Abstract
Purpose
ADP ribosylation factor guanine nucleotide exchange factors (ARFGEFs) are a family of proteins implicated in cellular trafficking between the Golgi apparatus and the plasma membrane through vesicle formation. Among them is ARFGEF1/BIG1, a protein involved in axon elongation, neurite development, and polarization processes. ARFGEF1 has been previously suggested as a candidate gene for different types of epilepsies, although its implication in human disease has not been well characterized.
Methods
International data sharing, in silico predictions, and in vitro assays with minigene study, western blot analyses, and RNA sequencing.
Results
We identified 13 individuals with heterozygous likely pathogenic variants in ARFGEF1. These individuals displayed congruent clinical features of developmental delay, behavioral problems, abnormal findings on brain magnetic resonance image (MRI), and epilepsy for almost half of them. While nearly half of the cohort carried de novo variants, at least 40% of variants were inherited from mildly affected parents who were clinically re-evaluated by reverse phenotyping. Our in silico predictions and in vitro assays support the contention that ARFGEF1-related conditions are caused by haploinsufficiency, and are transmitted in an autosomal dominant fashion with variable expressivity.
Conclusion
We provide evidence that loss-of-function variants in ARFGEF1 are implicated in sporadic and familial cases of developmental delay with or without epilepsy.
Similar content being viewed by others
INTRODUCTION
Vesicular trafficking is a critical process in eukaryotic cells. It allows numerous membrane-enclosed compartments to exchange proteins and lipids and is directly involved in countless cellular mechanisms including organelle biogenesis, cellular signaling, or membrane dynamics.1 Its paramount importance is shown by the substantial number of human disorders that derive from the dysfunction of interorganellar trafficking.2,3 Among those involved in vesicle formation are ADP ribosylation factors (ARF), which promote the coating of secretory vesicles in Golgi traffic and their molecular switches: guanine nucleotide exchange factors (GEFs) of the GBF1/BIG family.4 These proteins are composed of a Sec7 domain responsible for the catalysis of nucleotide exchange on ARFs.5 Among them is ADP ribosylation factor guanine nucleotide exchange factor 1 (ARFGEF1), also known as Brefeldin A-inhibited guanine nucleotide exchange protein 1 (BIG1). ARFGEF1 is a 200-kDa protein encoded by ARFGEF1 (MIM *604141), a 39-exon gene mapping to the chromosome 8q13 locus. This ubiquitously expressed small GTPase is highly conserved among mammals and eukaryotes4,6 and has been shown to be implicated in vesicle formation and to be essential for the maintenance of the Golgi apparatus’ structure and function.7,8 Its implication in cell trafficking between the Golgi apparatus and the plasma membrane has been shown to regulate axon elongation, neurite development and maintenance, and the polarization process.9
After initial studies implicated the 8q13 locus, ARFGEF1 was identified as the candidate gene within that interval that accounts for rolandic epilepsy,10 familial febrile convulsions11,12 and epileptic encephalopathy.13,14 Furthermore, following the discovery of an ARFGEF1 nonsense variant in a patient diagnosed with Lennox–Gastaut syndrome (LGS), Teoh and collaborators generated a haploinsufficient mouse model using CRISPR/Cas9 technology. The mice displayed developmental delay, altered cerebral and neuronal morphology, and had a high susceptibility to seizures.15 These findings were congruent with previous investigations in which BIG1-deficient mice showed altered axonal projection, delayed neural polarization, and had a smaller neocortex and hippocampus due to increased neuronal apoptosis.16
Our data therefore suggest ARFGEF1 as an interesting candidate gene for developmental delay and epilepsy.
MATERIALS AND METHODS
In silico analysis
ARFGEF1 sequences were all obtained from NCBI RefSeq database as FASTA files and aligned with EMBOSS Clustal Ω tool keeping its default settings.17 Human and yeast sequences were also pairwise-aligned with the EMBOSS water tool keeping its default settings. Modeling of the structure was performed using the resolved fragment 3LTL available at the RCSB Protein Data Bank (https://www.rcsb.org/). All structures were modeled using PyMol version 2.3.4 (https://pymol.org/). Different scripts were written to generate the structures. After calculation, all were rendered with shaders on and saved as png files. The PyMol Wizard and Protein Contact Potential tools were used to generate the putative mutant structures and to render the false red/blue charge-smoothed surface representation respectively.
Splicing reporter minigenes analysis
In this study, we used the reference transcript NM_006421.4.
To evaluate the impact on splicing of the NM_006421.4:c.2392G>A variant, we performed splicing minigene reporter assays using the pCAS2 vector based on a previously described protocol.18 This functional assay is based on the comparative analysis of the splicing pattern of wild-type and mutant after transfection of the pCAS2 plasmid in human cell lines. Briefly, the exon of interest and its intronic flanking regions were amplified from a DNA sample, using a high-fidelity polymerase (CloneAmp HiFi PCR, Takara). The resulting fragment was introduced into the pCAS2 plasmid according manufacturer’s instructions using the In-Fusion HD Cloning kit (CloneAmp HiFi PCR, Takara) and was transformed into the kit’s competent cells. Plasmids were purified (NucleoSpin Plasmid Mini kit, Macherey–Nagel) and were Sanger sequenced between the cloning sites. Next, minigenes representing the wild-type and the mutant allele (1 μg/well) were transfected into a human cell line using the FuGENE 6 transfection reagent (Promega). Total RNA was isolated 24 hours after transfection using the NucleoSpin RNA Plus kit (Macherey–Nagel) according to the manufacturer’s instructions. The concentration of total RNA was measured on a NanoDrop spectrophotometer (NanoDrop Technologies). Reverse transcription was performed with the ProtoScript II First Strand cDNA Synthesis Kit (New England Biolabs). Polymerase chain reaction (PCR) was performed on complementary DNA (cDNA) with the Platinum DNA Polymerase (Thermo Fisher Scientific) according to the PCR amplicons (PCR conditions and protocol details are available upon request). PCR amplicons were verified by electrophoresis (agarose gel or Caliper LabChip) and Sanger sequenced to evaluate the splicing effect.
Cell cultures, proteins extraction, and western blot analysis
Cell cultures, protein extraction, and western blot analyses were performed as previously described.19 Detailed information is available in Supplemental methods.
RESULTS
Patient cohort and clinical features
At the time of the first evaluation, individual 1 was a 5-year-old boy with global developmental delay and without any relevant family history. Pregnancy and delivery had been uneventful. He presented with severe language delay and was speaking only a couple of words at the age of 5 years. Clinical examination showed mild gait impairment suggestive of subtle cerebellar ataxia. Brain magnetic resonance image (MRI) performed at the age of 6 years revealed a thin corpus callosum. Psychometric evaluations revealed moderate intellectual disability. He had mild facial dysmorphism with a wide mouth, high forehead, and low-set ears. Trio exome sequencing (proband and both parents) identified a de novo guanine to adenine transition, lying within a CG dinucleotide, in the ARFGEF1 gene (NM_006421.4:c.2392G>A), and predicted to result in a missense variant (p.[Asp798Asn]) within the Sec7 domain at a genomic position showing intolerance to amino acid change (MetaDome nonsynonymous over synonymous ratio dN/dS score of 0.22)20 (Supplemental Fig. 1) and associated with deleterious in silico scores (Combined Annotation Dependent Depletion [CADD]21 = 34; PolyPhen-222 = 1.0; Genomic Evolutionary Rate Profiling [GERP]23 = 4.87).
Through an international collaboration facilitated by GeneMatcher,24 Decipher, and RD-connect25 platforms, we gathered 12 additional individuals from 10 unrelated families with rare likely pathogenic ARFGEF1 variants (Supplemental Table 1) identified by exome sequencing. These affected individuals displayed overlapping clinical features including developmental delay, intellectual disability, and behavioral problems with or without epilepsy (Supplemental Table 2). Overall, 6/11 variants occurred de novo and 3/11 were inherited from an affected or mildly affected father. Interestingly, Sanger sequencing results indicated that paternal variants were germline rather than mosaic (Supplemental Fig. 2). This observation was corroborated by the fact that in one family (the family of individuals 8 and 9) a paternal aunt also carried the variant. In 2/11 cases, the mode of inheritance could not be established due to lack of parental DNA.
Of note, 2 of 11 variants had been observed once in the gnomAD cohort: NM_006421.4:c.4033C>T/p.(Arg1345*) and NM_006421.4:c.3697C>T/p.(Gln1233*) but both were absent from healthy gnomAD controls (individuals who were not selected as a case in a case/control study of common disease) and one (p.[Gln1233*]) was absent in the gnomAD non-neurological cohort (individuals who were not ascertained for having a neurological condition in a neurological case/control study).
Interestingly, 12/13 (92%) individuals were males and only 1/13 (8%) was a female (Chi-squared = 9.3077, df = 1, p value = 0.002282; Fig. 1a). Twelve individuals were children with a mean age on referral of 8.6 years (Table 1). All individuals were born following uneventful pregnancies and delivered without perinatal complications with normal Apgar scores. Delivery was at term for 12/13 individuals and at 34 weeks of gestation for one individual. Only one individual was born to consanguineous parents. Two children (15%) had a birth weight under the 5th percentile with normal height and occipital–frontal circumference (OFC) while birth parameters were within normative values for the other 11 individuals.
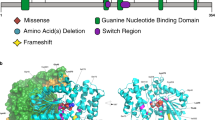
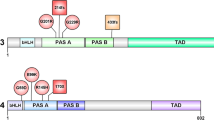
(a) Family trees from individuals with ARFGEF1 variants. Squares represent males and circles females. Individuals with heterozygous ARFGEF1 variants are indicated with a +, while − indicates absence of a variant. UN DNA unavailable for family segregation of the variant. Index cases are shown with a black arrow. (b) Schematic representation of ARFGEF1 protein showing the variants identified in this study. All domains are positioned and adapted as previously described4. Missense variants are shown in black and truncating in red.
All individuals had developmental delay of variable degree. First symptoms were noticed at a mean age of 22 months old, and before 3 years of age for all individuals. Almost all individuals (12/13) had predominant language delay, which was often severe, with 7/12 having a vocabulary of only few words at 6 years of age. Gross and fine motor skills were impaired for most children (12/13 and 10/13, respectively) but with slow progressive improvement following adequate rehabilitation. Of note, all children were ambulatory by 3 years of age. Behavioral problems were frequent (12/13 individuals) and often severe with variable occurrence of autism spectrum disorders, anxiety, aggressivity, anger bursts, stereotypies, attention disorders, and psychomotor agitation. Ten individuals had intellectual disability (ID) of moderate (4/10) or mild (6/10) degree. None of the children displayed severe ID and only one child had intellectual abilities at the lower limit of normal (borderline IQ).
Associated clinical features included hyperopia (4/13), astigmatism (3/13), frequent otitis media (2/13), strabismus (1/13), occipital meningocele (1/13), interstitial lung disease with bronchiectasis and chronic atelectasis (1/13), and alcohol addiction in the only adult patient (1/13). Most individuals (8/13) displayed facial dysmorphisms that were mostly mild. Some dysmorphic features were shared among the individuals including large (5/8) and low-set (4/8) ears, wide mouth (3/8), high forehead (3/8), bulbous nasal tip (3/8), and thin long facies (2/8) (Fig. 2). We performed a computer assisted facial visualization26 from all available photographs in order to generate a “typical” ARFGEF1 patient’s face using the Facer program.27 These analyses suggested that morphological features were mild in ARFGEF1-related disorders and highlighted some of the morphological features identified in the individuals: high forehead, wide mouth, and bulbous nasal tip (Fig. 2). Most recent available growth parameters were within normal ranges for 9 individuals (69%) with 3 displaying macrocephaly (between +2.1 and 2.5 SD deviation) and 1 displaying a small head circumference (−3.4 SD).
Neurological examinations were mostly unremarkable or limited to signs related to motor development delay while cerebellar ataxia was noted in two individuals and hypotonia in another two individuals. Of note, one patient displayed an action tremor and another presented with clumsiness and a balance disorder, raising the possibility of a mild underlying cerebellar ataxia in these two additional individuals. One patient presented with left laryngeal paralysis and limited left eye abduction. Six individuals were followed for epilepsy, while a seventh individual was treated with valproic acid but could not specify whether it was for neurologic or psychiatric reasons. Many seizure types were observed including tonic–clonic seizures (4/6), atonic seizures (2/6), myoclonic seizures (2/6), focal motor seizures (1/6), and typical absence seizures (1/6). Three patients were suspected to have LGS (one with generalized drug-resistant epilepsy with slow spike and wave on electroencephalogram (EEG), and two others who had generalized tonic–clonic seizures along with myoclonic and atonic seizures but without drug resistance and for whom access to EEG was not possible, and one had childhood absence epilepsy. Brain MRIs were available for ten individuals and revealed abnormalities in eight of them including white matter T2-weighted signal hyperintensity compatible with hypomyelination (3/8), type 1 Arnold–Chiari malformation (1/8), low-set cerebellar tonsils (1/8), thin corpus callosum (1/8), and meningocele with periventricular heterotopia and disrupted superior cerebellar vermis (1/8) (Fig. 3).
Brain MRI of individual 3 carrying the NM_006421.4:c.3592-2A>G variant (a–j), and of individual 2 carrying the p.(Arg1774*) variant (k–o). T1-weighted sagittal (a), axial T1 (b–e) and T2 fluid-attenuated inversion recovery (FLAIR) (k–m), and coronal (f–I and n–o) brain imaging. Note the nodular periventricular heterotopia (white arrows), occipital meningocele (blue arrow), and disrupted cerebellar vermis (red arrows) for patient 3 and the bilateral and diffuse signal hyperintensities of the subcortical white matter, especially of the posterior regions (yellow arrows) for individual 2.
Family history was unremarkable in seven individuals (54%), while six (46%) had at least one relative with a history of developmental delay, learning disorder, behavioral problems, or milder phenotypes with only difficulties at school requiring special education or classroom accommodations (Fig. 1). Furthermore, almost half (6/13) of the individuals inherited the variant from a parent who either presented with a milder version of their child’s phenotype, or for whom retrospective phenotyping revealed a history of difficulties at school, learning disorder, or a need for special education. For example, patient 7 is a child with severe drug-resistant epilepsy. Detailed clinical characterization following his molecular diagnosis revealed that his father, from whom he inherited the variant, had a self-limited childhood onset epilepsy. In individuals 8 and 9, revisiting the family history revealed that their father had global developmental delay, with marked behavioral problems (aggressivity and temper tantrums) resulting in difficulties at school. For individuals 10 and 11, further questioning revealed that their father had mild developmental delay and borderline intellect. Additionally, two of individuals 8’s and 9’s paternal cousins displayed prominent difficulties at school, behavioral problems, and delayed development requiring speech therapy; however, none of them had genetic testing as no diagnostic procedure was initiated. Yet, the familial segregation study revealed that the mother of these two relatives (individuals 8’s and 9’s paternal aunt) carried the variant and displayed intellectual abilities at the lower limit of normal. Thus, we suspect that these two cousins also carry the familial ARFGEF1 variant, although their biological samples were not available for confirmation.
Molecular findings and functional studies
Our cohort consisted of nine loss-of-function (LoF) variants (six stop-gain and three frameshift), one variant impacting a splice site (Supplemental Table 1, Fig. 1), and one missense variant. Given that all of the LoF variants identified in our cohort introduced premature stop codons before the last exon or upstream to the 50–55 nucleotides preceding the last exon–intron junction, all LoF variants were predicted to result in nonsense-mediated mRNA decay (NMD)28 rendering the locus haploinsufficient.
ARFGEF1 is a gene under high mutational constraint (Supplemental Fig. 1). The gnomAD missense Z score was 5.37, with two regional missense constraint regions corresponding to amino acid positions 482–1,301 and 1,302–1,850 with a calculated score of 0.37 (p value = 3.39 × 10–21) and 0.67 (p value = 2.38 × 10–5) respectively (source: Decipher https://decipher.sanger.ac.uk/, UniProt29 accession Q9Y6D6 https://www.uniprot.org). The probability of LOF intolerance (pLI) score is 1, with an observed/expected (o/e) ratio equal to 0.08 and an upper bound confidence interval equal to 0.14 (source gnomAD30 v2.1.1, https://gnomad.broadinstitute.org/).
These observations suggested that the mechanism responsible for ARFGEF1-related disorders is haploinsufficiency. However, a functional assessment of the impact of the only missense variant identified in our cohort on pre-mRNA splicing or protein stability was required to corroborate our hypothesis. Splicing prediction using the Alamut Visual v.2.15 software predicted that no obvious splicing defect results from the c.2392G>A transition. However, this variant was predicted to create a new SRp40 Exonic Splice Enhancer (ESE) (Supplemental Fig. 3). We thus performed functional assays in order to investigate the experimentally observed functional effect of the c.2392G>A variant.
To that end, we performed a minigene reporter assay using the pCAS2 vector as previously described (Supplemental Fig. 4).18 This assay allows a rapid screening of aberrant splicing events driven by DNA variants, cloned within a reporter vector containing two exons (named A and B) derived from the human SERPING1/C1NH gene, separated by an intron with BamHI and MluI cloning sites, and under the CMV-promoter-driven transcriptional control. This assay showed that the c.2392A allele induced exon skipping in approximately 22% of the reporter transcripts, corresponding to a 2.4-fold increase as compared to the clones carrying the wild-type c.2392G allele associated with 9% baseline skipped transcripts (Supplemental Fig. 4) suggesting that a splicing effect of this variant was unlikely. Then, we obtained a fibroblast cell line from patient 1 and performed targeted splicing analysis, using next-generation sequencing, on a PCR-derived amplicon obtained from retrotranscribed RNA. These analyses allowed us to exclude any possible splicing defect associated with c.2392G>A in vitro (Supplemental Fig. 5). Of note, these experiments also revealed that the variant allelic frequency (VAF) of c.2392G>A in retrotranscribed RNA was close to 50%, indicating a biallelic expression of ARFGEF1 in fibroblasts (Supplemental Fig. 6). Using the same approach, we also investigated the splicing defect associated with the de novo c.3592–2A>G variant in a blood sample obtained from patient 3. As expected, the variant triggered exon 26 skipping, inducing the loss of 152 nucleotides and resulting in NMD (Supplemental Fig. 7).
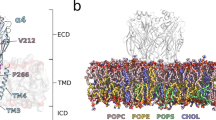
Structurally, the polypeptide surrounding the Asp798 residue is conserved across species (Fig. 4). In silico three-dimensional modeling of the Asp798 residue within the Sec7 domain, based on the human resolved fragment (PDB 3LTL), allowed us to predict the corresponding steric changes induced by the Asp798Asn variant (Fig. 4a) in support of a possible impact on protein misfolding or catalytic activity.
(a) The resolved fragments available in the RSCB Protein Data Bank are shown in the cartoon together with the position of the discussed mutation from leucine 695. (b) Alignment of the sequence region surrounding the variation in S. cerevisiae, S. pombe, C. elegans, M. musculus, D. melanogaster and H. sapiens. Sequences were fetched from NCBI RefSeq database. Fasta file combining the extracted protein sequences were then subjected to EMBOSS Clustal tool with default settings33. For mutation at position 798, the alignment shows a highly conserved region spanning on 18 amino acids in all 6 aligned sequences. The aspartic acid in position 798 represent the wild-type situation in all considered sequences including the yeast S. cerevisiae. (c) Modeling of mutation calculated in PyMol. The resolved fragment 3LTL was loaded with PyMol. The sequence spanning from proline 730 to lysine 885 was analyzed and modelled. The identied F, H and J loops of ARFGEF1 are rendered in red, orange and blue respectively. Asp798 was highlighted and is shown to probably interact with two tyrosines (Tyr825 and Tyr829) belonging to loop H (left panel). (d) Protein contact potential calculated in PyMol exhibits the charges modification when the mutation is simulated in the structure (from proline 730 to lysine 885). The top two insets show an enlargement of the aspartic acid 798 region. In the wild-type situation (WT), the area appears largely colored in red, while after simulating the Asp798Asn replacement, the same region charges are exhibiting a large channel colored in blue (dotted ovals). Similarly, 90° rotation in both horizontal and vertical directions of the structure give view on the rendered simulation of the Met884Val replacement as shown in the two bottom insets. The charge modification observed exhibits a clear shift (dotted circles). Black stars indicate the position of the amino acids subject to mutation Asp798 (top star), Met884 (bottom star). (e) western blot from individual 1 (p.(Asp798Asn)) and individuals 8 and 9’s father (p.(Gln648*)). Note the reduced levels of protein as compared to control samples (CTRL). Vinculin is used as a loading control.
Using an ARFGEF1-specific antibody, we then performed western blot analyses on protein extracts obtained from fibroblast cell lines obtained from patient 1 and the symptomatic father of Individuals 2 and 3 carrying the variant NM_006421.4:c.2392G>A; p.(Gln648*), and unrelated and healthy control individuals (Fig. 4b). Interestingly, both cell lines derived from affected individuals revealed comparable reduced levels of protein expression when compared with healthy controls, indicating that c.2392G>A (p.[Asp798Asn]) resulted in protein instability and degradation.
Taken together, these data suggest that the pathogenic effect of these variants is compatible with haploinsufficiency.
DISCUSSION
Although high-density single-nucleotide polymorphism (SNP) array,10 linkage analysis,11 and exome sequencing meta-analysis14 had previously linked the ARFGEF1 locus to increased risk for rolandic epilepsy10 and epileptic encephalopathy,14 there had been no substantial evidence to support an association between ARFGEF1 and high penetrance of a neurodevelopmental disorder conforming to Mendelian inheritance expectations. More recently, one patient with LGS harboring a truncating variant (p.Cys1455*) in ARFGEF1 has been reported by Teoh and collaborators,15 requiring additional observations in order to confirm its implication as a disease-causing gene. Here, we describe a cohort of 13 affected individuals gathered through an international collaboration that harbor likely pathogenic variants in ARFGEF1 identified via exome sequencing.
Interestingly, while almost half (6/13) of the individuals were followed for epilepsy, all of them displayed developmental delay with impaired motor skills, speech delay, or both (Table 1). When analyzing the cohort as a whole, features of this disorder including mild motor delay, mild or moderate intellectual disability, severe speech delay, and numerous behavioral problems such as attention disorders, psychomotor agitation, anxiety, or autism spectrum disorders. Regarding epilepsy, five different seizure types were observed in our cohort, ranging from focal motor seizures to generalized tonic–clonic seizures and although EEGs were unavailable for some individuals, two patients were suspected to have LGS and a third one was formally diagnosed with LGS (EEG available) leading to up to half of the individuals with epilepsy to display a LGS phenotype. ARFGEF1 thus appears as a possible gene contributing to LGS.
This increased susceptibility to seizures may be related to ARFGEF1’s involvement in cellular trafficking of GABAA receptors. Indeed, previous work has shown that ARFGEF1 is a binding partner of GABAA receptors, and that depletion of ARFGEF1, either through small interfering RNA (siRNA)31 or haploinsufficient mouse models,15 led to a decrease in GABAA receptors at the neuronal surface, which in turn might lead to impaired neuronal inhibition and higher seizure susceptibility. This point may prove critical in the future management of epileptic patients carrying pathogenic variants in ARFGEF1 and will require special attention from physicians when using GABA-based drugs, whose efficiency might be reduced in such individuals.
As shown by our genetic and functional data, the pathophysiology of variants in ARFGEF1 is consistent with haploinsufficiency. This is consistent with previously published functional studies, particularly in yeast, which showed that all protein domains are critical for the overall functioning of the protein.32 Based on the molecular data from our study, the variable clinical features and clinical severity observed in different individuals in our cohort do not correlate with the localization of the variants along the gene. However, identification of further individuals carrying deleterious variants might bring additional information about the clinical spectrum associated with ARFGEF1-related dysfunction.
Family pedigrees and histories show ARFGEF1-related disorders display an autosomal dominant mode of inheritance (Fig. 1a). As no rare copy-number nor additional likely pathogenic variants were identified in our cohort to account for intergenerational variability, variants in this gene seems to be subject to highly variable expressivity. Indeed, as explained above, one third of the children had inherited the variant from an apparently healthy parent for whom the question of a genetic neurodevelopmental disorder had not been raised during the initial assessment but only once the ARFGEF1 variant had been identified as part as the familial segregation analysis (reverse phenotyping, Fig. 1). This point is not particularly surprising as mild difficulties at school or in learning in general can sometimes be tolerated. The situation of the cousins of individuals 8 and 9 was particularly illustrative of this and also shows how two individuals carrying expected pathogenic variants in ARFGEF1 can be found in gnomAD, including one in the non-neurological cohort.
Regarding the findings on brain MRI, the most common abnormality was a delayed myelination shown by subcortical white matter signal hyperintensity. This is consistent with what was observed in mice that displayed altered neurodevelopment with delayed neural polarization and increased neural apoptosis.15,16 One individual—patient 3 with the focal motor seizures—displayed periventricular nodular heterotopies (PH), a feature that echoes the brain abnormalities observed in a Turkish family with ARFGEF2 (periventricular heterotopia with microcephaly, autosomal recessive; OMIM 608097) pathogenic variants who displayed severe developmental delay, microcephaly, early-onset refractory epilepsy, bilateral nodular periventricular heterotopia, and frequent infections,33 and a phenotype that partially overlaps the one observed in our cohort. Two hypotheses have been put forward to explain PH in ARFGEF2-related disorders. First, BIG2 inhibition has been shown to increase phosphorylation of Filamin A (a protein well known to be associated with PH34), thus disrupting cell intrinsic neuronal migration, which favors PH.35 Second, BIG2 has been shown to be involved in neuron migration through (1) regulation of actin dynamics and (2) its critical importance for vesicle and membrane trafficking, mechanisms that are fundamental regulators of proliferation and migration during human cerebral cortical development.35,36 We may thus hypothesize that ARFGEF1 pathogenic variants could also generate PH as BIG1 has also been shown to be involved in intracellular vesicle trafficking,7 axon elongation,9 and actin dynamics/directed migration.36
Surprisingly, our cohort mostly consisted of male individuals (12/13) and the ascertained inherited variants were inherited from mildly symptomatic fathers who carried Sanger-confirmed heterozygous variants. This appears surprising as no clear difference is observed between male and female individuals regarding tissue expression of ARFGEF1 according to databases such as GTEx37 or the human protein atlas.38 Furthermore, our RNA and protein expression data show that ARFGEF1 expression is biallelic in clinically accessible tissues (i.e., skin-derived fibroblasts and whole blood) although we cannot exclude that it might be selectively imprinted in other tissues during development. Thus, no clear biological explanation appears to account for this unbalanced sex ratio, although members of the ARF family have been implicated in asymmetrical cell division in female meiosis in the mouse39,40 and potentially provide an explanation for this observation. Further studies and reports should help untangle whether a real sex-dependent incidence of the disorders exists or whether this happened in our cohort simply by chance.
To conclude, we describe a cohort of 13 individuals with de novo or inherited likely pathogenic ARFGEF1 variants affected with developmental disorders of varying degree, offering a description of clinical phenotypes associated with this gene. We provide evidence that ARFGEF1 should be considered as a new gene responsible for intellectual disability, developmental delay, and syndromic epilepsy.
Data availability
Data are available upon request.
References
Muro, S. Alterations in cellular processes involving vesicular trafficking and implications in drug delivery. Biomimetics. 3, 19. https://doi.org/10.3390/biomimetics3030019 (2018).
Aridor, M. & Hannan, L. A. Traffic jams II: an update of diseases of intracellular transport. Traffic. 3, 781–790, https://doi.org/10.1034/j.1600-0854.2002.31103.x (2002).
Aridor, M. Visiting the ER: the endoplasmic reticulum as a target for therapeutics in traffic related diseases. Adv. Drug Deliv. Rev. 59, 759–781, https://doi.org/10.1016/j.addr.2007.06.002 (2007).
Wright, J., Kahn, R. A. & Sztul, E. Regulating the large Sec7 ARF guanine nucleotide exchange factors: the when, where and how of activation. Cell. Mol. Life Sci. 71, 3419–3438, https://doi.org/10.1007/s00018-014-1602-7 (2014).
Cherfils, J. et al. Structure of the Sec 7 domain of the Arf exchange factor ARNO. Nature. 392, 101–105, https://doi.org/10.1038/32210 (1998).
Mansour, S. J., Skaug, J., Zhao, X. H., Giordano, J., Scherer, S. W. & Melancon, P. p200 ARF-GEP1: a Golgi-localized guanine nucleotide exchange protein whose Sec7 domain is targeted by the drug brefeldin A. Proc. Natl. Acad. Sci. USA 96, 7968–7973, https://doi.org/10.1073/pnas.96.14.7968 (1999).
Boal, F. & Stephens, D. J. Specific functions of BIG1 and BIG2 in endomembrane organization. PLoS One. 5, e9898. https://doi.org/10.1371/journal.pone.0009898 (2010).
Zhao, X., Lasell, T. K. R. & Melançon, P. Localization of large ADP-ribosylation factor-guanine nucleotide exchange factors to different Golgi compartments: evidence for distinct functions in protein traffic. Mol. Biol. Cell. 13, 119–133, https://doi.org/10.1091/mbc.01-08-0420 (2002).
Zhou, C. et al. BIG1, a brefeldin A-inhibited guanine nucleotide-exchange protein regulates neurite development via PI3K-AKT and ERK signaling pathways. Neuroscience. 254, 361–368, https://doi.org/10.1016/j.neuroscience.2013.09.045 (2013).
Addis, L. et al. Identification of new risk factors for rolandic epilepsy: CNV at Xp22.31 and alterations at cholinergic synapses. J. Med. Genet. 55, 607–616, https://doi.org/10.1136/jmedgenet-2018-105319 (2018).
Wallace, R. H., Berkovic, S. F., Howell, R. A., Sutherland, G. R. & Mulley, J. C. Suggestion of a major gene for familial febrile convulsions mapping to 8q 13-21. J. Med. Genet. 33, 308–312, https://doi.org/10.1136/jmg.33.4.308 (1996).
Piro, R. M., Molineris, I., Ala U. & Di Cunto, F. Evaluation of candidate genes from orphan FEB and GEFS+ loci by analysis of human brain gene expression atlases. PLoS One. 6, e23149, https://doi.org/10.1371/journal.pone.0023149 (2011).
Appenzeller, S. et al. De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am. J. Hum. Genet. 95, 360–370, https://doi.org/10.1016/j.ajhg.2014.08.013 (2014).
Takata, A. et al. Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy. Nat. Commun. 10, 2506, https://doi.org/10.1038/s41467-019-10482-9 (2019).
Teoh, J. J. et al. Arfgef1 haploinsufficiency in mice alters neuronal endosome composition and decreases membrane surface postsynaptic GABAA receptors. Neurobiol. Dis. 134, 104632, https://doi.org/10.1016/j.nbd.2019.104632 (2020).
Teoh, J. J. et al. BIG1 is required for the survival of deep layer neurons, neuronal polarity, and the formation of axonal tracts between the thalamus and neocortex in developing brain. PLoS One. 12, 1–24, https://doi.org/10.1371/journal.pone.0175888 (2017).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641, https://doi.org/10.1093/nar/gkz268 (2019).
Soukarieh, O. et al. Exonic splicing mutations are more prevalent than currently estimated and can be predicted by using in silico tools. PLoS Genet. 12, 1–26, https://doi.org/10.1371/journal.pgen.1005756 (2016).
Da Costa, R. et al. Neutralization of HSF1 in cells from PIK3CA-related overgrowth spectrum patients blocks abnormal proliferation. Biochem. Biophys. Res. Commun.530, 520–526. https://doi.org/10.1016/j.bbrc.2020.04.146 (2020).
Wiel, L., Baakman, C., Gilissen, D., Veltman, J. A., Vriend, G. & Gilissen, C. MetaDome: pathogenicity analysis of genetic variants through aggregation of homologous human protein domains. Hum. Mutat. 40, 1030–1038, https://doi.org/10.1002/humu.23798 (2019).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894, https://doi.org/10.1093/nar/gky1016 (2019).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 76, 7.20.1–7.20.41. https://doi.org/10.1002/0471142905.hg0720s76 (2013).
Cooper, G. M., Stone, E. A., Asimenos, G., Green, E. D., Batzoglou, S. & Sidow, A. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–13, https://doi.org/10.1101/gr.3577405 (2005).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 928–930, https://doi.org/10.1002/humu.22844 (2015).
Lochmüller, H. et al. RD-Connect, NeurOmics and EURenOmics: Collaborative European initiative for rare diseases. Eur. J. Hum. Genet. 26, 778–785, https://doi.org/10.1038/s41431-018-0115-5 (2018).
Ferry, Q. et al. Diagnostically relevant facial gestalt information from ordinary photos. Elife. 3 e02020, https://doi.org/10.7554/eLife.02020.001 (2014).
GitHub. johnwmillr/Facer: Simple face averaging in Python. https://github.com/johnwmillr/Facer (2020).
Nagy, E. & Maquat, L. E. A rule for termination-codon position within intron-containing genes: When nonsense affects RNA abundance. Trends Biochem. Sci. 23, 198–199, https://doi.org/10.1016/S0968-0004(98)01208-0 (1998).
Bateman, A. et al. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212, https://doi.org/10.1093/nar/gku989 (2015).
Karczewski, K. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 581, 434–443, https://doi.org/10.1038/s41586-020-2308-7 (2020).
Li, C. et al. BIG1, a brefeldin A-inhibited guanine nucleotide-exchange factor, is required for GABA-gated Cl- influx through regulation of GABA A receptor trafficking. Mol. Neurobiol. 49, 808–819, https://doi.org/10.1007/s12035-013-8558-8 (2014).
Ramaen, O. et al. Interactions between conserved domains within homodimers in the BIG1, BIG2, and GBF1 Arf guanine nucleotide exchange factors. J. Biol. Chem. 282, 28834–28842, https://doi.org/10.1074/jbc.M705525200 (2007).
Sheen, V. L. et al. Mutations in ARFGEF2 implicate vesicle trafficking in neural progenitor proliferation and migration in the human cerebral cortex. Nat. Genet. 36, 69–76, https://doi.org/10.1038/ng1276 (2004).
Liu, W., Yan, B., An, D., Xiao, J., Hu, F. & Zhou, D. Sporadic periventricular nodular heterotopia: classification, phenotype and correlation with Filamin A mutations. Epilepsy Res. 133, 33–40, https://doi.org/10.1016/j.eplepsyres.2017.03.005 (2017).
Zhang, J., Neal, J., Lian, G., Shi, B., Ferland, R. J. & Sheen, V. Brefeldin A-inhibited guanine exchange factor 2 regulates Filamin a phosphorylation and neuronal migration. J. Neurosci. 32, 12619–12629, https://doi.org/10.1523/JNEUROSCI.1063-12.2012 (2012).
Le, K., Li, C. C., Ye, G., Moss, J. & Vaughan, M. Arf guanine nucleotide-exchange factors BIG1 and BIG2 regulate nonmuscle myosin IIA activity by anchoring myosin phosphatase complex. Proc. Natl. Acad. Sci. USA. 34, E3162–E3170. https://doi.org/10.1073/pnas.1312531110 (2013).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585, https://doi.org/10.1038/ng.2653 (2013).
Uhlen, M. et al. Tissue-based map of the human proteome. Science. 347, 1260419–1260419, https://doi.org/10.1126/science.1260419 (2015).
Duan, X., Zhang, H. L., Pan, M. H., Zhang, Y. & Sun, S. C. Vesicular transport protein Arf6 modulates cytoskeleton dynamics for polar body extrusion in mouse oocyte meiosis. Biochim. Biophys. Acta Mol. Cell. Res. 1865, 455–462, https://doi.org/10.1016/j.bbamcr.2017.11.016 (2018).
Wang, S., Hu, J., Guo, X., Liu, J. X. & Gao, S. ADP-ribosylation factor 1 regulates asymmetric cell division in female meiosis in the mouse1. Biol. Reprod. 80, 555–62, https://doi.org/10.1095/biolreprod.108.073197 (2009).
Acknowledgements
We thank the families and patients for taking part in the study. We thank the University of Burgundy Centre de Calcul (CCuB) for technical support and management of the informatics platform, and the GeneMatcher platform for data sharing. We thank the Centre de Ressources Biologiques Ferdinand Cabanne (CHU Dijon) for sample biobanking. This work was supported by grants from Dijon University Hospital, the ISITE-BFC (PIA ANR) and the European Union through the FEDER programs. Also supported in part by the US National Institutes of Health, National Human Genome Research Institute (NHGRI) to the Baylor Hopkins Center for Mendelian Genomics (UM1HG006542). This work was supported by the National Institute for Health Research (NIHR) Manchester Biomedical Research Centre. The Deciphering Developmental Disorders (DDD) study presents independent research commissioned by the Health Innovation Challenge Fund (grant number HICF-1009–003). This study makes use of DECIPHER (http://decipher.sanger.ac.uk), which is funded by the Wellcome. See Nature PMID 25533962 or www.ddduk.org/access.html for full acknowledgement. D.M. is also supported by a Medical Genetics Research Fellowship Program through the United States National Institute of Health (T32 GM007526–42). Several authors of this publication are members of the European Reference Network for Developmental Anomalies and Intellectual Disability (ERN-ITHACA) A.J., A-S.D-P., and A.V. are supported by Solve-RD. The Solve-RD project has received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement number 779257. J.E.P. was supported by NHGRI K08 HG008986.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.V., C.T.-R., L.F., L.D. Data curation: Q.T., A.V., Y.D., P.G. Investigation: Q.T., D.M., M.W., S.M., B.I., B.C., S.C., R.T., M.I., A.S., A.M., T.D., A.J., S.B., J.D., N.S., F.A., F.AZ., P.A., E.E., J.M. L.B., D.R., F.TM-T. A-L.B., P.C., N.M., A-S.D-P., C.P. Methodology: A.V., L.D., M.C., S.N., T.B., T.G., J.G. Supervision: Q.T., A.V., D.M. Visualization: T.G., Q.T., A.V, Writing - original draft: Q.T., A.V., L.F., C.T.-R. Writing – review & editing: all authors.
Corresponding authors
Ethics declarations
Ethics declaration
All affected individuals or their legal representative gave informed consent for the sequencing procedures and the publication of their results along with clinical and molecular data. Special consent forms were signed authorizing publication of pictures when relevant. The study was performed within the framework of the GAD (“Génétique des Anomalies du Développement”) collection and approved by the appropriate institutional review board of Dijon University Hospital (DC2011‐1332).
Competing interests
J.R.L. has stock ownership in 23andMe, is a paid consultant for Regeneron Genetics Center, and is a co-inventor on multiple United States and European patents related to molecular diagnostics for inherited neuropathies, eye diseases, and bacterial genomic fingerprinting. The Department of Molecular and Human Genetics at Baylor College of Medicine receives revenue from clinical genetic testing conducted at Baylor Genetics (BG) Laboratories. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Thomas, Q., Gautier, T., Marafi, D. et al. Haploinsufficiency of ARFGEF1 is associated with developmental delay, intellectual disability, and epilepsy with variable expressivity. Genet Med 23, 1901–1911 (2021). https://doi.org/10.1038/s41436-021-01218-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01218-6
This article is cited by
-
Golgipathies reveal the critical role of the sorting machinery in brain and skeletal development
Nature Communications (2022)