Abstract
Purpose
It is critical to identify putative causal targets for SARS coronavirus 2, which may guide drug repurposing options to reduce the public health burden of COVID-19.
Methods
We applied complementary methods and multiphased design to pinpoint the most likely causal genes for COVID-19 severity. First, we applied cross-methylome omnibus (CMO) test and leveraged data from the COVID-19 Host Genetics Initiative (HGI) comparing 9,986 hospitalized COVID-19 patients and 1,877,672 population controls. Second, we evaluated associations using the complementary S-PrediXcan method and leveraging blood and lung tissue gene expression prediction models. Third, we assessed associations of the identified genes with another COVID-19 phenotype, comparing very severe respiratory confirmed COVID versus population controls. Finally, we applied a fine-mapping method, fine-mapping of gene sets (FOGS), to prioritize putative causal genes.
Results
Through analyses of the COVID-19 HGI using complementary CMO and S-PrediXcan methods along with fine-mapping, XCR1, CCR2, SACM1L, OAS3, NSF, WNT3, NAPSA, and IFNAR2 are identified as putative causal genes for COVID-19 severity.
Conclusion
We identified eight genes at five genomic loci as putative causal genes for COVID-19 severity.
Similar content being viewed by others
INTRODUCTION
The coronavirus disease 2019 (COVID-19) pandemic represents a huge public health burden globally. Earlier research has revealed that specific molecular targets are essential for SARS coronavirus 2 (SARS-CoV-2) to enter into human cells [1]. Remdesivir, which blocks such targets, is approved by the US Food and Drug Administration to treat COVID-19. However, currently there remains no effective treatment for COVID-19. Therefore, there is a critical need to uncover additional causal molecular targets for COVID-19. A better characterization of targets can guide drug repurposing for identifying new uses of existing drugs. The fatality rate of COVID-19 is predominantly driven by those patients with severe respiratory failure who are hospitalized [2]. Causal molecular targets that can guide drug repurposing options are thus anticipated to be causally related to COVID-19 severity. However, such causal targets are quite difficult to identify due to the limitations of conventional studies and insufficient biological understanding of human genes.
One strategy to potentially reduce limitations of conventional study designs and identify candidate associated genes is to apply gene-level association tests that aggregate potential regulatory effects of genetic variants on genes [3,4,5,6,7]. Due to the random assortment of genetic alleles transferred from parent to offspring at the time of gamete formation, this approach focusing on genetically predicted gene expression should be less susceptible to selection bias, confounding effects, and reverse causation [8]. In the past several years, we and others have developed novel statistical methods in such transcriptome-wide association studies (TWAS) [3,4,5,6,7, 9]. The conventional TWAS design aims to develop genetic prediction models for gene expression using statistical methods, and further apply the gene expression prediction models to genome-wide association study (GWAS) data sets of the diseases of interest to identify genes with genetically predicted expression and associate them with the diseases. Applying such methods, we and others have conducted TWAS of multiple human diseases and identified multiple disease related genes [3, 5, 8,9,10,11].
Besides the conventional TWAS design, there are opportunities to develop novel integrative analyses by incorporating additional epigenetic and functional information. For example, DNA methylation interacts between genome and environment and is established to play an important role in the etiology of multiple diseases. It is known that DNA methylation could potentially regulate expression of genes. In several methylome-wide association studies (MWAS), we found that specific CpG sites could influence disease risk by regulating the expression of disease target genes [12, 13]. In earlier work, we have also shown that integrating information on enhancer–promoter interactions can improve statistical power for gene-level association tests [9, 14]. Built upon these works, we recently developed a novel gene-level association testing method, cross-methylome omnibus (CMO), by integrating genetically regulated DNA methylation in promoters, enhancers, and the gene body to identify disease related genes [15]. As demonstrated in our recent work, through simulation analyses and applied analyses of brain imaging–derived phenotypes and Alzheimer disease, CMO achieves high statistical power while well controlling for the type I error rate [15]. Importantly, CMO could reproducibly identify additional Alzheimer disease–associated genes that are not able to be identified by competing methods. This suggests that the novel method of CMO can be a complementary method for TWAS.
Despite the productivity of TWAS design using conventional methods (e.g., TWAS or S-PrediXcan) and novel methods (e.g., CMO) in identifying novel disease-associated genes, it is worth noting that such identified associated genes do not necessarily infer causality [16]. Aligned with other reports, although TWAS is useful for prioritizing causal genes, false positive findings cannot be avoided for some of the identified associations [16]. There are several potential reasons that could induce these, such as correlated expression across individuals, correlated predicted expression, and shared variants [16]. One strategy that can potentially prioritize causal genes in TWAS analyses is fine-mapping. Recently, we and others have developed several methods for fine-mapping in TWAS [17,18,19]. Focusing on a method we recently developed, fine-mapping of gene sets (FOGS), we find that FOGS adequately controls for type I error rates under various scenarios and performs better than competing methods, including FOCUS and p value ranking of TWAS results [17, 19]. Specifically, FOGS could achieve a higher area under the receiver operating characteristic (ROC) curve (AUC), identify more causal genes at the same false positive rate, and yield a smaller number of false positives at the same true positive rate [19].
Herein, we conducted a comprehensive multistage integrative multiomics study leveraging the data from COVID-19 patients and controls included in the COVID-19 Host Genetics Initiative (HGI) [20]. We first applied the CMO method to generate a list of promising genes associated with COVID-19 severity for discovery (comparing 9,986 hospitalized patients versus 1,877,672 population controls). We further applied the conventional S-PrediXcan method to characterize associations of predicted expression of these genes with COVID-19 severity. For associated genes, we further evaluated their associations with another COVID-19 phenotype, comparing very severe respiratory confirmed COVID versus population controls. Finally, we applied the FOGS fine-mapping method to determine the most likely causal genes for severe COVID-19 outcome. In our primary analyses, we focused on blood tissue to capture the systematic pattern of the body. It is also known that the immune system plays an important role in the host response to viral infection. By focusing on blood tissue we can well capture the effects of genes acting in immune related pathways. We also analyzed lung tissue as another likely target tissue for COVID-19 in our S-PrediXcan analyses.
MATERIALS AND METHODS
Genetic association data sets for COVID-19 severity in primary analyses
For evaluation of the association with COVID-19 severity, we used summary statistics data of the most recent version of GWAS analyses from the COVID-19 HGI (Release 5 [January 2021]) [20]. Detailed information on participating studies, quality control, and analyses has been provided on the COVID-19 HGI website (http://www.covid19hg.org/results/). Informed consent was obtained from all subjects. In brief, for discovery analyses comparing hospitalized patients and population controls, data (B2_ALL_eur) from 9,986 hospitalized COVID-19 patients and 1,877,672 population controls from studies in Biobanque Quebec COVID19, Columbia University COVID19 Biobank, Estonian Biobank, Geisinger Health System, Latvia COVID-19 research platform, UCLA Precision Health COVID-19 Biobank, 24Genetics, Amsterdam UMC COVID study group, Determining the Molecular Pathways and Genetic Predisposition of the Acute Inflammatory Process Caused by SARS-CoV-2, COVID19-Host(a)ge, GEN-COVID, reCOVID, deCODE, Million Veterans Program, 23andMe, Bonn Study of COVID19 genetics, FHoGID, Ancestry, The Genetic Predisposition to Severe COVID-19, Genomic, FinnGen, Genetic Modifiers for COVID-19 Related Illness, and UK Biobank were used. Hospitalized COVID-19 cases represented patients with (1) laboratory confirmed SARS-CoV-2 infection (RNA and/or serology based) and (2) hospitalization due to corona-related symptoms. Controls represent those that are not cases. The included subjects are Europeans only, to ensure the homogeneous population structure for the analyses. Only variants with imputation quality > 0.6 were retained. A fixed-effect meta-analysis of individual studies was performed with inverse variance weighting.
CMO test
Details of the CMO method have been described elsewhere [15]. CMO is an integrative gene-level test for identifying associated genes that may impact the trait of interest through DNA methylation pathways. Briefly, three main steps are involved. First, CMO links CpG sites located in enhancers, promoters, and the gene body to a target gene, considering that DNA methylation in enhancers and promoters may also play important roles in gene regulation. Importantly, CMO integrates comprehensive enhancer–promoter interaction information from a comprehensive database called GeneHancer and links CpG sites that are located in the enhancers, promoters, and the gene body to their target genes [21]. Second, by leveraging comprehensive blood DNA methylation genetic prediction models that were developed using a large reference data set involving 4,008 subjects [22], CMO tests associations between genetically regulated DNA methylation of each CpG site and COVID-19 severity using several widely used weighted gene-based tests, including burden, sum of squared score (SSU), and Aggregated Cauchy Association Test (ACAT) tests. The methylation prediction models were developed focusing on 151,729 CpG sites with a significant methylation quantitative trait locus (mQTL), and the lasso method was applied with genetic variants (i.e., single-nucleotide polymoprhisms [SNPs]) closer than 250 kb to each CpG site as potential predictors [22]. Because the optimal test depends on the underlying truth, which is unknown in practice, to maximum statistical power, we apply a Cauchy combination test to combine the results from burden, SSU and ACAT tests [23]. Third, CMO applies a Cauchy combination test to combine statistical evidence from multiple CpG sites for each target gene to determine the associations of target gene–COVID-19 severity. A Benjamini–Hochberg false discovery rate (FDR) of < 0.05 was used to adjust for multiple comparisons.
S-PrediXcan test for candidate genes identified from CMO test
To better characterize the candidate genes identified from the CMO test, we further conducted analyses using the orthogonal and complementary S-PrediXcan method to evaluate associations of their genetically predicted expression with COVID-19 severity [24]. We first leveraged comprehensive blood gene expression genetic prediction models that were developed using a reference data set involving subjects as included in the version 8 of the Genotype-Tissue Expression (GTEx) [25]. A modified cross-tissue UTMOST framework was used to build gene expression genetic models.[26, 27] In brief, SNPs within 1 Mb upstream and downstream of each gene body were included as candidate predictor variables in the model. The residual of the normalized gene expression (TPM) was used for model development after adjustment of age, sex, sequencing platform, the first five principal components (PCs), and probabilistic estimation of expression residuals (PEER) factors. The effect sizes were assessed by minimizing the loss function with a LASSO penalty on the columns (within-tissue effects) and a group LASSO penalty on the rows (cross-tissue effects). The group penalty term implemented sharing of the information from feature (SNP) selection across all the involved tissues. The original model training was modified by unifying the hyperparameter pairs to avoid the overestimation of the prediction performance [26, 27]. The details for the S-PrediXcan method are described elsewhere [24]. Briefly, the associations of genetically predicted gene expression with COVID-19 severity were estimated based on genetic prediction model weights, summary statistics of genetic variants with COVID-19 severity, and a variant correlation (linkage disequilibrium [LD]) matrix. We also tested the associations by leveraging lung tissue gene expression models developed using the same modified UTMOST method [27].
We further evaluated associations of identified genes with another COVID-19 phenotype. Briefly, we compared very severe respiratory confirmed COVID versus population controls by leveraging data sets of A2_ALL_eur (Europeans; 5,101 cases and 1,383,241 controls). S-PrediXcan was used to infer the gene–phenotype associations. We did not compare hospitalized COVID-19 patients versus nonhospitalized COVID-19 patients considering that only a relatively small sample size was available, which may induce insufficient power (B1_ALL_eur data set for Europeans: 4,829 cases and 11,816 controls). We did not investigate the data set of C2_ALL_eur (Europeans; 38,984 cases and 1,644,784 controls), which compared COVID-19 patients versus population controls. This is because the outcome of COVID-19 susceptibility would be difficult to interpret, as this may only reflect whether or not an individual was exposed to the SARS-CoV-2 virus.
FOGS fine-mapping analysis to determine putative causal genes for COVID-19 severity
To determine the most likely causal genes for COVID-19 severity, we conducted FOGS fine-mapping analysis for the genes supported by both CMO and S-PrediXcan analyses. Details for FOGS have been described in our earlier publication [19]. In brief, two steps are involved. First, a conditional analysis with ridge regression is conducted to account for the effects of other variants/genes in the locus of interest. Second, FOGS integrates genetic prediction model weights and conditional Z-scores by an adaptive test to maintain high statistical power.
RESULTS
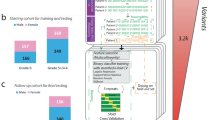
The overall study design flow is presented in Fig. 1. The description of several data sets used in this study is included in Supplementary Table 1. Based on the CMO test (Supplementary Table 2; Supplementary Figure 1), we identified significant associations of 76 genes with COVID-19 severity comparing hospitalized patients and population controls at FDR < 0.05 (Table 1). Interestingly, some of these genes tend to be implicated in immunological pathways (Table 1). Of these genes, there were also significant associations between genetically predicted expression in blood tissue of nine genes and COVID-19 severity comparing hospitalized patients and population controls (Table 2). Through analyzing another outcome comparing very severe respiratory confirmed patients versus controls, eight of them (except for CCR5) were validated at P < 0.10 (Table 2). Based on fine-mapping through FOGS, all these genes at five loci were determined to be putative causal genes. Plots showing associations of SNPs with COVID-19 severity (B2 outcome) at the locus of each of the identified putative causal genes were shown in Supplementary Figures 2–9. Positive associations between predicted expression levels in blood tissue and COVID-19 severity were detected for XCR1, CCR2, and OAS3. Conversely, associations between lower predicted expression levels in blood tissue and increased COVID-19 severity were identified for SACM1L, NSF, WNT3, NAPSA, and IFNAR2. In analyses of lung tissue gene expression prediction models, although for several of these genes there was no prediction model developed, for the three genes with models available (CCR2, WNT3, and IFNAR2), consistent associations were observed as well (Table 3).
Firstly, we applied cross methylome omnibus (CMO) test and leveraged data from The COVID-19 host genetics initiative (HGI) comparing 9,986 hospitalized COVID-19 patients and 1,877,672 population controls, in which we identified 76 candidate genes. Secondly, we evaluated associations using the complementary S-PrediXcan method and leveraging blood gene expression prediction models, from which nine genes showed an association. Thirdly, we assessed associations of the identified genes with another COVID-19 phenotype, comparing very severe respiratory confirmed COVID vs population controls, and eight of the genes showed consistent associations. We further applied FOGS fine-mapping method which confirms these eight genes as putative causal genes. Finally, additional analyses of lung tissue predicted gene expression confirm associations of these genes with COVID-19 severity.
DISCUSSION
This is one of the earliest studies to comprehensively evaluate the associations of genes across the genome with COVID-19 severity using genetic instruments combined with different layers of functional information. After careful assessment including fine-mapping analysis, we identified eight putative causal genes for COVID-19 severity, namely, XCR1, CCR2, and SACM1L on chromosome 3; OAS3 on chromosome 12; NSF and WNT3 on chromosome 17, NAPSA on chromosome 19; and IFNAR2 on chromosome 21. Our multistage study provides new information to improve our understanding of putative causal targets for SARS-CoV-2, which could be useful for further drug repurposing efforts. The identification of additional therapeutic strategies holds the promise of reducing the public health burden of COVID-19.
Literature supports potential functional roles of several of the identified genes. XCR1, CCR2, and SACM1L locate at locus 3p21.31. XCR1 is thought to mediate chemokine signaling pathways for inflammatory regulation, leukocyte chemotaxis, as well as immunopathies inducing lung injury [28]. Previous work suggested that this gene was critical for the advancement of influenza virus infection [29]. CCR2 is known to promote chemotaxis of monocyte/macrophage towards inflammation sites [30]. It has been reported that the canonical ligand for CCR2 is highly expressed in bronchoalveolar lavage fluid from lung tissue of COVID-19 patients during mechanical ventilation [31], and circulating MCP1 levels are related to more severe disease [32]. Another study reported that SACM1L expression was significantly changed in response to top candidate drugs from L1000 and SARS-CoV-2 settings [33]. Furthermore, the genetic locus harboring rs17713054 was identified to be coaccessible with the promoter region of several genes including SACM1L in lung single cells [34]. In the earlier GWAS of the Severe Covid-19 GWAS Group, rs11385942 at this locus showed a significant association with COVID-19 severity at the genome-wide level (P < 5×10−8) [35]. Our work suggested that XCR1, CCR2, and SACM1L could potentially be the causal genes at this locus. A more recent GWAS of critical illness in COVID-19 reported a novel variant rs10735079 at chr12q24.13 in a gene cluster encoding antiviral restriction enzyme activators including OAS3 [30]. In another study, it was also identified that a Neandertal haplotype that is protective against severe COVID-19 contains all or parts of three genes including OAS3. Interestingly, the SNPs showing the most significant associations are in OAS3 [36]. IFNAR2 at chromosome 21 encodes type I interferon (IFN-α/β), which is known to play a key role in human antiviral immunity [37]. Previous work reported that probes tagging this gene showed pleiotropic association with hospitalized COVID-19 [38]. Some of the genes suggested by CMO test but not following S-PrediXcan analyses may also warrant further investigation. For 42 of the genes, their genetic expression prediction models were not established using the modified UTMOST modeling strategy. For the S-PrediXcan analyses, the odd ratios reported in this study were for genetically predicted expression but not actual expression levels. Further functional validation to better understand the exact roles of these genes is needed.
A previous study reported likely causal links of IFNAR2, TYK2, and CCR2 with COVID-19 critical illness [30]. In the current study, we also identified IFNAR2 and CCR2. In another study analyzing an earlier version of COVID-19 HGI data (version 4), genes IFNAR2 and CCR2 were identified with allelic imbalance evidence at COVID-19 GWAS risk variants (unpublished data). IFNAR2 was also associated with migraine and throat pain (unpublished data). The genetically predicted expression of IFNAR2 was further identified to be inversely associated with creatine kinase. In this study, XCR1 and OAS3 were also implicated as likely susceptibility genes for COVID-19 severity, which was consistent with our findings. In the COVID-19 HGI main manuscript (unpublished data), it was identified that the COVID-19 associated variants modified the expression of OAS1/OAS3/OAS2 (12q24.13) and IFNAR2/IL10RB (21q22.11) in lung. Overall, besides identifying literature reported genes, in this work we also identified several novel putative causal genes for COVID-19.
There are several potential limitations in our study. First, due to the nature of COVID-19 HGI, it is possible that although all are required to meet the phenotype definition (e.g., be hospitalized COVID-19 patients), the included cases in different substudies are not completely homogeneous. For example, the criteria for COVID-19 patients’ hospitalization could be different across studies/regions, thus measurement errors could exist. Second, in our analyses, we were not able to comprehensively adjust for underlying cardiovascular and metabolic factors that are reported to be related to COVID-19 [39]. While a majority of implicated genes (except for OAS3 [40]) have not been reported to be associated with cardiovascular and metabolic factors according to GWAS Catalog, alleviating the concern of pleiotropy, further work with adjustment of such variables is needed to validate our findings. Third, in the data sets used in our analyses, information about the infection status of SARS-CoV-2 in the control participants was limited. By using the general population as controls, severe COVID-19 cases are actually compared with a large cohort of individuals who may or may not develop severe COVID-19 upon exposure to the virus. However, the presence of susceptible subjects in the control group, if any, is expected to only bias the results toward the null. Future work using cleaner controls would be necessary to better characterize the relationship. Fourth, the current study focuses on Europeans, the ethnic group with the largest available sample size. It would be critical to conduct analyses focusing on other ethnic groups, to enhance the generalizability of findings of such work. Currently, the available sample size of GWAS of COVID-19 in non-European populations is relatively small. For example, in the COVID HGI, for the B2 outcome, data are available for only 257 cases of Latinos, 60 cases of Arabs, 948 of Admixed Americans, 790 of Africans, 186 of South Asians, and 1,414 of East Asians. The power for such analyses would be relatively low. Additional work for sex specific analyses would be needed as well. We currently do not have the data available for sex specific analyses. Finally, besides the outcomes evaluated in the current study, analyses using brain tissue gene expression models could be helpful for characterizing factors related to the neurological symptoms of COVID-19. The available data in the COVID HGI may not be appropriate for testing this, as neurological symptoms may manifest in mildly symptomatic COVID-19 individuals. Future work leveraging cleaner disease phenotype is needed for testing this.
In conclusion, in a large scale multiphase integrative multiomics study with complementary methods, we identified eight putative causal genes at five loci for COVID-19 severity. Such findings will be very meaningful for guiding future drug repurposing efforts aiming to reduce the COVID-19 public health burden.
Data availability
All data and methods used in the analysis are described or included in this article and the electronic supplementary information. The COVID-19 HGI GWAS summary statistics are deposited at http://www.covid19hg.org/results/r5/. The blood methylation prediction models are available at http://bbmri.researchlumc.nl/atlas/#data. The data for running the CMO test are available at http://www.zenodo.org/record/4475935#.YItjZi2cY0w. The blood and lung tissue gene expression prediction models are available at http://www.zenodo.org/record/3842289#.YHqHcehKiUk.
Code availability
Access to the custom code may be requested from the corresponding authors.
References
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271–80.e8.
Richardson S, Hirsch JS, Narasimhan M, Crawford JM, McGinn T, Davidson KW et al. Presenting characteristics, comorbidities, and outcomes among 5700 patients hospitalized with COVID-19 in the New York city area. JAMA. 2020;323:2052–9.
Wu C, Pan W. Integration of methylation QTL and enhancer-target gene maps with schizophrenia GWAS summary results identifies novel genes. Bioinformatics. 2019;35:3576–83.
Hu Y, Li M, Lu Q, Weng H, Wang J, Zekavat SM, et al. A statistical framework for cross-tissue transcriptome-wide association analysis. Nat Genet. 2019;51:568–76.
Wu C, Pan W. Integrating eQTL data with GWAS summary statistics in pathway-based analysis with application to schizophrenia. Genet Epidemiol. 2018;42:303–16.
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52.
Gamazon ER, Wheeler HE, Shah KP, Mozaffari SV, Aquino-Michaels K, Carroll RJ, et al. A gene-based association method for mapping traits using reference transcriptome data. Nat Genet. 2015;47:1091–8.
Wu L, Wang J, Cai Q, Cavazos TB, Emami NC, Long J, et al. Identification of novel susceptibility loci and genes for prostate cancer risk: A Transcriptome-wide Association Study in over 140,000 European descendants. Cancer Res. 2019;79:3192–204.
Wu C, Pan W. Integration of enhancer-promoter interactions with GWAS summary results identifies novel Schizophrenia-associated genes and pathways. Genetics. 2018;209:699–709.
Wu L, Shi W, Long J, Guo X, Michailidou K, Beesley J, et al. A transcriptome-wide association study of 229,000 women identifies new candidate susceptibility genes for breast cancer. Nat Genet. 2018;50:968–78.
Zhong J, Jermusyk A, Wu L, Hoskins JW, Collins I, Mocci E, et al. A Transcriptome-Wide Association Study (TWAS) identifies novel candidate susceptibility genes for pancreatic cancer. J Natl Cancer Inst. 2020;112:1003–12.
Yang Y, Wu L, Shu XO, Cai Q, Shu X, Li B, et al. Genetically predicted levels of DNA methylation biomarkers and breast cancer risk: data from 228 951 women of European descent. J Natl Cancer Inst. 2020;112:295–304.
Yang Y, Wu L, Shu X, Lu Y, Shu XO, Cai Q, et al. Genetic data from nearly 63,000 women of European descent predicts DNA methylation biomarkers and epithelial ovarian cancer risk. Cancer Res. 2019;79:505–17.
Furlong EEM, Levine M. Developmental enhancers and chromosome topology. Science. 2018;361:1341–5.
Wu C, Bradley J, Li Y, Wu L, Deng HW. A gene-level methylome-wide association analysis identifies novel Alzheimer’s disease genes. Bioinformatics 2021. https://doi.org/10.1093/bioinformatics/btab045.
Wainberg M, Sinnott-Armstrong N, Mancuso N, Barbeira AN, Knowles DA, Golan D, et al. Opportunities and challenges for transcriptome-wide association studies. Nat Genet. 2019;51:592–9.
Mancuso N, Freund MK, Johnson R, Shi H, Kichaev G, Gusev A, et al. Probabilistic fine-mapping of transcriptome-wide association studies. Nat Genet. 2019;51:675–82.
Barfield R, Feng H, Gusev A, Wu L, Zheng W, Pasaniuc B, et al. Transcriptome-wide association studies accounting for colocalization using Egger regression. Genet Epidemiol. 2018;42:418–33.
Wu C, Pan W. A powerful fine-mapping method for transcriptome-wide association studies. Hum Genet. 2020;139:199–213.
COVID-19 Host Genetics Initiative. The COVID-19 Host Genetics Initiative, a global initiative to elucidate the role of host genetic factors in susceptibility and severity of the SARS-CoV-2 virus pandemic. Eur J Hum Genet 2020;28:715–8.
Fishilevich S, Nudel R, Rappaport N, Hadar R, Plaschkes I, Stein TI, et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database 2017;2017:bax028.
Baselmans BML, Jansen R, Ip HF, van Dongen J, Abdellaoui A, van de Weijer MP, et al. Multivariate genome-wide analyses of the well-being spectrum. Nat Genet. 2019;51:445–51.
Liu YXJ. Cauchy combination test: a powerful test with analytic p-value calculation under arbitrary dependency structures. J Am Stat Assoc. 2020;115:393–402.
Barbeira AN, Dickinson SP, Bonazzola R, Zheng J, Wheeler HE, Torres JM, et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat Commun. 2018;9:1825.
Consortium GT. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369:1318–30.
Liu D, Zhou D, Sun Y, Zhu J, Ghoneim D, Wu C, et al. A Transcriptome-Wide Association Study identifies candidate susceptibility genes for pancreatic cancer risk. Cancer Res. 2020;80:4346–54.
Zhou D, Jiang Y, Zhong X, Cox NJ, Liu C, Gamazon ER. A unified framework for joint-tissue transcriptome-wide association and Mendelian randomization analysis. Nat Genet. 2020;52:1239–46.
McCoy K, Peterson A, Tian Y, Sang Y. Immunogenetic Association underlying severe COVID-19. Vaccines. 2020;8:700.
Vastrad B, Vastrad C, Tengli A. Identification of potential mRNA panels for severe acute respiratory syndrome coronavirus 2 (COVID-19) diagnosis and treatment using microarray dataset and bioinformatics methods. 3 Biotech. 2020;10:422.
Pairo-Castineira E, Clohisey S, Klaric L, Bretherick AD, Rawlik K, Pasko D, et al. Genetic mechanisms of critical illness in Covid-19. Nature. 2021; 591:92–8.
Zhou Z, Ren L, Zhang L, Zhong J, Xiao Y, Jia Z, et al. Heightened innate immune responses in the respiratory tract of COVID-19 patients. Cell Host Microbe. 2020;27:883–90.e2.
Zhao Y, Qin L, Zhang P, Li K, Liang L, Sun J, et al. Longitudinal COVID-19 profiling associates IL-1RA and IL-10 with disease severity and RANTES with mild disease. JCI Insight. 2020;5:e139834.
El-Hachem N, Eid E, Nemer G, Dbaibo G, Abbas O, Rubeiz N, et al. Integrative transcriptome analyses empower the anti-COVID-19 drug arsenal. iScience. 2020;23:101697.
Wang A, Chiou J, Poirion OB, Buchanan J, Valdez MJ, Verheyden JM, et al. Single-cell multiomic profiling of human lungs reveals cell-type-specific and age-dynamic control of SARS-CoV2 host genes. Elife. 2020;9:e62522.
Severe Covid-19 GWAS Group, Ellinghaus D, Degenhardt F, Bujanda L, Buti M, Albillos A, et al. Genomewide Association Study of Severe Covid-19 with respiratory failure. N Engl J Med. 2020;383:1522–34.
Zeberg H, Paabo S. A genomic region associated with protection against severe COVID-19 is inherited from Neandertals. Proc Natl Acad Sci USA 2021;118:e20263091.
Duncan CJ, Mohamad SM, Young DF, Skelton AJ, Leahy TR, Munday DC, et al. Human IFNAR2 deficiency: lessons for antiviral immunity. Sci Transl Med. 2015;7:307ra154.
Liu D, Yang J, Feng B, Lu W, Zhao C, Li L. Mendelian randomization analysis identified genes pleiotropically associated with the risk and prognosis of COVID-19. J Infect. 2021;82:126–32.
Williamson EJ, Walker AJ, Bhaskaran K, Bacon S, Bates C, Morton CE, et al. OpenSAFELY: factors associated with COVID-19 death in 17 million patients. Nature 2020; 10.1038/s41586-020-2521-4.
Liu M, Jin HS, Park S. Protein and fat intake interacts with the haplotype of PTPN11_rs11066325, RPH3A_rs886477, and OAS3_rs2072134 to modulate serum HDL concentrations in middle-aged people. Clin Nutr. 2020;39:942–9.
Acknowledgements
We would like to acknowledge COVID-19 Host Genetics Initiative for making their COVID-19 GWAS summary statistics to be available.
Funding
This research is supported by University of Hawaii Cancer Center and Florida State University. D.L. is partially supported by the Harbin Medical University Cancer Hospital. Y.S. is partially supported by the Department of Education of Fujian Province, P. R. China.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics Declaration
This study was reviewed by the University of Hawaii Institutional Review Board (2019-00402). All individuals participating in the COVID HGI study properly signed the informed consent according to the participating studies’ Institutional Review Board. All data were de-identified before the analysis of the current study.
Role of the Funder/Sponsor
The funding organization had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; or decision to submit the manuscript for publication.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Wu, L., Zhu, J., Liu, D. et al. An integrative multiomics analysis identifies putative causal genes for COVID-19 severity. Genet Med 23, 2076–2086 (2021). https://doi.org/10.1038/s41436-021-01243-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01243-5
This article is cited by
-
Next-generation sequencing of host genetics risk factors associated with COVID-19 severity and long-COVID in Colombian population
Scientific Reports (2024)
-
SARS-CoV-2 rapidly evolves lineage-specific phenotypic differences when passaged repeatedly in immune-naïve mice
Communications Biology (2024)
-
Colocalization of expression transcripts with COVID-19 outcomes is rare across cell states, cell types and organs
Human Genetics (2023)
-
SUMMIT: An integrative approach for better transcriptomic data imputation improves causal gene identification
Nature Communications (2022)