Abstract
Purpose
To delineate the genotype–phenotype correlation in individuals with likely pathogenic variants in the CLTC gene.
Methods
We describe 13 individuals with de novo CLTC variants. Causality of variants was determined by using the tolerance landscape of CLTC and computer-assisted molecular modeling where applicable. Phenotypic abnormalities observed in the individuals identified with missense and in-frame variants were compared with those with nonsense or frameshift variants in CLTC.
Results
All de novo variants were judged to be causal. Combining our data with that of 14 previously reported affected individuals (n = 27), all had intellectual disability (ID), ranging from mild to moderate/severe, with or without additional neurologic, behavioral, craniofacial, ophthalmologic, and gastrointestinal features. Microcephaly, hypoplasia of the corpus callosum, and epilepsy were more frequently observed in individuals with missense and in-frame variants than in those with nonsense and frameshift variants. However, this difference was not significant.
Conclusions
The wide phenotypic variability associated with likely pathogenic CLTC variants seems to be associated with allelic heterogeneity. The detailed clinical characterization of a larger cohort of individuals with pathogenic CLTC variants is warranted to support the hypothesis that missense and in-frame variants exert a dominant-negative effect, whereas the nonsense and frameshift variants would result in haploinsufficiency.
Similar content being viewed by others
INTRODUCTION
The clinical consequences of pathogenic CLTC variants have been described in 14 individuals who presented with neurodevelopmental disability accompanied by variable phenotypic features.1,2,3 DeMari et al. identified a de novo frameshift variant in the CLTC gene in a 3-year-old girl with intellectual disability (ID) and dysmorphic features, postulating haploinsufficiency of CLTC as the pathogenetic mechanism.2 More recently, 11 additional variants in CLTC were described in 13 individuals with ID, of whom five also had epilepsy.1,3,4 The effects of the DNA variants have neither been studied on the protein nor on the cellular level.
CLTC encodes clathrin heavy chain 1 (CHC1), a component of clathrin that consists of a polymer of three clathrin heavy chains and three clathrin light chains, which form a triskelion. This structure enables the formation of curved envelopes of hexagons and pentagons that cover the cytoplasmic face of clathrin-coated vesicles.5 These specialized organelles are involved in intracellular trafficking of receptors and endocytosis of a variety of macromolecules,6 as well as in the stabilization of kinetochore fibers in the mitotic spindle that aid congression of chromosomes to the metaphase plate.7 DeMari et al. noted that CLTC is highly expressed in the developing brain and probably plays a role in neuronal transmission by facilitating the recycling and/or release of vesicles at the presynaptic termini of neurons.2 Experiments in Drosophila melanogaster showed that clathrin heavy chain (chc) is critical for synaptic vesicle recycling, further supporting a specific role for CHC1 in brain development.8
Here, we expand the phenotype of individuals with de novo CLTC variants, and delineate the genotype–phenotype correlation in individuals with likely pathogenic variants in the CLTC gene.
MATERIALS AND METHODS
We identified 11 previously unreported individuals with likely pathogenic variants in CLTC. Two were followed in the Department of Human Genetics at Radboud University Medical Center (Nijmegen, The Netherlands) and nine were ascertained through GeneMatcher.9 This study was approved by the institutional review board Commissie Mensgebonden Onderzoek Regio Arnhem-Nijmegen (2018–4733). Written informed consent was obtained for all individuals involved including permission to publish photographs. We characterized their phenotypes retrospectively. Additionally, we describe the clinical characteristics of two individuals whose CLTC variants, p.(Asp1207del) and p.(Gln1539*), were previously reported by Lelieveld et al.4 DNA extracted from peripheral blood was analyzed using diagnostic exome sequencing or ID gene panel sequencing including the CLTC gene, by standard procedures.4,10
We used MetaDome11 to generate a tolerance landscape, a regional genetic tolerance plot based on the ratio of observed missense and synonymous (dN/dS) variants from gnomAD r2.02 12 found in the protein-coding region of CLTC. The tolerance score is corrected for the sequence composition of the protein-coding region based on the total possible missense and synonymous variants and computed as a sliding window of 21 residues over the entirety of the protein. To investigate the effect of the variants on the structure and function of the clathrin coat complex, we used a combination of the fully determined N-terminal WD40 domain (PDB file 5ODS) and a C-alpha only structure of the full-length clathrin triskelion (PDB file 3IYV) for computer-assisted molecular modeling.13 We employed the Yet Another Scientific Artificial Reality Application (YASARA)/WHAT IF Twinset for structural alignments and subsequent analysis of the structures.14
Fisher’s exact test, using a two-sided p value, was carried out to compare proportions of clinical features between groups of individuals with missense variants and in-frame deletions with nonsense and frameshift CLTC variants. If a specific variant, i.e., p.(Pro890Leu) and p.(Asp1207)del, was found in multiple individuals, these were viewed as replicates and counted as one data point for the statistical analysis. After conducting Bonferroni correction for multiple testing, a p value of <0.0125 was considered significant.
RESULTS
Three nonsense, two frameshift, two missense, and three splice site variants and three in-frame deletions were identified in CLTC (Fig. 1a; Table S1) in seven males and six females (median age: 14 years; range: 8 months to 41 years). Variants were confirmed to be de novo, except for p.(Ser1494Cysfs*32) identified in individual 12. This variant was not detected in the healthy mother or sister, and could not be studied in the father, who was not available for further investigation. None of the variants were present in gnomAD.12 All missense variants had a Combined Annotation Dependent Depletion (CADD) score above 25.15 Furthermore, CLTC is depleted from truncating and missense variants in healthy individuals (probability of loss of function in tolerance [pLi] = 1.00 and missense z-score = 8.09, respectively)12 suggestive for such variants being under strong purifying selection. As the pLI and missense z-score provide an indication over the entire gene, we generated a tolerance landscape with MetaDome that displays regional tolerance or intolerance to missense and synonymous variation. The CLTC missense variants and in-frame deletions are all located at positions that are intolerant to variation (Fig. S1). Altogether, these data support the pathogenicity of the CLTC variants.
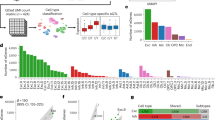
(a) Distribution of de novo variants in the CLTC gene in a schematic representation of the structure of the CLTC gene (NM_004859.3. 2; NP_004850.1; Uniprot: https://www.uniprot.org/uniprot/Q00610). Indicated are the globular terminal domain (TD) (residues 2–479), the flexible linker (residues 480–523), and the heavy chain arm (residues 524–1675) with clathrin heavy chain repeats (CHCR), which is involved in binding a clathrin light chain (CLC binding; residues 1213–1522) and contains the trimerization domain (TD; residues 1550–1675). Variants identified in this study are indicated in bold and variants located in a splice site are indicated in italics. (b) 3D structure of the clathrin coat complex (PDB file 3iyv) showing one centrally located CLTC protein (in gray) with the abovementioned domains and with the position of de novo in-frame CLTC variants in the protein (red points). Eight additional clathrin heavy chains, interacting with the centrally located CLTC protein, are shown in different colors (blue, green, and purple). Note the interaction of the C-terminal helix (trimerization domain) with two other heavy chains (blue/green). Smaller clathrin light chains, which are located close to the proximal domain of the heavy chain, are shown in yellow. (c) 3D structure close-ups of the clathrin complex displayed in (b), showing specific domains of the centrally located protein encoded by the CLTC gene (in gray) with de novo in-frame CLTC variants (in red), surrounded by clathrin heavy chains (in blue, green, and purple) and clathrin light chains (in yellow). (c1) Close-up of the distal segment of the heavy chain arm, showing that in-frame variants occur in an area that is in close interaction with the other heavy chains. An abnormal or loss of interaction with other heavy chains is predicted to affect the local structure of the clathrin coat complex. (c2) Close-up of the proximal segment of the heavy chain arm, showing that variants occur nearby a clathrin light chain and/or the KR-loop residues 1161–1165 (shown in orange), which is considered relevant for the regulation of the interactions with the light chain (Wilbur et al.16 Dev Cell 2010;18(5):841–848), likely resulting in an abnormal or loss of interactions with both heavy and light chains. Panels (c1) and (c2) were made using PDB file 3iyv.
The de novo CLTC variants are scattered throughout the gene. The missense variants and in-frame deletions of one to three amino acid residues are found in the part of the protein that contains the CHCR repeats, which is a long area that comprises seven repeats spanning amino acid residues 537–1566. The 3D structure of the CHC1 protein showed that the de novo missense CLTC variants do not cluster in three-dimensional proximity (Fig. 1b). However, these variants may affect the interaction with other clathrin heavy and light chains as CHC1 interacts with three light chains to form clathrin, ultimately changing the structure of the clathrin-coated vesicle (Fig. 1c).16 The nonsense and frameshift variants are all expected to result in nonsense-mediated messenger RNA (mRNA) decay.
All individuals presented with motor and/or speech delay (n = 13). All had ID that varied from mild to severe (Tables S2, S3). Neuropsychiatric features, noted in the majority of individuals (9/12), included attention deficit–hyperactivity disorder (ADHD) (4/9) and autism (3/10). Epilepsy (for the specific type, see Table S2) occurred in less than half of the individuals (5/13); had onset in the neonatal period, early infancy, or adulthood; and was fully controlled with antiepileptic drugs. Additional neurologic problems were observed, such as hypotonia (7/10), ataxia (2/8), and spastic gait (1/10). The majority of individuals had structural brain abnormalities (8/10) (Fig. 2a–f), which consisted of hypoplasia of the corpus callosum (a thin, small corpus callosum, but with all segments present), dilated Virchow–Robin spaces, asymmetric or dilated ventricles, frontal atrophy, pontocerebellar atrophy, and abnormal myelination. Hypoplasia of the corpus callosum occurred in five of these eight individuals (Fig. 2c–f). Microcephaly was observed in three of 13 individuals, all of whom had hypoplasia of the corpus callosum. Subtle craniofacial abnormalities were observed in all individuals (Fig. 2g–n). Facial dysmorphisms included a long face with a high, narrow forehead; wide nasal bridge; wide-set and long palpebral fissures; dysplastic ears that were large, protruding, and/or low-set; a deep philtrum consisting of prominent philtral ridges giving rise to an exaggerated groove in the midline; a wide mouth with thick lower vermilion; and large upper central incisors. Rarely, mild facial asymmetry was noticed. Ophthalmologic abnormalities were common (8/12), including strabismus (4/8) and abnormalities of refraction (4/8). Finally, gastrointestinal abnormalities were also diagnosed in more than 50% of individuals (7/13).
(a–f) Brain magnetic resonance imaging (MRI). The structure of the corpus callosum was normal in (a) individual 3 at 5 years (T1) and (b) individual 4 at 23 months, while hypoplasia of the corpus callosum (white arrows) was observed in (c) individual 5 at 2 months (T2), (d) individual 6 at 11 years (T2), (e) individual 9 at 4 years (T1), and (f) individual 12 at 2.5 years. (g–n) Craniofacial dysmorphisms, such as long face with high forehead, large ears, long palpebral fissures, midface hypoplasia, bulbous tip of the nose, deep philtrum, wide mouth, and large upper central incisors (individuals are ordered by increasing age at clinical photograph).
We hypothesized that the nonsense and frameshift variants have a different mode of action than the missense and in-frame variants. This might result in phenotypic differences. Therefore, we compared the proportion of severe ID, epilepsy, microcephaly, and hypoplasia of the corpus callosum in six individuals with unambiguous missense variants or in-frame deletions in CLTC with the proportions in ten individuals with proven de novo nonsense or frameshift variants. Of note, in-frame variants possibly affecting splicing were excluded from this analysis. Although differences did not reach significance, severe ID, epilepsy, microcephaly, and hypoplasia of the corpus callosum were more frequent in the group of individuals with in-frame variants than in individuals with nonsense and frameshift variants (3/6 vs. 0/10, 4/6 vs. 1/9, 2/6 vs. 0/9, 2/6 vs. 0/9, respectively; Fisher’s exact p values > 0.05; Tables S1 and S4). Of note, while the clinical triad epilepsy, microcephaly, and hypoplasia of the corpus callosum occurred in two of six individuals with likely pathogenic in-frame CLTC variants, it was not observed in any of the ten individuals with a frameshift variant (Fisher's exact p value = 0.125).
DISCUSSION
We describe the clinical manifestations observed in 13 individuals with 11 novel and 2 recurrent de novo variants in CLTC. Considering these 13 and the 14 previously reported individuals,1,2 a variable phenotype was observed, ranging from ID and subtle, but consistent, dysmorphic facial features to ID associated with microcephaly, brain abnormalities, and epilepsy. Notably, hypoplasia of the corpus callosum was the most common structural cerebral anomaly identified. Cerebral palsy (ataxic, hypotonic, or spastic) affecting gait was also recurrently seen. Neuropsychiatric features, including ADHD and/or autistic behavior, were observed in more than half of the individuals. Although recognizable, facial features are rather nonspecific. While previously rarely reported, vision abnormalities were common in our cohort.
The phenotypic variability observed in individuals with de novo CLTC variants is associated with allelic heterogeneity. A more severe phenotype, comprising severe ID, microcephaly, hypoplasia of the corpus callosum, and epilepsy was more frequently reported in individuals with missense and in-frame CLTC variants than with nonsense and frameshift variants. In particular, the triad microcephaly, callosal hypoplasia, and epilepsy occurred in two individuals with a missense variant or an in-frame deletion. Although the difference was not significant, this suggests that different types of pathogenic variants in CLTC may lead to different mechanisms of action. We proposed that truncating variants lead to a loss-of-function effect resulting in milder clinical manifestations, while missense and in-frame variants may contribute to a dominant-negative effect resulting in more severe clinical features. The suggestion of Hamdan et al.1 that “truncating de novo variants affecting the C terminus of CLTC tended to be associated with hypotonia, global developmental delay, and ID” falls within our more comprehensive hypothesis. Other additional genetic and nongenetic factors may eventually further explain the variability in the severity of the phenotype observed in these two groups of individuals. For instance, the four individuals with the de novo p.(Pro890Leu) variant presented with ID without epilepsy or callosal hypoplasia. Another example is given by the individual with the p.(Asp1207del) variant. Besides severe ID and callosal hypoplasia, he also had cerebral visual impairment, which may be associated with a likely pathogenic variant in the candidate UHMK1 gene.17 Since almost all CLTC variants described to date are private, we envision that the detailed clinical characterization of a larger cohort of individuals with pathogenic CLTC variants will be needed to further prove our hypothesis.
The variable expressivity associated with allelic heterogeneity in CLTC poses challenges to genetic counseling and warrants a comprehensive clinical evaluation and long-term multidisciplinary follow-up of these individuals, involving pediatricians, neurologists, psychiatrists, ophthalmologists, and medical geneticists, among others. While more severely affected individuals manifest moderate/severe ID associated with the clinical triad microcephaly, hypoplasia of the corpus callosum, and epilepsy, the less severely affected individuals may manifest mild ID/learning disability without epilepsy or brain abnormalities. Although not confirmed so far, the existence of individuals with inherited variants cannot be excluded, due to the mild phenotype associated with some likely pathogenic CLTC variants.
In summary, the clinical implications of pathogenic CLTC variants constitute a phenotypic spectrum, varying from mild ID to severe ID, microcephaly, hypoplasia of the corpus callosum, and epilepsy. An individual's phenotype seems to be more severe when associated with pathogenic missense and in-frame variants, likely leading to a dominant-negative effect, and less severe when associated with variants resulting in nonsense-mediated mRNA decay. Taken together, our findings suggest that there is a genotype–phenotype correlation in CLTC-related ID and neurologic/neuroradiologic abnormalities and corroborate the importance of precise clinical characterization of additional individuals to firmly establish this association.
References
Hamdan FF, Myers CT, Cossette P, et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am J Hum Genet. 2017;101:664–685.
DeMari J, Mroske C, Tang S, Nimeh J, Miller R, Lebel RR. CLTC as a clinically novel gene associated with multiple malformations and developmental delay. Am J Med Genet A. 2016;170A:958–966.
Manti F, Nardecchia F, Barresi S, et al. Neurotransmitter trafficking defect in a patient with clathrin (CLTC) variation presenting with intellectual disability and early-onset parkinsonism. Parkinsonism Relat Disord. 2019;61:207–210.
Lelieveld SH, Reijnders MR, Pfundt R, et al. Meta-analysis of 2,104 trios provides support for 10 new genes for intellectual disability. Nat Neurosci. 2016;19:1194–1196.
Dodge GR, Kovalszky I, McBride OW, et al. Human clathrin heavy chain (CLTC): partial molecular cloning, expression, and mapping of the gene to human chromosome 17q11-qter. Genomics. 1991;11:174–178.
Kaksonen M, Roux A. Mechanisms of clathrin-mediated endocytosis. Nat Rev Mol Cell Biol. 2018;19:313–326.
Royle SJ, Bright NA, Lagnado L. Clathrin is required for the function of the mitotic spindle. Nature. 2005;434:1152–1157.
Kasprowicz J, Kuenen S, Miskiewicz K, Habets RL, Smitz L, Verstreken P. Inactivation of clathrin heavy chain inhibits synaptic recycling but allows bulk membrane uptake. J Cell Biol. 2008;182:1007–1016.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–930.
Neveling K, Feenstra I, Gilissen C, et al. A post-hoc comparison of the utility of sanger sequencing and exome sequencing for the diagnosis of heterogeneous diseases. Hum Mutat. 2013;34:1721–1726.
Wiel L, Baakman C, Gilissen D, Veltman JA, Vriend G, Gilissen C. MetaDome: pathogenicity analysis of genetic variants through aggregation of homologous human protein domains. Hum Mutat. 2019;40:1030–1038.
Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291.
Fotin A, Cheng Y, Sliz P, et al. Molecular model for a complete clathrin lattice from electron cryomicroscopy. Nature. 2004;432:573–579.
Krieger E, Koraimann G, Vriend G. Increasing the precision of comparative models with YASARA NOVA—a self-parameterizing force field. Proteins. 2002;47:393–402.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–315.
Wilbur JD, Hwang PK, Ybe JA, et al. Conformation switching of clathrin light chain regulates clathrin lattice assembly. Dev Cell. 2010;18:841–848.
Bosch DG, Boonstra FN, de Leeuw N, et al. Novel genetic causes for cerebral visual impairment. Eur J Hum Genet. 2016;24:660–665.
Acknowledgements
We thank all individuals and their families for the invaluable contribution to this study. T.S.B. was supported by the Netherlands Organisation for Scientific Research (ZonMW Veni, grant 91617021) and by a National Alliance for Research on Schizophrenia & Depression (NARSAD) Young Investigator Grant from the Brain & Behavior Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nabais Sá, M.J., Venselaar, H., Wiel, L. et al. De novo CLTC variants are associated with a variable phenotype from mild to severe intellectual disability, microcephaly, hypoplasia of the corpus callosum, and epilepsy. Genet Med 22, 797–802 (2020). https://doi.org/10.1038/s41436-019-0703-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0703-y
Keywords
This article is cited by
-
A novel de novo CLTC variant altering RNA splicing causes fetal developmental abnormalities
BMC Medical Genomics (2023)
-
Continuous high-frequency deep brain stimulation of the anterior insula modulates autism-like behavior in a valproic acid-induced rat model
Journal of Translational Medicine (2022)
-
Novel CLTC variants cause new brain and kidney phenotypes
Journal of Human Genetics (2022)