Abstract
Purpose
This report describes the return of sequencing results to low-income Latino participants recruited through a Federally Qualified Health Center (FQHC). We describe challenges in returning research results secondary to social determinants of health and present lessons learned to guide future genomic medicine implementation studies in low-resource settings.
Methods
Five hundred Latino adults (76% women) consented to research sequencing for a predetermined panel of actionable genes. Providers and staff from the FQHC were engaged to align processes with the practice and a community advisory board grounded the project in the local community.
Results
A pathogenic/likely pathogenic variant was present in 10 participants (2%). Challenges in return of results included the time lag (582 ± 53 days) between enrollment and returning actionable results, difficulty reaching participants, missed appointments, low health literacy, lack of health insurance, and reconciling results with limited information on family history. Return of one actionable result was deferred due to acute emotional distress secondary to recent traumatic life events.
Conclusion
The social determinants of health influence the implementation of genomic medicine in low-income populations in low-resource settings. Considering nonbiological factors that contribute to disparities will be necessary to better appreciate how genomic medicine may fit within the context of health equity.
Similar content being viewed by others
INTRODUCTION
The Electronic Medical Records and Genomics (eMERGE) Network is a consortium of institutions funded by the National Human Genome Research Institute (NHGRI) to facilitate research in genomic medicine, including discovery, clinical implementation, and public health implications.1 At the forefront of the rapidly evolving field of genomic medicine is how to handle returning sequencing results.2 As such, in the most recent eMERGE Network funding cycle (phase III), the Network selected a panel of medically actionable genes for next-generation sequencing with a goal of returning pathogenic/likely pathogenic (P/LP) variants in these genes to participants.3
Mayo Clinic has been a member of the eMERGE Network since its inception. In phase III the Mayo Clinic genomic medicine implementation project, the Return of Actionable Results Empirical (RAVE) study, proposed to return actionable as well as neutral or “negative” research results (i.e., no P/LP findings) to participants. The RAVE study involved a collaboration with Mountain Park Health Center (MPHC), a Federally Qualified Health Center (FQHC) in Phoenix, Arizona. FQHCs are federally funded community health centers that provide comprehensive primary care and preventive services in medically underserved and low-income areas as a means to improve access to care and reduce health disparities.4 The Arizona RAVE study enrolled Latino participants through the Sangre Por Salud (SPS) Biobank, a collaboration between Mayo Clinic and MPHC that is governed by both institutions and designed to facilitate research with an underrepresented population.5 As part of the SPS Biobank, MPHC patients agreed to share data, biospecimens, and contact information with Mayo Clinic to support genomic medicine research studies. The SPS biobank was developed using principles of community-based participatory research6 and is guided by a community advisory board (CAB) comprising MPHC patients and community members.7 The CAB grounds the research activities in the local context and fosters transparency, broader engagement, and shared decision making.
Although biomedical research continues to shed light on disease predisposition attributable to genetic factors, the widening of disparities in some preventable diseases underscores the complexity of interactions that lead to a disproportionate burden of disease among low-income and minority populations.8,9 A compelling body of literature demonstrates that social determinants of health (SDoH) represent root causes of health disparities that are particularly operational in low-income and minority communities.10 SDoH are the conditions in which people are born, grow, live, work, and age. They include, among other things, unequal distribution of resources, poverty, access to health care, transportation, unstable housing, and health literacy.11 Additional SDoH that are operational within MPHC’s patient population include fragmented health-care interactions, food insecurity, exposure to violence/trauma, and social isolation/stigma related to sexual orientation, immigration status, or history of incarceration.
Populations with health disparities remain underrepresented in research in general12 and even more so in genomic research,13 which may limit the translation of genomic research findings within underserved communities. Beyond the inclusion of minority populations, it is critical that genomic medicine research be conducted within the broader context of SDoH to better integrate the complex, multifactorial nature of health disparities.14 However, current implementation science frameworks for genomic medicine have only recently begun to acknowledge the importance of SDoH and appreciate how health-care systems and settings that serve low-income patients will be critical for translating research findings into improved outcomes.15
Despite the importance of including diverse populations in genomic medicine research and the opportunity to engage with FQHCs to meet this need, there remains a gap in our understanding of how to handle return of results (RoR) from underserved populations within community-based settings. Therefore, the current report describes the return of research results from genome sequencing to low-income Latino participants within the FQHC setting. In addition, we collated the challenges faced in effectively returning those results, many of which were directly related to SDoH. Lastly, we offer lessons learned that may be useful for facilitating the implementation of genomic medicine through a lens of health equity.
MATERIALS AND METHODS
Design
The processes, workflow, recruitment, and enrollment of the 500 participants have been described in detail elsewhere.16 Briefly, primary care providers (PCPs) and staff from MPHC were engaged through a series of meetings and presentations to ensure that the processes for returning research results were consistent with those of the practice and that provisions for supporting providers and follow-up management were considered. The CAB was engaged to ensure that RAVE activities were responsive to the cultural, contextual, language, literacy, and socioeconomic factors of the local Latino population. The CAB reviewed written materials, provided input on recruitment scripts, and weighed in on approaches to contact participants once results were available. An iterative process with the Mayo Clinic institutional review board (IRB) led to approvals for English and Spanish documents and procedures that occurred outside of the academic institution. A list of potentially eligible individuals who participated in the SPS Biobank (N = 1621) with phenotypes of interest (hyperlipidemia and/or colorectal neoplasia) was generated. From this list, 500 participants provided written informed consent for sequencing their DNA, returning results, and scanning results into their electronic health record (EHR). As previously presented,16 enrolled participants differed from the nonenrolled sample with respect to age, insurance, country of birth, and full-time employment status but did not differ in sex, language preference, or educational attainment.
Sequencing
The DNA from 500 participants was sent to Baylor College of Medicine Human Genome Sequencing Center, a CLIA certified laboratory, for targeted sequencing using the eMERGEseq panel.17 The sequencing laboratory generated individual clinical reports that were subsequently reviewed by a variant curation group at Mayo Clinic before disclosure to participants. Variants of uncertain clinical significance were not provided to investigators or participants.
Returning research results
A letter was mailed to participants who had no pathogenic findings (Supplemental Fig. 1). Within a week of the mailing, the bilingual study coordinator phoned to confirm that the letter was received and inquired if there were any remaining questions and offered the option to discuss results with the medical geneticist (N.M.L.) who was a member of the research team. A copy of the letter along with the full sequencing report were scanned into the participant’s EHR.
For those with actionable research results, defined as a P/LP variant, initial contact was made by phone to inform the participant and schedule the RoR visit with the medical geneticist. In addition to the phone call, a certified letter (Supplemental Fig. 2) was mailed to participants. An alert was placed in the EHR to notify MPHC providers and staff that an actionable result was available for return. The alert did not disclose specifics of the result but was intended to facilitate contact between participants and the research team through the health center. The CAB was instrumental in crafting both letters to facilitate comprehension.
During the participant’s visit with the medical geneticist, the purpose of the RAVE study was reviewed, their consent for RoR was reaffirmed, risks and benefits of receiving research results were discussed, the participant’s medical profile and family history were reviewed, and information regarding the findings was provided. This information included medical implications, inheritance patterns, implications for relatives, and recommendations from guidelines/consensus statements on optimal management. Participants were given the option to have the research result confirmed in an independent CLIA certified lab on a fresh blood specimen. A behavioral health provider from MPHC was available to meet with individuals immediately following the genetics consultation and evaluate the need for psychosocial support. The medical geneticist subsequently met with the participant’s PCP to offer clinical decision support and written resources related to the actionable genetic finding. Sequencing results and the genetic consultation notes were scanned into the EHR. A follow-up visit with the participant’s PCP was scheduled through regular clinical channels at MPHC. Participants with actionable findings were contacted by phone by the research coordinator one week and one month following RoR as a check-in; at this time, the research coordinator also offered to connect participants with MPHC staff or the medical geneticist if desired.
RESULTS
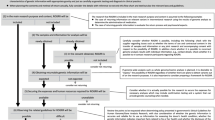
Of the 500 participants sequenced, 486 did not have any P/LP variants while 10 were identified as having an actionable finding (4 individuals were withdrawn due to technical reasons). There were no sociodemographic differences between those with and without actionable results (Table 1). Letters were mailed to the 486 participants without pathogenic findings and 926 follow-up phone calls were made to confirm receipt. No participant requested consultation with the medical geneticist to discuss their negative result.
The ten individuals identified with P/LP variants (Table 2) required 32 outreaches (calls and letters) to schedule the RoR with the medical geneticist. The average time between enrollment and recontacting participants with actionable results was 582 ± 53 days. Two participants were “no-shows” for their RoR visits on two separate occasions despite confirming with the research coordinator on the evening prior to their visit. These visits were eventually completed where it was noted that transportation issues, child care, and last-minute work opportunities contributed to missed appointments. Of the ten individuals with actionable results, eight were female, eight preferred Spanish over English, only one had more than a high school education, eight were uninsured at the time of RoR, and the majority (6/10) had not seen their PCP at MPHC within the previous 12 months (Table 3). Only one participant was found to have an actionable result (in LDLR) related to their enrollment phenotype (hyperlipidemia). Follow-up medical recommendations varied from advanced breast magnetic resonance imaging (MRI) to electrocardiograms. Given the number of uninsured patients and the lack of access to specialty care, universal adherence to recommended guidelines was challenging.
The research coordinator was unable to make contact with one participant (pathogenic variant in LDLR) who was eventually re-engaged after an unrelated medical visit at an MPHC site not involved with the study. The treating provider noted the alert placed in the EHR and facilitated re-engagement of the participant with the study team to return the research sequencing result. The RoR visit with the medical geneticist was scheduled within a week of recontact at the site where the participant was receiving care. This particular experience underscores the reach of MPHC as an integrated health-care model that serves a mobile population of more than 90,000 patients through seven community clinics within a coordinated system.
A recurring challenge in making recommendations was the lack of information on family history. Family members were often geographically displaced, lost at young ages, and/or did not have access to regular health care. As a result, family histories were incomplete and in some cases unknown. Examples include actionable results in which a positive family history might be expected, but the family and personal medical history was entirely unremarkable (BRCA1, SCN5A, PALB2, TP53). Whether this represented nonpenetrance, de novo variant, nonpaternity, or simply an unknown family history is unclear. For some, lack of family history information made it difficult to establish optimal management recommendations. For example, a BRCA2 pathogenic variant was found in a man in his 40s who was uninsured. P/LP variants in BRCA2 are associated with hereditary breast and ovarian cancer syndrome in women, as well as male breast cancer, pancreatic cancer, and prostate cancers. In his family, six cancers were reported (two breast, one colon) but the primary sites for the other three were unknown. Screening for pancreatic cancer is not routine and recommendations are predicated on family history. The participant was subsequently educated on breast self-examination by the PCP. During a follow-up call from the research coordinator, he reported hypervigilance in performing multiple daily self-examinations and he desired a mammogram but found that free mammography programs in the community were only available for women. He was referred back to the PCP by the study team to have follow-up questions answered and additional education on what to look for during self-examinations.
In contrast to incomplete family histories, nonconforming family histories offered additional challenges. A pathogenic variant in TP53 was found in a woman in her late 30s. P/LP variants in this gene lead to one of the most highly penetrant cancer predisposition syndromes, Li–Fraumeni syndrome, characterized by high rates of childhood cancers and lifelong very high risks for adult-onset cancers. The participant reported a large family (7 children and 15 nieces/nephews) that was unaffected by any form of cancer except for her mother with possible uterine cancer in her 40s. Discordance between genetic findings and clinical presentation is not altogether unique but in those without insurance and limited financial resources, the decision to screen per guidelines is associated with additional considerations.
The lack of apparent penetrance in genes considered to have high penetrance (e.g., BRCA1, TP53) raised concerns about technical errors. To address this concern, confirmatory testing was offered at no cost to participants with P/LP variants (of those that were requested, all actionable research results were confirmed). The opportunity for confirmation as well as payment for such testing were not in the original research plan nor had resources been put aside for this. Therefore, the research team worked with MPHC to establish a mechanism for collecting and sending samples to the independent lab with the costs absorbed through the investigator’s discretionary funds. This option was reassuring to participants as well as the practice.
Even when family histories conformed to sequencing results, low health literacy and fragmented interactions with the health-care system presented challenges. The 50-year-old woman with a LP FBN1 variant (which can cause Marfan syndrome) reported that her father died of a ruptured aorta, her sister underwent a recent aorta replacement, and a very tall, thin brother died suddenly at age 25 due to an unclear cause. The participant also reported congenital dislocation of her lenses (an FBN1/Marfan feature), early bilateral total hip replacements, and being told she had an “enlarged” heart. Despite the family history and known phenotypic characteristics associated with Marfan syndrome, the condition had never reportedly been mentioned to the family. After explanations by the geneticist and the PCP, the participant seemed to make the connection between the research result and her family and medical history. However, in subsequent interactions, it was apparent that true digestion of this information required additional explanation. She asked multiple questions of how her genetic findings could explain her and her family’s medical history. Comprehension became clear when the participant expressed concern about family members who have not yet experienced FBN1-related problems. She wanted her family members tested and requested education materials related to Marfan syndrome to distribute to relatives.
A particularly challenging situation encountered was the participant in his 40s with a LP variant where disclosure of the finding was deferred. During the RoR visit, he was despondent and tearful, sharing that he had sustained several recent very significant traumas/losses in his life. Though eager to talk, he asserted that he was not in a good place to absorb another psychological burden. RoR was deferred and a behavioral health consultation was provided in real time by MPHC. Five of the ten participants with actionable results had stress or anxiety previously identified within their EHR problem list and seven of the ten participants with P/LP results were seen by a behavioral health provider following RoR.
DISCUSSION
Most research on returning clinically actionable genetic results has been conducted in high-resource, academic environments.18 Although that work has provided useful guidance for best practices under highly controlled conditions, whether such guidance can be readily translated to primary care settings is unclear.19 This is particularly relevant in the context of low-resourced practices and underrepresented communities where there is a disproportionate burden of preventable diseases.20 To address this gap, the Arizona RAVE study set out to bring genomic medicine to a large FQHC providing primary care to a population with many barriers to obtaining health care. To embed the project within the clinical setting, staff from MPHC were integrated as key members of the research team and PCPs were engaged throughout the project. To ground the project in the larger community, a CAB was leveraged for feedback and advice.
Through the process and collaboration with MPHC, we were able to appreciate how SDoH may influence implementation of genomic medicine in a low-resource setting. Although there was considerable enthusiasm and support at MPHC for the project, poverty, lack of insurance, language barriers, disconnected phones, transportation issues, low health literacy, lack of family history information/loss of contact and early deaths of key family members, and limited access to medical specialists were challenges experienced during the conduct of the RAVE study.
While the optimal approach for returning negative results remains elusive,21 there was perceived value from MPHC and the CAB in conveying these results to participants. To address health literacy concerns we sought input from our CAB to simplify and clarify the notification letter. Providers at MPHC also recommended that the letter encourage patients to discuss their results with their PCP. While there were no requests for additional information or consultation with the geneticist, it is unknown whether receiving negative results by mail is an effective approach. It is possible that receiving negative results may have been misinterpreted as a “clean bill of health” by participants who were recruited for specific risk phenotypes and may have other medical diagnoses.
More complex than returning negative results was the return of actionable results in this setting. There was a lengthy delay between enrolling in RAVE and recontacting participants with actionable results. The sources for delays may be inherent to the research process and included the necessity to transfer specimens from the Mayo biobank in Phoenix, Arizona to the Mayo biobank in Rochester, Minnesota (parent eMERGE site). From there, they were cataloged and shipped to the core sequencing laboratory in Houston, Texas where the samples were in a queue behind other eMERGE sites. Once sequencing results were available from the laboratory, multiple attempts were required to re-establish contact with participants to schedule the RoR visit. Following contact, considerable effort was required to coordinate the in-person consultation given the need to align schedules of the participant, the medical geneticist, the coordinator (who also was the interpreter), a behavioral health provider, and the business office/registration as well as to reserve clinical space in the busy practice setting. To facilitate scheduling, appointments were offered after hours and on weekends. Despite this flexibility, the importance of the results, and confirming with participants one day prior, we did observe several missed appointments. Recognizing that hard-to-reach populations are underrepresented in genomic research, additional considerations by researchers and funding agencies that desire to extend genomic medicine to vulnerable populations include providing participants with an estimated timeframe for when to expect results, dedicating appropriate effort and resources for maintaining contact with individuals between enrollment and RoR, prioritizing the rapid return of medically actionable genomic results, and offering transportation and child care during visits.
From a health system perspective, no individual with medical genetics expertise existed at MPHC and concern was raised by providers that they would need guidance on how best to manage patients with uncommon genetic conditions. Although the genes and associated conditions reported as actionable may be familiar to geneticists and subspecialists in academic medical centers, this may not be the case for primary care providers in community practices. To address this deficiency, the medical geneticist met one-on-one with the PCP of each participant to review and interpret results, offer clinical decision support, and discuss management guidelines. While such an approach is not scalable, some form of clinical decision support will be necessary to facilitate implementation of genomic medicine in primary care and utilizing EHR-based tools has been proposed as a natural leverage point.22 Currently, such tools are not available at MPHC so integrating provider suggestions into EHR-based clinical decision support could enhance its utility and further tailor precision medicine to the practice setting.23 More broadly, coupling clinical decision support with education in clinical genomics will enhance appropriate application of information into primary care. With a finite list of genes, it is likely feasible for research initiatives to develop materials that will prepare PCPs who are tasked with managing an occasional patient in their practice. However, even if clinical decision support is optimized, the ability to pay for follow-up testing, access to specialists, and cascade testing of family members represent additional challenges that warrant additional consideration.
Despite the deliberated approach, close collaboration with MPHC, and community engagement strategies, we remain uncertain of how to assess the risk-to-benefit ratio of the study or the long-term potential for sustainability.24 While some participants with actionable results expressed appreciation for receiving information that could facilitate prevention strategies, we did not systematically evaluate this in an objective manner. A recent meta-analysis found that returning sequencing results was not associated with adverse psychological harms (e.g., increased anxiety or depression) with some populations reporting positive psychological effects.25 However, whether these findings can be extended to individuals with pre-existing behavioral/mental health disorders such as the individual in our study who deferred RoR warrants further consideration and research. Given the disproportionate burden of violence, crime, and adverse experiences in low-income communities,26 many FQHCs integrate behavioral health into their primary care practice.27 The option of having behavioral health providers available immediately following the consultation with the medical geneticist was suggested by MPHC providers early in the development of the project and further highlights the benefits of engaging with stakeholders who are closely connected to community needs.
Providers at MPHC expressed enthusiasm for the opportunity to integrate genomic medicine into their practice but whether this improves (or hinders) their ability to deliver care is not determined. Nonetheless, MPHC leadership continues to be supportive of genomic medicine despite the uncertainty it may bring to their practice. MPHC is collaborating on additional precision medicine research projects and recently identified precision medicine as a priority area in their strategic plan. However, outside of research studies, avenues to pay for genomic testing in the current health system have not been identified. From a broader perspective, the CAB remains optimistic about this research area despite the challenges ahead for those with actionable findings. Their optimism is based on the potential long-term benefits to the family members, particularly children, of affected individuals and the community at large who are underrepresented in research broadly and genomic research in particular. The CAB commented that advances in science could be more readily translated to disparate populations if those populations were involved early in the spectrum of translational research, including discovery. These sentiments are supported by an extensive body of literature highlighting the benefits of community-based participatory research,28 which may be a useful approach for informing and facilitating precision medicine research initiatives in underrepresented communities.29
Notwithstanding best efforts to balance scientific rigor with real-world application, there are several limitations that are worthy of comment. First, the relatively small sample size of the cohort limits generalizability. The lack of long-term follow-up of participants limits our understanding of the impact of receiving results (both negative and actionable) on individual health, health behaviors, and the health of family members. In the context of negative results, there is the potential that the letter provided a false reassurance of health. Lastly, the narrow focus on returning medically actionable research results precludes the ability to understand how returning nonactionable pathogenic variants and variants of unknown significance would be received.
The lessons learned from our experience are offered in Box 1 and suggest that the broader implementation of genomic medicine into practice should be considered within the context of structural factors as well as the SDoH. Further, engaging with primary care providers and the community at large can enhance the relevancy and potential impact of genomic medicine research. Integrating contextual factors, stakeholder input, and the resources to operationalize prevention/management recommendations will help further refine the definition of actionable results to individuals.30 Programmatic efforts to increase diversity in genomic medicine research must be forward thinking in their approaches so that advances in precision medicine are equitably realized across populations and settings. In low-resourced and underserved communities, this is challenging as preventive care is perceived as a luxury that may not be prioritized over other pressing needs. Therefore, precision medicine research initiatives in health disparity populations should appropriately balance genomic discoveries with the ethical, legal, and social implications of implementing such innovations across diverse practice settings.
References
Gottesman O, Kuivaniemi H, Tromp G, et al. The Electronic Medical Records and Genomics (eMERGE) Network: past, present, and future. Genet Med. 2013;15:761–771.
In: Downey AS, Busta ER, Mancher M, Botkin JR, eds. Returning Individual Research Results to Participants: Guidance for a New Research Paradigm. Ch. 2. Washington, DC: National Academic Press (US); 2018. p. 59–92
Kullo IJ, Olson J, Fan X, et al. The Return of Actionable Variants Empirical (RAVE) Study, a Mayo Clinic Genomic Medicine Implementation Study: design and initial results. Mayo Clin Proc. 2018;93:1600–1610.
Taylor J. Fundamentals of community health centers: NHPF background paper. Washington, DC: National Health Policy Forum; 2004.
Shaibi G, Singh D, De Filippis E, et al. The Sangre Por Salud Biobank: facilitating genetic research in an underrepresented Latino community. Public Health Genomics. 2016;19:229–238.
Jones L, Wells K. Strategies for academic and clinician engagement in community-participatory partnered research. JAMA. 2007;297:407–410.
Newman SD, Andrews JO, Magwood GS, Jenkins C, Cox MJ, Williamson DC. Community advisory boards in community-based participatory research: a synthesis of best processes. Prev Chronic Dis. 2011;8:A70.
Beckles GL, Chou CF. Disparities in the prevalence of diagnosed diabetes—United States, 1999–2002 and 2011–2014. MMWR Morb Mortal Wkly Rep. 2016;65:1265–1269.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34.
Woolf SH, Braveman P. Where health disparities begin: the role of social and economic determinants—and why current policies may make matters worse. Health Aff (Millwood). 2011;30:1852–1859.
Braveman P, Egerter S, Williams DR. The social determinants of health: coming of age. Annu Rev Public Health. 2011;32:381–398.
George S, Duran N, Norris K. A systematic review of barriers and facilitators to minority research participation among African Americans, Latinos, Asian Americans, and Pacific Islanders. Am J Public Health. 2014;104:e16–e31.
Sirugo G, Williams SM, Tishkoff SA. The missing diversity in human genetic studies. Cell. 2019;177:26–31.
West KM, Blacksher E, Burke W. Genomics, health disparities, and missed opportunities for the nation’s research agenda. JAMA. 2017;317:1831–1832.
Horowitz CR, Orlando LA, Slavotinek AM, et al. The Genomic Medicine Integrative Research Framework: a conceptual framework for conducting genomic medicine research. Am J Hum Genet. 2019;104:1088–1096.
Shaibi GQ, Kullo IJ, Singh DP, et al. Developing a process for returning medically actionable genomic variants to Latino patients in a Federally Qualified Health Center. Public Health Genomics. 2018;21:77–84.
Fossey R, Kochan D, Winkler E, et al. Ethical considerations related to return of results from genomic medicine projects: the eMERGE Network (Phase III) experience. J Pers Med. 2018;8:E2.
Fabsitz RR, McGuire A, Sharp RR, et al. Ethical and practical guidelines for reporting genetic research results to study participants: updated guidelines from a National Heart, Lung, and Blood Institute working group. Circ Cardiovasc Genet. 2010;3:574–580.
Mikat-Stevens NA, Larson IA, Tarini BA. Primary-care providers’ perceived barriers to integration of genetics services: a systematic review of the literature. Genet Med. 2014;17:169.
Roberts MC, Mensah GA, Khoury MJ. Leveraging implementation science to address health disparities in genomic medicine: examples from the field. Ethn Dis. 2019;29 Suppl 1:187–192.
Butterfield RM, Evans JP, Rini C, et al. Returning negative results to individuals in a genomic screening program: lessons learned. Genet Med. 2019;21:409–416.
Overby CL, Kohane I, Kannry JL, et al. Opportunities for genomic clinical decision support interventions. Genet Med. 2013;15:817–823.
Downing GJ, Boyle SN, Brinner KM, Osheroff JA. Information management to enable personalized medicine: stakeholder roles in building clinical decision support. BMC Med Inform Decis Mak. 2009;9:44.
Roberts MC, Clyne M, Kennedy AE, Chambers DA, Khoury MJ. The current state of funded NIH grants in implementation science in genomic medicine: a portfolio analysis. Genet Med. 2019;21:1218–1223.
Robinson JO, Wynn J, Biesecker B, et al. Psychological outcomes related to exome and genome sequencing result disclosure: a meta-analysis of seven Clinical Sequencing Exploratory Research (CSER) Consortium studies. Genet Med. 2019;21:2781–2790.
Sumner SA, Mercy JA, Dahlberg LL, Hillis SD, Klevens J, Houry D. Violence in the United States: status, challenges, and opportunities. JAMA. 2015;314:478–488.
Lardiere MR, Jones E, Perez M. National Association of Community Health Centers. Assessment of behavioral health services in Federally Qualified Health Centers. 2011. http://www.nachc.org/wp-content/uploads/2015/06/BHReport.pdf.
Oetzel JG, Wallerstein N, Duran B, et al. Impact of participatory health research: a test of the community-based participatory research conceptual model. Biomed Res Int. 2018;2018:7281405.
Riley WT, Nilsen WJ, Manolio TA, Masys DR, Lauer M. News from the NIH: potential contributions of the behavioral and social sciences to the precision medicine initiative. Transl Behav Med. 2015;5:243–246.
Laberge AM, Richer J, Ravitsky V. Toward broader genetic contextualism: genetic testing enters the age of evidence-based medicine. Am J Bioeth. 2019;19:77–79.
Acknowledgements
This project was supported through the eMERGE Network and funded by NHGRI through grant U01HG6379 to the Mayo Clinic. Additional support was provided by the Mayo Clinic Center for Individualized Medicine. We thank Dr. Faiz Naioom and the Mountain Park Health Center providers and staff for their support. We are indebted to the Sangre Por Salud Biobank Community Advisory Board for their contributions, support, and dedication to ensuring that the needs of the local community are considered in genomic medicine research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Shaibi, G.Q., Kullo, I.J., Singh, D.P. et al. Returning genomic results in a Federally Qualified Health Center: the intersection of precision medicine and social determinants of health. Genet Med 22, 1552–1559 (2020). https://doi.org/10.1038/s41436-020-0806-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0806-5
Keywords
This article is cited by
-
Benefits and concerns of expanded carrier screening: what do pregnant Latina women in Texas think?
Journal of Community Genetics (2023)
-
Implementing genomic screening in diverse populations
Genome Medicine (2021)
-
Disclosure of clinically actionable genetic variants to thoracic aortic dissection biobank participants
BMC Medical Genomics (2021)
-
Participation in genetic research among Latinx populations by Latin America birth-residency concordance: a global study
Journal of Community Genetics (2021)
-
Genetic testing and results disclosure in diverse populations: what does it take?
Genetics in Medicine (2020)