Abstract
Purpose
We describe the pathogenic variant spectrum and identify predictors of positive results among men referred for clinical genetic testing for prostate cancer.
Methods
One thousand eight hundred twelve men with prostate cancer underwent clinical multigene panel testing between April 2012 and September 2017. Stepwise logistic regression determined the most reliable predictors of positive results among clinical variables reported on test requisition forms.
Results
A yield of 9.4–12.1% was observed among men with no prior genetic testing. In this group, the positive rate of BRCA1 and BRCA2 was 4.6%; the positive rate for the mismatch repair genes was 2.8%. Increasing Gleason score (odds ratio [OR] 1.19; 95% confidence interval [CI] 0.97–1.45); personal history of breast or pancreatic cancer (OR 3.62; 95% CI 1.37–9.46); family history of breast, ovarian, or pancreatic cancer (OR 2.32 95% CI 1.48–3.65); and family history of Lynch syndrome–associated cancers (OR 1.97; 95% CI 1.23–3.15) were predictors of positive results.
Conclusion
These results support multigene panel testing as the primary genetic testing approach for hereditary prostate cancer and are supportive of recommendations for consideration of germline testing in men with prostate cancer. Expanding the criteria for genetic testing should be considered as many pathogenic variants are actionable for treatment of advanced prostate cancer.
Similar content being viewed by others
INTRODUCTION
Germline pathogenic variants (PVs) in cancer predisposition genes are reported in 7.3% to 11.8% of aggressive prostate cancer (PC) cases, including genes associated with homologous repair deficiency (HRD) (e.g., BRCA1, BRCA2, ATM, BRIP1, CHEK2, NBN, BARD1, RAD51C, MRE11A, and PALB2), and mismatch repair (MMR) deficiency (e.g., MLH1, MSH2, MSH6, and PMS2).1,2 Most of these genes have clear management guidelines for early cancer detection and risk reduction, which may benefit the patient and family members. Their relationship to PC screening and management has garnered recent interest.3 Previous reports suggest that PVs in BRCA1/2 confer increased risk for PC associated with poor survival and younger age at diagnosis; HOXB13 PVs also are associated with a young age at diagnosis.4,5 Men at increased risk for aggressive or earlier-onset disease may choose more aggressive screening or earlier intervention.6 Men with HRD or MMR-deficient metastatic prostate tumors may also benefit from targeted therapeutics, such as pembrolizumab, platinum therapies, or PARP inhibitors.7,8,9
Despite improved understanding of the prevalence of PVs among men with PC, it remains unclear which men will most benefit from genetic testing. Historically, the Hopkins criteria, i.e., ≥3 affected first-degree relatives, affected relatives in three successive generations, or ≥2 relatives affected at 55 years or younger, provided a working definition of hereditary PC (HPC), but there is little evidence that HPC is associated with DNA repair gene variants. Recent data suggest that aggressive disease or family history of other cancers may be a better predictor for germline PVs in men with PC.1,2 In 2017, expert consensus guidelines recommended genetic testing for men with HPC; men with ≥2 close unilinear relatives with a cancer associated with hereditary breast and ovarian cancer (HBOC) or Lynch syndrome (LS), men with metastatic castration-resistant PC, and men with somatic (PVs) identified via tumor testing.10 However, these testing guidelines are based on limited evidence. Thus, there is an urgent need to determine a more robust way of identifying men with PVs so they may benefit from early screening/intervention and therapeutic options. This study evaluates predictors of germline PV status in a large cohort of men with a personal history of PC who underwent clinical genetic testing to help inform clinical testing guidelines.
MATERIALS AND METHODS
Study population
Study participants included men with PC who underwent hereditary cancer multigene panel testing (MGPT) between April 2012 and September 2017 at a clinical diagnostic laboratory (Ambry Genetics) (n = 1878). Men with a BRCA1/2 PV reported in their family prior to testing were excluded (n = 66), leaving a total of 1812 individuals in the analyzed cohort. Patients who had prior genetic screening or testing (N = 150), including BRCA1/2 or LS germline testing, immunohistochemical screening for MMR deficiency in tumors, and other somatic testing, were analyzed separately to minimize bias in PV detection rates; 1662 men had no prior testing. Demographic and clinical data including age at testing, ethnicity, age of diagnosis, Gleason score, metastatic status, and personal and family history of cancer were collected through retrospective review of test requisition forms and other clinical documentation (e.g., pedigrees and consult notes) provided to the laboratory.
Laboratory methods
Depending on the type of clinical tests ordered, men underwent analysis of up to 67 cancer susceptibility genes. The frequencies of each gene tested are described in Table S1. Sanger or next-generation sequencing analysis was performed for all coding domains and well into the flanking 5’ and 3’ ends of all introns and untranslated regions, along with gross deletion/duplication analysis of covered exons and untranslated regions. Exceptions included GREM1, EPCAM, and MITF, for which analysis was limited to alterations known to be associated with disease (Table S1).
Variant classification
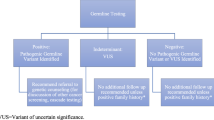
Variants were interpreted using a five-tier variant classification protocol (pathogenic variant [PV]; variant, likely pathogenic [VLP]; variant of unknown significance [VUS]; variant, likely benign [VLB]; and benign), based on published recommendations from the American College of Medical Genetics and Genomics and the International Agency for Research on Cancer.11,12,13 All identified alterations were deposited in ClinVar.
Statistical analysis
Descriptive statistics for the PC cohort are summarized as median (interquartile range [IQR]) for continuous and percentages for categorical characteristics. Logistic regression estimated odds ratios (OR) (95% confidence interval [CI]) for univariate associations between PV carrier status and personal and family history characteristics, by gene and for the combined set of PC predisposition genes (ATM, BRCA1, BRCA2, CHEK2, EPCAM, HOXB13, MLH1, MSH2, MSH6, NBN, PALB2, PMS2, RAD51D, and TP53). Stepwise logistic regression determined the most informative set of predictors from the following: continuous age at PC diagnosis; Ashkenazi Jewish ethnicity; personal history of non-PC; personal history of breast or pancreatic cancer; family history (presence of first- or second-degree relative) of cancer; >1 first or second-degree relative with PC; >1 first- or second-degree relative with breast, ovarian, or pancreatic cancer; >1 first- or second-degree relative with LS-related cancer (colorectal, endometrial, gastric, ovarian, pancreatic, small bowel, urothelial, kidney, or bile duct cancer); and Gleason score. The stepwise selection procedure aimed to minimize Akaike’s information criterion (AIC), allowing a maximum of 1000 backward and forward steps. All analyses were conducted in R V3.3.3.
RESULTS
Demographics
The study cohort was primarily Caucasian (70%; Table 1). The median age at testing was 66 (IQR 59, 73) years and the median age of PC diagnosis was 60 (IQR 54, 66) years. Forty-two percent had a personal history of other cancers, including colorectal (11.1%), breast (5.8%), and pancreatic (6.1%). Most individuals (92.4%) had a family history of cancer in at least one close (first, second, or third degree) relative; with 52.0% having a history of breast cancer and 50.6% PC. Thirty-six percent of men had more than one relative with breast, ovarian, or pancreatic cancer and 31.6% had more than one family member with a LS-related cancer. Gleason scores were available for approximately half of PC cases. Of the men with available Gleason scores, 17.7% were ≤6; 43.2% were equal to 7; and 39.1% were 8–10.
Germline genetic test results
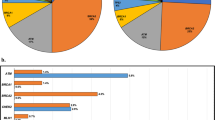
Among men with no prior genetic testing, the yield of a 14-gene hereditary prostate cancer panel ProstateNext was 9.4%, with 26/277 testing positive for a PV. The yield of all other MGPTs combined was 12.1%, with 168/1385 testing positive (Table S2). Among candidate prostate cancer genes, PV frequencies were highest for BRCA2 (3.8%), ATM (2.7%), and CHEK2 (2.5%). PVs were also seen in MSH2, HOXB13, BRCA1, MSH6, PMS2, PALB2, TP53, NBN, MLH1, and EPCAM. No PVs were detected in RAD51D (Fig. 1, Table S3). Among all men with no prior genetic testing, the pooled frequency of PV in therapeutically actionable genes (BRCA1/2 and MMR genes) was 7.4%. Nine men who underwent MGPT (0.6%) had PVs in genes not currently associated with PC; all had clinical history consistent with the alteration identified (Table S4). Fourteen men in the entire cohort (0.7%) were found to have more than one PV (Table S5). Among men with prior genetic testing, results for 34/40 were consistent with previous findings, such as confirmation of somatic PVs in the germline or PVs that were concordant with tumor immunohistochemistry (IHC) results; 6/40 were found to have discordant results with tumor testing or additional PVs identified on expanded testing that were not detected with initial limited testing (Figure S1).
Predictors of positive test results
Univariate analysis
The type of panel (ProstateNext vs. all other panels) used for testing was not associated with positive or negative results (p = 0.10). There was no significant difference in median (IQR) age at PC diagnosis for men testing positive versus negative (59 [11] vs. 60 [12] years; p = 0.32) (Table 2). Having a personal history of another cancer in addition to PC was significantly associated with a positive test result, (OR 1.35; 95% CI 1.03–1.75; p = 0.027). Having a first-degree relative with PC was not significant (p = 0.59), nor was having any family history of PC (p = 0.56). Similarly, men who satisfied Hopkins criteria were not significantly more likely to have a positive result (p = 0.78). However, having more than one family member with breast, ovarian, or pancreatic cancer was significantly associated with positive result status (OR 1.46; 95% CI 1.12–1.90; p = 0.005), as was having more than one family member with a LS-related cancer (OR 1.33; 95% CI 1.01–1.74; p = 0.040). Positive result status was marginally associated with higher Gleason score (OR per unit: 1.17; 95% CI 0.99–1.38; p = 0.07), although Gleason information was missing for 49.4% of the cohort. In this cohort, metastatic status was not significantly associated with a positive result (p = 0.47); however, this information was only available for 13.1% of the cohort.
Multivariate analysis
To assess predictors of positive results, a subset of the cohort was analzyed, comprised of men with an available Gleason score who did not have prior genetic testing whose testing included all 14 of these genes (n = 524). In a multivariate model, the most informative predictors of a positive result were Gleason score (OR 1.19; 95% CI 0.97–1.45; p = 0.096); personal history of breast or pancreatic cancer (OR 3.62; 95% CI 1.37–9.46; p = 0.008); >1 family member with breast, ovarian, or pancreatic cancer (OR 2.32; 95% CI 1.48–3.65; p = <0.001); and >1 family member with a LS-related cancer (OR 1.97; 95% CI 1.23–3.15; p = 0.004) (Table 3). The multivariable adjusted OR for Gleason score represents the comparison per 1 unit increase in Gleason score (i.e., men with a Gleason score of 7 were 19% more likely to test positive than men with a Gleason score of 6). Similarly, men with a personal history of breast or pancreatic cancer were more than three times as likely to test positive than men without these cancers; men with more than one family member with breast, ovarian, or pancreatic cancer were 2.3 times as likely to test positive compared with men without this family history; and men with more than one family member with a LS-related cancer were nearly twice as likely to test positive compared with men who do not have this family history. Ninety-five percent of men with PVs reported at least one of these informative predictors of a positive result.
DISCUSSION
The findings from this clinical laboratory genetic testing cohort demonstrate a 9.4–12.1% yield of PVs in men with PC who elected testing and identified predictors of positives result status to include increasing Gleason score; personal history of breast or pancreatic cancer; family history of breast, ovarian, or pancreatic cancer; and family history of LS -associated cancers. The ProstateNext yield of 9.4% was comparable with that previously reported by Pritchard et al. (11.8%) in a 16-gene panel targeted for genes presumed to be associated with PC1 in a cohort of men with metastatic PC and in a more recently reported cohort in selected DNA repair genes by Giri et al. (10.9%).14 The present yield of other MGPT (12.1%) is lower than recently reported by Nicolosi et al. (17.2%);15 however, Nicolosi et al. may have overestimated the PV frequency because the MGPT utilized included up to 80 genes, individuals with prior known familial PVs were not excluded, monoallelic MUTYH PVs were included, and the authors included the CFTR gene in their data set, which has a variant that is present in 1/24 European Caucasians.15 When restricting to genes relevant to PC screening and advanced disease (e.g., HOXB13 and HRD and MMR genes), findings from this study are similar to Nicolosi et al.
Among PC patients referred for genetic testing, BRCA2, ATM, CHEK2, and HOXB13 are the most commonly mutated genes. Here, 26/1501 men (1.7%) with no prior genetic testing whose testing included the LS genes (MLH1, MSH2, MSH6, PMS2, or EPCAM) were found to have a PV in one these MMR genes, and 66/1662 men (4.0%) with no prior testing were found to have BRCA1 or BRCA2 PVs. Men with advanced PC who are found to have PVs in the HRD or MMR genes may benefit from targeted therapeutic agents, such as pembrolizumab, platinum therapies, or PARP inhibitors.7,8,9 Positive men with PC may benefit from identifying increased risk for additional cancers, identifying risks to family members, and understanding the cause of their cancer diagnosis.
Stepwise regression analysis was used to identify factors for identifying PC patients with a PV. Factors associated with an increased risk for a positive result include increasing Gleason score; personal history of breast or pancreatic cancer; family history of breast, ovarian, or pancreatic cancer; and family history of LS-associated cancers. These findings are consistent with previous studies and current genetic testing recommendations.1,10 Recent expert consensus guidelines reflect that germline testing should be considered in men with aggressive disease, somatic PVs, or family history suggestive of HBOC or LS.10 Although a recent publication found no correlation between Gleason score or family history of HBOC or LS-related cancers and positive result status, our present methodology of employing multivariate analysis including only individuals who were tested for all 14 genes included on a prostate gene–specific panel eliminates confounders that were not previously accounted for.15 Recently, Nicolosi et al. proposed universal testing among all men with PC, citing an inability to find reliable criteria to predict which men will most benefit from genetic testing.15 Although the approach of universal testing proposed by Nicolosi et al. is appealing with respect to increased identification of at-risk patients and simplicity in application, additional studies are needed to assess the diagnostic yield and clinical utility of testing men with clinically localized low-risk disease and no other personal/family history suggestive of inherited cancer predisposition relative to targeted testing strategies.
Despite the potential therapeutic benefit of identifying a PV, we found that on average, there was a six-year delay between time of PC diagnosis and genetic testing. Arguably, targeted therapies for HRD and MMR-deficient tumors have been recently developed and may not have been available for most men at the time of their diagnosis. Going forward, the timing of genetic testing may become more integrated with treatment planning for PC. While the present study focuses on men with PC, ideally genetic testing will identify at-risk men before they are diagnosed so that genetic information can be used for surveillance and clinical decision making. Further analysis of unaffected men is warranted.
There are several limitations that deserve mention, including the fact that this cohort represents men clinically selected for genetic testing; 40.0% of this cohort had multiple primary cancers, indicating high threshold for genetic testing. Further, a clinical laboratory cohort is likely to give an overestimation burden of positive results; therefore, additional studies are needed to determine whether these predictors remain informative in an unselected group of men with PC. Men were tested with one of several multigene panel tests, which could potentially influence the PV frequencies we report, as shown in Table 4. However, our primary analyses limited to individuals tested for the same set of 14 PC susceptibility genes yielded no significant difference in the rate of PVs detected by panel type. This study was also limited by the data provided on the test requisition forms at the time of testing. While results from a recent study of predominately women with a history of breast cancer demonstrate that clinical history reported on test requisition forms is of comparable quality to clinic notes for most probands and their close relatives, it is unknown if the requisition data are similarly valid for prostate cancer and Gleason score reporting.16 Additional analysis of the association of metastatic status with positive genetic result status may provide further insight, as metastatic status was available for only 13.1% of the cohort, and we did not find a significant association with positive result status, which differs from previous reports.1,2 While the present findings regarding Gleason score were consistent with previous reports, Gleason scores were available for only 51.1% of the cohort.
The present results support multigene panel testing as the primary genetic testing approach for hereditary PC and confirm Gleason score, personal history of breast or pancreatic cancer, and family history of cancers related to HBOC and LS as informative predictors for positive genetic test results. These results provide generalizability of previous findings and support current recommendations for consideration of germline testing in men with PC.
References
Pritchard CC, Mateo J, Walsh MF, et al. Inherited DNA-repair gene mutations in men with metastatic PC. N Eng J Med. 2016;375:443–453.
Leongamornlert D, Saunders E, Dadev T, et al. Frequent germline deleterious mutations in DNA repair genes in familial PC cases are associated with advanced disease. Br J Cancer. 2014;110:1663–1672.
Columbus G. Gomella gives predictions for PC advances in 2018. Oncolive. https://www.onclive.com/onclive-tv/dr-gomella-on-screening-recommendations-in-prostate-cancer.
Leao RRN, Price AJ, Hamilton J. Germline BRCA mutation in male carriers—ripe for precision oncology? PC Prostatic Dis. 2018;21:48–56.
Witte JS, Mefford J, Plummer SJ, et al. HOXB13 mutation and PC: studies of siblings and aggressive disease. Cancer Epidemiol Biomarkers Prev. 2013;22:675–680.
Ballentine Carter H, Albertsen PC, Barry MJ, et al. Early detection of PC: AUA guideline. J Urol. 2013;190:419–426.
Kaufman B, Shapira-Frommer R, Schmutzler RK, et al. Olaparib monotherapy in patients with advance cancer and a germline BRCA11/2 mutation. J Clin Oncol. 2015;33:244250.
Mateo J, Carreira S, Sandhu S, et al. DNA-repair defects and olaparib in metastatic PC. N Engl J Med. 2015;373:1697–1708.
Imyanitov EN, Moiseyenko VM. Drug therapy for hereditary cancers. Hered Cancer Clin Pract. 2011;91:5.
Giri VN, Knudsen KE, Kelly WK, et al. Role of genetic testing for inherited PC risk: Philadelphia PC Consensus Conference 2017. J Clin Oncol. 2018;36:414–424.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Plon SE, Eccles DM, Easton D, et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat. 2008;29:282–291.
Pesaran T, Karam R, Huether R, et al. Beyond DNA: An integrated and functional approach for classifying germline variants in breast cancer genes. Int J Breast Cancer. 2016;2016:2469523.
Giri VN, Hegarty SE, Hyatt C, et al. Germline genetic testing for inherited prostate cancer in practice: implications for genetic testing, precision therapy, and cascade testing. Prostate. 2019;79:333–339.
Nicolosi P, Ledet E, Yang S, et al. Prevalence of germline variants in PC and implications for current genetic testing guidelines. JAMA Oncol. 2019;5:523–528.
LaDuca H, McFarland R, Gutierrez S, et al. Quality of clinician-reported cancer history when ordering genetic testing. JCO Clin Cancer Inform. 2018;2:1–11.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The following individuals are current or previous employees of Ambry Genetics, a Konica Minolta Company: M.P., Y.T., P.R., A.J.S., K.A., S.G., M.J., J.S.D., H.L., and M.H.B. The other authors declare no conflicts of interest.
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. A waiver was provided for this research project by the Western Institutional Review Board.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pritzlaff, M., Tian, Y., Reineke, P. et al. Diagnosing hereditary cancer predisposition in men with prostate cancer. Genet Med 22, 1517–1523 (2020). https://doi.org/10.1038/s41436-020-0830-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0830-5
Keywords
This article is cited by
-
Are population level familial risks and germline genetics meeting each other?
Hereditary Cancer in Clinical Practice (2023)
-
Prostate cancer in transgender women: considerations for screening, diagnosis and management
British Journal of Cancer (2023)
-
Performance of clinical risk scores and prediction models to identify pathogenic germline variants in patients with advanced prostate cancer
World Journal of Urology (2023)
-
Mammalian tumor-like organs. 1. The role of tumor-like normal organs and atypical tumor organs in the evolution of development (carcino-evo-devo)
Infectious Agents and Cancer (2022)
-
Inherited risk assessment and its clinical utility for predicting prostate cancer from diagnostic prostate biopsies
Prostate Cancer and Prostatic Diseases (2022)