Abstract
Purpose
To examine the overall genomic copy-number variant (CNV) landscape of Chinese pediatric patients with developmental disorders.
Methods
De-identified chromosomal microarray (CMA) data from 10,026 pediatric patients with developmental disorders were collected for re-evaluating the pathogenic CNV (pCNV) yields of different medical conditions and for comparing the frequency and phenotypic variability of genomic disorders between the Chinese and Western patient populations.
Results
The overall yield of pCNVs in the Chinese pediatric patient cohort was 21.37%, with variable yields for different disorders. Yields of pCNVs were positively associated with phenotypic complexity and intellectual disability/developmental delay (ID/DD) comorbidity for most disorders. The genomic burden and pCNV yield in neurodevelopmental disorders supported a female protective effect. However, the stratification analysis revealed that it was seen only in nonsyndromic ID/DD, not in nonsyndromic autism spectrum disorders or seizure. Furthermore, 15 known genomic disorders showed significantly different frequencies in Chinese and Western patient cohorts, and profiles of referred clinical features for 15 known genomic disorders were also significantly different in the two cohorts.
Conclusion
We defined the pCNV yields and profiles of the Chinese pediatric patients with different medical conditions and uncovered differences in the frequency and phenotypic diversity of genomic disorders between Chinese and Western patients.
Similar content being viewed by others
INTRODUCTION
Copy-number variants (CNVs) that alter the structure and function of the human genome have been identified as a common cause of human genetic diseases,1,2 particularly for severe pediatric conditions such as congenital anomalies,3 intellectual disability/developmental delay,3 autism,4 and epilepsy.5 For example, 25.7% of children with developmental delay harbor deleterious CNVs larger than 400 Kb.6 Detection of CNVs by chromosomal microarray (CMA) has been widely implemented in genetic research and clinical diagnostic laboratories and has been recommended as the first-tier genetic testing for pediatric developmental disorders since 2010.3,7 Large CNV databases, both for the general population, such as the Database of Genomic Variants (http://dgv.tcag.ca/dgv/app/home) and gnomAD-SV (https://gnomad.broadinstitute.org), and for patient populations such as the Simons Simplex Collection (SSC, https://www.sfari.org/) for autism and Deciphering Developmental Disorders (DDD, https://www.ddduk.org/) have played critical roles in aiding the interpretation of clinical significance of CNVs and understanding of the penetrance of recurrent pathogenic CNVs.6,8,9,10
To date, most such CNV aggregation efforts have been based on data generated from Western populations, mainly of European ancestry. Significant CNV differences have been reported in different populations,2 but there are very limited data for Asian or Chinese patient populations as most such studies have been based on small cohorts or emanated from a single center’s data,11,12,13,14 hindering comprehensive comparison. We set out to analyze the CNV profiles of over 10,000 Chinese pediatric patients from multiple centers and to evaluate the yield of clinically pathogenic CNVs in different pediatric medical conditions. We examined the differences in CNV profiles between the Chinese and Western populations with developmental disorders to explore the genomic diversity associated with human diseases. Finally, we evaluated three disease-related CNVs by leveraging de novo evidence.
MATERIALS AND METHODS
Ethics statement
The study was approved by the ethics committee of the respective institutions (Capital Institute of Pediatrics, Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region), which allowed us to perform aggregate analysis using de-identified clinical CMA data. Additional informed consent was obtained from the parents of some individuals to publish their detailed clinical information.
CNV detection and classification
Three different CMA platforms were used: Affymetrix HD chip, Agilent 244k/180k/60k commercial or custom-designed chip, and Illumina SNP chip. CNV calling followed the recommended settings of the respective platform (Table S1). CNVs below or near the recommended cutoff size were validated by quantitative polymerase chain reaction (qPCR), multiplex ligation-dependent probe amplification (MLPA), or gap PCR to avoid false positives. For a few complex CNVs, we also used genome sequencing (60×) for validation (Supplemental methods). We consulted both public databases (the DGV and gnomAD-SV databases) and our in-house database for CNV population frequency. We also consulted one Chinese control CNV database involving 500 healthy individuals with high-density CMA data to identify Chinese-specific benign CNVs.15 Only CNVs that were believed to be clinically relevant by each laboratory were included in this study. For established haploinsufficient or triplosensitive genes, we consulted the ClinGen dosage map database. All CNVs were manually reviewed and interpreted as “uncertain significance” (VUS CNV) and “pathogenic/likely pathogenic” (pCNV) by experienced geneticists (Y.S., X.Chen., J.W., and H.Y.) following American College of Medical Genetics and Genomics (ACMG) guidelines.16
Extraction of the frequencies and phenotypic profiles of genomic disorders from the Western patient cohort
The frequencies of known genomic imbalances in the Western patient population was derived from an aggregated CMA study where 73% of patients were diagnosed with neurodevelopmental disorders (NDDs),8 which is very similar to this Chinese patient population. Only CNVs with more than ten patients in Western or Chinese cohorts were chosen for frequency comparisons. The clinical features of 72 genomic imbalances were described in one recent aggregated Western patient cohort.17 The composition of this cohort was similar to that of our CMA cohort. We manually extracted the frequencies of the referred features for each genomic disorder and compared them with the frequencies in our Chinese cohort. Genomic disorders with more than ten independent patients in each cohort were chosen for referred phenotype comparisons.
Statistical analysis
The analyses were completed with Python or R. The qualitative comparisons between groups were conducted by Fisher’s exact test with false discovery rate (FDR) correction. All comparisons between subgroups were adjusted to p < 0.05 by Bonferroni. The numerical comparisons between groups were tested by Mann–Whitney U-test with Bonferroni correction. Samples with aneuploidy were excluded from all statistical analysis.
RESULTS
Characteristics of the Chinese pediatric cohort with developmental disorders
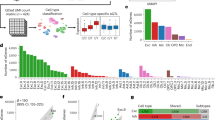
A total of 10,026 pediatric patients from seven regions of China were included in this study (Fig. S1). The median age of enrolled children was 24 months (range: 0–18 years) and young children (≤3 years) accounted for the majority (63.5%). Sixteen types of referral symptoms were listed and the top five common referral phenotypes (Table S2) were intellectual disability/developmental delay (ID/DD, 63.6%), hypotonia (12.5%), autism spectrum disorder (ASD, 11.9%), seizure/epilepsy (10.6%), and congenital heart disease (CHD, 9.0%). Overall, 72.28% patients (7247/10,026) presented with diverse NDDs including ID/DD, ADHD, ASD, or seizure/epilepsy. The phenotypic composition of this cohort is similar to those in previous Western CMA studies.6,8 The male to female ratio in this cohort was 1.58:1, but significant male skewness was seen only among those with ASD (ratio = 3.05), attention deficit–hyperactivity disorder (ADHD) (ratio = 4.18), and sex/urogenital disorder (ratio = 2.66) (Table S2).
Of 10,026 patients, 8,873 (88.50%) had a detailed clinical description, allowing for phenotype complexity and comorbidity analysis; 54.8% of patients exhibited an isolated phenotype, 29.9% of patients presented with two phenotypes, 11.3% with three, and 3.2% with four or more phenotypes (Table S2). ID/DD was a predominant comorbidity of the referred phenotypes and its co-occurrence with the top five phenotypes ranged from 59.29% (CHD) to 90.93% (micro/macrocephaly) (Table S3).
Whole-genome CNV profile of the Chinese pediatric cohort
After excluding 244 samples with aneuploidy (mostly involving chromosome 21 or chromosome X), a total of 2,572 clinically relevant CNVs (size range: 1028–154,880,970 bp; mean = 9.82 Mb; 1,725 deletions and 847 duplications, Table S4, first outer circle of Fig. S2A) from 2,230 patients were collected for further analysis. About half of these CNVs (51.8%) were <5 Mb in size, and deletions were generally smaller than duplications (mean 7.25 Mb vs. 15.10 Mb, p = 9.80E-18). These CNVs were enriched on chromosomes 7, 15, 22, and X (second circle of Fig. S2A); 93.4% of these CNVs were pCNV after reclassification (Fig. S2B), and the enrichment on chromosomes 7, 15, 22, and X was also evident for pCNVs. The size of pCNVs was generally larger than VUS CNVs (mean 10.44 Mb vs. 1.06 Mb, p = 1.50E-61), and the fraction of deletions among pCNV was much higher than in VUS CNV (68.4% vs. 47.6%). This CNV distribution is consistent with the notion that the human genome is less tolerant of deletion than duplication.18
Three hundred twenty-seven patients (3.26%) carried two or more CNVs. Among them, 160 patients (48.92%) carried deletion and duplication involving the terminal regions of two chromosomes; this scenario was most likely due to a parental balanced translocation (interchromosomal rearrangement). Fifty-three patients (16.20%) carried deletion and duplication on the same chromosome (intrachromosomal rearrangement), and 28 of these (52%) were due to a parental pericentric inversion because the CNVs occurred at both ends of the same chromosome. The breakpoints of these samples with inter-and intrachromosomal arrangements (Table S5) were unevenly distributed across chromosomes and along each chromosome (inner circus plots of Fig. S2A). Interchromosomal rearrangements were observed most often on chromosome 9 (39 CNVs), chromosome 4 (23) and chromosome 18 (22), whereas intrachromosomal rearrangements were more concentrated on chromosomes X (32) and 18 (13). Beside the terminal regions, we did not observe any specific genomic hotspots.
Yields of pCNV and their correlations with phenotypes
With exclusion of aneuploidy, there were 2,090 patients carrying pCNV, producing an average diagnostic yield of 21.37% (2,090/9,782). Patients with one of the following phenotypes had high CNV yields (Table 1): sex/urogenital malformation (41.71%), CHD (39.72%), craniofacial malformation (35.52%); those with one of these four phenotypes had relatively lower yields: ADHD (15.17%), ASD (16.89%), myopathy (17.91%), and preterm birth/low birth weight (LWB) (18.26%). For nonsyndromic patients (those exhibiting a single phenotype), the highest yields were among those with metabolic disorder (25.49%) and ID/DD (21.13%) while the lowest yields were among those with preterm birth/LWB (9.74%) and ASD (5.21%). Overall, syndromic patients (those exhibiting two or more phenotypes) had a higher yield of pCNVs than nonsyndromic patients, and this difference became significant among those with the following eight pediatric phenotypes: ID/DD (28.50% vs. 21.13%, p = 3.94E-10), ASD (23.73% vs. 5.21%, p = 1.16E-16), seizure/epilepsy (23.58% vs. 10.51%, p = 6.74E-06), CHD (43.91% vs. 17.27%, p = 1.73E-07), preterm birth/LBW (23.39% vs. 9.74%, p = 0.0001), endocrine/short stature (28.38% vs. 14.67%, p = 0.0003), craniofacial disorder (39.12% vs. 20.34%, p = 0.0168), and sex/urogenital disorder (52.74% vs. 15.31%, p = 6.33E-10).
The yield of pCNV was positively correlated with phenotypic complexity (Fig. S3). Patients with three or more phenotypes exhibited a 2.35- to 5.36-fold greater pCNV yield than those with single phenotypes (Table 2). In addition, the yield of pCNV was significantly correlated with onset age, as infants or young children (<3 years) exhibited a 1.22- to 1.33-fold greater pCNV yield compared with older children (>6 years).
ID/DD is the most common comorbid phenotype involved with other phenotypes in this cohort. The pCNV yields for nonsyndromic patients with ASD, seizure/epilepsy, and CHD (Table 1) were significantly lower than the yields for syndromic patients plus ID/DD comorbidity only (ASD: 5.21% vs. 16.58%, p = 1.79E-06; seizure/epilepsy: 10.51% vs. 18.21%, p = 0.0354; CHD: 17.27% vs. 46.25%, p = 5.81E-06); the differences were not significant for syndromic patients without ID/DD comorbidity (ASD: not relevant due to small sample size; seizure/epilepsy: 10.51% vs. 12.50%, p = 1; CHD: 17.27% vs. 28.57%, p = 0.175). For sex/urogenital malformation, syndromic patients had a higher yield than nonsyndromic patients, regardless of ID/DD (40.38% vs. 15.31%, p = 0.0071) or non-ID/DD comorbidity (36.11% vs. 15.31%, p = 0.0199).
pCNV profile and sex bias among NDD patients
The overall CMA diagnostic yield (excluding aneuploidy) for patients with NDDs was 23.13% (1643/7102). Among subphenotypes, isolated ID/DD had a significantly higher yield than isolated ASD (21.13% vs. 5.21%, p = 4.73E-18) and isolated seizure/epilepsy (21.13% vs. 10.51%, p = 1.25E-05, Table 1). Although there were more male patients in this NDD cohort, the CNV burden among females was significantly higher than for males in terms of autosomal overall CNV size (10.33 Mb vs. 8.45 Mb, p = 4.29E-10), affected gene count per person (209 vs. 172, p = 0.0002) (Fig. 1a) and autosomal pCNV yield (25.66% vs. 19.33%, p = 8.50E-10, Fig. 1b). Parental CNV testing for 132 CNVs in 123 NDD patients (78 boys and 45 girls) (Fig. 1c) revealed that the numbers of maternal CNVs and pCNVs were 2.36 and 2.42 times those of paternal CNVs. More maternal CNVs were seen in boys than in girls (33.7% vs. 12.7%), supporting the “female protective model” suggested by a Western NDD cohort.19
(a) The genome-wide and autosomal genomic CNV burden (size and gene count) in 4,288 boys and 2,627 girls with NDDs. Female patients carried larger CNVs (10.33 Mb vs. 8.45 Mb, p < 0.01) and more genes (209 vs. 172, p < 0.01) than males. The line and diamond in the solid box represent the median and mean of size and gene count with 25th and 75th percentiles labeled. *p < 0.05, **p < 0.01. (b) Left: The autosomal pCNV yield in male and female patients with NDDs. Higher yield was seen for girls compared with boys (25.66% vs. 19.33%, p < 0.01). Right: The pCNV yield in male and female patients with different subphenotypes of nonsyndromic NDDs (statistical power was not calculated for attention deficit–hyperactivity disorder (ADHD) due to small sample). Significantly higher pCNV yield was observed in girls for nonsyndromic intellectual disability/developmental delay (ID/DD) (23.66% vs. 17.37%, p < 0.01), but not for nonsyndromic autism spectrum disorder (ASD) (5.41% vs. 4.35%, p = 1), or seizure/epilepsy (13.59% vs. 7.10%, p = 0.18). *p < 0.05, **p < 0.01. NS not significant. (c) The inheritance status of 132 clinically relevant CNVs and 118 pCNVs in 124 patients with NDDs. The rate of de novo, maternal, and paternal inheritance was 63.6%, 25.8%, and 10.6% respectively for all CNVs, and the rate became 68.6%, 22.03%, and 9.4% respectively for pCNVs.
More importantly, when we further stratified the nonsyndromic NDD patients based on their phenotype (Fig. 1b and Table S6), we discovered that this sex bias was reflected in autosomal pCNV yield only for nonsyndromic ID/DD (23.66% vs. 17.37%, p = 4.18E-4), but not for nonsyndromic ASD (5.41% vs. 4.35%, p = 1) or seizure/epilepsy (13.59% vs. 7.10%, p = 0.18), suggesting that a “female protective effect” is limited to ID/DD.
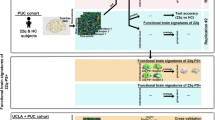
Comparison of the frequency and phenotypic penetrance of genomic disorders between Chinese and Western patient populations
Among the 19 known recurrent clinically relevant regions (Table S7), the top five recurrent deletions in the Chinese pediatric cohort were 15q11.2-q13.1 (Prader–Willi/Angelman syndrome, PWS/AS), 7q11.23 (Williams–Beuren syndrome, WBS), 22q11.2 (DiGeorge syndrome, DGS), 16p11.2 and 17p11.2 (Smith–Magenis syndrome, SMS)/1q21 region. The top five recurrent duplications were 15q11.2-q13.1, 16p13.11, 17p11.2 (Potocki–Lupski syndrome, PLS), 22q11.2 and 7q11.23 region. Among the 16 nonrecurrent regions (Table S8), the top five genomic disorders were 1p36 deletion followed by 22q13.3 deletion (Phelan–McDermid syndrome), 4p16.3 deletion (Wolf–Hirschhorn syndrome), 2q37 deletion, and 9q34.3 deletion.
When compared with their occurrence in the Western patient population (Fig. 2a, b),8 two recurrent genomic disorders including PWS/AS (2.07% vs. 0.14%, p = 1.4687E-80) and WBS (1.27% vs. 0.21%, p = 6.7700E-32) were significantly more frequent in the Chinese patient cohort, whereas six others, including 15q13.2 deletion (0.01% vs. 0.22%, p = 2.0678E-06) and duplication (0.01% vs. 0.10%, p = 0.0129), 16p12.1 deletion (0.01% vs. 0.17%, p = 6.9249E-05), 1q21 GJA5 duplication (0.06% vs. 0.17%, p = 0.0397), 16p11.2 duplication (0.06% vs. 0.21%, p = 0.0036), 22q11.2 duplication (0.10% vs. 0.33%, p = 0.0002) were significantly less frequent. 17q21.31 deletion was not detected in a single case in the Chinese cohort (0% vs. 0.11%, p = 0.0009). The other recurrent genomic disorders showed a similar frequency in both populations. Most nonrecurrent genomic disorders had a higher frequency in Chinese than in Western patients,8 with six achieving statistical significance (Fig. 2c) including 1p36 deletion (0.52% vs. 0.21%, p = 5.0436E-05), 2q37 deletion (0.21% vs. 0.09%, p = 0.0282), 4p16.3 deletion (0.31% vs. 0.11%, p = 0.0007), 9q34 deletion (0.19% vs. 0.07%, p = 0.0110), 15q26 deletion (0.14% vs. 0.04%, p = 0.0089), and Pitt–Hopkins syndrome (0.13% vs. 0.05%, p = 0.0441).
Only copy-number variants (CNVs) smaller than 10 Mb were included and genomic disorders of sex chromosomes were not compared. Both recurrent regions (including deletion and duplication) and nonrecurrent regions with at least 20 samples or CNVs with significant difference between the two cohorts are shown. Only the CNVs with 80% overlap were included. Nine small pathogenic deletions including one ELN deletion, seven UBE3A/SNRPN/NDN deletions, and one NSD1 deletion were not included. *p < 0.05, **p < 0.01. (a,b) The frequencies of 18 recurrent regions (a, deletion and b, duplication) related to human genomic disorders in two cohorts. (c) The frequencies of 16 nonrecurrent genomic disorders in the two cohorts. (d) The clinical feature spectrum of 15 genomic disorders in the two cohorts. Only genomic disorders with at least ten samples in each cohort are shown. DD/ID developmental delay/intellectual disability, MCA multiple congenital anomaly.
Large CMA cohort data also allowed for overall comparative analysis of clinical presentation for a list of genomic disorders in our Chinese cohort and the reported Western CMA cohort. We extracted the referred features of each genomic disorder from the Western population,17 and compared their frequencies with our Chinese cohort, to reflect the phenotypic difference of genomic disorders in these genetically and geographically distinct populations. In total 759 Chinese patients and 1,549 Western patients with 15 known genomic disorders were used for phenotypic profile analysis (Table S9). As shown in Fig. 2d, ID/DD was significantly more frequent as a phenotype in the Chinese cohort for series of disorders, including PWS/AS (77% vs. 48%, p = 0.0003), WBS (90% vs. 51%, p = 1.32E-08), DGS (61% vs. 36%, p = 0.0034), 1p36 deletion (79% vs. 56%, p = 0.0408), and 1q21.1 deletion syndrome (94% vs. 52%, p = 0.0143), as were the cardiac anomalies for WBS (33% vs. 16%, p = 0.0187), DGS (36% vs. 20%, p = 0.0348), 1p36 deletion (23% vs. 1%, p = 0.0001), and Wolf–Hirschhorn syndrome (52% vs. 12%, p = 0.0472). Other phenotypes also showing higher frequency in the Chinese cohort included ASD for Prader–Willi/Angelman duplication (52% vs. 12%, p = 0.0007), Phelan–McDermid syndrome (46% vs. 14%, p = 0.0179), and 17p11.2 deletion (41% vs. 3%, p = 0.0290); facial feature for WBS (47% vs. 28%, p = 0.0142) and 9q34 deletion syndrome (74% vs. 16%, p = 0.0159); seizure/epilepsy for DGS (22% vs. 6%, p = 0.0030) and 16p11.2 deletion (62% vs. 8%, p = 3.20E-09); speech and language delay for PWS/AS (12% vs. 0%, p = 0.0091), WBS syndrome (14% vs. 1%, p = 0.0033), Prader–Willi/Angelman duplication (44% vs. 1%, p = 2.50E-06), and 1p36 deletion (17% vs. 1%, p = 0.0013). Conversely, the multiple congenital anomaly (MCA) phenotype was significantly less frequent in the Chinese cohort for WBS (0% vs. 17%, p = 1.21E-05), DGS (0% vs. 26%, p = 2.35E-07), and 1p36 deletion (0% vs. 12%, p = 0.0408).
From the view of genomic disorders, Fig. 2d shows that five features of WBS, four features of 1p36 deletion and DGS, two features of PWS/AS, Prader–Willi/Angelman duplication and 1q21.1 deletion syndrome had significantly different frequencies in the two patient populations.
Identification of novel NDD-related genomic disorders/genes
We leveraged de novo evidence from our patient cohort and other publications or databases to identify three potential novel genomic disorders related to NDDs.
1q22-q23.1 (MEF2D) duplication
There is one de novo 1q22-q23.1 duplication in this Chinese cohort and there are three de novo and one parentally inherited duplication reported in Western patients (Fig. 3a). These five patients presented with diverse NDDs (Table S10). All patients except patient 292478 carried only this candidate CNV. The smallest overlapping region of these de novo duplications is chr1:156240271-156662371. The overlapping region contains the following genes: SMG5, TMEM79, CCT3, RHBG, MIR9-1/MIRN9-1, MEF2D, NAXE, BCAN, NES. Among these, only MEF2D and CCT3 are expressed in brain. MEF2D (OMIM 600663) belongs to the myocyte enhancer factor-2 family of transcription factors, and high expression of MEF2D was demonstrated in the cerebellum and cerebrum of developing and adult mouse brain.20 Overexpression of MEF2D was reported in the brain of patients with Parkinson disease.21 Another member of the myocyte enhancer factor-2 family, MEF2C (OMIM 613443) is a known causative gene for diverse NDDs by a haploinsufficiency mechanism.22 Minimal evidence for MEF2C triplosensitivity was reported in the ClinGen database. The current data suggest the 1q22-q23.1 region including MEF2D as a novel triplosensitive region.
Western patients are labeled by PMID ID or DECIPHER ID followed by patient ID and inheritance status. DN de novo, NA inheritance was not available, M maternal, P paternal. (a) Five MEF2D duplications including four de novo ones were detected from Chinese and Western patients. The smallest overlapped region of four de novo duplications is marked by the dashed red rectangle. (b) Twenty-eight CREBBP duplications including 21 de novo ones were detected from Chinese and Western patients. Among them, two duplications cover only CREBBP (QT5770, PMID23063576_P1, red arrow) in its entirety and three de novo duplications contain CREBBP and TRAP1 (PMID19833603_P10, PMID23063576_P2, DECIPHER303277, yellow arrow). The entire region of CREBBP and TRAP1 is marked by the dashed red rectangle. (c) Six PAX6 duplications including three de novo ones were detected from Chinese and Western patients. The overlapped region covers all of PAX6. (d) Upper panel shows a 919-Kb 11p13 duplication in a patient covering all of PAX6 and WT1. Middle panel shows the complex duplication–triplication arrangement after genome sequencing. Paternal chromosome contains three copies of PAX6 and RCN1, two copies of WT1, and one copy of ELP4. Bottom panel shows the precise junction sequence of 11p13 duplication. Red, yellow, brown, and green boxes represent ELP4, PAX6, RCH1, and WT1.
16p13.3 (CREBBP) duplication
The duplication of the 16p13.3 region has been proposed as a novel genomic disorder.23,24 We identified six duplications including two de novo cases involving the entire CREBBP from this Chinese cohort. We compiled the genomic coordinates and phenotypes of a total of 21 de novo cases involving CREBBP (Fig. 3b), and the smallest overlapping region is chr16:3761912-4005521, containing the entire CREBBP gene. Our Chinese patients with CREBBP duplications presented with manifestations similar to those reported,23,24 such as variable NDD phenotypes, facial dysmorphism, and thumb/halluces malformation (Table S10). Following the new ClinGen CNV interpretation guidelines,16 the triplosensitivity (TS) score of the 16p13.3 region covering CREBBP increased to 3, supporting it as a duplication disorder. However, there are only two duplications that involve only CREBBP (the red arrow in Fig. 3b), and the TS score of CREBBP is 1, so we cannot yet conclude that CREBBP is a TS gene.
11p13 (PAX6) triplication
PAX6 (OMIM 607108) pathogenic variant causes aniridia,25 contiguous gene deletion at 11p13 involving PAX6 and WT1 genes causes WAGR syndrome (OMIM 194072), and deletion of the PAX6 enhancer region containing ELP4 also causes aniridia as well as language impairment and ASD.26 Furthermore, duplications of 11p13 encompassing PAX6 and WT1 had been reported to be associated with ocular malformation and NDDs.27,28 We identified a novel copy-number gain at 11p13 in a boy and his father. The copy-number gain was de novo in the father. Both the boy and father presented with ocular phenotypes including microphthalmia and microcornea. The boy also exhibited neurodevelopmental problems such as autism and intellectual disability while the father did not have neurodevelopmental issues. Neither had Wilms tumor. Genome sequencing (Fig. 3d) and gap PCR (Fig. S4) revealed a complex tandem rearrangement at this locus, resulting in four intact copies of PAX6 and RCN1, three intact copies of WT1, and two intact copies of ELP4 in the genomes of the boy and his father. We reviewed five reported patients with 11p13 duplication in which three were de novo duplications covering all of PAX6, and compared their detailed clinical information and genomic coordinates of the duplication in each case (Fig. 3c). All 11p13 individuals exhibited diverse ocular abnormality and NDDs, but no patient presented with aniridia or Wilms tumor (Table S10). The clinical presentation of our subjects with four copies of PAX6 was not more severe than those with PAX6 duplication. Following the new ClinGen guideline,16 the TS score of the 11p13 region including PAX6 increased to 3, supporting it as a duplication disorder whose smallest overlapping region is chr11:31804340-31857737. Currently there is not sufficient evidence to fully support PAX6 as the TS gene since no duplication that involves only PAX6 has been described; nevertheless, we propose PAX6 as the most likely candidate gene responsible for this TS region.
DISCUSSION
The contribution of CNV to pediatric developmental conditions has been extensively evaluated and summarized in Western populations.3,6,7,8,9,10 It has also been explored in small cohorts of Chinese patients with specific phenotypes, such as CHD,14 ID/DD,11,12 ASD,29 and short stature,30 but the overall genomic CNV landscape of Chinese pediatric patients with developmental disorders has not been systematically evaluated. Meanwhile, previous studies have explored the diagnostic yield of some specific phenotypes without comprehensive comorbidity stratification, so it is not clear whether the index phenotype or the comorbidity affected the pCNV yield. This Chinese cohort study represents a comprehensive effort integrating over 10,000 pediatric CMA cases ascertained from multiple centers across China, covering 16 different pediatric disorders. This has allowed not only exploration of the CNV profiles in the Chinese pediatric patient population, but also comparison of the frequency and phenotypic diversity of known genomic disorders in comparison with the Western population. This study has revealed some interesting findings, many of which warrant further analysis.
First, we confirmed different degrees of pCNV contribution to different nonsyndromic disorders. Higher contributions were detected for ID/DD, craniofacial malformation, and CHD than for ASD. In the Western cohort, the deleterious CNV burden for cardiovascular and craniofacial phenotypes was reportedly higher than for ASD.6 A separate study involving 143,000 individuals referred for genetic testing also confirmed that highest pCNV yields were in pediatric NDDs and cardiology.31 In addition, we also examined the CNV involvement for some underinvestigated phenotypes, such as sex/urogenital disorder, hypotonia, and preterm birth/LWB. Our study revealed that for individuals presenting with hypotonia, a 20% diagnostic yield of CMA was detected, which increased to 30% when hypotonia was known to be comorbid with other phenotypes. A recent study involving a small number of children with developmental delay revealed a very high pCNV rate among individuals with hypotonia, and more patients in the CNV (+) group had hypotonia than those in the CNV (–) group.32 Thus, the contribution of CNV for hypotonia per se is worthy of further investigation. The contribution of CNV to preterm birth/LWB has not been previously studied. Our CNV study revealed a 10% CNV burden for preterm birth/LWB. Previous genome sequencing had revealed a significantly increased de novo variant burden in preterm babies compared with full-term controls.33 However, we did not identify overlapping genes/loci between our study and the genome sequencing study; nevertheless, both single-nucleotide variant (SNV) and CNV studies merit further exploration.
Compared with those reported for Western populations, even after excluding the aneuploid (2%), our study revealed a seemingly higher pCNV diagnostic yield (21.37%) for Chinese patients with developmental disorders.3,6,9 For example, Miller et al. reviewed 33 studies, involving a total of 21,698 patients tested by CMA, which revealed an average diagnostic yield of 12.2%.3 A higher diagnostic yield for typical conditions was also observed in our cohort compared with some Western cohorts (39.72% vs. 25% for CHD, 23.13% vs.14.2% for NDDs, 20.08% vs. 8% for seizure/epilepsy).5,8,34 There are several potential reasons for the higher yield in this Chinese cohort. First, most reported diagnostic rates in the Western population were from patients with no detectable abnormality by karyotyping. According to one Western CMA cohort with 5,110 patients, imbalanced translocation/pericentric inversion accounts for 1.2% of the tested sample.35 Whereas in our cohort, 829 patients with pathogenic CNVs would have been detected by karyotyping (>7 Mb), including 178 imbalanced translocation/pericentric inversion cases (1.8%). Second, Chinese physicians are more conservative and prefer to order testing for cases with severe phenotype due to pressure for a positive finding, which is supported by the fact that in this cohort, 40% of recruited Chinese patients express two or more phenotypes. Phenotypic severity is positively correlated with yield of pathogenic imbalances.17 These two factors in patient recruitment may also explain the high pCNV yields in other small Chinese patient cohorts (25.8–28%).11,12 When we repeated the pCNV yield analysis for all phenotypes after subtracting samples with pCNV larger than 7 Mb, the diagnostic rate in our Chinese cohort dropped to 14.08% (1,261/8,953), a figure compatible with those reported in Western patient populations.3,6 With these characteristics of the Chinese cohort in mind, we believe our data are compatible with what was reported for the Western population and the results derived from this analysis are comparatively meaningful.
Among more than 100 known genomic disorders, 72 are known to be associated with NDDs.17 As a main consequence of genomic imbalance, ID/DD comorbidity is believed to significantly contribute to the CNV diagnostic yield. For example, CHD comorbid with ID/DD or ASD had a significantly increased CNV detection rate compared with CHD without those comorbid features (22.7% vs. 4.3%).14 Seizure/epilepsy comorbid with ID had a higher CNV detection rate than isolated epilepsy (28% vs. 3%).36 In our cohort, we observed a significantly increased CNV yield for the four main pediatric disorders (CHD, ASD, seizure/epilepsy, and sex/urogenital disorder) when they were comorbid with DD/ID. Our data further support the notion that the majority of pathogenic CNVs detected in those pediatric disorders are attributable to ID/DD. This is particularly intriguing in regard to the contribution of CNV to ASD. Our data suggest that the CNV contribution to nonsyndromic ASD is quite low and pathogenic CNVs in patients with syndromic ASD are more likely reflective of its ID/DD components. This notion is also supported by a recent exome sequencing study of pathogenic SNV.37 The CNV contribution to nonsyndromic ASD is worthy of further scrutiny.
A female protective model has been proposed for ID/DD and ASD based on significantly increased deleterious autosomal CNVs and SNVs in female probands compared with male probands, and on significantly increased maternal CNVs compared with paternal CNVs in the probands with DD/ID.19,38 Our results revealed that both autosomal CNV burden (count and size) and pCNV yield in female probands with NDDs were significantly greater than in male probands. These findings are meaningful because the “female protective model” was validated in an independent large data set from another different population. However, interestingly, our study revealed that the sex bias was only seen for nonsyndromic ID/DD, not for nonsyndromic ASD and seizure/epilepsy, suggesting that the female protective model could be limited to the ID/DD phenotype, rather than for ASD directly. The female protective effect was raised for ASD from one Western study based on the finding that female ASD probands had a higher frequency of de novo events (11.7% vs. 7.4%, p = 0.16) and more affected genes in the CNVs (15.5 vs. 2, p = 0.05) than male probands,38 although the differences were not strongly significant. Moreover, their study did not stratify ASD into syndromic and nonsyndromic, whereas our finding that did not support the female protective model was based on the data from nonsyndromic ASD. These intriguing findings merit further analysis by studying syndromic and nonsyndromic ASD separately and combining both CNV and SNV data in the future.
Some well-established genomic disorders, such as Smith–Magenis syndrome and PWS/AS, are known to have high penetrance and distinct phenotypes across populations, so studying the frequency of these genomic disorders in our patient cohort can help us to predict their prevalence in general population. One Chinese postnatal CMA cohort with 3,096 patients showed that PWS/AS (2.9%), 16p11.2 deletion (2.9%), DGS (0.61%), 16p13.11 duplication (0.42%), Phelan–McDermid syndrome (0.35%), WBS (0.29%), as well as PWS/AS duplication (0.29%) were the top six genomic disorders.13 Other small Chinese cohorts also reported that PWS/AS (5.25%, 63/1,199) and WBS (3.17%, 38/1,199) were most frequent genomic disorders in the Chinese ID/DD population.11,12 In this study, we defined the frequency of 54 clinically relevant CNVs (deletion and duplication at 19 recurrent loci and 16 nonrecurrent loci) in a large Chinese pediatric cohort and confirmed the eight top genomic disorders: PWS/AS syndrome (2.07%), WBS (1.27%), DGS (0.73%), 1p36 deletion (0.52%), Phelan–McDermid syndrome (0.38%), 16p11.2 deletion (0.35%), Wolf–Hirschhorn syndrome (0.31%), and PWS/AS duplication (0.26%). This fundamental information is useful for clinical management and the general prevalence prediction of common genomic disorders in the Chinese population. Considering that in Guangxi province there is a Zhuang ethnic minority, the CNV profiles of 499 Zhuang samples were compared with those of the other Chinese samples. No over- or underrepresented CNV was seen in this ethnic group. In the future, larger samples with diverse minority groups are need to explore the potential for ethnic differences in the CNV profiles.
Furthermore, our study suggests that the frequency of 15 clinically relevant CNVs, including nine syndromic disorders and six genomic disorders with variable expressivity, can differ between populations of distinct ancestry and geography based on the significantly different frequencies in the Chinese and Western cohorts. Eight of the syndromic genomic disorders (1p36 deletion, 2q37 deletion, Wolf–Hirschhorn syndrome, WBS, 9q34 deletion, PWS/AS, 15q26 deletion, Pitt–Hopkins syndrome), excepting 17q21.31 deletion syndrome, showed significantly higher frequency in the Chinese patient population. Conversely, six genomic disorders with variable expressivity showed significantly lower frequency (1q21 GJA5 duplication,15q13.2 deletion and duplication, 16p12.1 deletion, 16p11.2 duplication, 22q11.2 duplication) in the Chinese patient population. One large Western pediatric CMA cohort had mentioned that WBS was underrepresented according to its frequency.6 Another Western CMA patient cohort (5,100 patients) reported a frequency of 0.35% and 0.31% for PWS/AS and WBS respectively, which is lower than DGS (0.55%) and 16p11.2 deletion (0.47%).35 These results support population-specific differences in the prevalence of common genomic disorders although further validation is needed.
CNVs occur at genomic rearrangement hotspots via nonallelic homologous recombination (NAHR) or nonhomologous end-joining (NHEJ),2 so the different frequency of genomic disorders between Chinese and Western populations suggests a different genomic architecture. This hypothesis was also supported by the CNV burden on specific chromosomes. Chromosomes 15, 16, 17, and 22 have been reported as autosomal rearrangement hotspots in Western populations,35 whereas in our Chinese patient population it was chromosomes 7, 15, 16, and 22 but not chromosome 17 that harbored more CNVs. Another Chinese ID/DD cohort also reported that chromosomes 7, 15, and 22 but not chromosome 17 were highly enriched for clinically relevant CNVs.11 For chromosome 15, we have confirmed that two CNV hotspots (PWS/AS and 15q13.2 region) showed differential frequency in the two distinct populations. PWS/AS duplication has been reported to be more frequent in a Chinese ASD cohort than in a Western ASD study (5/546 vs. 1/500).29 The underlying mechanism for such differences has been uncovered for some loci. For example, the 17q21.31 deletion that causes Koolen–de Vries syndrome, which as a recurrent event accounts for 0.1–0.3% of ID/DD patients in Westerners,39 is associated with the 900-kb inversion polymorphism (called H2 haplotype) whose frequency in Europeans is 20%.39 The H2 haplotype is absent from the Asian population; consequently, no 17q21.31 imbalances have been reported in Asian populations or in our large Chinese cohort. Although we have evidence to believe the observed differences are real, replicate studies will be needed to confirm the population differences observed across genomic disorders.
The phenotypic spectrum and penetrance of genomic disorders between genetically and geographically distinct populations is so far unknown, particularly for genomic disorders with variable expression. In our Chinese cohort and the reported Western CMA cohort, only referred clinical conditions were recorded, and insufficient phenotypic evaluation in these studies could affect the penetrance of certain phenotypes. Herein, we compared the average frequency of referred clinical conditions in patients with 15 genomic disorders, to reflect the possible diversity of phenotype for same genomic disorder. Although not definitive evidence, the different frequencies of clinical features in these 15 genomic disorders nonetheless suggests the existence of substantial phenotypic heterogeneity for genomic disorders across the two distinct populations. In the future, detailed phenotype–genotype investigations with larger sample sizes are needed, as exemplified by the Atlas of Human Malformation Syndromes in Diverse Populations project of the US National Institutes of Health (NIH) (https://research.nhgri.nih.gov/atlas/).
Finally, we reanalyzed the clinical significance of some de novo CNVs in our Chinese cohort. We confirmed triplosensitivity for two NDD-related regions (16p13.3 and 11p13), and provided initial evidence for implicating particular genes in this pathogenicity based on gene function and patients’ consistent phenotypes.
In summary, our study explored the CNV profiles of the Chinese pediatric patient population across different medical conditions, and uncovered frequency differences and phenotypic diversity associated with human diseases across Chinese and Western patient populations. The findings of this study can help the interpretation and understanding of the clinical significance and correlations of CNVs and point to a number of directions for future studies.
Data availability
The CNV data sets used for the study are available from the corresponding author upon request.
References
Lee, C., Iafrate, A. J. & Brothman, A. R. Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat. Genet. 39(7 Suppl), S48–S54 (2007).
Itsara, A. et al. Population analysis of large copy number variants and hotspots of human genetic disease. Am. J. Hum. Genet. 84, 148–161 (2009).
Miller, D. T. et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am. J. Hum. Genet. 86, 749–764 (2010).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science. 316, 445–449 (2007).
McTague, A., Howell, K. B., Cross, J. H., Kurian, M. A. & Scheffer, I. E. The genetic landscape of the epileptic encephalopathies of infancy and childhood. Lancet Neurol. 15, 304–316 (2015).
Cooper, G. M. et al. A copy number variation morbidity map of developmental delay. Nat. Genet. 43, 838–846 (2011).
Shen, Y. et al. Clinical genetic testing for patients with autism spectrum disorders. Pediatrics. 125, e727–e735 (2010).
Coe, B. P. et al. Refining analyses of copy number variation identifies specific genes associated with developmental delay. Nat. Genet. 46, 1063–1071 (2014).
Kaminsky, E. B. et al. An evidence-based approach to establish the functional and clinical significance of copy number variants in intellectual and developmental disabilities. Genet. Med. 13, 777–784 (2011).
Firth, H. V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Fan, Y. et al. Chromosomal microarray analysis in developmental delay and intellectual disability with comorbid conditions. BMC Med. Genomics 11, 49 (2018).
Wang, R. et al. Application of chromosome microarray analysis in patients with unexplained developmental delay/intellectual disability in South China. Pediatr. Neonatol. 60, 35–42 (2019).
Cheng, S. S. W. et al. Experience of chromosomal microarray applied in prenatal and postnatal settings in Hong Kong. Am. J. Med. Genet. C Semin. Med. Genet. 181, 196–207 (2019).
Geng, J. et al. Chromosome microarray testing for patients with congenital heart defects reveals novel disease causing loci and high diagnostic yield. BMC Genomics 15, 1127 (2014).
Zhang, C. et al. CNVbase: batch identification of novel and rare copy number variations based on multi-ethnic population data. J. Genet. Genomics 44, 367–370 (2017).
Riggs, E. R. et al. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet. Med. 22, 245–257 (2020).
Girirajan, S. et al. Phenotypic heterogeneity of genomic disorders and rare copy-number variants. N. Engl. J. Med. 367, 1321–1331 (2012).
Ruderfer, D. M. et al. Patterns of genic intolerance of rare copy number variation in 59,898 human exomes. Nat. Genet. 48, 1107–1111 (2016).
Jacquemont, S. et al. A higher mutational burden in females supports a “female protective model” in neurodevelopmental disorders. Am. J. Hum. Genet. 94, 415–425 (2014).
Ikeshima, H., Imai, S., Shimoda, K., Hata, J. & Takano, T. Expression of a MADS box gene, MEF2D, in neurons of the mouse central nervous system: implication of its binary function in myogenic and neurogenic cell lineages. Neurosci. Lett. 200, 117–120 (1995).
Yang, Q. et al. Regulation of neuronal survival factor MEF2D by chaperone-mediated autophagy. Science. 323, 124–127 (2009).
Le Meur, N. et al. MEF2C haploinsufficiency caused by either microdeletion of the 5q14.3 region or mutation is responsible for severe mental retardation with stereotypic movements, epilepsy and/or cerebral malformations. J. Med. Genet. 47, 22–29 (2010).
Demeer, B. et al. Duplication 16p13.3 and the CREBBP gene: confirmation of the phenotype. Eur. J. Med. Genet. 56, 26–31 (2013).
Thienpont, B. et al. Duplications of the critical Rubinstein-Taybi deletion region on chromosome 16p13.3 cause a novel recognisable syndrome. J. Med. Genet. 47, 155–161 (2010).
Jordan, T. et al. The human PAX6 gene is mutated in two patients with aniridia. Nat. Genet. 1, 328–332 (1992).
Davis, L. K. et al. Pax6 3’ deletion results in aniridia, autism and mental retardation. Hum. Genet. 123, 371–378 (2008).
Aalfs, C. M. et al. Tandem duplication of 11p12-p13 in a child with borderline development delay and eye abnormalities: dose effect of the PAX6 gene product? Am. J. Med. Genet. 73, 267–271 (1997).
Aradhya, S., Smaoui, N., Marble, M. & Lacassie, Y. De novo duplication 11p13 involving the PAX6 gene in a patient with neonatal seizures, hypotonia, microcephaly, developmental disability and minor ocular manifestations. Am. J. Med. Genet. A. 155A, 442–444 (2011).
Guo, H. et al. Genome-wide copy number variation analysis in a Chinese autism spectrum disorder cohort. Sci. Rep. 7, 44155 (2017).
Hu, G. et al. Copy number variations in 119 Chinese children with idiopathic short stature identified by the custom genome-wide microarray. Mol. Cytogenet. 9, 16 (2016).
Truty, R. et al. Prevalence and properties of intragenic copy-number variation in Mendelian disease genes. Genet. Med. 21, 114–123 (2019).
Park, K. B. et al. Effects of copy number variations on developmental aspects of children with delayed development. Ann. Rehabil. Med. 43, 215–223 (2019).
Li, J., Oehlert, J., Snyder, M., Stevenson, D. K. & Shaw, G. M. Fetal de novo mutations and preterm birth. PLoS Genet. 13, e1006689 (2017).
Zaidi, S. & Brueckner, M. Genetics and genomics of congenital heart disease. Circ. Res. 120, 923–940 (2017).
Catusi, I. et al. Testing single/combined clinical categories on 5110 Italian patients with developmental phenotypes to improve array-based detection rate. Mol. Genet. Genomic Med. 8, e1056 (2019).
Mullen, S. A. et al. Copy number variants are frequent in genetic generalized epilepsy with intellectual disability. Neurology. 81, 1507–1514 (2013).
Coe, B. P. et al. Neurodevelopmental disease genes implicated by de novo mutation and copy number variation morbidity. Nat. Genet. 51, 106–116 (2018).
Levy, D. et al. Rare de novo and transmitted copy-number variation in autistic spectrum disorders. Neuron. 70, 886–897 (2011).
Koolen, D. A. et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat. Genet. 38, 999–1001 (2006).
Acknowledgements
This work was supported by grants from CAMS Innovation Fund for Medical Sciences (2016‐I2M‐1‐008), the Beijing Natural Science Foundation (7202019 to X. Chen), the Chinese National Nature Science Fund (31671310 to X. Chen, 81873633 to Y.S.), the Eastern Scholar Fund, the Guangxi Bagui Scholar Fund, National Key Research and Development Program (2018YFC1002501), the Major Research Plan of the Provincial Science and Technology Foundation of Guangxi (AB16380214) to Y.S., Capital Health Research and Development of Special (2020-1-4071 and 2020-2-1131), and the Innovation Project of Beijing Municipal Human Resources and Social Security Bureau to X. Chen. The authors thank all involved families who contributed their de-identified CMA data to research, and are particularly grateful for the subset who collaborated on follow-up, providing detailed clinical information. The authors thank members of the collaborating clinical laboratories for performing the microarray experiments.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.S., X.Chen. Writing: Y.S., X.Chen, H.Y., J.F.G. Review and editing: Y.S., X.Chen, J.F.G. CMA data and collection: H.Y., S.S., J.L., J.S., R.Y., Shun Zhang. CNV curation and first-round pathogenic interpretation: J.L., S.S., R.Y. CNV second-round pathogenic interpretation H.Y., J.W., X.Chen, and Y.S. Data statistics analysis: Z.L. Western CNV literature review, the comparison between Chinese and Western cohorts for CNV frequency and referred phenotype: S.S., Z.L., H.Y. Clinical information collection: H.Y., Q.C., Z.G., Y.Zhu, X.W., L.Liu, J.Z., H.Li, H.Q., Y.Lin, H.Z., M.Y., M.M., L.Z., D.Z., H.W., H.Lv, Y.Liu, and L.Liang. CMA testing and validation: S.S., J.S., C.L., Shujie Zhang, W.Li, W.Lu, Y.Zhang, H.X., F.L., Q.W., B.X., C.G. Resources and software support and intelligent suggestion: Y.Liu and X.Cui.
Corresponding authors
Ethics declarations
Ethics Declaration
The study was approved by the ethics committee of the respective institutions (Capital Institute of Pediatrics, Maternal and Child Health Hospital of Guangxi Zhuang Autonomous Region), which allowed us to perform aggregate analysis using de-identified clinical CMA data. Additional informed consents were obtained from the parents of some individuals to publish their detailed clinical information.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yuan, H., Shangguan, S., Li, Z. et al. CNV profiles of Chinese pediatric patients with developmental disorders. Genet Med 23, 669–678 (2021). https://doi.org/10.1038/s41436-020-01048-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-01048-y
This article is cited by
-
A cohort study of neurodevelopmental disorders and/or congenital anomalies using high resolution chromosomal microarrays in southern Brazil highlighting the significance of ASD
Scientific Reports (2024)
-
Genetic subtypes and phenotypic characteristics of 110 patients with Prader-Willi syndrome
Italian Journal of Pediatrics (2022)