ABSTRACT
Purpose
We investigated the diagnostic and clinical performance of trio exome sequencing (ES) in parent–fetus trios where the fetus had sonographic abnormalities but normal karyotype, microarray and, in some cases, normal gene-specific sequencing.
Methods
ES was performed from DNA of 102 anomalous fetuses and from peripheral blood from their parents. Parents provided consent for the return of diagnostic results in the fetus, medically actionable findings in the parents, and identification as carrier couple for significant autosomal recessive conditions.
Results
In 21/102 (20.6%) fetuses, ES provided a positive-definitive or positive-probable diagnosis. In 10/102 (9.8%), ES provided an inconclusive-possible result. At least 2/102 (2.0%) had a repeat pregnancy during the study period and used the information from the study for prenatal diagnosis in the next pregnancy. Six of 204 (2.9%) parents received medically actionable results that affected their own health and 3/102 (2.9%) of couples received results that they were carriers for the same autosomal recessive condition.
Conclusion
ES has diagnostic utility in a select population of fetuses where a genetic diagnosis was highly suspected. Challenges related to genetics literacy, variant interpretation, and various types of diagnostic results affecting both fetal and parental health must be addressed by highly tailored pre- and post-test genetic counseling.
Similar content being viewed by others
INTRODUCTION
Congenital anomalies affect 2–4% of all infants and are responsible for 21% of perinatal deaths.1 Prenatal ultrasound diagnosis of fetal structural anomalies has become routine and karyotype and chromosomal microarray are recommended as first-tier tests from amniocentesis or chorionic villus sampling (CVS) when a fetal structural abnormality is identified. Gene-specific targeted sequencing panels can be considered if first-tier tests are negative, depending on the clinical findings. However, limited prenatal phenotypic information with ultrasound reduces the ability to choose an appropriate panel.
Our group and others have shown that there is an improved diagnostic yield with exome sequencing (ES) when fetal abnormalities are diagnosed prenatally and standard genetic testing including chromosomal microarray and targeted gene panels are negative.2,3,4,5 However, there are unique challenges specific to prenatal sequencing. The biggest challenge is due to limited prenatal phenotype information, which limits the ability to interpret the variants. The highest yield of ES is in fetuses with multiple anomalies, thus, this technology will likely become incorporated into prenatal care as the cost of sequencing continues to decrease. In addition, it is becoming apparent that prenatal phenotypes differ greatly from postnatal descriptions of the same syndrome.6 Thus, ES is expanding our knowledge of prenatal presentations of various disorders as well as enabling identification of novel candidate genes critical to human development.6,7 To develop best practices related to prenatal ES, it is important to address several questions, such as what types of results to report and what should comprise standard elements of the informed consent form for the mother and father.
Given that our center is studying and addressing many of the critical issues related to a clinical workflow and because laboratories and clinicians are already offering prenatal ES as part of clinical care in the evaluation of fetal anomalies, our objective was to determine the diagnostic yield in our cohort of anomalous fetuses, describe the workflow related to prenatal ES, and report on secondary findings in the parents in our cohort. Our overall goal is to provide a comprehensive approach to prenatal exome sequencing for potential integration into clinical practice. We previously published a pilot study on the first 15 families with severe fetal abnormalities not compatible with life2 but have since expanded our cohort to 87 more families (102 total) in both ongoing and noncontinuing pregnancies. The majority of the cohort (80%) have multiple abnormalities and were preselected for ES because the families were referred to us from providers with genetics expertise. We hypothesize that ES on our cohort will yield information about the spectrum of genetic variants that cause fetal abnormalities and expand our understanding of prenatal phenotypes. Finally, we make some suggestions regarding future studies to improve clinical implementation of prenatal ES.
MATERIALS AND METHODS
Mother–father–fetus trios in pregnancies complicated by a fetus with single or multiple congenital anomalies with normal or nondiagnostic chromosomal microarray were identified from the University of North Carolina (UNC) at Chapel Hill prenatal diagnosis clinics (Chapel Hill, NC and Raleigh, NC) and from various referring prenatal diagnosis clinics in the United States between July 2014 and February 2019. The results from the first 15 trios were previously published and included only fetuses with multiple anomalies highly suggestive of a genetic disorder.2 After the first 15 families were enrolled, we expanded enrollment to any pregnancy with a fetal anomaly (isolated or multiple). Approval from the University of North Carolina at Chapel Hill Institutional Review Board (IRB) (13-4084) was obtained prior to patient consent and enrollment. Inclusion criteria include the following: (1) isolated fetal anomaly or multiple fetal anomalies; (2) unknown diagnosis based on karyotype, microarray, and in some cases, gene-specific sequencing; (3) fetal and parental DNA available. Trios were identified prospectively and retrospectively, enabling us to obtain fetal specimens at various gestational ages. Prospectively, women pregnant with a singleton fetus with any major structural anomaly, (using the EUROCAT definitions)8 consistent with a genetic disorder were approached for participation after they made the decision to continue the pregnancy. In the case of noncontinuing pregnancies, the research study was not mentioned or offered until after the couple had made a decision to terminate the pregnancy. Retrospective identification of potential trios was accomplished by querying the UNC Perinatal Database to identify women with a history of fetal or neonatal death or termination of pregnancy who had not received an explanatory diagnosis by standard prenatal testing. We contacted women who previously indicated a desire to be recontacted if additional fetal testing options become available and who had fetal cells archived and available for DNA extraction for potential enrollment. Additional participants in the retrospective cohort were either self-referred or referred by a clinician aware of our current study recruitment. Once participants were enrolled, we collected parental blood and retrieved stored fetal samples for ES analysis. After the first seven trios were enrolled, we expanded enrollment to individuals not receiving care at UNC, using secure video teleconferencing to facilitate counseling, consent, and results discussion in nonlocal cases. The sample size of 102 trios was an exploratory convenience sample based on enrollment at the time that funding ended.
Mothers and fathers from both retrospective and prospective groups had pretest counseling about ES and the possible results that may be provided. Counseling points are summarized in Supplementary Table 2. Consent was obtained separately from mothers and fathers; the mother was informed about the possibility of ES revealing misattributed parentage. Participants were given the option to opt out at any time during the study. Because of the complexity of the genetic information that results from ES, consent and return of results were performed by a certified genetic counselor who was not involved in the patient’s clinical care to avoid bias and undue pressure on the patient to participate. All participants agreed to learn of (1) any diagnostic findings with potential to explain the fetal phenotype, (2) any medically actionable secondary findings in a parent that would have medical implications for that parent,9 and (3) carrier status for significant autosomal recessive conditions in which both parents are carriers. Diagnostic results were classified into seven categories (Supplementary Table 1). More than one result could be provided for a trio. Further details on workflow including DNA extraction, creating duplicate samples, and backup cultures are described in the supplementary material.
ES and variant analysis
We created ES libraries and exome capture from maternal, paternal, and fetal DNA samples as previously described10 and transferred them to the UNC High Throughput Sequencing Facility for sequencing using Illumina Hi-Seq 2500 or Hi-Seq 4000 instruments. We processed, mapped, and aligned raw-read data, and identified variants using a standard bioinformatics pipeline developed for the NCGENES project in collaboration with colleagues in the Department of Genetics and the Renaissance Computing Institute.11 Further information on bioinformatics analysis, quality checks, and workflow including how variants were classified is provided in the supplementary material.
Results were reviewed by a multidisciplinary committee of clinical and laboratory geneticists, a prenatal geneticist, genetic counselors, and a pediatric geneticist not involved in the patient’s clinical care. The committee reviewed all potentially reportable findings within two weeks of the primary analyst identifying the variant. Results considered clearly or possibly (including variants of uncertain significance [VUS] results) explanatory of the fetal phenotype were confirmed in a CLIA-certified molecular genetics clinical laboratory using Sanger sequencing of previously aliquoted and stored DNA samples. Positive results and inconclusive-possible results were reported to parents after Sanger confirmation. Parents were counseled against using VUS findings for prenatal diagnosis in subsequent pregnancies. All trios with nondiagnostic sequencing results were reanalyzed every 6–12 months to determine variant classification changes or to determine if newly identified genes were responsible for the fetal phenotypes. Parents were consented at enrollment to be recontacted if any variants previously reported to them were reclassified to benign or if variants that were not previously reported to them were found to be causative of the fetal phenotype.
All parental samples were analyzed for a small subset of medically actionable genes (e.g., BRCA1/2) using a medically actional list from an ongoing NCGENES project (National Institutes of Health [NIH]. Only variants variants classified as pathogenic or likely pathogenic from this gene list were reported.9,12,13,14,15,16,17 Parents also consented to return of carrier status for significant autosomal recessive conditions in which both parents are carriers of a pathogenic or likely pathogenic variant. All reported variants, whether diagnostic for the fetus or secondary results for the parents, were confirmed by Sanger sequencing in our CLIA-certified molecular genetics laboratory.
RESULTS
Demographics are reported in Table 1 with the majority of our cohort Caucasian and college-educated. All referred cases had sonographic abnormalities or sonographic abnormalities and a pregnancy history of suggestive of a genetic diagnosis (Supplementary Table 3).
One hundred two prenatal trio samples fulfilling designated inclusion criteria were received for prenatal ES. ES was performed on 99/102 trios and GS with in silico analysis of exons and immediately adjacent regions was performed on 3/102 trios to pilot GS prior to a larger scale study being implemented at our institution.
Cases were obtained from UNC–Chapel Hill (n = 38) as well as tertiary care centers and private practice outside of UNC–Chapel Hill (n = 56). There was sufficient DNA to perform trio ES on all 102 samples.
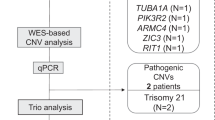
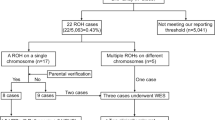
Of 102 cases, 31 (30.4%) families received results that were thought to explain the fetal findings after review by a multidisciplinary committee review; 21/102 (20.6%) were positive (positive-definitive [6/102; 5.9%] or positive-probable [15/102; 14.7%]). Ten of 102 (9.8%) were classified as inconclusive-possible according to our classification scheme (Table 2; Supplementary Table 1). Forty different contributing variants were reported in these 31 cases with various inheritance patterns, de novo being the most common in our cohort (Table 3). In 28/102 (27.5%) cases, there were recurrent pregnancies with similar phenotypes or another affected family member. In this cohort with a positive family history, fetal results were reported in 9/28 (32.1%) cases.
Thirteen of 102 fetuses had a single organ system apparently affected such as renal agenesis, intrauterine growth restriction (IUGR), anencephaly, or anhydramnios; 1 of these 13 cases (7.7%) received a reportable result from ES, which was a case of anhydramnios due to congenital abnormality of the kidneys and urinary tract. Eighty-nine of 102 fetuses (87.3%) had multiple anomalies identified by ultrasound. Supplementary Fig. 1 shows the frequency of the types of anomalies detected by ultrasound. Thirty of these 89 cases (33.7%) with multiple anomalies received a reportable result from ES.
In one case, a fetus was identified with a de novo variant in HRAS and compound heterozygous variants in HEXB and was therefore given two separate genetic diagnoses. We believed that the de novo variant in HRAS was responsible for the phenotypic findings on ultrasound. In one case, there was evidence of a copy-number variant from the sequencing data, enabling identification of an intragenic duplication, which was confirmed using fluorescence in situ hybridization (FISH) probes specific to the region. The intragenic duplication was in trans with a likely pathogenic variant in DYNC2H1, demonstrating the combined value of molecular and cytogenetic technologies in prenatal diagnosis.18 In one case initially reported as negative, further phenotypic information obtained on the mother was provided by the referring physician several years later, which enabled us to analyze the variants again and report a VUS in FOXC2 in both the mother and fetus.
In our study, in 10 of 102 cases (9.8%) fetal results were classified as inconclusive-possible using the definitions in Supplementary Table 1. Variants were Sanger confirmed in all of these cases so that results could be reported to the family for the purpose of recommending further clinical testing or for the purpose of gathering additional family history information to assist with interpretation. Over the course of 5 years, variants of uncertain significance in two families were reclassified to likely benign. The families were recontacted with this information. Neither family had used their results for prenatal testing in a subsequent pregnancy. These variants are not included in Table 2.
The turnaround time after receipt of trio samples was between 6 and 12 months, and was primarily limited by batching of samples (16 trios in a sequencer flow cell) to reduce sequencing cost. Parents of enrolled continuing pregnancies were informed that results would not be available prior to delivery. In one case where the results were available shortly after delivery and were likely to impact immediate newborn care, we obtained emergency IRB approval to verbally report a result of Gillespie syndrome prior to Sanger confirmation. Sanger subsequently confirmed the result. In this cohort, the rate of Sanger confirmation of the variants identified in our research sequencing lab was 46/46 (100%).
We are aware of three cases (PIEZO;WDR81; HEXB; see Table 2 for information on variants) in which families used the results of ES for prenatal testing by CVS or amniocentesis in a future pregnancy, allowing for an earlier diagnosis than would have been possible by ultrasound alone. One additional family used their likely pathogenic results for preimplantation genetic diagnosis with in vitro fertilization. It is possible that other families have used their results for prenatal diagnosis but have been lost to follow up after we provided the results.
Secondary findings were reported in 9/204 parents (4.4%) (Table 4) and include 6 cases of medically actionable autosomal dominant conditions in parents (6/204; 2.9%) and three couples (6 parents/204; 2.9%) who were carriers for the same recessive condition. Overall, in our cohort, 36/102 (35.3%) families in total received a reportable sequencing result, including those who received likely or possibly diagnostic fetal results and those who received a secondary finding. Four of 102 (3.9%) families received both a likely diagnostic fetal result and a secondary finding.
DISCUSSION
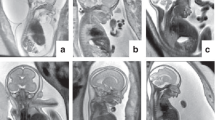
Our study provided an ES result to 30.4% of families in our cohort who did not receive a diagnosis with standard genetic testing (chromosomal microarray and in some cases gene-specific targeted panels). Our diagnostic yield is higher than other published prenatal ES series due to the highly selected nature of our cohort. However, our study adds to the current literature by providing information regarding the spectrum of genetic variants and associated prenatal phenotypes. As genomic technologies improve and are applied prenatally, our ability to refine fetal phenotypes will also inevitably improve.6 In our cohort, the following conditions had never been described prenatally: (1) scalp–ear–nipple syndrome (KCTD1); (2) cerebellar ataxia, mental retardation, and dysequilibrium syndrome 2 (WDR81); (3) Wiedemann–Steiner syndrome (KMT2A); and (4) spinocerebellular ataxia, Gillespie syndrome (ITPR1), and branchio-ocular facial syndrome (TFAP2A). In our fetal case of Costello syndrome, a massively enlarged liver was identified prenatally. To our knowledge, this has only been described in mice.19 In the case of branchio-ocular facial syndrome, absent radii has not been previously described in humans but knockout mice forTFAP2A show radial agenesis.20 These cases demonstrate how prenatal ES expands our understanding of fetal presentations of various disorders (Table 4: ultrasound phenotypes and genotypes).
Prenatal sequencing is improving the ability to identify conditions prenatally but also importantly identifies a gap in current prenatal practice of inadequate prenatal phenotyping. For prenatal sequencing to be optimized, efforts are needed to perform deep phenotyping of prenatal imaging studies and extract pertinent information from family and medical history. Postnatally, artificial intelligence is being leveraged for automated phenotyping and interpretation by accessing information from the electronic medical records allowing for rapid diagnosis in critically ill newborns.21 However, studies and efforts to improve deep prenatal phenotyping with automated interpretation to achieve faster turnaround time are needed to enable reproductive decision making and improve care in the prenatal and newborn period.
Our classification scheme (Supplementary Table 1) addressed the important issue of incorporating VUS in our definition of positive diagnoses as inconclusive-possible results. As sequencing becomes more integrated into prenatal practice, many variants classified as VUS may be reclassified as pathogenic or benign. In the prenatal setting, VUS pose a particular dilemma especially if parents want to use VUS results for reproductive planning purposes. Many infertility centers discourage the use of VUS for embryo selection prior to vitro fertilization. This issue further highlights the need for highly tailored genetic counseling to explain VUS results to avoid potential misuse. Over time, many VUS will be reclassified and more refined prenatal counseling will be possible with improved genotype–phenotype information.
Our study confirms findings from other published studies3,4,5 that trio ES improves diagnostic yield and expands use of prenatal ES to include information to parents regarding reporting of secondary findings. We demonstrate use of results for future pregnancy planning, identification of carrier couple status, and secondary medically actionable findings in parents. Our diagnostic yield is on the higher end of what has previously been reported in other studies; this may be stochastic variation or might be explained by the majority of patients in our study having multiple organ system abnormalities (79%) and referral from outside centers from providers with genetics expertise who prescreen fetal cases prior to referral.
Because the circumstances surrounding prenatal diagnostic genomic sequencing are different than in the postnatal setting and no guidelines currently exist for receipt of secondary findings in the prenatal setting,22 we plan to investigate whether there are differences in parental perceptions and preferences for learning about secondary findings from prenatal sequencing. This will guide providers on pre- and post-test counseling points to address and assessment of parental preferences with regard to types of results they desire. It will also be important to study psychological impact, decisional conflict, and test-related distress of providing secondary findings from trio ES. Given the complexity of the information and the many options available to receive results not related to the fetal findings (i.e., medically actionable findings in the parents, childhood onset disorders in the fetus, carrier couple findings, carrier findings in the fetus, etc.), more creative methods of counseling patients such as tablet-based decision aids23 may aid in decreasing decisional conflict related to the types of results parents ultimately choose to receive. Finally, as genome sequencing becomes more accessible due to reduced cost and our ability to interpret intronic regions of the genomic improves, it will be important to compare diagnostic yield of GS with the stepwise approach of chromosomal microarray followed by ES.
There are limitations of our study that will require expanded evaluation as genome-scale sequencing moves toward clinical use prenatally. This is a selected cohort in which cases with a high likelihood of genetic abnormality were referred for trio ES, so findings should be interpreted with caution in cohorts where the pretest probability of a genetic etiology may be lower. Our turnaround time was significantly lengthy due to financial considerations of research studies and the need to completely fill a flow cell with 16 trios prior to sequencing, an issue that impacts the timely utility and clinical applicability of the information in the current, affected pregnancy. Sanger confirmation of research results was an additional necessary step prior to reporting results to families contributing to increased turnaround time. However, patients were informed at consent that results would not be available for management in the current pregnancy.
Another limitation of our study is that married, college-educated Caucasian, non-Hispanic women with higher socioeconomic status self-selected into the study. New technologies can inadvertently create health disparities and poor access to new technologies for minority and lower socioeconomic status patients.24 A distrust of research and genomics is prevalent in some populations due to historical racial abuses in the United States. More data on implementation of genomics, health disparities, and patient perspectives from a broad and diverse population regarding integration of genomics in prenatal care and medicine in general is needed. Our group is planning focus groups with reproductive age women with low socioeconomic status and various ethnic backgrounds regarding views of use of genomics in prenatal care. The inclusion of families with various ethnic backgrounds is also critical to further our ability to accurately assess the significance of particular genetic variants. Genetic variants in minority populations are underrepresented in large databases and are often classified as VUS due to lack of information.
As sequencing is integrated into prenatal practice, it will be critical to maintain shared databases to improve interpretation and counseling for specific variants. Further research will be needed on parental understanding of testing and results, and the educational needs of families before and after results are delivered. If the return of secondary findings becomes standard, research regarding when to return those results in the prenatal setting is important. Returning these results separately from the fetal findings may allow time for adjustment to the fetal diagnosis. Finally, prenatal exome and genome sequencing is a relatively new and largely untapped method of identifying genes critical to human development since most genotype–phenotype information has been learned from pediatric or adult diagnoses. Prenatal sequencing may allow for identification of new genes and variants that are not compatible with survival and for the expansion of phenotypes of already known genes. Current imaging technologies must be improved and new methods of obtaining accurate prenatal phenotypic information must be developed so prenatal ES data can be optimally interpreted. Collaboration, both national and international, will be needed to move the field forward to improve prenatal care for families with a pregnancy affected with a birth defect.
Change history
18 June 2020
The original version of this Article contained an error in the spelling of the TP63 gene “NM_003722.4:c.1028G>C.; [p.Arg343Pro]" in Table 2, which was incorrectly given as "1028G>A.”. This has now been corrected in both the PDF and HTML versions of the Article.
References
Ely DMDA. Infant mortality in the United States, 2017: data from the period linked birth/infant death file. National Vital Statistics Reports. Hyattsville, MD: National Center for Health Statistics. 2019;68:10.
Vora NL, Powell B, Brandt A, et al. Prenatal exome sequencing in anomalous fetuses: new opportunities and challenges. Genet Med. 2017;19:1207–1216.
Lord J, McMullan DJ, Eberhardt RY, et al. Prenatal exome sequencing analysis in fetal structural anomalies detected by ultrasonography (PAGE): a cohort study. Lancet. 2019;393:747–757.
Petrovski S, Aggarwal V, Giordano JL, et al. Whole-exome sequencing in the evaluation of fetal structural anomalies: a prospective cohort study. Lancet. 2019;393:758–767.
Normand EA, Braxton A, Nassef S, et al. Clinical exome sequencing for fetuses with ultrasound abnormalities and a suspected Mendelian disorder. Genome Med. 2018;10:74.
Gray KJ, Wilkins-Haug LE, Herrig NJ, Vora NL. Fetal phenotypes emerge as genetic technologies become robust. Prenat Diagn. 2019;39:811–817.
Filges I, Friedman JM. Exome sequencing for gene discovery in lethal fetal disorders—harnessing the value of extreme phenotypes. Prenat Diagn. 2015;35:1005–1009.
EUROCAT. Malformation coding guides. 2017. http://www.eurocat-network.eu/aboutus/datacollection/guidelinesforregistration/malformationcodingguides.
Green RC, Berg JS, Grody WW, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15:565–574.
Lee K, Berg JS, Milko L, et al. High diagnostic yield of whole exome sequencing in participants with retinal dystrophies in a clinical ophthalmology setting. Am J Ophthalmol. 2015;160:354–363 e359.
Renaissance Computing Institute Technologies for Genomic Medicine. CANVAS and AnnoBot, solutions for genomic variant annotation. March 2014. http://www.renci.org/TR-14-04. Accessed 24 March 2014.
Berg JS, Adams M, Nassar N, et al. An informatics approach to analyzing the incidentalome. Genet Med. 2013;15:36–44.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Retterer K, Juusola J, Cho MT, et al. Clinical application of whole-exome sequencing across clinical indications. Genet Med. 2016;18:696–704.
Vora NLRS, Ralston SJ, Dugoff L, Kuller JA. Microarrays and next-generation sequencing technology: the use of advanced genetic diagnostic tools in obstetrics and gynecology. Obstet Gynecol. 2016;128:e262–e268.
Strande NT, Berg JS. Defining the clinical value of a genomic diagnosis in the era of next-generation sequencing. Annu Rev Genomics Hum Genet. 2016;17:303–332.
Berg JS, Foreman AK, O’Daniel JM, et al. A semiquantitative metric for evaluating clinical actionability of incidental or secondary findings from genome-scale sequencing. Genet Med. 2016;18:467–475.
Marchuk DS, Crooks K, Strande N, et al. Increasing the diagnostic yield of exome sequencing by copy number variant analysis. PLoS ONE. 2018;13:e0209185.
Figueiredo ML, Stein TJ, Jochem A, Sandgren EP. Mutant Hras(G12V) and Kras(G12D) have overlapping, but non-identical effects on hepatocyte growth and transformation frequency in transgenic mice. Liver Int. 2012;32:582–591.
Zhang J, Williams T. Identification and regulation of tissue-specific cis-acting elements associated with the human AP-2alpha gene. Dev Dyn. 2003;228:194–207.
Clark MM, Hildreth A, Batalov S, et al. Diagnosis of genetic diseases in seriously ill children by rapid whole-genome sequencing and automated phenotyping and interpretation. Sci Transl Med. 2019;11:eaat6177.
International Society for Prenatal Diagnosis; Society for Maternal and Fetal Medicine; Perinatal Quality Foundation. Joint Position Statement from the International Society for Prenatal Diagnosis (ISPD), the Society for Maternal Fetal Medicine (SMFM), and the Perinatal Quality Foundation (PQF) on the use of genome-wide sequencing for fetal diagnosis. Prenat Diagn. 2018;38:6–9.
Lewis MA, Paquin RS, Roche MI, et al. Supporting parental decisions about genomic sequencing for newborn screening: the NC NEXUS Decision Aid. Pediatrics. 2016;137 (Suppl 1):S16–S23.
West KM, Blacksher E, Burke W. Genomics, health disparities, and missed opportunities for the nation’s research agenda. JAMA. 2017;317:1831–1832.
Acknowledgements
The following funding is acknowledged: Clinical and Translational Science Award (CTSA) at UNC-CH TTR11403, Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) Building Interdisciplinary Research in Women's Health (BIRCWH) award 2K12HD00144116; National Human Genome Research Institute (NHGRI) HG006487, and NICHD K23 HD088742.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
N.L.V., K.G., and B.C.P. receive research supplies from Illumina for an unrelated project that is cofunded by NICHD and NHGRI (R01 HD055651). The other authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This work was presented at American College of Medical Genetics and Genomics; Seattle, Washington, April 2019.
Supplementary information
Rights and permissions
About this article
Cite this article
Vora, N.L., Gilmore, K., Brandt, A. et al. An approach to integrating exome sequencing for fetal structural anomalies into clinical practice. Genet Med 22, 954–961 (2020). https://doi.org/10.1038/s41436-020-0750-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0750-4
Keywords
This article is cited by
-
Prenatal whole-exome sequencing for fetal structural anomalies: a retrospective analysis of 145 Chinese cases
BMC Medical Genomics (2023)
-
Whole exome sequencing improves genetic diagnosis of fetal clubfoot
Human Genetics (2023)
-
Singleton exome sequencing of 90 fetuses with ultrasound anomalies revealing novel disease-causing variants and genotype–phenotype correlations
European Journal of Human Genetics (2022)
-
Impact of prenatal exome sequencing for fetal genetic diagnosis on maternal psychological outcomes and decisional conflict in a prospective cohort
Genetics in Medicine (2021)