Abstract
Purpose
To investigate the contribution of PALB2 pathogenic gene variants (PGVs, PALB2_PGV) and the CHEK2 c.1100delC (CHEK2_1100delC) PGV to familial breast and ovarian cancer, and PALB2_PGV associated breast cancer pathology.
Methods
Outcomes of germline PALB2_PGV and CHEK2_1100delC testing were recorded in 3,127 women with histologically confirmed diagnoses of invasive breast cancer, carcinoma in situ, or epithelial nonmucinous ovarian cancer, and 1,567 female controls. Breast cancer pathology was recorded in PALB2_PGV cases from extended families.
Results
Thirty-five PALB2 and 44 CHEK2_1100delC PGVs were detected in patients (odds ratio [OR] PALB2 breast–ovarian = 5.90 [95% CI: 1.92–18.36], CHEK2 breast–ovarian = 4.46 [95% CI: 1.86–10.46], PALB2 breast = 6.16 [95% CI: 1.98–19.21], CHEK2 breast = 4.89 [95% CI: 2.01–11.34]). Grade 3 ER-positive HER2-negative, grade 3 and triple negative (TN) tumors were enriched in cases with PALB2 PGVs compared with all breast cancers known to our service (respectively: 15/43, 254/1,843, P = 0.0005; 28/37, 562/1,381, P = 0.0001; 12/43, 204/1,639, P < 0.0001). PALB2_PGV likelihood increased with increasing Manchester score (MS) (MS < 15 = 17/1,763, MS 20–39 = 11/520, P = 0.04) but not for CHEK2_1100delC (MS < 15 = 29/1,762, MS 20–39 = 4/520). PALB2 PGVs showed perfect segregation in 20/20 first-degree relatives with breast cancer, compared with 7/13 for CHEK2_1100delC (P = 0.002).
Conclusion
PALB2 PGVs and CHEK2_1100delC together account for ~2.5% of familial breast/ovarian cancer risk. PALB2 PGVs are associated with grade 3, TN, and grade 3 ER-positive HER2-negative breast tumors.
Similar content being viewed by others
INTRODUCTION
Breast cancer is the most common form of cancer in women.1 It has been known for over 30 years that breast cancer predisposition can be inherited in an autosomal dominant manner with around 4% of cases on a population basis being compatible with the inheritance of a high-risk (circa 50–80% lifetime risk) pathogenic germline gene variant (PGV).2 The BRCA1 and BRCA2 genes were identified as such genes in 1994 and 1995 accounting for around 2% of incident breast cancers and with a combined population prevalence of 1 in 300–400.3 Although TP53 can cause a pattern consistent with dominantly inherited breast cancer it is usually associated with onset at extremely young ages, is rare (circa population prevalence 1 in 5,000) and more typically causes patterns of other malignancies, particularly sarcoma.4 Other extremely rare genes have been suggested as potential high-risk genes with lifetime risks of >40% such as CDH1, PTEN, and STK11, but definitive evidence based on case control data has been lacking due to their rarity (population prevalence ~1 in 50–100,000 for each).5 PALB2 was originally identified in 2007 as a moderate risk gene conferring only around a 2.3-fold relative risk;6 this was, however, based on identification in 10/923 BRCA1/2 negative breast cancer cases, but 0/1,083 controls. Therefore, the originally calculated relative risk was based on study of the families rather than the odds ratio (OR) generated from case–control analysis because this is usually inflated by using familial risk probands. Given the low predicted lifetime risk of only 20–30%, PALB2 testing was not widely utilised until it became included in gene panel testing. It was not until more than 7 years later in 2014 that penetrance analysis in 158 families suggested risks approaching high-risk with a 35% (95% CI: 26–46%) estimated risk by age 70 years.7 This risk was higher in the context of a close relative with breast cancer. This analysis was updated recently in 524 affected families with estimated risks to age 80 years of 53% (95% CI: 44–63%) for female breast cancer and 5% (95% CI: 2–10%) for ovarian cancer.8
In contrast to PALB2, the original assessment of CHEK2 as a moderate risk gene has been maintained from its discovery in 2002.5,9,10,11 Nonetheless, like PALB2, the risk increases when there is a positive family history of breast cancer.11 In the UK, while initially consensus was reached to include both CHEK2 and PALB2 in breast cancer gene panels in the UK,12 mainstreaming of tests to oncologists and other specialties in both breast and ovarian cancer now only includes PALB2 in addition to BRCA1 and BRCA2.13
In view of these changes in recommendations, namely the recent addition of PALB2, but not CHEK2, to UK genetic testing guidance, we report our single-center experience to date of germline PALB2 and CHEK2 c.1100delC; p.(Thr367MetfsTer15) (CHEK2_1100delC) testing in breast/ovarian cancer; these data will help provide evidence for future testing guidance.
MATERIALS AND METHODS
Patients
Women were eligible for this study if they had a histologically confirmed diagnosis of invasive or in situ breast cancer or an epithelial nonmucinous ovarian cancer and had undergone germline testing of BRCA1, BRCA2, PALB2, and CHEK2_c.1100delC for PGVs. Women were either referred for genetic testing to the Manchester Center for Genomic Medicine (MCGM) or the Family History Risk and Prevention Clinic (FHRPC) at the Nightingale Center, Wythenshawe Hospital (n = 2,603).14 In addition, 524 women were tested as part of the population based Predicting the Risk Of Cancer At Screening (PROCAS) study, in Greater Manchester.15 Demographic details of the study population are outlined in Table 1. Women without a breast cancer diagnosis (n = 1,567, aged 46–73 years), who were also recruited to the PROCAS study, were included as controls.
For pathology comparisons we included 1,843 women known to the FHRPC/MCGM who had developed breast cancer but did not have a PGV in the known breast cancer predisposition genes. We also included women with a PALB2 associated breast cancer in the extended families.
Clinical or research consent was given for extended testing of breast cancer associated genes (approval from the North Manchester Research Ethics Committee, reference 09/H1008/81 [PROCAS] and 08/H1006/77).
Genetic screening
For women that were seen through the MCGM and FHRPC, DNA was extracted from lymphocytes, whereas those recruited through PROCAS (including the controls) had DNA extracted from saliva. Those attending MCGM had DNA initially analyzed for PGVs in BRCA1/2 and CHEK2_c.1100delC (with the standard clinical panel expanded to include PALB2 in 2016) by a combination of next-generation sequencing and multiplex ligation-dependent probe amplification (MLPA). Testing of DNA in the cohort and controls all patients was performed by a combination of targeted sequencing, panel test, and exomes (Supplementary Table 1).
Variants were classified according to the American College of Medical Genetics and Genomics/Association for Molecular Pathology (ACMG/AMP) guidelines.16 Variants that were classified as likely pathogenic and pathogenic only are reported here and combined along with gene rearrangements as PGVs.
Tumor pathology information was obtained for each case when available through hospital records, and cancer registries as previously described.17 The probability of a BRCA1/2 PGV was determined using the Manchester score (MS) for each affected individual.18 This adds scores for each breast and ovarian cancer in a direct lineage with increasing scores for earlier age at onset and pathologies in the proband suggestive of BRCA1 such as high-grade serous ovarian cancer and triple negative breast cancer (TNT). A MS of 15–19 and 20–24 roughly equate to a 10% and 20% likelihood threshold for BRCA1/2 respectively with a score of ≥40 equivalent to a likelihood of >75%.18
Statistical analysis was undertaken using GraphPad Prism version 9.0.1 for Windows (GraphPad Software, San Diego, CA, USA) and GraphPad QuickCalcs.19
RESULTS
Pathogenic germline variants
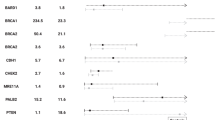
A total of 3,127 index cases with breast and/or ovarian cancer have undergone testing for PGVs of BRCA1, BRCA2, PALB2, and CHEK2_c.1100delC (Table 2). There were 35 (1.12%) with PGVs in PALB2 and 44 (1.41%) with CHEK2_c.1100delC. This compared to rates of 3/1,567 (0.19%, P = 0.0004) and 5/1,567 (0.32%, P = 0.0003) for PALB2 and CHEK2_c.1100delC respectively in the PROCAS controls, generating ORs of 5.90 (95% CI = 1.92–18.36) and 44.46 (95% CI = 1.86–10.46) respectively.
Testing of 302 cases with ovarian cancer detected two PALB2 PGVs, (c.2167_2168delAT; p.[Met723ValfsTer21] and c.3113G>A; (p.Trp1038Ter]) (OR = 3.47, 95% CI = 0.61–17.07, P = NS), and one with CHEK2_c.1100delC (OR = 1.04, 95% CI = 0.09–7.49, P = NS). Considering the breast cancer only index cases, there were 33 PALB2 PGVs and 43 CHEK2_c.1100delC PGVs (PALB2: OR = 6.16, 95% CI = 1.98–19.21, P = 0.0003; CHEK2: OR = 4.83, 95% CI = 2.01–11.34, P = 0.0001). Testing in the population based PROCAS breast cancer study identified PALB2 PGVs in 4/524 (0.76%) (OR = 4.01, 95% CI = 1.07–15.95, P = 0.071) and the CHEK2_c.1100delC PGV in 9/524 (1.71%) (OR = 5.46, 95% CI = 1.93–14.60, P = 0.0021), whereas testing of breast cancer index cases in the context of family history/early onset breast cancer only, detected PALB2 PGVs in 29/2,301 (1.26%) (OR = 6.65, 95% CI = 2.26–20.86, P = 0.0002) and the CHEK2 c.1100delC PGV in 34/2,301 (1.48%) (OR = 4.69, 95% CI = 1.88–11.12, P = 0.0002).
The mean age at diagnosis of first breast cancer for those with a PALB2 PGV was 49.8 years (median 50 years, SEM = 2.30, range = 24–77 years) and, for the CHEK2_c.1100delCPGV, 49.3 years (median 49.3 years, SEM 1.83, range 27–76 years).
Overall, there were three common PGVs in PALB2. There were ten instances of c.3113G>A; p.(Trp1038Ter) in cases and one in controls, and four each of c.3116del; p.(Asn1039llefsTer2) and c.3549C>G; p.(Tyr1183Ter), with neither occurring in controls (Table 3). Together, these accounted for >50% (20/38) of the PALB2 PGVs identified. Considering germline gene rearrangements and CNVs, three multiexon deletions and a translocation (46,XX,t[5;16][q33.1;p12.2]), of PALB2 were detected in the patient cohort.
PALB2 PGVs: receptor status and grade
Table 3 shows the MS, age, and pathology information for breast tumors where a germline PALB2 PGV was detected. Where pathology grade and receptor status were known, 8/25 breast tumors were triple negative (all grade 3). This was similar to the numbers and age at diagnosis of TNTs known to our service with a BRCA2 PGV (n = 9) (PALB2 PGV TNT: mean age 48.6 years, median 54.0 years, range 27–59 years; BRCA2 PGV TNT: mean age 46.8 years, median 48 years, range 33–55 years), but significantly fewer than for a BRCA1 PGV (P < 0.0001). While there was a trend toward increasing age of TNT diagnoses with a PALB2 PGV as compared with BRCA1 PGV associated TNTs (BRCA1 PGV TNT: mean = 40.4 years, median = 39 years, range = 22–77 years), this was not significant (P = 0.060).
To investigate the breast pathology where there is a PALB2 PGV further, we then considered all the patients with PALB2 associated breast cancers known to our service where a full histological record was available (n = 43), some of whom had genetic testing through other sources and so are not included in the 33 presented in Table 3. Here, the TNT phenotype occurred in 12/43 (27.9%) PALB2 PGVs, as compared with 11.1% (204/1843) of all breast cancers cases known to our service where full histology was known (OR = 3.11, 95% CI = 1.61–5.96, P = 0.002).
We then investigated other breast cancer subtypes and whether they were also enriched in heterozygotes for a PALB2 PGV. A grade 3 ER-positive HER2-negative phenotype occurred in 15/43 (34.9%) PALB2 PGVs, while accounting for 254/1843 (13.8%) breast cancers (OR = 3.35, 95% CI = 1.76-6.44, P = 0.0005).
For a grade 3 phenotype, regardless of receptor status, these again were overrepresented occurring in 28/37 (75.7%) individuals with a PALB2 PGVs as compared with 562/1381 (40.7%) of all invasive ductal carcinomas known to our service and testing negative for BRCA1, BRCA2, and PALB2 PGVs (OR = 4.53, 95% CI = 2.11-9.65, P < 0.0001).
Manchester score
To assess the probability of PGVs in BRCA1/2, MS was determined for all affected women (Table 2). While the likelihood of a PALB2 PGV increased with increasing MS, no such trend was seen for CHEK2_1100delC. Overall, the rates of CHEK2_1100delC were similar in BRCA1/2 negative cases at the lowest MS of <9 (12/759, 1.6%) compared with 4/520 with a MS = 20–39, 0.8%, P = NS) (Table 4). For PALB2 there was a significantly higher likelihood of a PGV for MS of 20–39 than for MS < 9 and MS < 15 (P < 0.05) (Table 4).
Segregation of PGVs with breast cancer in families
There was perfect segregation of PALB2 PGVs in all 20 first-degree relatives (FDRs) with breast cancer who were available for testing; many FDRs with breast cancer were deceased and unavailable for testing and not all living FDRs opted for testing. This compared to only 7/13 for CHEK2_1100delC (P = 0.002) families (Table 5). Although there was only 88.8% and 87.4% segregation in BRCA1 and BRCA2 respectively, this was not significantly different to PALB2.
DISCUSSION
This is the largest study looking at the prevalence of PALB2 PGVs and the CHEK2_1100delC PGV from a single genetics center in patients attending with a personal diagnosis of breast and/or nonmucinous ovarian cancer. We demonstrated PALB2 OR ≥ 6 and CHEK2 OR ≥ 4 for breast and breast–ovarian cancer, and found specific breast cancer pathology associations where a PALB2 PGV is present.
The timely importance of our study reflects the recent publication and implementation of the National Health Service England (NHSE) national test directory.13 In England, testing for only BRCA1, BRCA2, and PALB2 PGVs is offered for hereditary breast cancer. Despite CHEK2 and ATM also being recommended for inclusion by the UK Cancer Genetics Group,12 these genes have been omitted. Furthermore, recent publication of the BRIDGES case control study20 showed truncating variants of these five genes to be strongly associated (P < 0.0001) with breast cancer risk.
The association of germline BRCA1 and BRCA2 PGVs with high-risk breast cancer predisposition has been well recognized and clinical diagnostic testing of these genes has been offered for the past 23 years in Manchester. While PALB2 was identified as a cause of hereditary breast cancer in 2007,6 it is only more recently that PALB2 PGVs have been shown to be associated with a >50% lifetime breast cancer risk in females.7,8 Routine diagnostic testing of PALB2 has been offered in MCGM since 2016, with some samples having been analyzed previously through research studies with diagnostic laboratory confirmation of any PGVs identified.
We detected an excess of CHEK2_1100delC and PALB2 PGVs in our patients and this was significant across all subgroups, except for PALB2 PGVs in the population based PROCAS subgroup. This is in keeping with PGVs of PALB2 being relatively rare and associated with high-risk breast cancer predisposition, and so would be less likely to be detected in excess in our non-high-risk breast cancer cases.
Our analysis of MS and likelihood of a PALB2 PGV showed that a higher MS (20–29 and 20–39) is strongly correlated with the presence of a germline PALB2 PGV as compared with a lower MS (<15), the current NHSE threshold of diagnostic BRCA1/2 and PALB2 testing.21 Interestingly at very high MS, in our data set, the likelihood of a PALB2 PGV appears to tail off as compared with BRCA1/2; further data will be needed to see if this finding is replicated. These data confirm the validity and clinical utility of the MS in the prediction of high-risk single gene breast cancer predisposition. Although the MS was designed for BRCA1/2 PGV likelihood, as PALB2 is a high-risk gene and associated with increasing frequency as MS rises, MS is also a useful marker for PALB2 PGVs except at very high scores. The lack of association of MS with the CHEK2_1100delC PGV likely reflects the associated moderate risk breast cancer predisposition of CHEK2_1100delC. When working up families for breast cancer gene panel testing, our data does suggest that MS is not suitable for determining likelihood of the presence of the CHEK2_1100delC PGV.
We also detected two PALB2 PGVs in individuals with high-grade serous ovarian cancer. PALB2 PGVs are associated with a 5% lifetime risk of ovarian cancer8 and our data would suggest that the additional weighting within the MS for ovarian cancer (and not counting mucinous subtypes) is appropriate.
We confirmed the recently reported association of PALB2 PGVs with a tendency to the development of TNTs,22 accounting for 28% of breast tumors occurring in female PALB2 PGV heterozygotes known to our service. While this tumor phenotype is more typical of BRCA1 PGVs, TNTs occur in 16% of BRCA2 associated breast cancers and at increasing ages as compared with BRCA1.23 In our data set, the mean age of the TNTs associated with a PALB2 PGV (46.6 years) was similar to that for TNTs associated with a BRCA2 PGV (46.8 years) and, while tending toward an older age than for BRCA1 PGVs (40.4 years), this was not significant.
More striking was the association of grade 3 ER-positive HER2-negative tumors with a PALB2 PGV, comprising 35% of breast cancers in our PALB2 PGV cohort, which, to our knowledge, has not been previously demonstrated. While Hu et al.22 noted an OR of 5.2 for ER-positive HER2-negative breast tumors for PALB2 PGVs, their study did not account for tumor grade. Considering grade in particular, we also found a marked excess of grade 3 tumors occurring in 76% of all PALB2 PGV associated breast cancers. While our data may be biased toward referrals received where a woman has developed a high-grade breast cancer with or without a relevant family history, it is likely that such an association is real, although further data collection is needed to tease out these tumor phenotype correlations.
Our data would suggest that, for women who had a breast cancer where the histology is of a TNT or grade 3 ER-positive HER2-negative phenotype in association with a high MS, there may be merit it considering testing for germline PALB2_PGVs where BRCA1/2 testing was negative.
The similarities of the phenotypes associated with BRCA1/2 and PALB2 PGVs, and strong association with MS, likely reflect the combined BRCA1-PALB2-BRCA2 functional unit that facilitates the repair of double stranded DNA breaks using the high-fidelity homologous recombination repair pathway.24
Exploitation of this HR pathway has enabled the utilization of poly (ADP-ribose) polymerase (PARP) inhibitors in the drug treatment of ovarian cancers associated with germline and/or somatic BRCA1/2 PGVs and their investigation in clinical trials of advanced breast cancer. Given the functional interaction of PALB2 with BRCA1 and BRCA2, and the phenotypic similarities of the associated cancers presented within and by others,8 it is likely these agents will also be of utility in the treatment of PALB2 deficient cancers. In fact, a recent phase II trial investigating Olaparib in metastatic breast cancer showed an objective response rate of 82% where there was a germline PALB2 PGV; furthermore, 85% of the breast cancers were ER-positive HER2-negative (grade not given).25
What is less clear is, despite BRCA1, BRCA2, and PALB2 forming a functional unit, why BRCA1 and BRCA2 PGVs are much more common than those affecting PALB2, with BRCA1/2 PGVs detected in 7% of our patient cohort and PALB2 PGVs in 1%. While the reasons for these differing frequencies may not be clear, influencing factors may include the smaller size of PALB2 and, until recently, PALB2 PGVs have not been routinely screened for in either patient or population cohorts. Furthermore, BRCA1/2 PGVs are thought to be as frequent as 1 in 250–300 of the general population.26
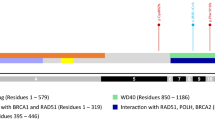
Of note we detected three recurrent PGVs of PALB2 in our cohort, c.3113G>A; p.(Trp1038Ter), c.3116del; p.(Asn1039llefsTer2) and c.3549; p.(Tyr1183Ter), together accounting for 50% of the individuals with PALB2 PGVs detected. These PGVs lie within the C-terminal WD40 domain of PALB2, which interacts with BRCA2. We note that for PALB2 PGVs reported by Yang et al.,8 while PGVs were distributed throughout PALB2, c.3113G>A was the most common PGV identified in breast/ovarian cancer detected in 61 families, with c.3549C>G detected in 19 families, and c.3116del in 9. Given our much smaller data set we cannot prove or disprove the possibility of a founder effect in our local population in the absence of haplotype studies; however, it may be that these loci represent PALB2 mutation hotspots, especially as the PALB2 founder PGVs reported are not localized to the 3’ end.27,28
Regarding the PALB2 del copy-number variants (CNVs) detected, both in this study and that reported by Yang et al.,8 again these all reside within the C-terminal WD40 domain that interacts with BRCA2. Thus, these data together may, in part, explain the similarities between the phenotypes seen with BRCA2 and PALB2 PGVs. Furthermore, it is likely that the genomic architecture toward the 3’ end of PALB2 predisposes to recombination events given this is where the CNVs and the balanced translocation we detected have been described. We note that our PALB2 testing strategy did not include CNVs for all our patients and so it is possible that further PALB2 CNVs remain to be identified in this cohort. Based on the data we report, we would recommend that any clinical testing strategy for PALB2 PGVs includes CNV analyses.
The association of CHEK2 PGVs with moderate risk breast cancer predisposition is well recognized. CHEK2 has a role in DNA repair, although at an earlier stage whereby it detects then determines the cellular response to DNA damage.29 We detected the CHEK2_1100delC PGV in 44/3,127 cases compared with just 5/1,567 controls reflecting the also increased OR attained from the much larger BRIDGES study.20 There has been considerable debate regarding the utility of including the moderate risk penetrance genes, CHEK2 and ATM, in breast cancer diagnostic genetic testing panels. The recent BRIDGES data would support their inclusion, given that, along with the high-risk breast cancer genes, BRCA1, BRCA2, and PALB2, truncating PGVs are associated with a significant increased breast cancer risk, all with OR > 2.20 Similar findings were reflected in the CARRIERS study30 although with an ATM OR < 2; possibly reflecting the classification of all variants identified, including missense, as being PGV only where classified as (likely) pathogenic in ClinVar.31 We do not have a comprehensive data set for ATM PGVs and therefore were not included in our analyses; but data from BRIDGES20 would suggest ATM ought to also be included in breast cancer diagnostic panels. For our future studies, we would seek to attain data for germline ATM PGVs and include in our comparisons.
Given truncating CHEK2 PGVs have been associated with a relative risk of breast cancer of 2.2,9 a high risk of contralateral disease,32 and are enriched in this cohort and others, this would substantiate inclusion of testing for CHEK2 PGVs in clinical diagnostic breast cancer genetic testing panels.
Considering segregation of PALB2 PGVs and CHEK2_1100delC within a family, we detected perfect concordance for PALB2 in FDR with breast cancer but not for CHEK2 with only 7/13 testing positive for the familial variant. This likely reflects the moderate risk predisposition of CHEK2_1100delC and that breast cancer risk arises from the combined contribution of single PGVs and polygenic risk score (PRS) with PRS having a greater contribution where the single PGV effect is lower. Recently it has been shown that the combination of a high-risk breast cancer PRS and CHEK2_1100delC equated to a breast cancer lifetime risk equivalent to that of a PALB2 PGV alone.33 These data suggest that CHEK2_1100delC should be incorporated into diagnostic breast cancer genetic testing panels as this knowledge of moderate risk PGVs, combined with a PRS, will become part of routine practice in breast cancer risk assessment.
In this single-center comprehensive study of germline PALB2 PGVs and the CHEK2_1100delC PGV in breast/ovarian cancer, we show a 1% detection rate for PALB2 PGVs, 1.4% for CHEK2_1100delC, and ~7% for BRCA1/2 PGVs. Breast cancers associated with PALB2 PGVs tended toward phenotypes seen with a BRCA2 PGV, namely both triple negative and high-grade ER-positive HER2-negative tumors. The detection of PALB2 and CHEK2 PGVs, in addition to PGVs detected in BRCA1/2, in breast/ovarian cancer is important for accurate risk assessment and the activation of subsequent cancer prevention and early detection strategies in the individual and their family.
Data availability
The data analyzed in this study are available from the corresponding author on request.
References
Ferlay, J. et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer. 144, 1941–1953 (2019).
Newman, B. et al. Inheritance of human breast cancer: evidence for autosomal dominant transmission in high-risk families. Proc. Natl. Acad. Sci. U.S.A. 85, 3044–3048 (1988).
Anglian Breast Cancer Study Group. Prevalence and penetrance of BRCA1 and BRCA2 mutations in a population-based series of breast cancer cases. Br. J. Cancer. 83, 1301–1308 (2000).
Lalloo, F. et al. Prediction of pathogenic mutations in patients with early-onset breast cancer by family history. Lancet. 361, 1101–1102 (2003).
Easton, D. F. et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 372, 2243–2257 (2015).
Rahman, N. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat. Genet. 39, 165–167 (2007).
Antoniou, A. C. et al. Breast-cancer risk in families with mutations in PALB2. N. Engl. J. Med. 371, 497–506 (2014).
Yang, X. et al. Cancer risks associated with germline PALB2 pathogenic variants: an international study of 524 families. J. Clin. Oncol. 38, 674–685 (2020).
Meijers-Heijboer, H. et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 31, 55–59 (2002).
Schutte, M. et al. Variants in CHEK2 other than 1100delC do not make a major contribution to breast cancer susceptibility. Am. J. Hum. Genet. 72, 1023–1028 (2003).
Weidner, A. E. et al. Breast cancer screening implications of risk modeling among female relatives of ATM and CHEK2 carriers. Cancer. 126, 1651–1655 (2020).
Taylor, A. et al. Consensus for genes to be included on cancer panel tests offered by UK genetics services: guidelines of the UK Cancer Genetics Group. J. Med. Genet. 55, 372–377 (2018).
NHS England. National Genomic Test Directory. Testing criteria for rare and inherited disease. https://www.england.nhs.uk/wp-content/uploads/2018/08/Rare-and-Inherited-Disease-Eligibility-Criteria-November-2020-21.pdf (2020).
Howell, A. et al. Long-term evaluation of women referred to a breast cancer family history clinic (Manchester UK 1987–2020). Cancers (Basel). 12, 3697 (2020).
Evans, D. G. et al. Improvement in risk prediction, early detection and prevention of breast cancer in the NHS Breast Screening Programme and family history clinics: a dual cohort study. (Southampton, UK, NIHR Journals Library, 2016).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Moran, A. et al. Risk of cancer other than breast or ovarian in individuals with BRCA1 and BRCA2 mutations. Fam. Cancer. 11, 235–242 (2012).
Evans, D. G. et al. Addition of pathology and biomarker information significantly improves the performance of the Manchester scoring system for BRCA1 and BRCA2 testing. J. Med. Genet. 46, 811–817 (2009).
GraphPad. https://www.graphpad.com/quickcalcs/ (2021).
Breast Cancer Association Consortium, Dorling, L., Carvalho, S. et al. Breast cancer risk genes—association analysis of rare coding variants in 34 genes in 60,466 cases and 53,461 controls. N. Engl. J. Med. 384, 428–439 (2021).
The National Institute for Health and Care Excellence. Clinical guideline. Familial breast cancer: classification and care of people at risk of familial breast cancer and management of breast cancer and related risks in people with a family history of breast cancer. https://www.nice.org.uk/guidance/cg164/evidence/full-guideline-pdf-190130941 (2021).
Hu, C. et al. The contribution of germline predisposition gene mutations to clinical subtypes of invasive breast cancer from a clinical genetic testing cohort. J. Natl. Cancer Inst. 112, 1231–1241 (2020).
Mavaddat, N. et al. Pathology of breast and ovarian cancers among BRCA1 and BRCA2 mutation carriers: results from the Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA). Cancer Epidemiol. Biomarkers Prev. 21, 134–147 (2012).
Ducy, M. et al. The tumor suppressor PALB2: inside out. Trends Biochem. Sci. 44, 226–240 (2019).
Tung, N. M. et al. TBCRC 048: phase II study of olaparib for metastatic breast cancer and mutations in homologous recombination-related genes. J. Clin. Oncol. 38, 4274–4282 (2020).
Grzymski, J. J. et al. Population genetic screening efficiently identifies carriers of autosomal dominant diseases. Nat. Med. 26, 1235–1239 (2020).
Behl, S. et al. Founder BRCA1/BRCA2/PALB2 pathogenic variants in French-Canadian breast cancer cases and controls. Sci. Rep. 10, 1–17 (2020).
Vagena, A. et al. PALB2 c. 2257C>T truncating variant is a Greek founder and is associated with high breast cancer risk. J. Hum. Genet. 64, 767–773 (2019).
Tung, N. & Silver, D. P. Chek2 DNA damage response pathway and inherited breast cancer risk. J. Clin. Oncol. 29, 3813–3815 (2011).
Hu, C. et al. A population-based study of genes previously implicated in breast cancer. N. Engl. J. Med. 384, 440–451 (2021).
National Center for Biotechnology Information. ClinVar. https://www.ncbi.nlm.nih.gov/clinvar/ (2021).
de Bock, G. H. et al. Tumour characteristics and prognosis of breast cancer patients carrying the germline CHEK2*1100delC variant. J. Med. Genet. 41, 731–735 (2004).
Mars, N. et al. The role of polygenic risk and susceptibility genes in breast cancer over the course of life. Nat. Commun. 1, 6383 (2020).
Acknowledgements
We thank the BRIDGES study investigators for their contribution to panel sequencing of a subset of our study participants. D.G.E., E.F.H., E.R.W., M.J.S., E.v.V., H.J.B., W.G.N., S.J.H., and A.H. are supported by the Manchester National Institute for Health Research (NIHR) Biomedical Research Center (IS-BRC-1215-20007). J.M.E. is funded by a postdoctoral research fellowship from the Health Education England Genomics Education Programme. The genotyping work was supported by the Biomedical Research Center and Prevent Breast Cancer (GA19-002).
Author information
Authors and Affiliations
Contributions
E.R.W., E.M.v.V., A.H., W.G.N., M.J.S., and D.G.E. conceptualized the study. E.M.v.V., J.M.E., N.L.B., G.J.B., H.S., and A.J.W. interpreted the sequencing analyses. E.R.W., E.M.v.V. and D.G.E. interpreted the data and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics Declaration
Participants provided clinical or research consent for extended testing of breast cancer associated genes (approval from the North Manchester Research Ethics Committee, reference 09/H1008/81 [PROCAS] and 08/H1006/77).
Competing interests
D.G.E. has received travel grants from AstraZeneca. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Woodward, E.R., van Veen, E.M., Forde, C. et al. Clinical utility of testing for PALB2 and CHEK2 c.1100delC in breast and ovarian cancer. Genet Med 23, 1969–1976 (2021). https://doi.org/10.1038/s41436-021-01234-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01234-6
This article is cited by
-
Cascade screening in HBOC and Lynch syndrome: guidelines and procedures in a UK centre
Familial Cancer (2024)