Abstract
Purpose
Fetal fraction (FF) is the percent of cell-free DNA (cfDNA) in the mother’s peripheral blood that is of fetal origin, which plays a pivotal role in noninvasive prenatal screening (NIPS). We present a method that can reliably estimate FFs by examining autosome single-nucleotide polymorphisms (SNPs).
Methods
Even at a very low sequencing depth, there are plenty of SNPs covered by more than one read. At those SNPs, we define read heterozygosity and demonstrate that the percent of read heterozygosity is a function of FF, which allows FF to be inferred.
Results
We first demonstrated the effectiveness of our method in inferring FF. Then we used the inferred FF as an informative alternative prior to computing Bayes factors to test for aneuploidy, and observed better power than the Z-test. In analysis of clinical samples, we were able to identify female–male twins thanks to the accurate FF inference.
Conclusion
Knowing FF improves efficacy of NIPS. It brings a powerful Bayesian method, allows “no call” for samples with small FFs, renders screening for XXY syndrome simpler, and permits an adaptive design to sequence at a higher depth for samples with small FFs.
Similar content being viewed by others
INTRODUCTION
Fetal fraction (FF) is the percent of cell-free DNA (cfDNA) in maternal peripheral blood that comes from the placenta of the fetus. FF plays a pivotal role in noninvasive prenatal screening (NIPS), which aims to examine whether a given chromosome is trisomy in the fetus. The American College of Medical Genetics and Genomics (ACMG) recommended in its position statement that all laboratories should include a clearly visible FF on the NIPS report, and all laboratories should establish and monitor analytical and clinical validations for FF.1 The lower limit of FF maintaining a reliable result is approximately 4%, and a low FF in maternal circulation was associated with an increased risk of fetal aneuploidies.2 Thus, ACMG recommends that no call due to low FF needs to be specified in the NIPS report.1
Given the importance of the FF in NIPS, many methods have been proposed to detect FF. Early methods examined DNA sequences from the Y chromosome, either using polymerase chain reaction (PCR)3 or massive parallel sequencing technology.4 Since methods based on the Y chromosome can only be applied to male fetuses, researchers focused efforts on developing methods that can apply to both male and female fetuses. Some used fragment size of cfDNA, where fetal cfDNA is generally shorter than maternal cfDNA;5 some explored the methylation differences between paternal and maternal cfDNA.6,7 However, these methods were not yet accurate enough for practical use. There are two very promising and elegant approaches that require no additional data. One explores the difference between DNA digestion of fetal and maternal cell-free DNA.8 The other explores the fact that fetal cfDNA sequences are not uniformly distributed along the genome,9 presumably because actively expressed genomic regions were digested faster than those dormant gene regions. But these two methods present difficulties for smaller FFs.
The most successful methods are those that utilize inheritance patterns in single-nucleotide polymorphism (SNP) markers. Early methods assumed both maternal and paternal genotypes are known, and at a set of loci where the father is AA and the mother is BB, one can deduce FF by counting reads carrying As and Bs at those loci from sequencing data of maternal cfDNA.10,11 A recent method only genotyped the mother.12 It first identified maternal homozygous loci, tallied and computed nonmaternal allele fractions at these loci from low-coverage sequencing data of maternal plasma, and used these to train a linear model to predict FF. High-depth targeted sequencing of maternal plasma was successfully used to determine FF.13,14 Because of high sequencing depth, minor allele frequencies can be reliably estimated, the informative maternal–fetal joint genotypes can be inferred (one homozygous, one heterozygous), and based on this, FF can be determined. These methods, however, are not completely satisfactory as they are either not accurate enough, not cost-effective, or too laborious.
In this paper we introduce a statistical method that can infer FF using low depth sequencing of maternal cfDNA. The precision of the FF inference can be greatly improved if we also sequence maternal white blood cell DNA (wbcDNA) at a low depth. Our method is based on the read heterozygosity (RHet). A SNP locus is called RHet if it is covered by at least two reads carrying different alleles. RHet is determined mainly by the unobserved maternal–fetal joint genotypes and the FF. Other contributing factors include the inbreeding coefficient of the mother, the inbreeding coefficient of the fetus, the sequencing error rate, and SNP allele frequencies of the reference population. Sequencing of maternal wbcDNA contributes to the FF inference in two ways: it can be used to infer maternal inbreeding coefficient, and we can combine reads from the maternal cfDNA and wbcDNA sequencing to infer a diluted FF from a higher coverage data set and scale back.
The traditional Z-test method for NIPS first estimates chromosomal dosages for a sample (assuming a euploid mother carrying a single fetus), then it calculates the deviation of the chromosomal dosages from the mean dosage of a set of euploid controls (euploid mothers each carrying a euploid fetus), and lastly it normalizes the deviation by the sample standard deviation of the euploid controls to obtain a Z-score. Samples with Z >3 are declared trisomy positive. The cutoff of 3 is chosen such that the false positive rate is about 0.001 if the Z-score is normally distributed. If the fetus is trisomy, then the deviation is expected to be the FF. Thus, the higher the FF, the higher the power to detect true trisomy.
When FF is known (denoted by h), a more powerful test can be developed. Z-test only compares the deviation of the centered chromosomal dosage from 0. Since for a trisomy sample the chromosomal dosage is expected to be h above the mean dosage of euploid controls, we can also compare how close the centered chromosomal dosage is to h. In addition to increased power to screen for trisomy, knowing FF brings several other benefits. First, if FF is too small, which is a major source of false negatives, we can declare “no call.” Second, testing aneuploidy of sex chromosomes, such as Klinefelter syndrome (47, XXY), Turner syndrome (45, X), and XYY syndrome, becomes a much simpler problem. Third, it allows us to develop an adaptive design to sequence at a higher depth for samples with small FFs.
MATERIALS AND METHODS
NIPS samples
Beginning 15 March 2016, we enrolled pregnant women who were undergoing routine obstetrical care at the Beijing Hospital. The institutional review board of the Beijing Hospital approved the study. All experiments were performed in accordance with relevant guidelines and regulations. Written informed consent was obtained from all patients. To be eligible for the study, pregnant women had to be at least 18 years of age and had to be carrying a fetus with a gestational age of at least 8 weeks. More information is in the Supplementary Material and Methods.
Naive model
An ideal scenario would be no sequencing errors, and both mother and the fetus have an inbreeding coefficient of 0. Denote FF by h, and at an arbitrary biallelic SNP (A and B) denote the frequency of allele A as p. Assuming no inbreeding, the Hardy–Weinberg equilibrium holds for each genotype of one individual, such that \(f_{AA} = p^2\), \(f_{AB} = 2p(1 - p)\), and \(f_{BB} = (1 - p)^2\). The distribution of the joint maternal–fetal genotypes and the A allele frequency of the mixture can be derived, as shown in Table 1.
Those SNPs covered by one read are ignored because of their likelihood containing no h. Suppose a SNP has coverage of 2, with counts of two alleles as (2, 0), (1, 1), and (0, 2). To evaluate likelihood for each count, we are conditioning on the joint genotype to obtain a weighted sum of binomial likelihood.
The read heterozygous (1,1) has the probability of \(p(1 - p)(1 + h - h^2)\). If we assume p is uniformly distributed and integrate out p, we have the expected proportion of read heterozygosity among SNPs covered by two reads as \(\frac{1}{6}(1 + h - h^2)\), which is a function of h.
Full model
Let p, q be the allele frequency of the A, B allele with p + q = 1, andF be the individual’s inbreeding coefficient. The standard model to account for inbreeding coefficient as a generalization of Hardy–Weinberg equilibrium15 is
The joint distribution of mother–fetus genotypes can be computed (Tables 2).
The only possible errors that can occur after bi-allelic ascertainment are A to B or B to A, and the sequencing errors are different from A to B (denote eAB) and from B to A (denoteeBA). Accounting for sequencing errors, we can write the allele frequency of the mixture for each instance of joint genotype (Table 3), from which the likelihood can be obtained by modeling reads covering a given locus as binomial sampling.
At SNP i covered by more than one read, denote \(G_i = (c_i^A,c_i^B)\) the counts of reference and alternative alleles. Let x be the frequencies of A; the binomial likelihood for \(G_i\) is
Denote G the collection of all \(G_i\) for i in an index set ofS. For each SNP i, we write out the binomial likelihood conditioning on the joint genotypes, weighted by the probability of the joint genotype, and sum together to obtain marginal likelihood
Multiplying the marginal likelihood for each SNP to obtain a composite likelihood
Finally, from nonpolymorphic markers covered by exactly one read, we can estimate the error spectrum used in the model (eAB and eBA for A and B in four nucleotides). We may optimize Eq. 5 to obtain ĥ.
RESULTS
Read heterozygosity informs FF
The rationale to infer FF from low depth sequencing data is detailed in the scientific facts and logic arguments in this section. First, even at a low coverage, there are sizable numbers of SNPs from the 1000 Genomes Project that are covered by more than one read. This can be deduced simply by assuming Poisson distribution at each locus and verified by real data (Supplementary Table S1). Second, at those SNPs covered by at least two reads, we can define read heterozygosity (and similarly read homozygosity). For example, at coverage of two, we may define (A, B) or (B, A) as read heterozygous, and (A, A) or (B, B) as read homozygous, where A and B are reference and alternative alleles respectively. Note that read heterozygosity, which involves sampling uncertainty, is different from the genotype heterozygosity. For example, at coverage of two, a genotype heterozygous AB has a 50% chance to produce read heterozygosity. Third, the percent of read heterozygosity is a function of FF (denoted by h). Under the naive model with ideal assumptions (see “Materials and Methods”), the percent of SNPs that are read heterozygous among SNPs covered by two reads is \(\frac{1}{6}(1 + h - h^2)\). Thus h can be inferred. Note, however, the naive model only has theoretical value and performs poorly in real data analysis, as real data violate most of its assumptions.
Statistical model to infer FF
For a given biallelic SNP, its allele frequency largely determines how likely we observe read heterozygosity. We identified three additional factors that affect read heterozygosity: the inbreeding coefficient of the mother F1, the inbreeding coefficient of the fetus F2, and the sequencing error ratee. Because a majority of the cfDNA comes from the mother, F1 determines the baseline of percent of read heterozygosity of a sample and is critical to the accuracy of FF inference. We therefore sequenced maternal wbcDNA and developed a statistical model to infer F1 (Supplementary Material and Methods). F2, on the other hand, contributes little to the percent of read heterozygosity, particularly when the FF is small. In practice, we can safely ignore F2 by setting it to 0. Sequencing errors produce more read heterozygosity than read homozygosity. The effect can be modeled and the sequencing error rate can be estimated for an individual sample from the nonpolymorphic part of genomes covered by a single read. At each marker covered by more than one read, we can write the single-marker likelihood, which involves parameters (h, F1, F2, and e) and data (counts of reference and alternative alleles and population allele frequencies). Multiplying all the single-marker likelihoods we obtain the composite likelihood, which is a function of h (with F1 and e inferred and plugged in and F2 set to 0). Maximizing the composite likelihood we obtain the maximum likelihood estimate of FF (see “Materials and Methods”).
Numerical results on FF inference
To demonstrate the effectiveness of our statistical method in inferring FF, we performed (in silico) simulations to mix reads from sequences of a mother and her son, conducted (in vitro) laboratory experiments to mix DNA of mothers and their male children, and reanalyzed real data from clinical NIPS samples with putative male fetus (in natura). These study designs allow us to compare fetal fractions estimated by our method against those estimated from the sex chromosomes. The details of FF estimates from sex chromosomes can be found in Supplementary Material and Methods, Supplementary Figs. S1 and S2, and reference.16
In the in silico experiments, we first mix reads from mothers and their children at different FF (Supplementary Material and Methods) to examine how F1 and F2 impact the inference of FFs. Fig. 1a, b demonstrates thatF1 has a large effect on FF and F2 has a negligible effect. We then focused on the variations of the FF estimates due to sampling. The data were simulated from mixing reads of two samples (a mother–child pair) in the 1000 Genomes Project (Supplementary Material and Methods). Figure 1c plotted the mean (of 100 replicates) with bar of the sample standard deviation of the inferred FF (y-axis) versus the true FF (x-axis). The sample standard deviation is 0.006 for FF = 0.02 and 0.008 for FF = 0.20. Lastly, we investigated how different sequencing depths affect the uncertainty of the FF estimates. Here we assume maternal wbcDNA was sequenced at the same depth as the cfDNA. We sampled and mixed reads to simulate 100 NIPS samples with FF = 0.10 at different sequencing depths. Violin plots in Fig. 1d confirmed that the higher sequencing depth, the smaller the variation of the FF estimates, and it appears that 0.5× would provide a good balance between sequencing depth and accuracy. The same simulations were done for FF = 0.04 and FF = 0.06 and similar patterns of variation were observed (Supplementary Fig. S3).
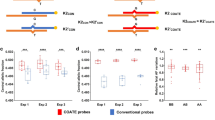
Performance in in silico mixtures. (a) Fetal fraction (FFs) inferred using the full model at the coverage of 0.5× (x-axis) versus FFs inferred using the full model but setting F1 = 0 (y-axis). (b) FFs inferred using the full model at the coverage of 0.5× (x-axis) versus FFs inferred using the full model but setting F2 = 0 (y-axis). (c) True FF (x-axis) versus inferred mean of 100 replicates (y-axis) ± sample standard deviation (not sd of the mean) from the in silico mixture experiments. (d) Variations of FF estimates at different coverage when the true FF is 0.1.
In the in vitro experiments, we used DNA from 11 mother–son pairs, mixed the DNA at different fetal proportions, sequenced the mixture at 0.5×, and inferred the FFs separately using sex chromosome dosages and read heterozygosity. Figure 2a compared the inferred FFs. The two sets of inferences are in high concordance to each other, with the coefficient of determination of \(R^2 = 0.987,\) and maximum absolute deviation being 0.014. Comparing both sets of inferences against the truth, however, showed that both estimates have slight upward biases and larger variations for large FFs (Fig. 2b, c). Experimental variation in DNA quantity measurements for mixing experiment is likely to be the explanation (Supplementary Material and Methods).
Performance in in vitro mixtures. (a) Fetal fractions (FFs) inferred using sex chromosomes (x-axis) versus those inferred using read heterozygosity (y-axis). (b) FFs inferred using read heterozygosity (y-axis) versus the truth (x-axis). (c) FFs inferred using sex chromosome (y-axis) versus the truth (x-axis). In panels (b) and (c) the true values are slightly jiggled for clarity.
In the in natura experiments, we used samples from patients who consented to participate in an ongoing study to improve methods of NIPS (Supplementary Material and Methods). The study was approved by the Institutional Review Board of Beijing Hospital and the DNA samples were de-identified. We first retrospectively selected 69 clinical samples who carry putative male fetus with FFs (obtained from sex chromosome dosages) ranging from 0.03 to 0.15 and we intentionally collected more samples of small FFs, resequenced their cfDNA and wbcDNA at 0.5×, and inferred FFs. Figure 3a compared FFs inferred from sex chromosomes with those inferred from the read heterozygosity. The overall pattern of this plot is highly similar to that of Fig. 2a, with exception of two outliers whose FFs inferred by read heterozygosity are about twice as large as those inferred by sex chromosomes (Fig. 3b). We hypothesize these two samples are female–male twins. Following the IRB protocol, we obtained anonymized patient data and confirmed that those two samples are indeed female–male twins and both pregnancies were results of in vitro fertilization (IVF). For samples with a single male fetuses, FFs inferred by our method are in high concordance with FFs inferred from sex chromosomes (coefficient of determination \(R^2 = 0.972\)) and the largest three absolute deviations are 0.017, 0.016, and 0.015.
Performance of fetal fraction (FF) inference in retrospective clinical noninvasive prenatal screening (NIPS) samples. These 69 samples with male fetus were selected to have a good representation of FFs, with emphasis on smaller ones. The inset histogram is the distribution of ratios between FFs inferred from read heterozygosity and FFs inferred from sex chromosomes. Note that the two outliers, marked by asterisks, were confirmed female–male twins.
To further evaluate our method in real data and examine the prevalence of IVF in our clinical samples, we randomly chose 443 clinical samples and performed the same data collection and statistical analysis as those 69 retrospect samples. Figure 4 demonstrated that our method works well. The 23 red dots (about 5% of 443) in Fig. 4 are IVF pregnancies with two embryos implanted. Along the blue line (y = 2x) are all red dots, indicating female–male twin pregnancies. Those vertical clusters of dots along the y-axis are female fetuses, and the dots along the diagonal line are male fetuses. For samples with a single male fetus, FFs inferred by our method are in high concordance with FFs inferred from sex chromosomes (\(R^2 = 0.971\)) and the largest three absolute deviations are 0.027, 0.019, and 0.013.
Performance of fetal fraction (FF) inference in clinical noninvasive prenatal screening (NIPS) samples. For each sample (represented by a dot), FFs estimated from sex chromosomes are on the x-axis and FFs estimated from read heterozygosity are on the y-axis. Dots marked in red (23/443) are twin pregnancies after in vitro fertilization (IVF). Samples with female fetuses are marked in gray, and samples with male fetuses are marked in black. The gray diagonal line is y = x and the blue line is y = 2x.
Incorporating priors on FFs to test for aneuploidy
The Z-test compares chromosomal dosages of a sample against a set of euploid controls to detect fetal trisomy.17 Because chromosomal dosage estimates have a heavier tail than the normal distribution (Supplementary Fig. S4), the Z-test tends to produce more false positives at a fixed threshold.16 By incorporating an informative null prior to account for the heavy tail, our Bayesian method can reduce false positives.16 With the knowledge of the FF, we can also incorporate the informative alternative prior to compute a Bayes factor to test for the aneuploidy (Supplementary Material and Methods). Intuitively, theZ-test only examines how far a chromosomal dosage is away from the euploid dosage; with the knowledge of the FF, we can also examine how close a chromosomal dosage is to the putative dosage by assuming fetal trisomy. We may use a normal prior for FF under the alternative hypothesis to capture the uncertainty of the FF estimates, which critically depends on the sequencing depth (Supplementary Tables S2 and S3). Note that the ability to incorporate informative priors (both under the null and under the alternative) is a deciding advantage of the Bayesian approach over the Z-test method, which is oblivious to the alternative hypothesis by design.
To compare powers between the Z-test and our Bayesian method that incorporates informative priors, we simulated slightly overdispersed chromosomal dosages under the null, and used this to decide the cutoff value for both Z and Bayesian methods (Supplementary Material and Methods). Then for each target FF denoted by Θ, we simulated chromosomal dosages from N(Θ, σ) and computedZ scores and Bayes factors, where σ depends on sequencing depth and different chromosomes or regions have different σs (Supplementary Tables S2 and S3). The power can be estimated by the percent of test statistics surpassing their respective cutoff values. Table 4 demonstrates that the Bayesian method outperforms the Z-test, particularly for more difficult situations (smaller sequencing depths and smaller FFs). More power simulation results using different priors can be found in Supplementary Tables S4 and S5.
DISCUSSION
We developed a statistical method to infer FF in NIPS, extensively studied its performance, and demonstrated that incorporating the knowledge of FF improves statistical power of NIPS. Our method makes use of read heterozygosity of SNP markers on autosomes and can be applied to samples with either female or male fetuses. The use of read heterozygosity, however, makes our method sensitive to maternal inbreeding coefficient. We therefore propose to sequence maternal wbcDNA in addition to cfDNA. Sequencing maternal wbcDNA brings several benefits. First, it allows us to infer the maternal inbreeding coefficient to better estimate FF. Second, one can mix sequencing reads from wbcDNA and cfDNA to infer a diluted FF under a higher sequencing depth, and the diluted FF can be used to estimate FF after appropriate scaling. Third, although we didn’t pursue here, we would like to note that maternal wbcDNA sequencing can improve NIPS by providing individual-specific reference.
Sequencing wbcDNA in addition to cfDNA bears extra cost. Our method, however, can make do without sequencing wbcDNA. One approach is to plug in the population average inbreeding coefficient, and fit the full model to obtain fetal fraction (the plug-in method). The other is to jointly fit the full model to infer both maternal inbreeding coefficient F1 and fetal fraction h. This can be done efficiently using an iterative method by fixingF1 to update h and then fixing h to update F1 until both converge (the iterative method). The plug-in method worked well for samples with modest inbreeding coefficient (say, between −0.03 and 0.03), but suffered significant bias for samples with extreme inbreeding coefficient. Such an example can be found in Supplementary Fig. S5 (left), in which the outlying sample has an inbreeding coefficient of 0.085. The iterative method, on the other hand, worked well for samples with extreme inbreeding coefficients, but produced a larger variation for samples with modest coefficients (Supplementary Fig. S5, middle). Naturally we combined the plug-in method and the iterative method via thresholding F1 (estimated from the iterative method), such that if \(\left| {F_1} \right| \, > \, 0.03\) we used the iterative method to estimate h and otherwise we used the plug-in method. Supplementary Fig. S5 (right) demonstrated the strength of the combined approach. More extensive numerical studies are warranted.
Our method is designed for nonadmixed samples, but it can be extended to work with admixed samples such as African Americans and Mexicans. The trick is to select a subset of SNPs to make the inference. The natural weight goes into the likelihood calculation for each SNP is p(1 − p) where p is the reference allele frequency. Taking African Americans as an example, we defined \({\rm{logr}} = {\rm{log}}_2\frac{{p_1(1 - p_1)}}{{p_2(1 - p_2)}}\) for each SNP where p1 and p2 are reference allele frequencies of a SNP in European and African populations respectively. We select SNPs whose logr are in a small range, e.g., (−1, 1), to make inferences. Since these SNPs are less ancestry informative, our Hardy–Weinberg assumption is arguably applicable to these SNPs. Supplementary Fig. S6 showed such SNPs are plenty (more than 60%) among SNPs whose minor allele frequencies are >0.01 between any pair of ancestral populations. We simulated cfDNA from African American samples with different fetal fractions (Supplementary Material and Methods), and used selected SNPs to infer their fetal fractions, using either European allele frequencies or African allele frequencies. Supplementary Fig. S7 showed that this approach works well, particularly when the two sets of estimates were averaged.
The knowledge of FF is critical to increase the efficacy of NIPS. One study suggests that samples with smaller FFs may have increased risk of fetal aneuploidy.2 Reporting a “no call" and referring the subject to an invasive test is an effective way to reduce false negatives among these samples. Alternatively, we can perform additional sequencing to increase the sequencing depth to a level that has sufficient power for a given FF. The similar adaptive design can be applied to screen microduplications and microdeletions, where the power tends to be much smaller than the whole chromosome trisomy screening (Table 4).
Because of the widespread practice of drastically reducing costs, NIPS is usually done at the raw sequencing depth of 0.1× in China. When the FF is as large as 6%, 0.1× appears to have sufficient power to screen for trisomy of whole chromosomes. When the FF is at 4%, however, the power is 83% at the type I error of 0.001, which is unsatisfactory given the high social and economic cost of false negatives (Table 4). The situation is much worse for using NIPS to screen for microdeletion and microduplication (NIPS+). Such screenings are done at the raw sequencing depth of 0.5× in China. At such a sequencing depth, our power simulation suggested that for a 5M region, the power is merely 36% at FF of 4%, and 74% at FF of 6% (Table 4). On the other hand, our data suggested that there are 2.2% clinical samples whose FF is <4% and 12.9% clinical samples whose FF is <6% (Supplementary Fig. S8). Therefore, it is imperative to increase sequencing depths for both NIPS and NIPS+ to guarantee the efficacy of the screening.
Code Availability
Software to infer fetal fraction and the source code can be downloaded from https://haplotype.org/download/hetFF.tar.gz. It is free for academic use and can be licensed for commercial use.
Change history
11 December 2019
After further review of the manuscript, the author decided to withdraw the NIH grant acknowledgment as the manuscript was not directly related to the specific aims of the grant. The PDF and HTML versions of the Article have now been modified accordingly.
References
Gregg AR, Skotko BG, Benkendorf JL, Monaghan KG, Bajaj K, Best RG, et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2016;18:1056–1065.
Norton ME, Jacobsson B, Swamy GK, Laurent LC, Ranzini AC, Brar H, et al. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med. 2015;372:1589–1597.
Lo YMD, Tein MSC, Lau TK, Haines CJ, Leung TN, Poon PMK, et al. Quantitative analysis of fetal DNA in maternal plasma and serum: implications for noninvasive prenatal diagnosis. Am J Hum Genet. 1998;62:768–775.
Chiu RWK, Akolekar R, Zheng YWL, Leung TY, Sun H, Chan KCA, et al. Non-invasive prenatal assessment of trisomy 21 by multiplexed maternal plasma DNA sequencing: large scale validity study. BMJ. 2011;342:c7401.
Yu SCY, Chan KCA, Zheng YWL, Jiang PY, Liao GJW, Sun H, et al. Size-based molecular diagnostics using plasma DNA for noninvasive prenatal testing. Proc Natl Acad Sci U S A. 2014;111:8583–8588.
Chan KA, Ding C, Gerovassili A, Yeung SW, Chiu RW, Leung TN, et al. Hypermethylated RASSF1A in maternal plasma: a universal fetal DNA marker that improves the reliability of noninvasive prenatal diagnosis. Clin Chem. 2006;52:2211–2218.
Nygren AOH, Dean J, Jensen TJ, Kruse S, Kwong W, van den Boom D, et al. Quantification of fetal DNA by use of methylation-based DNA discrimination. Clin Chem. 2010;56:1627–1635.
Straver R, Oudejans CBM, Sistermans EA, Reinders MJT. Calculating the fetal fraction for noninvasive prenatal testing based on genome-wide nucleosome profiles. Prenat Diagn. 2016;36:614–621.
Kim SK, Hannum G, Geis J, Tynan J, Hogg G, Zhao C, et al. Determination of fetal DNA fraction from the plasma of pregnant women using sequence read counts. Prenat Diagn. 2015;35:810–815.
Lo YMD, Chan KCA, Sun H, Chen EZ, Jiang PY, Lun FMF, et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci Transl Med. 2010;2:61ra91.
Liao GJW, Lun FMF, Zheng YWL, Chan KCA, Leung TY, Lau TK, et al. Targeted massively parallel sequencing of maternal plasma DNA permits efficient and unbiased detection of fetal alleles. Clin Chem. 2011;57:92–101.
Jiang P, Peng X, Su X, Sun K, Yu SCY, Chu WI, et al. FetalQuant(SD): accurate quantification of fetal DNA fraction by shallow-depth sequencing of maternal plasma DNA. NPJ Genom Med. 2016;1:16013.
Chu TJ, Bunce K, Hogge WA, Peters DG. A novel approach toward the challenge of accurately quantifying fetal DNA in maternal plasma. Prenat Diagn. 2010;30:1226–1229.
Zhang J, Li J, Saucier JB, Feng Y, Jiang Y, Sinson J, et al. Non-invasive prenatal sequencing for multiple Mendelian monogenic disorders using circulating cell-free fetal DNA. Nat Med. 2019;25:439–447.
Vieira FG, Fumagalli M, Albrechtsen A, Nielsen R. Estimating inbreeding coefficients from NGS data: impact on genotype calling and allele frequency estimation. Genome Res. 2013;23:1852–1861.
Xu H, Wang S, Ma LL, Huang S, Liang L, Liu Q, et al. Informative priors on fetal fraction increase power of the noninvasive prenatal screen. Genet Med. 2018;20:817–824.
Chiu RWK, Chan KCA, Gao Y, Lau VYM, Zheng W, Leung TY, et al. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci U S A. 2008;105:20458–20463.
Acknowledgements
This work was supported by the National Key Research and Development Program of China under contract number 2016YFC100707 awarded to USCI Medical Diagnostic Laboratory Inc. The majority of the work was done when Y.G. was a faculty member at Baylor College of Medicine and was supported in part by the US Department of Agriculture/Agriculture Research Service under contract number 6250-51000-057.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
Yongtao Guan is a consultant for Beijing USCI Medical Laboratory serving as its Chief Scientific Officer and owns its stocks and options. Beijing USCI Medical Laboratory contributed to fund the study, but played no role in designing experiment, analyzing data, and interpreting results.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dang, M., Xu, H., Zhang, J. et al. Inferring fetal fractions from read heterozygosity empowers the noninvasive prenatal screening. Genet Med 22, 301–308 (2020). https://doi.org/10.1038/s41436-019-0636-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0636-5
Keywords
This article is cited by
-
Estimation of cell-free fetal DNA fraction from maternal plasma based on linkage disequilibrium information
npj Genomic Medicine (2021)