2}}\), where \(\widehat \sigma _{{IBD}}^2\) is the estimated variance due to the IBD-GRM). Last, we estimated genetic variances due to LD-stratified and MAF-stratified imputed variant SNP-GRMs (\(\widehat h_{{SNP}}^2\)) as well as the IBD-GRM (\(\widehat h_{{IBD}}^2\)) in the same model, which we term GREML-IBD+LDMS. From previous work, we knew that GREML-LDMS underestimates variance attributable to the rarest CVs when using imputed data. We therefore wished to determine if the IBD-GRM could capture that missing heritability. To do this, we estimated 16 SNP-GRMs stratified into the above 4 MAF categories and 4 LD score quartiles using imputed genome-wide variants, and included these plus the IBD-GRM in the model (17 GRMs total). To determine if the IBD-GRM captured the genetic variance due to the rarest CVs, we also tested a model with 12 SNP-GRMs, removing the rarest MAF category described above, for a total of 13 GRMs in the analysis (three MAF categories × four LD score quartiles + 1 IBD-GRM). To impute, we first phased SNP data using SHAPEIT2 (Delaneau et al. 2013), imputed using minimac3 (Das et al. 2016), and retained variants with imputation R2 ≥ 0.3 (Yang et al. 2015). We used the HRC sequence data as our imputation reference panel after removing all target (8201 unrelated + relatives) individuals in the HRC reference panel, thereby assuring ~independence (no relatedness) between the target and reference panels. Additional details of the imputation procedure can be found in Evans et al. (2017). We estimated LD scores for the LD stratification using GCTA. In all cases we included 20 principal components (PCs; 10 from worldwide PC analysis and 10 from the specific subsample PC analysis) as continuous covariates, with sequencing cohort as a categorical covariate. We used unconstrained GREML (--reml-no-constrain option), which ensured unbiased estimation of the parameters (σ2G and σ2E), even if the true value is close to 0 by allowing estimates to be negative (Yang et al. 2017).
Estimating IBD-GRMs
The process of IBD segment identification is itself challenging, and several excellent discussions of the topic exist (Browning and Browning 2012, 2013b, 2013c; Bjelland et al. 2017), but here we focus on applying IBD information to estimate heritability, \(\widehat h_{{IBD}}^2\), using established IBD estimation methods. To mimic computationally phased SNP data with realistic phase errors, we first un-phased the sequence data for each data subset and then re-phased the Axiom array positions using SHAPEIT2 (Delaneau et al. 2013). We then used FISHR2 (Bjelland et al. 2017) to identify shared haplotype segments that are putatively IBD across all pairs of individuals within each of our four structure samples. FISHR2 first uses a modified version of GERMLINE (Gusev et al. 2009) to find candidate IBD segments. It then improves the accuracy of the segment endpoints by comparing an observed moving average of haplotype mismatches (potential phase or SNP call errors) for a given candidate IBD segment to (a) the distribution of haplotype mismatches in segments that are almost certainly IBD (the middlemost sections of very long IBD segments) and (b) the distribution of haplotype mismatches in segments that are almost certainly non-IBD (between random pairs of individuals at matched locations). FISHR2 truncates candidate segments when this moving average becomes more consistent with non-IBD than IBD. FISHR2 is more accurate than leading competitors at detecting long (>3 cM) IBD segments and is the only software that gives unbiased estimates of the true length of IBD segments (Bjelland et al. 2017). The parameters we used for FISHR2 were stringent (command line -err_hom 4 -err_het 1 –min_snp 128 –min_cm_initial 1 –min_cm_final 1 –window 50 –gap 100 -h_extend -w_extend –homoz -emp-ma-threshold 0.06 -emp-pie-threshold 0.015 -count.gap.errors TRUE), chosen to minimize false-positive IBD detection (Bjelland et al. 2017). We used an initial length threshold of 1 cM, but because longer IBD segments are more likely to share rare variants, we also identified segments of length greater than 2, 3, 4, 6, 9, and 12 cM. The FISHR2 parameters we used should lead to consistently low false-positive rates (<0.05) at all threshold lengths, and should lead to a sensitivity that increases as a function of the length of the true IBD segments, with a predicted sensitivity >.90 for IBD segments >3 cM (Bjelland et al. 2017). To reduce the influence of low recombination regions artificially extending segments (e.g., due to one or a few matching IBS SNPs that are far from the termini of true IBD segments), we windsorized genetic map positions by setting the maximum distance between adjacent markers to 0.2 cM, and used an initial 1 cM minimum IBD segment length threshold. To test how different IBD-identification methods would perform, we also applied IBDseq (Browning and Browning 2013c), which estimates the likelihood of IBD for individual markers between pairs of individuals, to a small subset of simulations as a comparison.
We then summed the length in Mb of all segments shared between each pair of individuals and divided by twice the length of the genome. This IBD-GRM then represents the estimated proportion of the genome, Dij, shared IBD between individuals i and j in the sample, similar to the Aij elements of the SNP-GRM. We created IBD-GRMs for each minimum segment cM length threshold. As recombination rate varies throughout the genome, we also tested whether an IBD-GRM based on the summed cM length of segments influences heritability estimates within the moderate and low stratification subsamples.
Investigations into population stratification effects
Population stratification refers to allele frequency differences between subpopulations in a sample. It is well known that when environmental factors cause mean differences between subpopulations, environmental variance can be misattributed to genetic variance, which can be mitigated or eliminated by including ancestry PCs as fixed effects in association or GREML analyses (Price et al. 2006; Yang et al. 2010). However, even in the absence of environmentally driven mean differences between subpopulations, such ancestry-level population stratification can lead to long-range (e.g., across chromosome) LD between rare CVs that biases estimates of h2 (Evans et al. 2017). We investigated this by estimating \(\widehat h_{{IBD}}^2\)across the four samples that varied by degree of ancestry stratification found across Europe (see Samples and Population Structure above) where no environmental effects based on ancestry were simulated.
Because we observed biases in \(\widehat h_{{IBD}}^2\) in the two most stratified samples (see Results section), we performed four additional tests to understand the cause and potential ways to mitigate these biases. First, to test whether bias observed in stratified samples was due to inadequate control of structure, we ran K-means clustering on the somewhat stratified subsample for K = 2 clusters, then ran PC analysis within each of the two clusters. We included the first 35 PCs within each cluster, for a total of 90 PCs (the original 20 plus 35 from each cluster). Because PC analysis was run within each cluster separately, we set the PC scores for the alternate cluster to 0 (the mean).
Second, we tested, within the stratified subsample, whether including 10 additional PCs from very rare variants could correct for the upward bias (Mathieson and McVean 2012). We used 150,000 randomly selected very rare SNPs from the WGS data and pruned for LD (plink2 command: --indep-pairwise 1000 400 0.2), leaving 129,710 variants for the PCA. As a comparison, we also estimated heritability with no covariates included.
Third, we estimated \(\widehat h_{{IBD}}^2\) for phenotypes in which all CVs were drawn from odd chromosomes using IBD-GRMs estimated only from the even chromosomes. The presence of uncontrolled cryptic relatedness or population structure can lead to cross-chromosome LD that inflates h2 estimates (Yang et al. 2011). We estimated the correlation of off-diagonal GRM elements between the IBD-GRMs from even chromosomes and those from odd chromosomes. We also examined the correlation between the off-diagonal elements from IBD-GRMs and the off-diagonal elements from GRMs built from very rare (MAC > 5 and MAF < 0.0025) and common (MAF > 0.05) sequence variants. This tested whether correlations between even and odd chromosome IBD-GRMs were stronger in more stratified subsamples, and whether the correlation with very rare variants was stronger with increasing minimum cM length of the IBD-GRM.
Finally, simultaneously fitting GRMs derived from each chromosome protects against cross-chromosome correlations induced by stratification or cryptic relatedness because the estimates of variance explained by one GRM are conditional on the other GRMs (Yang et al. 2011). However, because the variances of the off-diagonal elements in the IBD-GRMs were so small, models with 22 IBD-GRMs would not converge. Instead, we tested a two GRM model with one IBD-GRM estimated from the odd-numbered chromosomes and a second from the even-numbered chromosomes, which should partially address the effects of long-range LD (Speed et al. 2012).
Heritability of complex traits in the UK Biobank
We applied the IBD-based approaches to height and body mass index (BMI) data in the UK Biobank, a very large resource of ~500 K adults from the UK, genotyped using the Affymetrix Axiom array (Sudlow et al. 2015). The initial release includes ~150 K genotyped individuals, imputed using the combined UK10K/1000 Genomes reference panels. We used this resource previously, and full details on quality control can be found in Evans et al. (2017). We identified putative IBD segments as described above using FISHR2 and then calculating IBD-GRMs with minimum cM thresholds of 2, 3, 4, 6, 9, and 12 cM. We applied a relatedness cutoff of 0.05, and used individuals of European ancestry, resulting in a final sample size of ~120 K individuals included in the analysis (Fig. S2). We used GCTA to estimate variance components and included sex, UK Biobank assessment center, genotype measurement batch, and qualification (highest level of educational attainment) as categorical covariates, and the Townsend deprivation index, age at assessment, age at assessment squared, and the 15 PC scores from the UK Biobank as quantitative covariates. We compared these models using Akaike information criterion with sample size correction (AICc) (Burnham and Anderson 2002), and used this to determine if additional information was added by using an IBD-GRM.
Results
Simulated phenotypes—GREML-IBD
In our simulated genome sequence data, we found that 95% confidence intervals (CIs) of \(\widehat h_{{IBD}}^2\) estimates overlapped the true h2 when no vertically inherited shared environmental variance was present and when using the true IBD segments to construct the IBD-GRM (Fig. 1), suggesting unbiased estimates in this scenario. Using FISHR2-estimated IBD segments, with a 1 cM IBD length threshold, \(\widehat h_{{IBD}}^2\) underestimated the true h2, but increasing the length threshold led to unbiased estimates, where the 95% CI overlapped the simulated h2. This unbiasedness likely stemmed from the very low rate of false-positive long IBD segments, but also suggests that false negative IBD segments, which have a higher rate at long cM thresholds (Bjelland et al. 2017), do not influence \(\widehat h_{{IBD}}^2\). However, the presence of non-genetic, familial environmental variance led to drastically overestimated \(\widehat h_{{IBD}}^2\) whether using the true or FISHR2-estimated IBD segments to construct the IBD-GRM (Fig. 1).
Estimates of IBD-based heritability from forward-time simulated phenotypes, with GREML-SC using GRMs computed from the true IBD segments or FISHR2-estimated IBD segments with varying cM length thresholds. Mean and 95% CI shown from 70 replicates. Relatedness cutoff of 0.05 used. Shown are two sets of simulations, with and without non-genetic, vertically inherited shared environmental variance (f2), with either common (MAF > 0.05) or very rare (MAF < 0.0025) causal variants
In our simulations using real WGS data and FISHR2-estimated IBD segments, \(\widehat h_{{IBD}}^2\) estimates varied greatly when using a single IBD-GRM depending on the MAF range of the CVs in simulated phenotypes and the amount of stratification in the subsample (Fig. 2). In the two more homogeneous subsamples, \(\widehat h_{{IBD}}^2\) was at first underestimated, but increased and then stabilized with increasing IBD segment length threshold, similar to what we observed in simulated genome data. The 95% CI overlapped the true heritability (0.5) for all IBD thresholds >4 cM and for all CV MAF classes, suggesting that GREML-IBD produces unbiased estimates of h2 in relatively homogeneous samples when removing short, likely false-positive IBD segments. Results were similar for different relatedness thresholds (Figs S3 & S4) and for larger numbers of CVs (Fig. S5), although \(\widehat h_{{IBD}}^2\) appeared to be biased upwards in phenotypes with 10,000 common CVs and long IBD length thresholds in the low stratification subsample (Fig. S5). Precision of the estimates declined with longer IBD cM length thresholds, as shown by larger standard errors (Figs. S3-S5) and larger root mean square error (Fig. S6). We note that in tests of a different IBD detection method, IBDseq, the estimates were biased downward compared to those using FISHR2 (Fig. S7). This suggests that an alternative IBD detection method would not correct for the downward bias observed at shorter IBD length thresholds using FISHR2.
GREML-SC using an IBD-GRM. \(\widehat h_{{IBD}}^2\) estimates (mean ± 95% CI from 400 replicates). X axis indicates the IBD-shared haplotype length threshold for the IBD-GRM. Phenotypes with 1000 CVs randomly drawn from the MAF range specified in each panel. Different colors indicate degree of stratification in the sample. Relatedness cutoff of 0.05 used
In the two most stratified samples, we observed underestimates at short cM IBD thresholds, but upward biases at long cM IBD thresholds, particularly for the rarest CVs (\(\widehat h_{{IBD}}^2\) > 1). This bias remained when using higher or lower relatedness thresholds (Figs. S3–4), and with 10,000 CVs (Fig S5). Controlling for 70 additional PCs or with additional PCs from very rare variants did not correct for the upward bias in very rare CV phenotypes, though inclusion of PCs did correct for bias in common CV phenotypes (Fig. S8). Furthermore, this bias was not mitigated by summing genetic length (cM) of IBD segments for calculating the GRM rather than physical length (Fig. S9) nor when using a two GRM model, with one IBD-GRM calculated from even-numbered chromosomes and the second from odd-numbered chromosomes (Fig. S10-S11). Fitting a larger number of IBD-GRMs (e.g., one per chromosome) would better capture all the long-range correlations and might better mitigate the bias, but this approach is impractical for GREML-IBD in real data because the low variance of Dij creates estimation problems. Thus, stratification has strong impacts on GREML-IBD estimates of heritability that we were unable to control for.
To explore why stratification had such strong influences on \(\widehat h_{{IBD}}^2\), we first examined the correlations of off-diagonal GRM elements between the odd chromosome GRMs and even chromosome GRMs. Stratification clearly led to stronger long-range correlations, as did, in most subsamples, longer IBD thresholds for the GRM (Fig. S12). In the two least stratified subsamples, the correlation of even chromosome IBD-GRMs with odd chromosome WGS SNP-GRMs, estimated from either common or very rare WGS variants, was weak, and did not change drastically with increasing cM thresholds. There were stronger correlations overall in the two most stratified subsamples, especially between even chromosome IBD-GRMs and odd chromosome GRMs built from either IBD segments or from very rare WGS variants. Thus, stratification increased long-range correlations between Dij’s, such that Dij for a pair of individuals at one chromosome predicted rare variant sharing at other chromosomes, which can presumably lead to over-estimation of \(\widehat h_{{IBD}}^2\) due to rare CVs being redundantly tagged by IBD sharing.
In simulations with odd chromosome CVs and IBD-GRMs calculated from even chromosomes only, we observed upward biases in \(\widehat h_{{IBD}}^2\) estimates for long IBD thresholds that were particularly severe in stratified samples with rare odd-chromosome CVs (Fig. S13). This pattern of results was similar to the pattern observed in our primary simulations (Fig. 2), consistent with the explanation that the upward biases in \(\widehat h_{{IBD}}^2\) for rare CVs we observed at long IBD thresholds was due to long-range, redundant tagging of CVs in stratified samples. Note that the simulated h2 for the even chromosomes was 0. Because there is more recent common ancestry within than between subpopulations, there is more sharing of long IBD segments—and importantly more sharing of rare (recently arisen) causal variants. Consequently, due to stratification, long, shared IBD segments at one genomic location indicate sharing of rare variants across the genome. This redundant tagging of rare causal variants across the genome in stratified samples presumably leads to inflated \(\widehat h_{{IBD}}^2\). The same phenomenon has been described for \(\widehat h_{SNP}^2\)in the context of stratification (Yang et al. 2011; Speed et al. 2012), although the bias is less extreme and, because the variance of Aij elements is much greater than the variance of Dij, is more easily alleviated by fitting multiple GRM models.
Simulated phenotypes—GREML-SNPs + IBD
The second model we tested was GREML-SNPs + IBD, which included a common SNP-GRM and the IBD-GRM. For phenotypes with 1000 or 10,000 CVs, the total heritability (\(\widehat h_{{IBD}}^2\) + \(\widehat h_{SNP}^2\)=\(\widehat h_{Total}^2\)) was unbiased when using long IBD cM thresholds in the two least stratified subsamples regardless of the CV MAF range (Fig. S14, S15). However, \(\widehat h_{Total}^2\) was underestimated at shorter cM IBD thresholds, and increasingly overestimated with longer thresholds in the two most stratified samples for very rare CV phenotypes. As expected, partitioning the variance to each of the GRMs, GREML-SNPs + IBD attributed more of the phenotypic variance to the common SNP-GRM when the CVs were common, and more of the variance to the IBD-GRM when the CVs were rarer (Figs. S16-S17). For common CV phenotypes, the variance attributable to the common SNP-GRM was overestimated by ~20%, which is consistent with previous findings for a common SNP-GRM based on the Axiom array positions and occurs because CVs in the common bin have higher average MAF than the SNPs on the Axiom array (Evans et al. 2017). Interestingly, this overestimate was balanced by a negative variance estimate attributed to the IBD-GRM, such that the total estimated heritability was unbiased at ~0.5 at long cM IBD thresholds (Figs. S14-S17). Nevertheless, for very rare CV phenotypes, \(\widehat h_{{IBD}}^2\) was again underestimated, then overestimated in structured samples as the cM IBD threshold length was increased.
Simulated phenotypes—GREML-LDMS + IBD
Our third model included 16 imputed variant GRMs that were MAF-stratified and LD-stratified, and the IBD-GRM. We found that across subsamples, GREML-LDMS+IBD produced generally unbiased \(\widehat h_{{\rm Total}}^2\) with either 1000 CVs or 10,000 CVs across all CV MAF ranges when IBD thresholds >4 cM were applied (Figs. S18-S19). Partitioning the variance among GRMs revealed that for the rare and very rare CV phenotypes, the IBD-GRM explained a small amount of the variance, but was near-zero otherwise (Figs. S20-S21).
When we excluded the rarest MAF bin from the model, leaving 12 imputed variant GRMs plus the IBD-GRM, GREML-LDMS + IBD also produced generally unbiased \(\widehat h_{{\rm Total}}^2\) with either 1000 CVs or 10,000 CVs across all CV MAF ranges in subsamples with little or no stratification (Fig. 3, S22). However, with increased stratification, \(\widehat h_{{\rm Total}}^2\) was again overestimated for very rare CV phenotypes in the context of stratification. Partitioning the variance into that attributable to the LDMS imputed variant GRMs and the IBD-GRM showed that, in unstratified samples, most of the genetic variance was attributable to the LDMS GRMs for CV MAF ranges >0.0025 while the IBD-GRM captured the genetic variance for very rare CV MAF ranges (MAC > 5 to MAF < 0.0025) (Fig. 4, S23). While the variance attributed to the LDMS GRMs was never overestimated, that attributed to the IBD-GRMs at longer IBD thresholds was overestimated, resulting in total heritability estimates >1 for the rarest CV phenotypes in the presence of stratification (Fig. 4)
GREML-LDMS + IBD model. This model had 13 components, 12 LD and MAF-stratified GRMs using imputed genome-wide variants, and one GRM from IBD-shared haplotypes. Total h2 estimates are shown (mean ± 95% CI from 400 replicates). X axis indicates the different IBD-shared haplotype length thresholds for the IBD-GRM. Phenotypes with 1000 CVs randomly drawn from the MAF range specified in each panel. Different colors indicate degree of stratification in the sample. Relatedness cutoff of 0.05 used
GREML-LDMS + IBD. This model had 13 components, 12 LD and MAF-stratified GRMs using imputed genome-wide variants, and one GRM from IBD-shared haplotypes. Separate h2 estimates for each component are given by the symbols (mean ± 95% CI from 400 replicates). Note that the “Imputed LDMS” symbol represents the sum of the imputed LDMS GRM variance estimates. X axis indicates the different IBD-shared haplotype length thresholds for the IBD-GRM. Phenotypes with 1000 CVs randomly drawn from the MAF range specified in each panel. Different colors indicate degree of stratification in the sample. Relatedness cutoff of 0.05 used
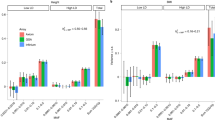
Total heritability estimates for three continuous traits in the UK Biobank. a GREML-IBD, which had a single IBD-GRM. b GREML-LDMS + IBD for two continuous traits in the UK Biobank. This model had 13 components, 12 LD and MAF-stratified GRMs using imputed genome-wide variants, and one GRM from IBD-shared haplotypes. Total h2 estimates are shown (±95% CI). X axis indicates the different IBD-shared haplotype length thresholds for the IBD-GRM. Relatedness cutoff of 0.05 used. Dashed lines represent, for comparison, the SNP-based estimates, using either GREML-SC (a) or GREML-LDMS (b). See Supplementary Table 1 for estimates
Real phenotypes from the UK Biobank
Using GREML-IBD, \(\widehat h_{{IBD}}^2\) for height (but not for BMI) increased with longer minimum shared haplotype length, did not stabilize at longer segment thresholds, and appeared upwardly biased, similar to what we observed in stratified samples in our simulations (Fig. 5a, Table S1). The 95% CIs increased with longer minimum IBD length, as expected given the lower variance in Dij at longer segment thresholds. For comparison, \(\widehat h_{{SNP}}^2\) estimates from approaches using only SNPs are also presented in Table S1.
Using either GREML-SNPs + IBD or GREML-LDMS + IBD, we found similar patterns of increasing \(\widehat h_{{IBD}}^2\) estimates with longer minimum IBD length for height, but the pattern was less extreme, and 95% CIs were generally smaller (Fig. 5b, Table S1). Results for GREML-LDMS + IBD either including the rarest MAF category or excluding it were similar: height \(\widehat h_{{IBD}}^2\) estimates increased from 0.75 to 1.1 across the range of minimum IBD lengths we examined. This increase in \(\widehat h_{{IBD}}^2\) was due to increasing estimates of variance attributable to the IBD-GRM rather than to the imputed variant SNP-GRMs (Fig. S24, Table S1). BMI \(\widehat h_{{IBD}}^2\) were again ~0.2–0.3, though at longer minimum IBD length thresholds the standard errors were large, and the 95% CI overlapped 0 (Table S1).
Interestingly, inclusion of the IBD-GRM in addition to the SNP-GRM or LDMS GRMs often improved model fit (Table S1). Likelihood ratio tests of GREML-SNP vs. GREML-SNP + IBD and GREML-LDMS vs. GREML-LDMS + IBD suggested that model fit when analyzing height, but not BMI, was improved by including the IBD-GRM. Furthermore, comparing AICc across all the models and thresholds, the lowest AICc was often found with shorter IBD minimum length thresholds. For instance, for height, the minimum AICc was found when using all LD-stratified and MAF-stratified imputed variant GRMs and the IBD-GRM with a 3 cM minimum IBD length threshold (Table S1), while AICc increased with longer length thresholds. Thus, while increasing the minimum length threshold led to unreasonable and uninterpretable total heritability estimates, at shorter IBD length thresholds, the inclusion of the IBD-GRM accounted for additional variance explained over using only GREML-SNP or GREML-LDMS. This may have reflected the effect of CVs that are not well captured by imputed variants.
Discussion
We present here the most thorough assessment to-date of an IBD-based heritability estimation approach. The interest in using IBD information in classically unrelated samples to estimate heritability arises from the potential to estimate the full narrow-sense heritability without the confounding of effects shared within families that can bias estimates when close relatives are used, and without the downward bias in estimation when CVs are rare or poorly tagged by SNPs. We demonstrated that GREML-IBD can produce unbiased heritability estimates in realistic whole-genome SNP data so long as there is little genetic stratification in the sample and with estimated IBD length thresholds >4 cM to account for IBD estimation errors.
While IBD-based approaches are appealing in principle, our study highlights three important drawbacks. First, stratification can bias heritability estimates upward, depending on the allele frequencies of CVs. The effect of stratification is strong when CVs are very rare, and is not controlled by inclusion of a large number of PC covariates, the typical approach to controlling such effects (Price et al. 2010), or even PCs derived from very rare variants (Mathieson and McVean 2012). Similar overestimates have been observed in a related method that used sharing at predefined, segregating haplotypes (Bhatia et al. 2016). Overestimates appear to stem from redundant tagging by long IBD segments across the genome in stratified samples, and from non-genetic shared environmental variance. Previous studies using IBD-based approaches (Zuk et al. 2012; Browning and Browning 2013a) used isolated, homogeneous populations, which should mitigate this source of bias. Our simulation results suggest somewhat less homogenous samples, such as those of general northern/western European ancestry, can be used to derive unbiased heritability estimates so long as there are no additional confounding factors.
Second, non-genetic shared environments can strongly bias \(\widehat h_{{IBD}}^2\) estimates upwards. Because long IBD segments identify pairs of individuals with relatively recent shared ancestry, shared environmental influence within families can be confounded with IBD sharing, driving up \(\widehat h_{{IBD}}^2\). In our simulations, we excluded closely related individuals (relatedness < 0.05), demonstrating that this confound is not alleviated by using only nominally unrelated individuals.
Third, the standard error (SE) and RMSE of \(\widehat h_{{IBD}}^2\) is large due to the very low variance in IBD sharing among unrelated individuals in large, non-founder populations. For example, for height in the UK Biobank when using GREML-LDMS + IBD, total heritability SE ≥ 0.053 for minimum IBD lengths ≥ 6 cM, largely due to the IBD-GRM variance component SE. However, using just the imputed variant GREML-LDMS approach SE = 0.015. Thus, while the GREML-LDMS + IBD may have accounted for more of the genetic variance, it did so with substantially lower precision. Very large sample sizes will be required to reach high levels of precision. Taken together, it seems unlikely the increased variance explained, arising from capturing rare CVs with IBD-based GRMs, outweighs the very large increase in standard errors and the increased potential for bias due to stratification or shared environmental variance.
Heritability of real complex traits
Our results from real UK Biobank data for height demonstrate the potential for additional biases of an IBD-based approach that were not captured in our simulation. The estimates of total heritability for height increased with minimum IBD cM length, and were much greater than other reported estimates (e.g., Yang et al. 2015; Evans et al. 2017). This was unexpected given that the stratification of the UK Biobank sample was similar to the unstratified subsets in our simulations, suggesting that stratification in the UK Biobank sample is not the cause of the upward bias in height \(\widehat h_{{IBD}}^2\).
Alternatively, vertically transmitted non-genetic effects, shared common environmental effects, and assortative mating may also confound estimates of \(\widehat h_{{IBD}}^2\). Estimates of \(\widehat h_{{PED}}^2\) using close relatives can be altered by these factors (Eaves et al. 1978; Martin et al. 1978; Coventry and Keller 2005; Zuk et al. 2012). Common environmental effects, which can induce similarity across extended pedigrees, would be confounded with IBD sharing, and are therefore a potential source of bias in IBD-based estimates. As demonstrated in our first set of forward-time simulations, f2 can indeed bias \(\widehat h_{{IBD}}^2\) estimates, and this is a potential explanation for why height \(\widehat h_{{IBD}}^2\) is unrealistically high. It is possible BMI does not have a similar influence of f2, which is why we observed reasonable BMI \(\widehat h_{{IBD}}^2\) estimates. Further work will be required to test this hypothesis, such as including various environmental matrices, sensu Xia et al. (2016). The use of lower relatedness thresholds may alleviate the problem, but lower relatedness thresholds would decrease the sample size and variance of IBD sharing and therefore further exacerbate the already high standard errors of these estimates. Rare variants are more differentially confounded by stratification than common variants, and typical approaches using PCA may not fully correct for such confounding (Mathieson and McVean 2012). Extremely rare SNPs, as with long IBD segments, will co-segregate along extended pedigrees, and future work must focus on the role of confounding between familial and environmental effects and rare variants or long IBD segments.
While we cannot conclude with certainty which factors led to the apparent bias in height \(\widehat h_{{IBD}}^2\), estimates of \(\widehat h_{{IBD}}^2\) for BMI were more stable and also in line with previous reports. They suggest that BMI h2 is roughly 0.25–0.3, with up to 5% of the total phenotypic variance due to very rare or otherwise poorly imputed variants that are captured by the IBD-GRM (see Table S1). As estimates from classical twin design studies range from 0.4–0.8, this suggests that much of the family based estimates are due to shared environment, assortative mating, or non-additive genetic variance, supported by extended twin design variance estimates (Coventry and Keller 2005; Keller and Coventry 2005). This also suggests that little unexplained variance remains for BMI, as estimates of BMI \(\widehat h_{{SNP}}^2\) from recent studies range from 0.21 (Locke et al. 2015) to 0.27 (Yang et al. 2015).
Our findings may also offer context to the observed heritability estimates reported by several other studies that used haplotype-based approaches. Browning and Browning (2013a) reported \(\widehat h_{{IBD}}^2\) for BMI of 0, with standard error of 0.16 (height was not measured), although their upper 95% CI estimate is not inconsistent with a true h2 of 0.25–0.3. This low estimate may simply be due to sampling variance, arising from the small number of individuals (5,402) in the Finnish sample they used, or to true heritability differences among populations. Zaitlen et al. (2013) used IBD among close relatives to derive estimates of \(\widehat h_{{IBD}}^2\) of 0.69 for height and 0.42 for BMI. As discussed by the authors, these estimates may be upwardly biased due to common environmental and non-additive genetic effects.
Conclusions
Identical-by-descent haplotypes in common between a pair of chromosomes capture sharing at all variants that existed along their length in the last common ancestor. The ability to estimate such IBD segments using SNP data means that there is potential to estimate narrow-sense heritability of traits. We conclude that IBD-based estimates can be used to obtain estimates of the near full narrow-sense heritability. However, IBD-based estimates are imprecise, very sensitive to stratification, and can be confounded by shared environmental variance, even in unrelated samples. Moreover, when we estimated \(\widehat h_{{IBD}}^2\) in real data, we observed biases that appeared similar to those that we had observed due to stratification and shared environments, which suggests that there are biases in real data that we were not able to adequately control. Taken together, these factors diminish the appeal of IBD-based approaches for estimating heritability, especially when compared to approaches that use imputed variants, such as GREML-LDMS. Nevertheless, until whole-genome sequence data is feasible for the large sample sizes required for h2 estimation from genotype data, IBD-based estimates may be able to capture the rarest CVs better than imputation. In particular, though larger and more diverse reference panels are becoming available, IBD-based approaches offer a method to capture rare genome-wide variants not represented in imputation reference panels and structural variation that remains difficult to capture even with whole-genome sequencing (Auton et al. 2015). Furthermore, isolated, homogeneous populations may also be the most advantageous for IBD-based heritability estimation due to the larger variance in IBD sharing, though extremely large sample sizes would be required to offset the lower precision in heritability estimates.
Data archiving
Data are from the Haplotype Reference Consortium (http://www.haplotype-reference-consortium.org/) and the UK Biobank (http://www.ukbiobank.ac.uk/).
References
Auton A, Abecasis GR, Altshuler DM, Durbin RM, Bentley DR, Chakravarti A et al. (2015) A global reference for human genetic variation. Nature 526:68–74
Bhatia G, Gusev A, Loh P-R, Finucane HK, Vilhjalmsson BJ, Ripke S et al. (2016). Subtle stratification confounds estimates of heritability from rare variants. bioRxiv. https://doi.org/10.1101/048181.
Bjelland DW, Lingala U, Patel P, Jones M, Keller MC (2017) A fast and accurate method for detection of IBD shared haplotypes in genome-wide SNP data. Eur J Hum Genet 25:617–624
Browning SR, Browning BL (2012) Identity by descent between distant relatives: detection and applications. Annu Rev Genet 46:617–633
Browning SR, Browning BL (2013a) Identity-by-descent-based heritability analysis in the Northern Finland Birth Cohort. Hum Genet 132:129–138
Browning BL, Browning SR (2013b) Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 194:459–471
Browning BL, Browning SR (2013c) Detecting identity by descent and estimating genotype error rates in sequence data. Am J Hum Genet 93:840–851
Browning SR, Thompson EA (2012) Detecting rare variant associations by identity-by-descent mapping in case-control studies. Genetics 190:1521–1531
Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Consortium SWG of the PG (2015) LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295. et al.
Burnham KP, Anderson DR (2002) Model selection and multi-model inference, 2nd edn. Springer, New York, NY
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ et al. (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4:7
Coventry WL, Keller MC (2005). Estimating the extent of parameter bias in the classical twin design: a comparison of parameter estimates from extended twin-family and classical twin designs. Twin Res Hum Genet. 8: 214–223.
Das S, Forer L, Schönherr S, Sidore C, Locke AE, Kwong A et al. (2016) Next-generation genotype imputation service and methods. Nat Genet 48:1284–1287
Delaneau O, Zagury J-F, Marchini J (2013) Improved whole-chromosome phasing for disease and population genetic studies. Nat Methods 10:5–6
Eaves LJ, Last KA, Young PA, Martin NG (1978) Model-fitting approaches to the analysis of human behaviour. Heredity 41:249–320
Evans LM, Tahmasbi R, Vrieze SI, Abecasis GR, Das S, Bjelland DW et al. (2017). Comparison of methods that use whole genome data to estimate the heritability and genetic architecture of complex traits. https://doi.org/10.1101/115527.
Falconer DS, Mackay TFC (1996). Introduction to quantitative genetics, Longman Limited: Burnt Mill, England.
Gusev A, Lowe JK, Stoffel M, Daly MJ, Altshuler D, Breslow JL et al. (2009) Whole population, genome-wide mapping of hidden relatedness. Genome Res 19:318–326
Hayes BJ, Visscher PM, Goddard ME (2009) Increased accuracy of artificial selection by using the realized relationship matrix. Genet Res 91:47–60
Keller MC, Coventry WL (2005) Quantifying and addressing parameter indeterminacy in the classical twin design. Twin Res Hum Genet 8:201–213
Lee SH, DeCandia TR, Ripke S, Yang J, Sullivan PF, Goddard ME et al. (2012) Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat Genet 44:247–250
Locke AE, Kahali B, Berndt SI, Justice AE, Pers TH, Day FR et al. (2015) Genetic studies of body mass index yield new insights for obesity biology. Nature 518:197–206
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates, Sunderland, MA
Martin NG, Eaves LJ, Kearsey MJ, Davies P (1978) The power of the classical twin study. Heredity 40:97–116
Mathieson I, McVean G (2012) Differential confounding of rare and common variants in spatially structured populations. Nat Genet 44:243–6
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A et al. (2016) A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet 48:1279–1283
Polderman TJC, Benyamin B, de Leeuw CA, Sullivan PF, van Bochoven A, Visscher PM et al. (2015) Meta-analysis of the heritability of human traits based on fifty years of twin studies. Nat Genet 47:702–709
Price AL, Helgason A, Thorleifsson G, McCarroll SA, Kong A, Stefansson K (2011). Single-tissue and cross-tissue heritability of gene expression via identity-by-descent in related or unrelated individuals. PLoS Genet 7: e1001317.
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick Na, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–9
Price AL, Zaitlen Na, Reich D, Patterson N (2010) New approaches to population stratification in genome-wide association studies. Nat Rev Genet 11:459–63
Speed D, Cai N, Consortium U, Johnson M, Nejentsev S, Balding D (2017) Re-evaluation of SNP heritability in complex human traits. Nat Genet 49:986–992
Speed D, Hemani G, Johnson MR, Balding DJ (2012) Improved heritability estimation from genome-wide SNPs. Am J Hum Genet 91:1011–1021
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J et al. (2015) UK Biobank: An Open Access Resource for Identifying the Causes of a Wide Range of Complex Diseases of Middle and Old Age. PLoS Med 12:1–10
Tahmasbi R, Keller MC (2016) Genetics and population analysis GeneEvolve: a fast and memory efficient forward-time simulator of whole-genome data. Bioinformatics 33:294–396
Tenesa A, Haley CS (2013) The heritability of human disease: estimation, uses and abuses. Nat Rev Genet 14:139–149
Visscher PM, Hill WG, Wray NR (2008). Heritability in the genomics era--concepts and misconceptions. Nat Rev Genet. 9: 255–266.
Visscher PM, Medland SE, Ferreira MaR, Morley KI, Zhu G, Cornes BK et al. (2006) Assumption-free estimation of heritability from genome-wide identity-by-descent sharing between full siblings. PLoS Genet 2:e41
Wakeley J (2009) Coalescent theory: an introduction. Roberts and Company, Greenwood Village, CO
Xia C, Amador C, Huffman J, Trochet H, Campbell A, Porteous D et al. (2016) Pedigree- and SNP-associated genetics and recent environment are the major contributors to anthropometric and cardiometabolic trait variation. PLoS Genet 12:e1005804
Yang J, Bakshi A, Zhu Z, Hemani G, Vinkhuyzen AAE, Lee SH et al. (2015) Genetic variance estimation with imputed variants finds negligible missing heritability for human height and body mass index. Nat Genet 47:1114–20
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR et al. (2010) Common SNPs explain a large proportion of the heritability for human height. Nat Genet 42:565–569
Yang J, Lee SH, Goddard ME, Visscher PM (2011) GCTA: A tool for genome-wide complex trait analysis. Am J Hum Genet 88:76–82
Yang J, Manolio TA, Pasquale LR, Boerwinkle E, Caporaso N, Cunningham JM et al. (2011) Genome partitioning of genetic variation for complex traits using common SNPs. Nat Genet 43:519–25
Yang J, Zeng J, Goddard ME, Wray NR, Visscher PM (2017) Concepts, estimation and interpretation of SNP-based heritability. Nat Genet 49:1304–1310
Zaitlen N, Kraft P, Patterson N, Pasaniuc B, Bhatia G, Pollack S et al (2013) Using extended genealogy to estimate components of heritability for 23 quantitative and dichotomous traits. PLoS Genet 9:e1003520
Zuk O, Hechter E, Sunyaev SR, Lander ES (2012) The mystery of missing heritability: Genetic interactions create phantom heritability. Proc Natl Acad Sci USA 109:1193–8
Acknowledgements
This work utilized the Janus supercomputer, which is supported by the National Science Foundation (award number CNS-0821794), the University of Colorado Boulder, the University of Colorado Denver, and the National Center for Atmospheric Research, and is operated by the University of Colorado Boulder. This research has been conducted using the UK Biobank Resource. We thank the participants of the individual HRC cohorts and the UK Biobank. We thank the Keller and Vrieze lab groups, the Institute for Behavioral Genetics, and Sean Caron. This study was funded by NIH R01MH100141 (MCK), NIH R01DA037904 and R01HG008983 (SV), NHMRC grants 1078037 (PMV) and 1113400 (PMV and JY), and Sylvia & Charles Viertel Charitable Foundation Senior Medical Research Fellowship (JY).
Haplotype Reference Consortium:
http://www.haplotype-reference-consortium.org/ Gonçalo Abecasis, David Altshuler, Carl A Anderson, Andrea Angius, Jeffrey C Barrett, Sonja Berndt, Michael Boehnke, Dorrett Boomsma, Kari Branham, Gerome Breen, Chad M Brummett, Fabio Busonero, Harry Campbell, Peter Campbell, Andrew Chan, Sai Chen, Emily Chew, Massimiliano Cocca, Francis S Collins, Laura J Corbin, Francesco Cucca, Petr Danecek, Sayantan Das, Paul I W de Bakker, George Dedoussis, Annelot Dekker, Olivier Delaneau, Marcus Dorr, Richard Durbin, Aliki-Eleni Farmaki, Luigi Ferrucci, Lukas Forer, Ross M Fraser, Timothy Frayling, Christian Fuchsberger, Stacey Gabriel, Ilaria Gandin, Paolo Gasparini, Christopher E Gillies, Arthur Gilly, Leif Groop, Tabitha Harrison, Andrew Hattersley, Oddgeir L Holmen, Kristian Hveem, William Iacono, Amit Joshi, Hyun Min Kang, Hamed Khalili, Charles Kooperberg, Seppo Koskinen, Matthias Kretzler, Warren Kretzschmar, Alan Kwong, James C Lee, Shawn Levy, Yang Luo, Anubha Mahajan, Jonathan Marchini, Steven McCarroll, Mark I McCarthy, Shane McCarthy, Matt McGue, Melvin McInnis, Thomas Meitinger, David Melzer, Massimo Mezzavilla, Josine L Min, Karen L Mohlke, Richard M Myers, Matthias Nauck, Deborah Nickerson, Aarno Palotie, Carlos Pato, Michele Pato, Ulrike Peters, Nicola Pirastu, Wouter Van Rheenen, J Brent Richards, Samuli Ripatti, Cinzia Sala, Veikko Salomaa, Matthew G Sampson, David Schlessinger, Robert E Schoen, Sebastian Schoenherr, Laura J Scott, Kevin Sharp, Carlo Sidore, P Eline Slagboom, Kerrin Small, George Davey Smith, Nicole Soranzo, Timothy Spector, Dwight Stambolian, Anand Swaroop, Morris A Swertz, Alexander Teumer, Nicholas Timpson, Daniela Toniolo, Michela Traglia, Marcus Tuke, Jaakko Tuomilehto, Leonard H Van den Berg, Cornelia M van Duijn, Jan Veldink, John B Vincent, Uwe Volker, Scott Vrieze, Klaudia Walter, Cisca Wijmenga, Cristen Willer, James F Wilson, Andrew R Wood, Eleftheria Zeggini, He Zhang
WHI Acknowledgment:
The WHI program is funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, U.S. Department of Health and Human Services through contracts HHSN268201600018C, HHSN268201600001C, HHSN268201600002C, HHSN268201600003C, and HHSN268201600004C. We thank the WHI investigators and staff for their dedication, and the study participants for making the program possible. A full listing of WHI investigators can be found at: http://www.whi.org/researchers/Documents%20%20Write%20a%20Paper/WHI%20Investigator%20Long%20List.pdf
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Evans, L.M., Tahmasbi, R., Jones, M. et al. Narrow-sense heritability estimation of complex traits using identity-by-descent information. Heredity 121, 616–630 (2018). https://doi.org/10.1038/s41437-018-0067-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-018-0067-0
This article is cited by
-
Evaluation of novel genomic markers for pedigree construction in an isolated population of Weddell Seals (Leptonychotes weddellii) at White Island, Antarctica
Conservation Genetics Resources (2022)
-
Rapid detection of identity-by-descent tracts for mega-scale datasets
Nature Communications (2021)
-
Across the great divide: pluralism and the hunt for missing heritability
Synthese (2021)
-
Genetic predictors of educational attainment and intelligence test performance predict voter turnout
Nature Human Behaviour (2020)
-
Detecting the dominance component of heritability in isolated and outbred human populations
Scientific Reports (2018)