Abstract
Purpose
Biallelic pathogenic variants in the mismatch repair (MMR) genes cause a recessive childhood cancer predisposition syndrome known as constitutional mismatch repair deficiency (CMMRD). Family members with a heterozygous MMR variant have Lynch syndrome. We aimed at estimating cancer risk in these heterozygous carriers as a novel approach to avoid complicated statistical methods to correct for ascertainment bias.
Methods
Cumulative colorectal cancer incidence was estimated in a cohort of PMS2- and MSH6-associated families, ascertained by the CMMRD phenotype of the index, by using mutation probabilities based on kinship coefficients as analytical weights in a proportional hazard regression on the cause-specific hazards. Confidence intervals (CIs) were obtained by bootstrapping at the family level.
Results
The estimated cumulative colorectal cancer risk at age 70 years for heterozygous PMS2 variant carriers was 8.7% (95% CI 4.3–12.7%) for both sexes combined, and 9.9% (95% CI 4.9–15.3%) for men and 5.9% (95% CI 1.6–11.1%) for women separately. For heterozygous MSH6 variant carriers these estimates are 11.8% (95% CI 4.5–22.7%) for both sexes combined, 10.0% (95% CI 1.83–24.5%) for men and 11.7% (95% CI 2.10–26.5%) for women.
Conclusion
Our findings are consistent with previous reports that used more complex statistical methods to correct for ascertainment bias. These results underline the need for MMR gene–specific surveillance protocols for Lynch syndrome.
Similar content being viewed by others
INTRODUCTION
Lynch syndrome (MIM 120435) is an inherited autosomal dominant condition predisposing to the development of primarily colorectal and endometrial cancer. It is caused by pathogenic variants in the mismatch repair (MMR) genes MLH1 (MIM *120436), MSH2 (MIM *609309), MSH6 (MIM *600678), and PMS2 (MIM *600259). Estimation of Lynch syndrome–associated cancer risk is challenging because until recently, testing for Lynch syndrome was based on clinical or family history criteria such as the Amsterdam II criteria and the (revised) Bethesda guidelines.1,2 Consequently the majority of known Lynch syndrome families were ascertained based on familial cancer history. In recent years there has been a shift toward universal screening of all colorectal and endometrial cancer patients for tumor hallmarks of Lynch syndrome.3,4 These hallmarks include aberrant immunohistochemistry for the MMR proteins and the presence of microsatellite instability.5,6 Furthermore, panel testing of cancer genes, including the MMR genes, is becoming standard practice and is also performed in families with a cancer history that does not necessarily include Lynch syndrome–associated cancers.7 Families identified through universal screening or panel testing may show lower penetrance for Lynch syndrome–associated malignancies, and Hampel et al. were among the first to notice that Lynch syndrome cancer risks are not as high as previously estimated based on analyses of families ascertained using existing guidelines.8 Appropriate surveillance measures for these newly identified families can only be established if risks can be estimated accurately.
Based on retrospective cohorts, current estimates of lifetime colorectal cancer risks for carriers of pathogenic variants in MLH1 and MSH2 are between 52% and 97%.9 Colorectal cancer risk estimates are lower for carriers of a pathogenic variant in MSH6 (22–36%) and lowest of all for PMS2 (11–20%).9,10,11,12 A recent study of a prospective cohort of pathogenic MMR variant carriers undergoing surveillance reported even lower risks, with colorectal cancer risks of 12% for MSH6 and 0% for PMS2, respectively.13 As in the general population, men with Lynch syndrome appear to have a higher colorectal cancer risk than women.14 In most studies, statistical approaches such as modified segregation analysis, exclusion of index cases, and genotype-restricted likelihood estimates have been used to correct for ascertainment bias, but these methods are complex and rely on specific assumptions, and it is difficult to prove that they do not lead to either under- or overestimation of true risk.14 Indeed, Vos et al. showed that a substantial proportion of the variation found in cancer risk estimation in selected hereditary breast cancer families, who show similar ascertainment patterns to Lynch syndrome families, can be explained by the different ascertainment correction method used.15 An alternative approach that minimizes the need for ascertainment bias correction is the selection of families in which the index patient has constitutional mismatch repair deficiency (CMMRD). This childhood cancer predisposition syndrome is caused by biallelic pathogenic variants in one of the MMR genes, most commonly in PMS2. The syndrome is characterized by the development of a broad spectrum of cancers, including hematological, central nervous system, and gastrointestinal neoplasia at a very young age. CMMRD patients may also show signs suggestive of neurofibromatosis type 1, most commonly café au lait macules.16 The CMMRD phenotype is so striking that the diagnosis is often suspected regardless of family history and in one report only 6 of 23 CMMRD patients (26%) had a family history of Lynch syndrome–associated cancers.17 Identification of a child with CMMRD means that both parents are likely to be heterozygous for a pathogenic MMR variant and are at risk for Lynch syndrome–associated malignancies; other family members may similarly be at risk. Because these families were identified due to the CMMRD phenotype rather than family history, they likely represent a near random sample of Lynch syndrome families.
Pathogenic variants in PMS2 were once considered rare and were thought to account for less than 5% of all Lynch syndrome cases.18,19 Nevertheless, germline pathogenic variants in PMS2 were found in a small yet significant proportion (at least 0.57%) of universally screened colorectal cancer cases,20 and recent insights suggest that the carrier frequency for pathogenic variants in PMS2 and MSH6 in the general population is actually much higher than for MLH1 and MSH2.21 The majority of CMMRD patients carry variants in PMS2, followed by MSH6, while MLH1 and MSH2 variants are rarely associated with CMMRD.16 One explanation for this phenomenon is that biallelic pathogenic variants in MLH1 and MSH2 may be embryonically lethal.22,23 However, a higher carrier frequency for variants in PMS2 and MSH6 may also (partly) explain differences in the frequency of pathogenic variants in the MMR genes among patients with CMMRD.
Here we report cumulative cancer risks in family members of CMMRD patients with variants in the PMS2 or MSH6 genes. This study will not only help in the counseling of family members of CMMRD patients, but also represents a novel approach to determining cancer risk in Lynch syndrome.
MATERIALS AND METHODS
Data collection
Families were collected through international collaborations with clinical genetics departments and consortia and by following up CMMRD families described in literature. Corresponding authors were contacted to collect (more) family data. Family structure was recorded and information was collected on each family member regarding gender, variant status, cancer status and age at cancer diagnosis, and last contact or death. A diagnosis of CMMRD was considered confirmed if pathogenic variants were identified or if strong indicators of CMMRD were identified (i.e., phenotype and inheritance pattern plus aberrant immunohistochemistry and/or microsatellite instability in non-neoplastic tissue and/or abnormal functional tests).24
As classified in the InSiGHT database (http://www.insight-database.org/classifications/), 31 unique class 4/5 pathogenic variants in PMS2 and 19 class 4/5 pathogenic variants in MSH6 were found in our cohort.25 Another 30 variants in PMS2 and 8 variants in MSH6 have not been officially classified to date, but were deemed either class 4 or 5 (i.e., [likely] pathogenic) by an expert in the field (H.M.v.d.K.) according to InSiGHT variant classification criteria. Twenty variants of uncertain significance (VUS), distributed over 18 families, were identified and included in the analyses (Tables S1–S4). Seven of the VUS were identified in trans with a (likely) pathogenic variant. Since the patients carrying these VUS displayed a CMMRD phenotype this argues in favor of a functional impact of the variants on protein function. Furthermore, six of the VUS were identified in previously published CMMRD patients (Tables S3 and S4) and as such these variants were considered the most probable cause of the phenotype in these patients. The remaining seven variants were all identified in patients with a CMMRD phenotype and were considered a probable cause of the phenotype by the reporting laboratory and clinicians.
Statistical analysis
Eligible first- and second-degree family members for the risk analysis were defined based on complete data describing gender, age at cancer diagnosis, last contact or death, and status as a (possible) carrier of the PMS2/MSH6 variant. Proven and obligate carriers as well as untested family members were included, whereas noncarriers, as confirmed by DNA analysis, were excluded. Known CMMRD patients were excluded from the analysis, as were (deceased) siblings of a CMMRD patient when they had a cancer within the CMMRD spectrum. In consanguineous families, family members with an unknown variant status, but a cancer diagnosis within the CMMRD cancer spectrum at a young age (i.e., <25 years of age) were considered to be homozygous carriers and were thus excluded from the risk analysis. The total number of colorectal and endometrial cancers is described for the total cohort as well as for the part of the cohort included in the risk analysis. To avoid a reporting bias due to distant relatives (distant family members may be more likely to be included in the pedigree if they were affected, while unaffected distant family members may go unreported), only first- and second-degree relatives of the index patients were included in the risk analyses. This approach was supported by both visual inspection of the pedigrees and by an otherwise unexplained increase in colorectal cancer frequency among more distant family members (data not shown, available upon request).
Colorectal cancer risk is reported as cumulative incidence at age 70, accounting for death and other cancer diagnoses as competing risks.26 Age at removal of a colon polyp was included as a censoring event because the likelihood of developing colorectal cancer is probably reduced after this preventive measure. Likewise, family members were censored at the development of any type of cancer, excluding basal cell carcinoma, because treatment of a cancer (e.g., by radiotherapy or chemotherapy) might influence future cancer risk.
To avoid testing bias, which may arise when the decision to undergo genetic testing is related to cancer status, we included untested family members in our study, weighted according to their genetic distance to confirmed carriers. Specifically, variant probabilities based on kinship coefficients were used as analytical weights in a Cox proportional hazard regression to model the hazard of developing colorectal cancer in the presence of competing events (death and other cancer diagnosis), and including sex as a covariate (for details see “Statistical Methods” in the Supplemental Data). For example, first-degree relatives of a confirmed carrier who were not tested were given a weight of 0.50, whereas second-degree relatives had a weight of 0.25. Confidence intervals (CIs) were obtained by bootstrapping at family level (1000 repetitions).
Medical ethical approval for this study was obtained through the ethics committee of Leiden University Medical Centre (reference number P14.090). Informed consent was not required because all data was collected anonymously.
RESULTS
After exclusion of the CMMRD cases, the PMS2 cohort included 1809 family members from 77 families and the MSH6 cohort consisted of 561 family members from 26 families.
Age at colorectal and endometrial cancer diagnosis
Sixty patients from 31 families were diagnosed with colorectal cancer in the total PMS2 cohort, and 16 women from 14 families were diagnosed with endometrial cancer after excluding the CMMRD cases. Age of colorectal cancer diagnosis within this cohort ranged from 36 to 80 years, with a median age of 60 years. Age at diagnosis was unknown for 17 colorectal cancer cases (Table 1). For the 16 endometrial cancer cases, the age at diagnosis ranged from 40 to 85, with a median of 61 years. Age was missing for only one of these cases.
Seventeen patients from 12 families were diagnosed with colorectal cancer in the total MSH6 cohort after exclusion of CMMRD cases. Age of colorectal cancer diagnosis in this cohort ranged from 42 to 58 years, with a median of 48 years (Table 1). There were five cases of endometrial cancer distributed over four families, with a median age at diagnosis of 54 years and an age range of 47 to 59 years.
Other cancers
While a range of other cancer types were reported in both the PMS2 and MSH6 cohort, low numbers did not allow risk analyses to be performed. The most commonly reported cancers were breast cancer, lung cancer, leukemia, and prostate cancer (Table 1 and Table S5).
Colorectal cancer risk
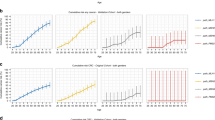
For individuals with CMMRD and variants in PMS2, 549 family members from 64 families were eligible for risk analysis; of these, 212 were confirmed or obligate carriers and the rest potential carriers. The estimated cumulative colorectal cancer risk at age 70 for heterozygous PMS2 variant carriers was 8.7% (95% CI 4.3–12.7%, Fig. 1) for both sexes combined, and was 9.9% (95% CI 4.9–15.3%) for men and 5.9% (95% CI 1.6–11.1%) for women. Endometrial cancer risk could not be estimated due to the low number of events (n = 8).
For MSH6, 148 family members from 24 families were eligible for risk analysis; of these 69 were confirmed or obligate carriers and the rest potential carriers. The cumulative colorectal cancer risk at age 70 for heterozygous MSH6 gene variant carriers was 11.8% (95% CI 4.5–22.7%, Fig. 2) for both sexes, and 10.0% (95% CI 1.8–24.5%) and 11.7% (95% CI 2.1–26.5%) for men and women, respectively. There were no cases of endometrial cancer that could be included in the risk analysis.
DISCUSSION
Using a new approach to establishing cancer risks in Lynch syndrome, we can confirm the low PMS2- and MSH6-associated colorectal cancer risks reported in previous studies that used ascertainment bias correction methods10,11,12,14 or prospective data.13,27 The main strengths of our approach were the reduction in clinical ascertainment bias by analyzing family members of CMMRD patients and the use of a competing risk analysis approach to avoid bias due to informative right censoring. Our results further indicate that gene-specific surveillance guidelines are needed to avoid subjecting carriers at low cancer risk to the invasive processes of surveillance, in some cases from an unnecessarily young age. The earliest age of colorectal cancer diagnosis was 36 and 42 years for PMS2 and MSH6, respectively, well above the age (20–25 years) at which surveillance is usually started for individuals with Lynch syndrome.28 This suggests that, in heterozygous carriers of PMS2 or MSH6 variants from families that do not meet clinical selection criteria for Lynch syndrome, surveillance could be started at a later age, e.g., at 35–40 years. Although current lifetime risk estimates are only slightly (2–3 times) elevated above the population risk of ~4%,29 there are indications (e.g., from the median age at diagnosis) that risk is elevated at younger ages, and a faster progression from precursor lesion to carcinoma cannot be excluded. Therefore, we do not recommend that surveillance be omitted based on the current data. Furthermore, large variation in penetrance has been observed in clinically ascertained families, indicating that other risk factors may influence risk. Together these considerations suggest that our risk estimates remain useful when counseling families who were not ascertained based on criteria such as the Amsterdam II criteria and the (revised) Bethesda guidelines, e.g., families with a CMMRD proband or with a pathogenic MMR variant identified as an incidental finding through exome sequencing. However, they should be used with caution in more severely affected families, for example when a family history fulfills the Amsterdam criteria.2
Unfortunately, both cohorts were too small to provide risk estimations for endometrial cancer. It is striking that there were only some cases of endometrial cancer in the total MSH6 cohort and none that could be included in the risk analysis, while the risk of endometrial cancer in MSH6 has been reported to be high.27 This may be partly due to the relatively low median age of 45 years (Table 1) of the cohort, while the youngest age at diagnosis of endometrial cancer was 47 years (Table 1).
There are some limitations to the current study. Firstly, genotype–phenotype correlations in Lynch syndrome and CMMRD have been proposed (although thus far no conclusive evidence has been yielded and some studies even show contradictory results).30,31,32,33,34,35 If correlations exist, variants with a milder phenotype might be overrepresented in a CMMRD cohort. For PMS2, age at cancer diagnosis and risk estimates were within the range of previous retrospective studies that corrected for ascertainment bias, indicating that we have not selected a cohort of (solely) low-risk PMS2 alleles.10,11,12 Cancer risk estimates and age at cancer diagnosis for MSH6 are similar to a study by Bonadona et al.,36 but risk estimates are slightly lower than those reported by Baglietto et al.37
A possible mechanism for a genotype–phenotype correlation could be nonsense-mediated messenger RNA (mRNA) decay. Nonsense-mediated decay (NMD) detects mRNAs with premature termination codons and initiates their degradation, preventing potential dominant negative effects from truncated proteins.38 Some variants, e.g., missense variants, are likely to escape NMD. To assess a possible role for NMD, we performed a stratified risk analysis that divided family members into groups based on whether their risk variant is expected to result in NMD, as described previously (Suerink et al.30). Family members were excluded from this analysis when no reliable prediction of NMD was available for the variant or if it was not known which variant segregated in which half of the family (maternal or paternal). This analysis produced no clear genotype–phenotype correlations and for both genes cases of colorectal cancer were seen in the NMD group as well as in the group with predicted retention of RNA expression. However, it should be noted that wide confidence intervals excluded detection of small differences (data available upon request). Whether risk stratification is possible based on genotype will require further study.
It could also be argued that a bias toward a milder phenotype is inherent to our cohort because those who die of cancer at a young age cannot have children with CMMRD. However, because both the parents and more distant relatives were included in the current analyses, it seems unlikely that this possible bias could have a major impact, particularly because the youngest age at colorectal cancer diagnosis within the total cohort was 36 years.
Another potential problem was testing bias, which arises because family members with cancer are more inclined to undergo genetic testing. We therefore used variant probabilities based on genetic distance to confirmed carriers as analytical weights in our statistical analysis, which also enabled inclusion of untested family members. By including obligate carriers in the analysis there is a risk of misidentifying someone as a possible carrier because the CMMRD patient may have had a de novo variant. However, de novo variants are rarely reported in Lynch syndrome (2.3% in a cohort described by Win et al.39) and a large proportion (55% and 50% for PMS2 and MSH6, respectively) of CMMRD index patients were homozygous for one variant and/or were from consanguineous families. Moreover, a major testing bias was not expected due to a low overall cancer risk and because a relatively large proportion of confirmed carriers were obligate carriers (45/212 [21%] for the PMS2 cohort and 21/69 [30%] for the MSH6 cohort) whose testing status is by definition uninfluenced by their phenotype. It is worth mentioning that while our approach avoids clinical ascertainment bias, the selection strategy results in a relatively young cohort, which implies large uncertainty in the incidence estimation at older ages, as reflected by the broad confidence intervals in Figs. 1 and 2.
A final limitation of our study that could impact the reliability of data is the fact that most cancer diagnoses in this cohort were based on the proband’s knowledge of family history rather than on medical records. Reassuringly, a 2011 study showed that the accuracy of reported colorectal cancer for first-degree family members was over 90% (ref. 40). Because we included only first- and second-degree family members, with family history reported by the parents in most cases, we expect a comparable accuracy rate in our risk analysis.
To complement and confirm the data presented here, we suggest a similar risk analysis should be performed in PMS2 and MSH6 families detected through universal screening of colorectal cancers for mismatch repair deficiency. These families will also be less affected with ascertainment bias.
In summary, we used an alternative approach to establish colorectal cancer risk in Lynch syndrome patients with PMS2 and MSH6 variants in CMMRD families. We confirmed this relatively low cancer risk relative to earlier, biased estimates of risk. These results underline the need for gene-specific surveillance protocols for PMS2- and MSH6-related Lynch syndrome families. Further investigations will be required to estimate the cancer risk for other Lynch syndrome–associated malignancies for PMS2 and MSH6, as well as estimating unbiased cancer risks estimates for carriers of pathogenic variants in MLH1 and MSH2.
References
Umar A, Boland CR, Terdiman JP, et al. Revised Bethesda Guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst. 2004;96:261–268.
Vasen HF, Watson P, Mecklin JP, Lynch HT. New clinical criteria for hereditary nonpolyposis colorectal cancer (HNPCC, Lynch syndrome) proposed by the International Collaborative group on HNPCC. Gastroenterology. 1999;116:1453–1456.
Leenen CH, Goverde A, de Bekker-Grob EW, et al. Cost-effectiveness of routine screening for Lynch syndrome in colorectal cancer patients up to 70 years of age. Genet Med. 2016;18:966–973.
Dillon JL, Gonzalez JL, DeMars L, et al. Universal screening for Lynch syndrome in endometrial cancers: frequency of germline mutations and identification of patients with Lynch-like syndrome. Hum Pathol. 2017;70:121–128.
Boland CR, Thibodeau SN, Hamilton SR, et al. A National Cancer Institute workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res. 1998;58:5248–5257.
Richman S. Deficient mismatch repair: read all about it (review). Int J Oncol. 2015;47:1189–1202.
Desmond A, Kurian AW, Gabree M, et al. Clinical actionability of multigene panel testing for hereditary breast and ovarian cancer risk assessment. JAMA Oncol. 2015;1:943–951.
Hampel H, Stephens JA, Pukkala E, et al. Cancer risk in hereditary nonpolyposis colorectal cancer syndrome: later age of onset. Gastroenterology. 2005;129:415–421.
Hampel H, de la Chapelle A. The search for unaffected individuals with Lynch syndrome: do the ends justify the means? Cancer Prev Res (Phila). 2011;4:1–5.
Senter L, Clendenning M, Sotamaa K, et al. The clinical phenotype of Lynch syndrome due to germ-line PMS2 mutations. Gastroenterology. 2008;135:419–428.
ten Broeke SW, Brohet RM, Tops CM, et al. Lynch syndrome caused by germline PMS2 mutations: delineating the cancer risk. J Clin Oncol. 2015;33:319–325.
Ten Broeke SW, van der Klift HM, Tops CMJ, et al. Cancer risks for PMS2-associated Lynch syndrome. J Clin Oncol. 2018;36:2961–2968.
Moller P, Seppala T, Bernstein I, et al. Cancer incidence and survival in Lynch syndrome patients receiving colonoscopic and gynaecological surveillance: first report from the prospective Lynch syndrome database. Gut. 2017;66:464–472.
Barrow E, Hill J, Evans DG. Cancer risk in Lynch syndrome. Fam Cancer. 2013;12:229–240.
Vos JR, Hsu L, Brohet RM, et al. Bias correction methods explain much of the variation seen in breast cancer risks of BRCA1/2 mutation carriers. J Clin Oncol. 2015;33:2553–2562.
Wimmer K, Kratz CP, Vasen HF, et al. Diagnostic criteria for constitutional mismatch repair deficiency syndrome: suggestions of the European consortium ‘care for CMMRD’ (C4CMMRD). J Med Genet. 2014;51:355–365.
Lavoine N, Colas C, Muleris M, et al. Constitutional mismatch repair deficiency syndrome: clinical description in a French cohort. J Med Genet. 2015;52:770–778.
Gill S, Lindor NM, Burgart LJ, et al. Isolated loss of PMS2 expression in colorectal cancers: frequency, patient age, and familial aggregation. Clin Cancer Res. 2005;11:6466–6471.
Peltomaki P. Deficient DNA mismatch repair: a common etiologic factor for colon cancer. Hum Mol Genet. 2001;10:735–740.
Truninger K, Menigatti M, Luz J, et al. Immunohistochemical analysis reveals high frequency of PMS2 defects in colorectal cancer. Gastroenterology. 2005;128:1160–1171.
Win AK, Jenkins MA, Dowty JG, et al. Prevalence and penetrance of major genes and polygenes for colorectal cancer. Cancer Epidemiol Biomarkers Prev. 2017;26:404–412.
Bakry D, Aronson M, Durno C, et al. Genetic and clinical determinants of constitutional mismatch repair deficiency syndrome: report from the constitutional mismatch repair deficiency consortium. Eur J Cancer. 2014;50:987–996.
Durno C, Boland CR, Cohen S, et al. Recommendations on surveillance and management of biallelic mismatch repair deficiency (BMMRD) syndrome: a consensus statement by the US Multi-Society Task Force on Colorectal Cancer. Gastroenterology. 2017;152:1605–1614.
Bodo S, Colas C, Buhard O, et al. Diagnosis of constitutional mismatch repair-deficiency syndrome based on microsatellite instability and lymphocyte tolerance to methylating agents. Gastroenterology. 2015;149:1017–29 e1013.
Thompson BA, Spurdle AB, Plazzer JP, et al. Application of a 5-tiered scheme for standardized classification of 2,360 unique mismatch repair gene variants in the InSiGHT locus-specific database. Nat Genet. 2014;46:107–115.
Putter H, Fiocco M, Geskus RB. Tutorial in biostatistics: competing risks and multi-state models. Stat Med. 2007;26:2389–2430.
Moller P, Seppala TT, Bernstein I, et al. Cancer risk and survival in path_MMR carriers by gene and gender up to 75 years of age: a report from the Prospective Lynch Syndrome Database. Gut. 2018;67:1306–1316.
de Vos tot Nederveen Cappel WH, Jarvinen HJ, Lynch PM, et al. Colorectal surveillance in Lynch syndrome families. Fam Cancer. 2013;12:261–265.
Noone AM, Howlader N, Krapcho M, et al. SEER cancer statistics review, 1975-2015. 2018. https://seer.cancer.gov/csr/1975_2015/.
Suerink M, van der Klift HM, Ten Broeke SW, et al. The effect of genotypes and parent of origin on cancer risk and age of cancer development in PMS2 mutation carriers. Genet Med. 2016;18:405–409.
Geary J, Sasieni P, Houlston R, et al. Gene-related cancer spectrum in families with hereditary non-polyposis colorectal cancer (HNPCC). Fam Cancer. 2008;7:163–172.
Peltomaki P, Gao X, Mecklin JP. Genotype and phenotype in hereditary nonpolyposis colon cancer: a study of families with different vs. shared predisposing mutations. Fam Cancer. 2001;1:9–15.
Perez-Cabornero L, Infante M, Velasco E, et al. Genotype-phenotype correlation in MMR mutation-positive families with Lynch syndrome. Int J Colorectal Dis. 2013;28:1195–1201.
Ryan NAJ, Morris J, Green K, et al. Association of mismatch repair mutation with age at cancer onset in Lynch syndrome: implications for stratified surveillance strategies. JAMA Oncol. 2017;3:1702–1706.
Li L, Hamel N, Baker K, et al. A homozygous PMS2 founder mutation with an attenuated constitutional mismatch repair deficiency phenotype. J Med Genet. 2015;52:348–352.
Bonadona V, Bonaiti B, Olschwang S, et al. Cancer risks associated with germline mutations in MLH1, MSH2, and MSH6 genes in Lynch syndrome. JAMA. 2011;305:2304–2310.
Baglietto L, Lindor NM, Dowty JG, et al. Risks of Lynch syndrome cancers for MSH6 mutation carriers. J Natl Cancer Inst. 2010;102:193–201.
Popp MW, Maquat LE. Nonsense-mediated mRNA decay and cancer. Curr Opin Genet Dev. 2018;48:44–50.
Win AK, Jenkins MA, Buchanan DD, et al. Determining the frequency of de novo germline mutations in DNA mismatch repair genes. J Med Genet. 2011;48:530–534.
Edwards E, Lucassen A. The impact of cancer pathology confirmation on clinical management of a family history of cancer. Fam Cancer. 2011;10:373–380.
Acknowledgements
We acknowledge Susan E. Andrew (Department of Medical Genetics, University of Alberta, Edmonton, Canada) and Kate Green (Division of Evolution and Genomic Medicine, Manchester Academic Health Science Centre [MAHSC], University of Manchester, St Mary’s Hospital, Manchester, UK) for providing data, and the Care for CMMRD (C4CMMRD) Consortium for providing data and a platform to discuss this study. The authors thank Medactie.com for help with editing of this paper. This work was supported by a grant from the Dutch Cancer Society (KWF UL 2012–5155).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
D.G.E. is supported by the Manchester NIHR Biomedical Research Centre (IS-BRC-1215-20007). K.W.J. is a full-time employee of Ambry Genetics. S.N. is funded by Foundation ARC 2017, Foundation Gustave Roussy, and Swiss Cancer League KFC-3985-08-2016. The other authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Suerink, M., Rodríguez-Girondo, M., van der Klift, H.M. et al. An alternative approach to establishing unbiased colorectal cancer risk estimation in Lynch syndrome. Genet Med 21, 2706–2712 (2019). https://doi.org/10.1038/s41436-019-0577-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0577-z
Keywords
This article is cited by
-
Tumorrisikosyndrome: Erkennen - Untersuchen - Betreuen
InFo Hämatologie + Onkologie (2021)