Abstract
Invasive species exert a serious impact on native fauna and flora and have been the target of many eradication and management efforts worldwide. However, a lack of data on population structure and history, exacerbated by the recency of many species introductions, limits the efficiency with which such species can be kept at bay. In this study we generated a novel genome of high assembly quality and genotyped 4735 genome-wide single nucleotide polymorphic (SNP) markers from 78 individuals of an invasive population of the Javan Myna Acridotheres javanicus across the island of Singapore. We inferred limited population subdivision at a micro-geographic level, a genetic patch size (~13–14 km) indicative of a pronounced dispersal ability, and barely an increase in effective population size since introduction despite an increase of four to five orders of magnitude in actual population size, suggesting that low population-genetic diversity following a bottleneck has not impeded establishment success. Landscape genomic analyses identified urban features, such as low-rise neighborhoods, that constitute pronounced barriers to gene flow. Based on our data, we consider an approach targeting the complete eradication of Javan Mynas across Singapore to be unfeasible. Instead, a mixed approach of localized mitigation measures taking into account urban geographic features and planning policy may be the most promising avenue to reducing the adverse impacts of this urban pest. Our study demonstrates how genomic methods can directly inform the management and control of invasive species, even in geographically limited datasets with high gene flow rates.
Similar content being viewed by others
Introduction
Biological invasions have far-reaching impacts on native biodiversity and cause lasting economic damage across many nations (Vitousek et al. 1997; Pimentel et al. 2005; Kumschick and Nentwig 2010). In human-modified landscapes, such as urban and suburban environments, the introduction of urban pests is often thought to exacerbate the decline and displacement of native species, which are generally slower to adapt to anthropogenic changes (Sala et al. 2000; Gurevitch and Padilla 2004). The challenge of eradication or management of biologically invasive species remains a problem that has hitherto been met only with mixed success by government agencies across the world.
Although genetic data have previously been used to inform management and eradication practices of pest species (Baker and Moeed 1979, 1987; Fleischer et al. 1991; Dlugosch and Parker 2008), the range of their inferences has been limited, possibly because of extremely small genetic sample sizes associated with traditional population-genetic methods which use markers such as microsatellites or single-locus DNA sequences. These limitations have been overcome with recent technological advances in sequencing technology (Baird et al. 2008; Emerson et al. 2010; Hohenlohe et al. 2010; Peterson et al. 2012) that have granted population geneticists a newly-accessible wealth of genomic SNP markers. The substantial increase of analyzable loci has conferred a significantly improved ability to infer demographic histories and study population connectivity and structure (Cristescu 2015; Rius et al. 2015) as compared to earlier studies that reported difficulties in reconciling genetic inferences with historical information using, for example, microsatellites (Estoup et al. 2010; Barun et al. 2013).
In the tropics, mynas of the songbird genus Acridotheres are some of the most successful invasive species (Feare and Craig 1998; Lowe et al. 2000; Wells 2010). Various governments have tried to eradicate and/or manage invasive myna populations with great difficulty and limited success. For example, the spread of the Common Myna A. tristis across the Australian continent has not been halted despite millions of dollars of funding and decades of effort and management (Martin 1996; Pell and Tidemann 1997; Grarock et al. 2012). Similarly, the introduced Javan Myna A. javanicus population in Singapore has been expanding despite years of active management by authorities in this small island city nation (719.1 km2) (Hails 1985; Nee 1989; Lee and Nee 1990; Chong et al. 2012).
The Javan Myna was first recorded on Singapore Island occurred between 1920 and 1921 (Lever 2010), but numbers remained relatively low as late as the 1960s (Ward 1968). However, by the mid-1980s, Javan Mynas outnumbered previously dominant Common Mynas 2.7-fold (Hails 1985). Ostensibly, this change had much to do with the urbanization of Singapore, which replaced large natural and agricultural areas with urban spaces especially between the 1950s and 1990s (Corlett 1992). Today, estimates place the burgeoning Javan Myna population at around 230,000 (Chong et al. 2012), an increase from the 2002 estimate of 139,000 (Lim et al. 2003).
The exact circumstances of the Javan Myna’s introduction remain unclear, and the size of the founding population is unknown. More importantly, uncertainty surrounds the levels of connectivity between different myna populations in Singapore and their ability to recolonize areas in which stringent management has successfully kept populations low. Specifically, one of the authorities’ most vexing management questions is whether Javan Mynas are widely dispersive and able to spread over large parts of the island within short periods of time, or whether they are predominantly resident in their small foraging and roosting home patches. The answer to this question has important direct management implications, as management responses to problematic population clusters of Javan Mynas would need to be designed differently depending on the birds’ intrinsic dispersal ability.
Previous research has tried to address questions about invasive mynas’ dispersal capabilities using radio-tracking (Yap 2003). However, currently available radio-tags for birds have a limited lifespan and such studies are only able to follow a limited number of individuals for a relatively short time, less than a generation in length. As a result, it is difficult for them to account for behavioral changes due to anthropogenic causes such as urban development. Furthermore, radio-tracking studies are unable to shed light on the spatial extent of multi-generational gene flow as a result of such dispersal. In contrast, Next-Generation Sequencing technology has made it possible to sample thousands of loci from across the genomes of dozens of individuals, making it possible to infer patterns of gene flow and reproductive connectivity among myna individuals across a large matrix of space, for example on the basis of powerful population-genomic and landscape-genomic methods, as well as coalescent simulations (Cristescu 2015; Keis et al. 2013; Peterson et al. 2012; Rius et al. 2015; Rollins et al. 2016).
The genomic approach is destined to be a preferred method in government authorities’ arsenal of tools to combat invasive species as it permits them to render their management approach more effective. For instance, if genomic data indicate a localized population structure with limited dispersal among clusters, targeted eradication such as culling may result in a complete eradication of the invasive species if well planned and carried out rigorously (Abdelkrim et al. 2005; Cook et al. 2010). Conversely, in the case of widespread connectivity across the target range, a complete eradication of the invasive species would be impossible to achieve, and authorities would be better off tailoring their approach to reducing numbers in particular focal areas of nuisance (Rollins et al. 2009). In such cases, city-wide education programs and laws discouraging and penalizing feeding or the provision of nesting opportunities would also be recommended courses of action.
In the present study, we showed that genome-wide sequence data from across dozens of individuals is able to reveal useful demographic information even in high gene-flow invasive systems within geographically-limited datasets. We sequenced a novel whole genome for the Javan Myna at 103 × coverage and generated thousands of genome-wide markers using a double-digest restriction enzyme-associated sequencing (ddRAD-Seq) approach from 78 Javan Mynas across Singapore. We assessed their population structure and level of connectivity at a microgeographic scale through the use of multiple contemporaneous population-genomic approaches. Additionally we inferred their demographic history, assessing whether the known trajectory of demographic expansion based on historic records and modern surveys (Ward 1968; Hails 1985; Lim et al. 2003; Lever 2010; Chong et al. 2012) has led to a corresponding increase in population-genetic diversity. We expect that knowledge about the Javan Myna’s spatio-ecological dynamics based on population-genomic insights and understanding of historical demography will not only aid in reaching sound management decisions, but also provide the basis for future work into genomic research into adaptive responses to urban lifestyle.
Materials and methods
Tissue sampling
All samples were collected in Singapore between January 2011 and August 2014. We obtained 105 Javan Myna samples (Supplementary Figure S1 and Supplementary Table S1) in the form of blood, liver, or breast muscle tissue from individuals collected either as (i) road kill, window strike, or cull victims, or (ii) with mist-nets (one location, Dairy Farm Nature Park, Singapore). In addition, we trapped a single individual for reference genome sequencing using a simple box-and-stick trap (Sunplaza Park, Singapore). All sampling protocols were in accordance with institutional ethics.
Reference genome sequencing and assembly
Genomic DNA was extracted from fresh breast muscle tissue from one individual of Javan Myna using the KingFisher™ Duo Prime Magnetic Particle Processor (Thermo Fisher Scientific, Waltham, MA, USA) and the KingFisher Cell and Tissue DNA Kit (Thermo Fisher Scientific), following the manufacturer´s protocol. Preparation of libraries, sequencing and the assembly of the de novo genome was performed by Science for Life Laboratory (SciLifeLab) in Stockholm. We constructed one standard library (short-insert-size, 180 bp) and two mate-pair libraries (5 and 8 kb). All libraries were sequenced on the Illumina HiSeq 2500 platform with a 2 × 126 setup in RapidHighOutput mode. After filtering out low quality and clonally duplicated reads, a total of 130 Gb (103×) was obtained for de novo assembly of the Javan Myna genome. Paired-end sequence data from the genomic DNA libraries were assembled using three short oligonucleotide analysis packages:ALLPATHS-LG (Gnerre et al. 2011), ABySS (Simpson et al. 2009), and SOAPdenovo (Li et al. 2010). Assembly quality and completeness were assessed by checking all read pair coverage information and supporting evidence. In addition, to evaluate the assembly correctness, we used feature-response curves (Vezzi et al. 2012) to plot regions of suspected mis-assemblies (features) against the coverage depth (Supplementary Figure S2). In this analysis, allowing a maximum number of suspected mis-assemblies (features), the y-axis reports the approximate genome coverage (%) achieved by all contigs (sorted in decreasing order by size) with a total number of errors/features equal to or less than the limit imposed. Only contigs longer than 1000 bp were used during validation. The ALLPATHS-LG assembly consistently presented the best assembly, generating better genome coverage with fewer suspect errors relative to the other two, and was used for subsequent analysis.
ddRAD-Seq library sequencing
DNA was obtained from liver tissue using the DNEasy Blood & Tissue Kit (Qiagen, Hilden, Germany), while DNA from all other tissue types was extracted using the Exgene Clinic SV Kit (GeneAll Biotechnology, Seoul, South Korea) following manufacturer’s protocols and overnight Proteinase K digestion to maximize DNA yield.
We used combinatorial indices and barcodes derived from Peterson et al. (2012) to uniquely identify each sequenced individual in our final multiplexed ddRAD-Seq library. Library preparation followed Tay et al. (2016) with the following modifications: (i) 150 ng of DNA from each individual were simultaneously double-digested with restriction enzymes and ligated to adapters, in triplicate, and (ii) the targeted fragment size range was 250–650 bp. We followed the original protocol and PCR-amplified size-selected DNA with 20 amplification cycles in triplicate before pooling replicate sample libraries and performing a second clean-up with size-selection to remove unannealed adapters and PCR primer dimers. Sample library fragment size distributions were checked using a Fragment Analyzer (Advanced Analytical Technologies, Ankeny, IA, USA), and final library concentrations were measured with a Qubit® 2.0 Fluorometer (Thermo Fisher Scientific). Eight samples were discarded due to improper fragment size selection (long tails > 650 bp) or insufficient total DNA concentrations in final libraries.
Equimolar volumes of the remaining 97 sample libraries were pooled to provide 20 times the base Illumina molar requirements and sequenced on one lane of an Illumina HiSeq 2500 machine at the Singapore Centre on Environmental Life Sciences Engineering (SCELSE, Singapore) to produce 150 bp paired end reads.
Quality filtering and demultiplexing
We used FastQC (Babraham Bioinformatics) to analyze sequence quality. Sequence read demultiplexing, cleanup, read-end truncation, and correction of single-nucleotide errors within barcodes was conducted using the program process_radtags in Stacks v1.34 (Catchen et al. 2011; Catchen et al. 2013). The final demultiplexed and quality filtered dataset consisted of almost 210 million reads, each 120 bp long. The number of reads per individual ranged from a minimum of 410,000 to a maximum of 4,006,000. As the individual with the fewest retained reads had less than half the number possessed by the next lowest (410,000 compared to 1,051,000), it was excluded from downstream analysis, leaving 96 individuals.
Reference genome alignment, SNP calling and quality filtering
We used BWA-MEM v0.7.12 (Li 2013) to index the genome assembly and subsequently aligned demultiplexed ddRAD-Seq reads against it. We then filtered out aligned reads with a MAPQ score lower than 20 using SAMtools v1.2 (Li et al. 2009). Reads were subsequently exported into bam format and sorted into coordinate order.
The pipeline ref_map.pl in Stacks v1.34 (Catchen et al. 2013) was used to call single nucleotide polymorphic markers (SNPs). To do this, three routines within the pipeline handled the data as follows. Using pstacks, we first grouped identical reference-aligned sequence reads into sets, accepting generated ‘stacks’ as assembled loci. After preliminary testing to explore effects of parameters on SNP yields, we used a minimum stack depth of 5 and excluded all stacks with lower coverage. Mean coverage depth for each stack was 19.25 calculated across all individuals. pstacks then proceeded to call SNPs for each individual using the default SNP model with a chi-square significance level of 0.05 to make heterozygous calls. Output from pstacks was used in cstacks to create a catalog of consensus loci and merge alleles based on genomic location. This was then used in sstacks as a ‘reference’ to map sample loci against, after which genomic location data was used to verify that (i) all loci matched a single catalog locus, (ii) all SNPs were accounted for in the catalog, and (iii) at least one catalog haplotype matched each of the query locus haplotypes. Loci which did not meet these criteria were removed. In all, the reference-aligned assembly produced around 196 000 loci.
We used the populations module in Stacks v1.34 to retain loci that were found in more than 90% of individuals. As a first step to reduce the effects of linkage disequilibrium in subsequent analyses, we only accepted one SNP from each locus using the –write_single_snp option provided. We retained 5921 SNPs after this step.
Investigation of population genetic structure
File format conversions necessary for all subsequent analyses were conducted using PGDSPIDER v2.0.7.4 (Lischer and Excoffier 2012).
We used PLINK v1.9 (Purcell et al. 2007; Chang et al. 2015) to retain only individuals with less than 10% missing data across all loci. We removed previously unfiltered loci under strong linkage disequilibrium by using a 25 bp window frame sliding 10 bp at a time. Pairwise SNPs with squared correlation (r 2) greater than 0.9 were greedily pruned using this approach, leaving 4735 SNPs from 78 individuals. We did not run a filter for SNPs under selection because our widely dispersed, non-equilibrium (recently introduced) dataset would have led to unacceptably high false positive rates (Lotterhos and Whitlock 2014). In any case, the patterns that emerge from our dataset argue against selection playing a main role (see Results).
We next performed a maximum likelihood estimation (Milligan 2003; Choi et al. 2009) of pairwise relatedness (r) in all retained samples using SNPRelate (Zheng et al. 2012). Only one pair of individuals (C24 and C158, r = 0.2349) was determined to be related at the equivalent level of half-siblings, but was not removed from the dataset because downstream exploration of the data did not reveal any clustering or grouping patterns closely associated with this pair alone.
We used GENALEX 6.5 (Peakall and Smouse 2006; Peakall and Smouse 2012) to perform principal coordinate analysis (PCoA) using calculated pairwise codominant genetic distances (Smouse and Peakall 1999), and to calculate expected heterozygosity (H e ), observed heterozygosity (H o ), and fixation index (F) (Hartl and Clark 1997).
We assessed population subdivision of Javan Mynas in Singapore using a model-based clustering approach implemented in STRUCTURE v2.3.4 (Pritchard et al. 2000). STRUCTURE runs were implemented without a priori hypotheses of cluster membership. We ran STRUCTURE from K 1 to 10 with ten iterations per K. For each iteration we implemented a burnin of 100,000 generations and MCMC for 500,000 generations. We used two methods to determine the most explanatory K-value for the dataset, starting by using the STRUCTURE Harvester Web v0.6.94 (Earl and vonHoldt 2012) implementation of the Evanno method (Evanno et al. 2005), which attempts to statistically calculate the most likely number of genotypic clusters (K) in the dataset. Subsequently, we also subjectively compared STRUCTURE plots across K-values to determine the most ecologically meaningful value of K. Results were averaged across replicates by evaluating individual ancestry coefficients (q values) with CLUMPP v1.1.2 (Jakobsson and Rosenberg 2007) using the Greedy option provided.
A second complementary, non-population model-based approach was conducted with NetView v.1.1, using network theory to construct networks of individuals in order to depict the connectivity and information flow within and between populations based on genetic similarity (Neuditschko et al. 2012; Steinig et al. 2016). In this analysis, network topologies are explored using a single user-defined threshold parameter, the number of mutual nearest neighbors (k). Fewer individuals are considered nearest neighbors at small values of k, leading to only genetically more similar individuals being connected and highlighting finer-scale structure in the dataset. Conversely, more individuals are connected at higher values of k, causing deeper community or population-wide differences to stand out in the resultant topology. To determine an appropriate k-value for our dataset, we first used multiple cluster detection algorithms (Fast-Greedy, Infomap, Walktrap) to visualize how the choice of k affects construction and structure of the mutual k-nearest neighbor graphs generated (Clauset et al. 2004; Pons and Latapy 2005; Wakita and Tsurumi 2007; Rosvall and Bergstrom 2008). Using the resultant k-selection plot as a guide (see Supplementary Figure S3a), we then explored population structure at different levels of genetic similarity by first focusing on more closely related samples at small values of k (k = 10) before moving to broader patterns of population structure in the dataset at higher values of k (k = 20, 30) (Neuditschko et al. 2012). We plotted the generated graphs using the Kamada-Kawai force-directed graph drawing algorithm (Kamada and Kawai 1989) as implemented in iGraph (Csardi and Nepusz 2006).
Investigation of population spatial structure
We conducted multivariate global spatial autocorrelation analysis in GENALEX 6.5 (Smouse and Peakall 1999; Peakall et al. 2003; Double et al. 2005; Peakall and Smouse 2006, 2012) to explore spatial patterns of genetic structure. We performed global autocorrelations to obtain evidence of fine scale population subdivision at a microgeographic scale and local spatial autocorrelations to obtain an estimate of genetic patch size. For each distance class, we also obtained an estimate of genetic similarity between individuals by calculating the autocorrelation coefficient r (Sokal and Wartenberg 1983; Smouse and Peakall 1999; Peakall et al. 2003; Peakall and Smouse 2006, 2012). Briefly, we first performed random permutations (999 permutations) and obtained 95% confidence intervals (CI) around r assuming no spatial autocorrelation (r p , null distribution). In a two tailed test we inferred significant global autocorrelation if observed r (r bs, observed distribution) fell outside this confidence interval. Further, within each distance class we tested for significance of r bs by performing bootstrapping (10,000 bootstraps). We considered r bs as significant whenever the bootstrap confidence intervals did not include zero.
To determine the appropriate range for which to apply this analysis, we first estimated the true extent of detectable non-random spatial structure by calculating r bs and its associated 95% CI and null hypothesis for the first interval of increasing distance classes in 1 km intervals, spanning the range of pairwise geographical distances following Peakall et al. (2003). Due to the cumulative nature of pairwise combinations available for each step of this analysis, we were able to estimate r p using 9999 permutations instead of 999. We considered the distance class at which r bs is no longer significant as the distance at which non-random genetic spatial structure ceases to be detectable and set this as the maximum distance (28 km) considered subsequently (see Fig. 4a). Finally, we explored spatial structure using 23 distance classes from 1 to 12 km at intervals of 0.5 km. Distance classes were chosen in this manner to explore how the value of the first x-intercept changed with increasing distance class size, a quantity often considered to represent the diameter of a genetic “patch” (Sokal and Wartenberg 1983; Smouse and Peakall 1999; Diniz-Filho and Telles 2002; Krauss and Koch 2004).
To test the presence of corridors and barriers to gene flow, we mapped the pairwise residuals of isolation-by-distance (IBD) using the distribution of residual dissimilarity (DResD) procedure from Keis et al. (2013). We used the pairwise genetic distance matrix calculated in GENALEX 6.5 as data input for the initial regression accounting for IBD, and ran three separate DResD analyses at distance scales informed by our calculated genetic patch size (13.5 km, see Results) in order to maximize the signal of any corridors or barriers present across the sampled range. We set 0.5 km as the minimum distance between points to exclude pairwise genetic distances of individuals from the same location. The three DResD analyses included genetic distance values for individuals (1) 0.5–13.5 km, (2) >13.5 km, and (3) >0.5 km apart. For each DResD analysis, we interpolated the residual variance on a 500 m by 500 m grid. We ran 1000 random iterations and 250 bootstraps to identify areas where residual values significantly deviated from the expected values derived from a null model of zero population structure.
Inference of historical demography
We explored three simple models of demographic history using Approximate Bayesian Computation (ABC) as implemented in DIYABC v2.1 (Cornuet et al. 2014) to determine the most likely population trajectory and its associated parameters (time of event, t, and effective population size, N e ; Fig. 1). We employed a common starting scenario where ancestral effective population size (N anc) first decreases at the time of introduction (t 2) to the effective population size during bottleneck (N bot), before undergoing one of three possible scenarios at t 1 following the bottleneck: (1) population expansion, i.e., a great increase in effective population size at t 1 to a present-day value (N exp) larger than N anc (N exp ≥ N anc), (2) population stability/recovery, i.e., a modest increase to a value (N rec) between N bot and N anc in magnitude (N anc ≥ N rec ≥ N bot), or (3) population contraction, i.e., a further decrease in effective population size to N con (N bot ≥ N con) (Fig. 1). Log-uniform prior distributions were set with the following ranges: 10 ≤ N anc ≤ 500,000; 2 ≤ N bot ≤ 1000; 2 ≤ N exp ≤ 500,000; 2 ≤ N rec ≤ 500,000; 2 ≤ N con ≤ 1000; 2 ≤ t 1 ≤ 10,000; 10 ≤ t 2 ≤ 10,000. A subset of 3808 SNPs that had a minimum allele frequency of at least 0.01 was considered for this analysis.
In a graphical representation of three demographic scenarios tested using an approximate Bayesian computational framework, ancestral populations with an unknown effective population size (N anc) first experience a population bottleneck (N bot) at time t 2, before undergoing either expansion (N exp; N exp ≥ N anc), stabilization/recovery (N rec; N anc ≥ N rec ≥ N bot), or contraction (N con; N bot ≥ N con) at time t 1 , and thereafter remain constant until sampled at the point of study. The width of each scenario line is relative to its effective population size
Coalescent simulations were first run with an even distribution across all scenarios (500,000 runs per scenario). For each simulation, we obtained the following one-sample summary statistics (SS): mean (Nei 1987) and variance of gene diversity across polymorphic loci, and mean gene diversity across all loci. We used the logistic regression estimate implemented in DIYABC to obtain the posterior probabilities of each scenario (Cornuet et al. 2008, 2010). This method estimates the posterior probability of the closest-to-observed 1% of simulated datasets (15,000 datasets) by subjecting the SS of each simulation to a polychotomous logistic regression. We chose the optimal scenario as the one with the highest posterior probability value with a non-overlapping 95% CI. To evaluate confidence in the optimal scenario, we analyzed 500 pseudo-observed datasets (PODs) closest to the observed dataset and calculated the posterior predictive error. Finally, we evaluated whether the model-posterior distribution combination generated by our optimal scenario was better able to reproduce our observed dataset than competing scenarios. Using the Model Check option in DIYABC, we ran principal component analysis (PCA) on the SS from two sets of 1000 PODs simulated from the prior and posterior predictive parameter distributions of each scenario.
We estimated the posterior distributions of demographic parameters for the optimal scenario using the 1% closest-to-observed simulated datasets by performing a logit transformation of parameters and subsequent local linear regressions. In order to evaluate our confidence in parameter estimates, we used the estimated parameter distributions from 500 PODs simulated for the optimal scenario to calculate the mean and median relative bias, as well as the square root of mean square error. This provided a more relevant analysis of parameter estimate accuracy compared to drawing parameters for simulation from across the wide prior distribution space (Cornuet et al. 2014).
Finally, we estimated present day effective population size independently with the heterozygote excess (Zhdanova and Pudovkin 2008) and molecular coancestry (Nomura 2008) methods as implemented in NeEstimator v2 (Do et al. 2014) using a minimum 0.05 allele frequency.
Results
Novel genome assembly
We sequenced and assembled a de novo genome for the Javan Myna from 130 Gb (103 × coverage). Of the three assemblies generated, the ALLPATHS-LG assembly was consistently better across several standard contiguity metrics (Table 1). It produced the fewest scaffolds (4312), the highest proportion of scaffolds longer than 1000 bp, and the highest N50 statistic (5.4 Mb) among the three, indicating assemblies with the highest connectivity. Ranking of assemblies using a feature-response curve clearly indicated that the ALLPATHS-LG assembly outperformed the rest at reconstructing the genome with fewer features or errors (Supplementary Figure S2). Our genome assembly compares favorably to a number of recent avian genome assemblies, particularly those of other songbirds (Table 2).
Population genetic structure
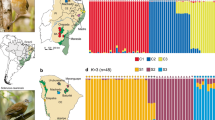
The Singaporean Javan Myna population is near panmictic, with several analyses providing evidence for this result (Fig. 2). PCoA showed a tight cluster containing the majority of individuals and several outliers (Fig. 2a) when the first two axes explaining the most variation in the dataset were plotted. PCoA outliers did not share a common geographic location, instead comprising the individuals with the highest levels of missing data. These individuals remained as outliers even when the next two sets of axes explaining the most variation were plotted (not shown). Running a PCoA without these outliers did not reveal any hidden geographic pattern in the data (Supplementary Fig. S4).
Population subdivision was explored with several complementary analyses. a Principle coordinate analysis (PCoA) plot using codominant genetic distances. Percentage of total variation explained by each axis shown in brackets. The spread of points along axis 1 reflects a trend in missing data, although total missing data remains < 10% per individual. b Averaged results of ten iterations per K-value in STRUCTURE for K = 1 to 3. A strong panmictic signal is evident in the dataset, with no additional population subdivision revealed as K increases. c Kamada-Kawai force-directed depictions of mutual k-nearest neighbor network graphs for k = 10 show a network with high interconnectivity within a single cluster
Although the optimal number of K-clusters in STRUCTURE was determined to be K = 2 according to the Evanno method (Evanno et al. 2005) (Supplementary Fig. S5), we found that exploration of plots at K values above K = 1 did not reveal any subdivision across Singapore (Fig. 2b). This discrepancy is partially explained by the fact that an optimal cluster value of K = 1 cannot be detected with the Evanno method. Additional clusters at higher values of K were assigned in horizontal tranches at almost equal proportions across individuals and may or may not reflect disparate contributions from ancestral populations at uniform levels across Singapore.
Mutual k-nearest neighbor network graphs generated in NetView further corroborate these broad results of population genetic structure. When drawn at a low k-value (k = 10, Fig. 2c) to attempt separating the most genetically similar individuals into distinct communities (Neuditschko et al. 2012; Steinig et al. 2016), a pattern of high interconnectivity in a single large cluster emerged regardless. Further exploration at increasing k-values (k = 20, 30, Supplementary Fig. S3) showed a centralizing trend to the pattern of connectivity between individuals. No consistent geographical patterns were detectable in networks at any value of k upon inspection.
We estimated observed heterozygosity H o (x̅ = 0.151, SE = 0.003), expected heterozygosity H e (x̅ = 0.139, SE = 0.002), and fixation index F (x̅ = −0.017, SE = 0.003). We observed that mean H o was higher than H e , and that fixation index F was negative and significantly different from zero (95% CI = [−0.023, −0.011]).
Population spatial structure
Results from spatial autocorrelation tests revealed the presence of positive genetic spatial structure up to a maximum of 28 km (Fig. 3a). Single distance class correlograms were generated by global spatial autocorrelation across a range of distance classes from 1 km to 12 km at 0.5 km increments. Nearly all distance classes analyzed displayed a long distance cline, indicating isolation by distance in the dataset (Diniz-Filho and Telles 2002) (Fig. 3b). We determined the genetic patch size to be 13.50 km (SD = 2.3056, SE = 0.4808, 95% CI = [12.50, 14.49]) based on x-intercept values across all analyzed distance classes.
In these correlograms, the autocorrelation coefficient r is denoted by blue lines and bounded by a 95% CI determined by bootstrapping. It is considered significant when it lies above or below the 95% CI about the null hypothesis of no genetic spatial structure in the dataset, as denoted by the red lines (U: upper bound, L: lower bound). a The true extent of detectable spatial genetic autocorrelation is estimated by calculating the correlation coefficient r for the first interval of increasing distance classes, and is around 28 km. b In a representative 5.5 km even distance class correlogram, the trajectory of the autocorrelation coefficient (r, blue solid line) gradually goes from being positively significant to negatively significant, indicative of isolation by distance in the dataset (Diniz-Filho and Telles 2002). The average x-intercept distance across all distance classes tested in this study was 13.50 km (SD = 2.3056, SE = 0.4808, 95% CI = [12.50, 14.49])
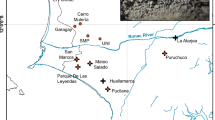
At above-patch distances (>13.5 km), DResD analysis indicated significantly higher interpolated residual values than expected from the null model in the central to southwest parts of Singapore (Fig. 4). These residual values had good bootstrap support, and suggest the presence of a barrier to gene flow. Comparatively, no significant residual values were detected across Singapore for analysis at below-patch distances (0.5–13.5 km) (Supplementary Fig. S6a), while significantly high residual values were also detected in the universal dataset within a smaller area in central and southern Singapore (Supplementary Fig. S6b).
Distribution of residual dissimilarity (DResD) plots overlaid on map of Singapore for combinations with minimum pairwise distance of 13.5 km. The intensity of pink and blue background colors indicate resistance to gene flow from high to low, respectively (intensity corresponds to residual deviation values in legend). The solid blue line denotes statistically significant areas of the analysis (P < 0.05), while the solid red lines denote areas with significant post hoc statistical power (based on 1000 iterations). Sampling localities are depicted as yellow dots. Significant barriers to gene flow are detectable in the central to southwest area of Singapore Island. Black, dark gray, and light gray areas represent government-recognized nature reserves, designated public parks, and unmanaged forested vegetation respectively
Demographic history
ABC analysis strongly supported the population stability/recovery scenario (Table 3). We estimated a posterior predictive error of 0.262 for the logistic scenario choice. All summary statistics were significantly discriminatory between the scenarios tested (Supplementary Table S2).
We also estimated the times of events (t 1 , t 2 ) and N e for the ancestral (N anc ), founding (N bot ), and current (N rec ) populations (Table 4) for the most probable demographic scenario. As the distributions of all posterior parameter estimates were right-skewed, we considered the modes of the distributions to be the best estimates of the actual parameters. Present day estimates of N e corresponded well with those from NeEstimator in the low single or double digits (Table 4).
In short, we inferred that Javan Mynas underwent a population bottleneck within the last century based on the magnitude of t 2 (149 generations, Table 4) and considering that Javan Mynas breed at least once a year in Singapore (Nee 1989). At the same time, N e across Singapore dropped to less than ten during this bottleneck and has subsequently barely increased up to the present day. Bias and error estimates for posterior parameter distributions for the optimal scenario were generally close to zero and are available in Supplementary Table S3.
Discussion
Novel avian genome
In this study, we sequenced the first Acridotheres myna genome at 103 × coverage. This novel genome will be essential for future research on the genomic origins of the Javan Myna’s remarkable plasticity and unique life-history adaptations that have enabled it to become such a successful colonizer and pest species. This also represents the closest available reference genome to date for SNP mapping in the Common Myna, another important invasive species in many parts of the world, and additional sturnid model species. In this study, we expected a large proportion of linked loci due to the recent bottleneck undergone by the study population, and the new genome proved invaluable by allowing the removal of over ~1000 linked SNPs which would have otherwise biased population-genomic signals revealed by subsequent analyses.
Genetic connectivity of Javan Mynas across Singapore
Our genome-wide marker set, based on over ~4700 neutral SNPs, reveals a general lack of population subdivision across Singapore, as demonstrated by a suite of different population-genomic methods ranging from STRUCTURE to network plots and principal component analysis (Fig. 2). This general homogeneity is suggestive of a single introduction event, given its recent timing according to known records.
In order to further shed light on the gene flow patterns of Javan Mynas across local scales spanning only a few kilometers, we turned to other approaches measuring the mynas’ “genetic patch size”. This refers to the diameter of an idealized circular area within which individuals are not genetically independent. In other words, any individual sampled within the estimated patch is more likely to be related to another individual sampled inside the area and less likely to be related to individuals sampled outside than would be expected by chance (Sokal and Wartenberg 1983). This information is a crucial prerequisite to determining whether a complete eradication approach is viable (in the case of non-dispersive, localized populations) or whether range-wide mitigation measures accompanied by targeted control in focal areas of nuisance is a better choice for resource investment (in the case of pronounced connectivity) (Abdelkrim et al. 2005; Rollins et al. 2009). Using genome-wide SNPs and spatial autocorrelation, we computed a genetic patch size for Javan Mynas across Singapore of ~13.5 km (95% CI = [12.50, 14.49]) (Fig. 3b).
The genomic estimate of Javan Myna patch size is easily reconciled with life history information about this species in two main ways. Previous studies have shown that hundreds to thousands of individuals roost communally every night and then disperse to their individual foraging areas during daytime (Nee et al. 1990; Yap et al. 2002). In these roosting colonies, individuals socialize and find breeding partners from a potential pool of mates, leading to a population structure in which there is a spatial autocorrelation commensurate with the average approximate distance between roost sites. In addition, a radiotracking study of seven Javan Mynas in Singapore previously revealed a maximum ranging distance from roost sites between 100 m and 2 km (Yap 2003). This information about the typical ranging behavior of the species is several times less than our estimate for patch size and therefore in agreement, since a genetic patch is reflective of genetic relatedness in a network of inter-related individuals as opposed to that of an individual and its nearest neighbors.
Barriers to gene flow at a fine geographic scale
Our genomic data was further able to detect a barrier to gene flow in the central to southwestern areas of Singapore (Fig. 4). This barrier was statistically significant only in analyses that included pairwise combinations across distances larger than the calculated genetic patch size (13 km), and had the clearest signals when combinations below this distance were excluded. The areas in which the barrier is located are relatively less densely populated by humans, dominated by a mix of low-rise housing estates and parkland, and have correspondingly fewer open-air eateries, which are otherwise common across Singapore (Singapore Department of Statistics 2016a, b).
Refuse from human eateries is a major but ephemeral source of food for Javan Mynas, and mynas in areas with a high density of eateries typically forage dynamically over a large range in order to optimize feeding efficiency across short-lived high-reward sites (Nee 1989). Conversely, a lower density of open-air eateries decreases overall fluctuation in food availability, which is in turn likely to decrease territorial plasticity and dynamism as colonies defend stable foraging grounds (Kark et al. 2007; Hulme-Beaman et al. 2016). We hypothesize that territorial mynas may therefore form barriers for genetic connectivity when they defend their territories and reject new membership of extraterritorial mynas. Further investigation into how refuse management practices and urban planning create behavioral changes leading to genetic barriers within the detected sites will inform authorities seeking to limit the population growth and connectivity of Javan Mynas on the island. Additionally, the presence of the genetic barrier suggests that division of the island into distinct management units on either side of the barrier may be warranted.
Previous work involving genetic spatial autocorrelation and determination of patch size have generally made use of genotype data from traditional population genetic methods either for populations sampled across areas many times larger than the present study area, or species with limited dispersal capabilities relative to study area (Smouse and Peakall 1999; Diniz-Filho and Telles 2002; Peakall et al. 2003; Krauss and Koch 2004; Double et al. 2005; Beck et al. 2008). In this study, we extended the utility of this technique by showing an example of how genetic patch size determination using genomic data can directly inform the design of downstream landscape genomic analyses. This approach allows us to elucidate underlying spatial connectivity, population structure, and barriers to gene flow in invasive populations, even over small geographic scales, directly informing management and control measures.
Recovery of population-genetic diversity lags far behind population expansion
Almost a century after its initial introduction to Singapore and peninsular Malaysia, the Javan Myna has become the dominant introduced avian exotic (Lim et al. 2003) in these areas. Field censuses in Singapore estimated the local population at over 200 000 birds in 2012 (Chong et al. 2012), following a steady and increasingly aggressive expansion, especially over the last few decades (Ward 1968; Hails 1985; Lim et al. 2003; Lever 2010; Chong et al. 2012).
However, our ABC analysis combined with N e estimates from other sources indicates that effective population size of Singapore’s Javan Mynas has barely increased since the point of introduction (Table 4). These results concur with summary statistics corresponding to a recently bottlenecked population that has yet to reach a new equilibrium (see Results) (Cornuet and Luikart 1996). However, the true extent of the Javan Myna’s recent recovery may be masked as recent expansions are visible in genetic data mostly through an increase in rare variants that are only detectable using very high sample sizes at great coverage (Keinan and Clark 2012).
Our ABC estimate of a founding effective population size (N bot) in the single digits is reflective of the mode of introduction for this species. Javan Mynas would have been brought into Singapore as a stowaway on ships from Java, or more likely as part of the songbird trade, rather than for food or pest control as in other species such as the Rock Pigeon (Columba livia) or the Common Myna (Gibson-Hill 1950; Hails 1985; Lim et al. 2003). As a result, population founders were likely to be birds that managed to escape their cages. The situation in Singapore may therefore mirror the circumstances surrounding the introduction of Common Mynas to Durban, where a known number of a few founding members resulted in a correspondingly low effective population size even nearly a century post-introduction (Baker and Moeed 1987).
While severe genetic bottlenecks have traditionally been thought to inhibit invasion success by limiting the invading population’s ability to respond to selective pressures (Planes and Lecaillon 1998; Kinziger et al. 2011), the successful establishment of Javan Mynas in Singapore despite a persistently low effective population size appears to contradict this assumption and may instead indicate a bridgehead scenario (Lombaert et al. 2010; Lawson Handley et al. 2011). In such a scenario, an invasive species first establishes itself in an introduced range and evolves post-introduction adaptations that allow it to be a successful invader (Keller and Taylor 2008; Bock et al. 2015). Invasive populations that undergo this “evolutionary shift” are then able to subsequently colonize other areas without slowing down despite repeated bottlenecks. Our results are in good agreement with this concept, and we postulate that Javan Mynas in Singapore underwent an evolutionary shift in the last century around the time when their numbers began to rise dramatically. If correct, Javan Mynas from Singapore are now preadapted to invade other regional cities given the opportunity, without regard for further population bottlenecks. Their recent colonization of new urban areas in southern Thailand and Borneo is consistent with this scenario (Sontag 1998; Eaton et al. 2016).
Implications for management
One important goal of invasion biology is the characterization of dispersal in invasive systems (Lawson Handley et al. 2011). While estimates using gene flow are one widely-employed method of doing so, invasive systems are often not at migration-drift equilibriums. Additionally, many invasive populations relevant to management are confined to small, high-impact areas such as urban cities, which makes the detection of gene flow problematic. In this study on introduced Javan Mynas in Singapore, we sequenced the first Acridotheres myna genome and were able to elucidate key parameters describing historical demography and fine-scale population connectivity at small geographic scales in an otherwise homogenous non-equilibrium invasive system. We expect that our analytical approach will be widely emulated in other urban invasive systems in the future, eventually informing local management.
The demonstration of a relatively large genetic patch size (~13.5 km) and genetic connectivity across Singapore calls into doubt the possibility of a complete eradication of this species. Given the size of Singapore Island, which is ~50 km from east to west and ~27 km from north to south, our patch size estimate is relatively large, corroborating that a complete eradication effort, based on a succession of more localized eradication programs, is not feasible (Abdelkrim et al. 2005; Rollins et al. 2009). Indeed, the large patch size indicates that even local control of mynas will likely be an uphill challenge, since keeping priority areas free of mynas will require regular maintenance clearing of birds within a ~7 km radius to prevent recolonization. As a result, cost-benefit studies will have to be carefully considered when prioritizing myna-free zones.
Even with extraordinary effort and at extreme cost, a near-complete eradication across Singapore Island would likely be followed by a re-bounding through an influx of re-invasions from the uncontrolled population in nearby peninsular Malaysia. The Malaysian population (itself invasive) is thought to be derived from Singaporean birds, and its nearest occurrence to Singapore is less than 2 km as measured across the Johor Straits (Wells 2010). Instead, the preferred approach by Singaporean authorities will need to aim at mitigation of nuisance in areas that are especially problematic, coupled with a long-term program to curb the entire population by reducing nesting opportunities and food sources often provided by an unwitting public that is unaware of the negative effects of their well-meant actions.
Importantly, we were able to detect areas serving as genetic barriers and link them to features in the urban landscape that are associated with lower gene flow and presumably higher territoriality of Javan Mynas. We recommend that authorities consider designation of separate management units for different parts of the island and put the new knowledge about the position of specific gene flow barriers to use in management practices.
Future investigations into the genetic basis of life history traits that render Javan Mynas and their widespread cousins, the Common Mynas, so successful in colonizing new habitats will be able to make use of the novel genome, allowing us to understand the origins of traits that equip avian species for survival in anthropogenic environments.
Data archiving
The Javan Myna genome sequence is available under the accession number PEJO01000000 (GenBank). Other DNA sequences used in this study are available under NCBI Sequence Read Archive study number SRP121087 (BioProject PRJNA415335). Supplementary information is available at Heredity’s website.
References
Abdelkrim J, Pascal M, Calmet C, Samadi S (2005) Importance of assessing population genetic structure before eradication of invasive species: examples from insular Norway rat populations. Conserv Biol 19(5):1509–1518
Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA et al. (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers. PLoS ONE 3(10):e3376
Baker AJ, Moeed A (1979) Evolution in the introduced New Zealand populations of the common myna, Acridotheres tristis (Aves: Sturnidae). Can J Zool 57(3):570–584
Baker AJ, Moeed A (1987) Rapid genetic differentiation and founder effect in colonizing populations of common mynas (Acridotheres tristis). Evolution 41(3):525–538
Barun A, Niemiller ML, Fitzpatrick BM, Fordyce JA, Simberloff D (2013) Can genetic data confirm or refute historical records? The island invasion of the small Indian mongoose (Herpestes auropunctatus). Biol Invasions 15(10):2243–2251
Beck NR, Peakall R, Heinsohn R (2008) Social constraint and an absence of sex-biased dispersal drive fine-scale genetic structure in white-winged choughs. Mol Ecol 17(19):4346–4358
Bock DG, Caseys C, Cousens RD, Hahn MA, Heredia SM, Hübner S et al. (2015) What we still don’t know about invasion genetics. Mol Ecol 24(9):2277–2297
Catchen J, Amores A, Hohenlohe P, Cresko W, Postlethwait JH (2011) Stacks: building and genotyping loci de novo from short-read sequences. G3: Genes, Genomes, Genet 1(3):171–182
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013) Stacks: an analysis tool set for population genomics. Mol Ecol 22(11):3124–3140
Chang C, Chow C, Tellier L, Vattikuti S, Purcell S, Lee J (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4(1):7
Choi Y, Wijsman EM, Weir BS (2009) Case-control association testing in the presence of unknown relationships. Genet Epidemiol 33(8):668–678
Chong KY, Teo S, Kurukulasuriya B, Chung YF, Rajathurai S, Lim HC et al. (2012) Decadal changes in urban bird abundance in Singapore. The Raffles Bulletin of Zoology 25(1):189-196
Clauset A, Newman MEJ, Moore C (2004) Finding community structure in very large networks. Phys Rev E 70(6):066111
Cook AJ, Poncet S, Cooper APR, Herbert D, Christie D (2010) Glacier retreat on South Georgia and implications for the spread of rats. Antarct Sci 22(3):255–263
Corlett RT (1992) The ecological transformation of Singapore, 1819-1990. J Biogeogr 19(4):411–420
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144(4):2001–2014
Cornuet JM, Pudlo P, Veyssier J, Dehne-Garcia A, Gautier M, Leblois R et al. (2014) DIYABCv2.0: a software to make approximate Bayesian computation inferences about population history using single nucleotide polymorphism, DNA sequence and microsatellite data. Bioinformatics 30(8):1187–1189
Cornuet JM, Ravigné V, Estoup A (2010) Inference on population history and model checking using DNA sequence and microsatellite data with the software DIYABC (v1.0). BMC Bioinforma 11(1):1–11
Cornuet JM, Santos F, Beaumont MA, Robert CP, Marin JM, Balding DJ et al. (2008) Inferring population history with DIYABC: a user-friendly approach to approximate bayesian computation. Bioinformatics 24(23):2713–2719
Cristescu ME (2015) Genetic reconstructions of invasion history. Mol Ecol 24(9):2212–2225
Csardi G, Nepusz T (2006) The igraph software package for complex network research. Inter Complex Syst 1695(5):1–9
Diniz-Filho JAF, Telles MPDC (2002) Spatial autocorrelation analysis and the identification of operational units for conservation in continuous populations. Conserv Biol 16(4):924–935
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17(1):431–449
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimatorv2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14(1):209–214
Double MC, Peakall R, Beck NR, Cockburn A (2005) Dispersal, philopatry, and infidelity: dissecting local genetic structure in superb fairy-wrens (Malurus cyaneus). Evolution 59(3):625–635
Earl DA, vonHoldt BM (2012) STRUCTURE Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361
Eaton JA, van Balen S, Brickle NW, Rheindt FE (2016) Birds of the Indonesian Archipelago: Greater Sundas and Wallacea. Lynx Edicions, Barcelona, Spain
Ellegren H, Smeds L, Burri R, Olason PI, Backstrom N, Kawakami T et al. (2012) The genomic landscape of species divergence in Ficedula flycatchers. Nature 491(7426):756–760
Emerson KJ, Merz CR, Catchen JM, Hohenlohe PA, Cresko WA, Bradshaw WE et al. (2010) Resolving postglacial phylogeography using high-throughput sequencing. Proc Natl Acad Sci 107(37):16196–16200
Estoup A, Baird SJE, Ray N, Currat M, Cornuet JM, Santos F et al. (2010) Combining genetic, historical and geographical data to reconstruct the dynamics of bioinvasions: application to the cane toad Bufo marinus. Mol Ecol Resour 10(5):886–901
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620
Feare C, Craig A (1998) Starlings and mynas. A&C Black, London
Fleischer RC, Williams RN, Baker AJ (1991) Genetic variation within and among populations of the common myna (Acridotheres tristis) in Hawaii. J Hered 82(3):205–208
Gibson-Hill CA (1950) Myna matters. Malay Nat J 5(2):59–75
Gibson L, Yong DL (2017) Saving two birds with one stone: solving the quandary of introduced, threatened species. Front Ecol Environ 15(1):35–41
Gnerre S, MacCallum I, Przybylski D, Ribeiro FJ, Burton JN, Walker BJ et al. (2011) High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc Natl Acad Sci USA 108(4):1513–1518
Grarock K, Tidemann CR, Wood J, Lindenmayer DB (2012) Is it benign or is it a pariah? Empirical evidence for the impact of the common myna (Acridotheres tristis) on Australian birds. PLoS ONE 7(7):e40622
Gurevitch J, Padilla DK (2004) Are invasive species a major cause of extinctions? Trends Ecol Evol 19(9):470–474
Hails CJ. (1985) Studies on problem bird species in Singapore: 1. Sturnidae (mynas and starlings). Official report to the Ministry of National Development, Singapore, pp 97
Hartl DL, Clark AG (1997) Principles of population genetics, 3rd edn. Sinauer Associates, Inc, Sunderland, Massachusetts, Vol 116
Hillier LW, Miller W, Birney E, Warren W, Hardison RC, Ponting CP et al. (2004) Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 432(7018):695–716
Hohenlohe PA, Bassham S, Etter PD, Stiffler N, Johnson EA, Cresko WA (2010) Population genomics of parallel adaptation in threespine stickleback using sequenced RAD tags. PLoS Genetics 6(2):e1000862
Hulme-Beaman A, Dobney K, Cucchi T, Searle JB (2016) An ecological and evolutionary framework for commensalism in anthropogenic environments. Trends Ecol Evol 31(8):633–645
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14):1801–1806
Jarvis ED, Mirarab S, Aberer AJ, Li B, Houde P, Li C et al. (2014) Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346(6215):1320–1331
Kamada T, Kawai S (1989) An algorithm for drawing general undirected graphs. Inf Process Lett 31(1):7–15
Kark S, Iwaniuk A, Schalimtzek A, Banker E (2007) Living in the city: can anyone become an ‘urban exploiter’? J Biogeogr 34(4):638–651
Keinan A, Clark AG (2012) Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336(6082):740–743
Keis M, Remm J, Ho SYW, Davison J, Tammeleht E, Tumanov IL et al. (2013) Complete mitochondrial genomes and a novel spatial genetic method reveal cryptic phylogeographical structure and migration patterns among brown bears in north-western Eurasia. J Biogeogr 40(5):915–927
Keller SR, Taylor DR (2008) History, chance and adaptation during biological invasion: separating stochastic phenotypic evolution from response to selection. Ecol Lett 11(8):852–866
Kinziger AP, Nakamoto RJ, Anderson EC, Harvey BC (2011) Small founding number and low genetic diversity in an introduced species exhibiting limited invasion success (speckled dace, Rhinichthys osculus). Ecol Evol 1(1):73–84
Krauss SL, Koch JM (2004) Methodological insights: Rapid genetic delineation of provenance for plant community restoration. J Appl Ecol 41(6):1162–1173
Kumschick S, Nentwig W (2010) Some alien birds have as severe an impact as the most effectual alien mammals in Europe. Biol Conserv 143(11):2757–2762
Lawson Handley LJ, Estoup A, Evans DM, Thomas CE, Lombaert E, Facon B et al. (2011) Ecological genetics of invasive alien species. BioControl 56(4):409–428
Lee PG, Nee K (1990) The status of birds in Singapore - a brief perspective. In: Chou LM, Ng PKL (eds) Essays in Zoology, Papers Commemorating the 40th Anniversary of the Department of Zoology. National University of Singapore, Singapore
Lever C (2010) Naturalised Birds of the World. Bloomsbury Publishing, London, UK
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:13033997.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079
Li R, Zhu H, Ruan J, Qian W (2010) De novo assembly of human genomes with massively parallel short read sequencing. Genome Research 20(2):265-272
Lim HC, Sodhi NS, Brook BW, Soh MCK (2003) Undesirable aliens: factors determining the distribution of three invasive bird species in Singapore. J Trop Ecol 19(06):685–695
Lischer HEL, Excoffier L (2012) PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28(2):298–299
Lombaert E, Guillemaud T, Cornuet JM, Malausa T, Facon B, Estoup A (2010) Bridgehead effect in the worldwide invasion of the biocontrol harlequin ladybird. PLoS ONE 5(3):e9743
Lotterhos KE, Whitlock MC (2014) Evaluation of demographic history and neutral parameterization on the performance of F(ST) outlier tests. Mol Ecol 23(9):2178–2192
Lowe S, Browne M, Boudjelas S, De Poorter M (2000) 100 of the world’s worst invasive alien species: a selection from the global invasive species database. The Invasive Species Specialist Group (ISSG), a specialist group of the Species Survival Commission (SSC) of the World Conservation Union (IUCN), Auckland, New Zealand
Martin WK (1996) The current and potential distribution of the common myna Acridotheres tristis in Australia. Emu 96(3):166–173
Milligan BG (2003) Maximum-likelihood estimation of relatedness. Genetics 163(3):1153–1167
Nee K (1989) Comparative behavioural ecology of the mynas, Acridotheres tristis (Linnaeus) and A. javanicus (Cabanis) in Singapore. PhD thesis, National University of Singapore
Nee K, Sigurdsson J, Hails C, Counsilman J (1990) Some implications of resource removal in the control of mynas (Acridotheres spp.) in Singapore. Malayan Nature. Journal 44(2):103–108
Nei M (1987) Molecular Evolutionary Genetics.. Columbia University Press, New York, NY, USA
Neuditschko M, Khatkar MS, Raadsma HW (2012) NetView: a high-definition network-visualization approach to detect fine-scale population structures from genome-wide patterns of variation. PLoS ONE 7(10):e48375
Nomura T (2008) Estimation of effective number of breeders from molecular coancestry of single cohort sample. Evolut Appl 1(3):462–474
Peakall R, Ruibal M, Lindenmayer DB (2003) Spatial autocorrelation analysis offers new insights into gene flow in the australian bush rat. Ratt Fuscipes Evol 57(5):1182–1195
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28(19):2537–2539
Pell AS, Tidemann CR (1997) The ecology of the common myna in urban nature reserves in the Australian Capital Territory. Emu 97(2):141–149
Peterson BK, Weber JN, Kay EH, Fisher HS, Hoekstra HE (2012) Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS One 7(5):e37135
Pimentel D, Zuniga R, Morrison D (2005) Update on the environmental and economic costs associated with alien-invasive species in the United States. Ecol Econ 52(3):273–288
Planes S, Lecaillon G (1998) Consequences of the founder effect in the genetic structure of introduced island coral reef fish populations. Biol J Linn Soc 63(4):537–552
Poelstra JW, Vijay N, Bossu CM, Lantz H, Ryll B, Müller I et al. (2014) The genomic landscape underlying phenotypic integrity in the face of gene flow in crows. Science 344(6190):1410–1414
Pons P, Latapy M (2005) Computing communities in large networks using random walks. In: Yolum P, Güngör T, Gürgen F, Özturan C (eds) Computer and Information Sciences - ISCIS 2005: 20th International Symposium, Istanbul, Turkey, October 26-28, 2005. Proceedings. Springer, Berlin, Heidelberg, p 284–293
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559-575
Rius M, Bourne S, Hornsby HG, Chapman MA (2015) Applications of next-generation sequencing to the study of biological invasions. Curr Zool 61(3):488–504
Rollins LA, Woolnough AP, Wilton AN, Sinclair RON, Sherwin WB (2009) Invasive species can’t cover their tracks: using microsatellites to assist management of starling (Sturnus vulgaris) populations in Western Australia. Mol Ecol 18(8):1560–1573
Rollins LA, Woolnough AP, Fanson BG, Cummins ML, Crowley TM, Wilton AN et al. (2016) Selection on mitochondrial variants occurs between and within individuals in an expanding invasion. Mol Biol Evol 33(4):995–1007
Rosvall M, Bergstrom CT (2008) Maps of random walks on complex networks reveal community structure. Proc Natl Acad Sci 105(4):1118–1123
Sala OE, Chapin FS, Armesto JJ, Berlow E, Bloomfield J, Dirzo R et al. (2000) Global Biodiversity Scenarios for the Year 2100. Science 287(5459):1770–1774
Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJM, Birol İ (2009) ABySS: A parallel assembler for short read sequence data. Genome Res 19(6):1117–1123
Singapore Department of Statistics (2016a) Planning Areas/Subzones in Singapore (Year2016). Singapore, Singapore
Singapore Department of Statistics (2016b) Singapore Residents by Planning Area/Subzone and Type of Dwelling, June 2000 - 2016. Singapore, Singapore
Smouse PE, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82(5):561–573
Sokal RR, Wartenberg DE (1983) A test of spatial autocorrelation analysis using an isolation-by-distance model. Genetics 105(1):219–237
Sontag WA (1998) Devastated, damaged, and fully intact forest habitats and the Sturnidae family in a dry lowland evergreen biome in Southeast Thailand. Nat Hist Bull Siam Soc 46:43–53
Steinig EJ, Neuditschko M, Khatkar MS, Raadsma HW, Zenger KR (2016) NetView P: A network visualization tool to unravel complex population structure using genome-wide SNPs. Mol Ecol Resour 16(1):216–227
Tay YC, Chng MWP, Sew WWG, Rheindt FE, Tun KPP, Meier R (2016) Beyond the Coral Triangle: high genetic diversity and near panmixia in Singapore’s populations of the broadcast spawning sea star Protoreaster nodosus. Royal Soc Open Sci 3(8):160253
Vezzi F, Narzisi G, Mishra B (2012) Reevaluating Assembly Evaluations with Feature Response Curves: GAGE and Assemblathons. PLoS ONE 7(12):e52210
Vitousek PM, Mooney HA, Lubchenco J, Melillo JM (1997) Human Domination of Earth’s Ecosystems. Science 277(5325):494–499
Wakita K, Tsurumi T (2007) Finding Community Structure in Mega-scale Social Networks. In: Proceedings of the ACM Conference on World Wide Web, Banff, AB, Canada, p 1275-1276
Ward P (1968) Origin of the avifauna of urban and suburban Singapore. Ibis 110(3):239–255
Warren WC, Clayton DF, Ellegren H, Arnold AP, Hillier LW, Kunstner A et al. (2010) The genome of a songbird. Nature 464(7289):757–762
Wells DR (2010) The birds of the Thai-Malay peninsula, Vol 2. Bloomsbury Publishing, London
Wood JP, Dowell SA, Campbell TS, Page RB (2016) Insights into the introduction history and population genetic dynamics of the Nile monitor (Varanus niloticus) in Florida. J Hered 107(4):349–362
Yap CAM (2003) A study of the changes in the range sizes of white-vented mynas in Singapore. Raffles Bull Zool 51(1):159–164
Yap CAM, Sodhi NS, Brook BW (2002) Roost characteristics of invasive mynas in Singapore. J Wildl Manag 66(4):1118–1127
Zhang G, Li C, Li Q, Li B, Larkin DM, Lee C et al. (2014) Comparative genomics reveals insights into avian genome evolution and adaptation. Science 346(6215):1311–1320
Zhdanova OL, Pudovkin AI (2008) Nb_HetEx: a program to estimate the effective number of breeders. J Hered 99(6):694–695
Zheng X, Levine D, Shen J, Gogarten SM, Laurie C, Weir BS (2012) A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28(24):3326–3328
Acknowledgements
GWL, QT, and FER were funded through an Agri-Food and Veterinary Authority of Singapore grant (grant number R-154-000-692-490). Additionally, KMG and FER received funding from a Singapore Ministry of Education Tier I grant (R-154-000-658-112). BC acknowledges salary by SEABIG (grant numbers: R-154-000-648-646 and R-154-000-648-733). PGPE and MI were supported by the Swedish Research Council (grant number 621-2013-5161 to PGPE and grant number 621-2014-5113 to MI). We were also supported by the National Natural Science Foundation of China (grant number 31772441 to SW and FER). The authors would like to acknowledge support from Science for Life Laboratory, the National Genomics Infrastructure (NGI) and Uppmax for providing assistance in massive parallel sequencing and computational infrastructure. We also thank Keren Resha Sadanandan, Nathaniel Shengrong Ng, Ywee Chieh Tay, Elize Ying Xin Ng, Chui Shao Xiong, David Jian Xiong Tan, Emilie Sidonie Cros, and Mahathir Humaidi for help with laboratory work, helpful discussion, and constructive comments on early drafts of the manuscript. We also thank colleagues at the Jurong Bird Park for sample collection. Finally, we thank the National Environment Agency of Singapore, the National Parks Board of Singapore, and the Lee Kong Chian Natural History Museum for access to their respective collections for samples.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
B Chattopadhyay and KM Garg contributed equally to this work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Low, G., Chattopadhyay, B., Garg, K. et al. Urban landscape genomics identifies fine-scale gene flow patterns in an avian invasive. Heredity 120, 138–153 (2018). https://doi.org/10.1038/s41437-017-0026-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-017-0026-1
This article is cited by
-
Aquatic invasive species specialists’ perceptions on the importance of genetic tools and concepts to inform management
Biological Invasions (2022)
-
The conservation value of admixed phenotypes in a critically endangered species complex
Scientific Reports (2020)
-
Vegetation on and around large-scale buildings positively influences native tropical bird abundance and bird species richness
Urban Ecosystems (2019)