Abstract
Purpose
To provide a detailed electroclinical description and expand the phenotype of PIGT-CDG, to perform genotype–phenotype correlation, and to investigate the onset and severity of the epilepsy associated with the different genetic subtypes of this rare disorder. Furthermore, to use computer-assisted facial gestalt analysis in PIGT-CDG and to the compare findings with other glycosylphosphatidylinositol (GPI) anchor deficiencies.
Methods
We evaluated 13 children from eight unrelated families with homozygous or compound heterozygous pathogenic variants in PIGT.
Results
All patients had hypotonia, severe developmental delay, and epilepsy. Epilepsy onset ranged from first day of life to two years of age. Severity of the seizure disorder varied from treatable seizures to severe neonatal onset epileptic encephalopathies. The facial gestalt of patients resembled that of previously published PIGT patients as they were closest to the center of the PIGT cluster in the clinical face phenotype space and were distinguishable from other gene-specific phenotypes.
Conclusion
We expand our knowledge of PIGT. Our cases reaffirm that the use of genetic testing is essential for diagnosis in this group of disorders. Finally, we show that computer-assisted facial gestalt analysis accurately assigned PIGT cases to the multiple congenital anomalies–hypotonia–seizures syndrome phenotypic series advocating the additional use of next-generation phenotyping technology.
Similar content being viewed by others
INTRODUCTION
Glycosylphosphatidylinositol (GPI) is a glycolipid that anchors proteins to the cell membrane.1 The synthesis of GPI-anchored proteins (GPI-APs) is important for protein processing and function.2,3 It also plays a crucial role in embryogenesis, immune response, and neurogenesis.4,5,6,7 GPI synthesis and GPI-AP modification are mediated by at least 29 genes and loss-of-function pathogenic variants in 19 of these genes have been described to cause neurological impairments including seizures, intellectual disability (ID), developmental delay (DD), and multiple congenital anomalies8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23 (http://www.iembase.org/nosology/n-browse.asp).
PIGT (MIM 610272) encodes phosphatidylinositol-glycan biosynthesis class T, which is a subunit of the heteropentameric GPI transamidase complex that facilitates the attachment of GPI anchors to proteins.24,25 In 2013, Kvarnung et al. identified a homozygous PIGT pathogenic variant in four patients from a consanguineous Turkish family with multiple congenital anomalies–hypotonia–seizures syndrome 3 (MCAHS3) (MIM 615398).21 Subsequently, compound heterozygous pathogenic variants in PIGT were identified in seven unrelated families with a similar clinical presentation.26,27,28,29 Among these seven families were two brothers recently described by Skauli et al. who presented with typical features of MCAHS3, but in addition, pyramidal tract signs.26 The predominant presentation for the 13 patients described so far is that of an epileptic encephalopathy including profound ID, hypotonia, cortical visual impairment, and nystagmus plus cortical/cerebellar atrophy.
Here we describe 13 additional patients from eight unrelated families with PIGT pathogenic variant presenting with seizures, ID, and congenital anomalies. We delineate the electroclinical features of PIGT-CDG and expand the phenotypic spectrum. In addition, we show that some of the mild facial dysmorphic features constitute a recognizable entity given that automated facial image analysis correctly assigns five of the novel patients to the MCAHS phenotypic disease entity.
MATERIALS AND METHODS
Patient analysis and variant identification
All patients were subjected to next-generation sequencing (NGS)-based panel or exome sequencing approaches and were contributed by clinicians from epilepsy and genetic centers in Europe and the United States as well as from the Deciphering Developmental Disorders (DDD) Study in the United Kingdom (Supplementary Methods and Supplementary File S-1). The patients and their families underwent detailed clinical examinations. Medical files, magnetic resonance images (MRIs), electroencephalograms (EEGs), and NGS data were carefully reviewed.
Standard protocol approvals, registrations, and patient consents
The study was approved by the following local ethics committees: ethics committee of Region Zealand in Denmark (RJ-91); ethics committee at the Kiel University’s Faculty of Medicine, Germany (A 115/02, A 116/02); ethics committee of the University of Hospital Bonn Medical Center; and the UK Research Ethics Committee (10/ H0305/83; Cambridge South REC, and GEN/284/12; Republic of Ireland REC). All probands or, in case of minors, their parents or legal guardians gave informed consent (including for the use of photos). The clinical information has been collected from hospital records of the patients and their family members.
Automated facial gestalt analysis
The resemblance between the face of a patient and the representation of a syndrome class can be quantified by the gestalt score obtained from Face2Gene. Here we used the gestalt scores of the top 30 syndromes suggested by Face2Gene’s DeepGestalt30 system as vectors for an indirect representation of each case in a high-dimensional vector space (Supplemental Fig. S-1, D). We used 100 published cases from five cohorts (PIGA [MCAHS2], PIGN [MCAHS1], PIGT [MCAHS3], PIGV [HPMRS1], and PGAP3 [HPMRS4], described in Knaus et al.31) and the five novel PIGT cases to create a 215-dimensional syndrome space. A linear principal component analysis (PCA) was performed on the vector matrix to reduce representation to two and three dimensions, respectively (Fig. 2 and Supplementary Figure S-1E). The relationship between the different cohorts was compared by relative distance measure (Supplemental Fig. S-1F). Confounder effects were analyzed by distance consistency measure (data not shown).
RESULTS
We recruited 13 patients with homozygous or compound heterozygous variants in PIGT from research and diagnostic programs in Europe and United States. The pedigrees of the families are shown in Fig. 1. Mining the available literature, we were able to identify 13 additional previously reported PIGT patients.21,26,27,28,29 A short overview of the clinical, MRI, and genetic features of the 13 novel patients and of the 13 previously published PIGT-CDG cases is provided in Supplementary Table 1 (see Supplementary Table S-4 for a comprehensive overview).
Phenotypic analysis
All 13 novel patients had either profound (patients 1–2, 6–9, and 11–13) or severe (patients 3–5 and 10) intellectual disability and epilepsy with neonatal or infantile onset. The median age at seizure onset was six months (range 1 day–18 months). All patients had myoclonic and/or tonic seizures, often with apnea, sometimes evolving to bilateral tonic–clonic seizures. Subtle focal seizures were also described (patients 1–2, 8–9, and 11–12). Fever-associated seizures were reported in all patients.
The symptoms varied from profound ID and severe drug resistant, neonatal, or infantile onset epilepsy (patients 1–2, 6–9, and 11–13) with recurrent episodes of convulsive status epilepticus (patients 1–2), to severe ID and treatable epilepsy with later age at onset (8–18 months) (patients 3–5 and 10).
The interictal EEG was available in 12 of 13 patients (no data available for patient 8) and was severely abnormal in 11 of them (see Supplementary Table 1). One patient had a normal EEG at epilepsy onset (patient 8) and no further EEG controls were available. In patients with neonatal epilepsy onset (patients 1–2, 12, and 13), the EEG showed initially a burst suppression pattern and at follow up (patients 1–2) was characterized by background slowing, with frequent multifocal spike and slow waves. Patients with early infantile epilepsy onset and severe drug resistant epilepsy (patients 6, 8, 9, and 11) might have a normal EEG at epilepsy onset (patients 8–9) and develop background slowing, focal theta–delta activity in the frontotemporal regions or in the posterior quadrants, and frequent multifocal spike and slow waves at follow up. The EEG of patients with treatable epilepsy with later age at onset (patients 3–5 and 10) showed only background slowing (patient 10) or background slowing with sporadic (patients 3–5) epileptiform abnormalities in the frontotemporal regions.
Cortical visual impairment was diagnosed in five patients (patients 1, 2, 4, 5, 7, and 11). None of the patients were diagnosed with hearing loss, and they all exhibited normal alkaline phosphatase, plasma calcium, plasma phosphate, and parathyroid hormone values. Congenital heart defects were identified in two individuals and both resolved spontaneously: a persistent foramen ovale (PFO) in one patient (patient 8) and the combination of a PFO and an atrial septal aneurysm in the other (patient 6).
Patients 2 and 7 both died due to pneumonia at 11 and 15 months of age, respectively, and patient 12 died at six months of age following a cardiac arrest. Patient 3 was operated on at the age of two years for a suprasellar adamantinomatous craniopharyngioma.
All patients shared similar craniofacial features that included a high forehead with bitemporal narrowing, a depressed nasal bridge, a short anteverted nose, distinct and prominent philtrum, full cheeks, and an open mouth consistent with general hypotonia. Scalp hair, eyebrows, and eyelashes were sparse in 8 of the 13 patients (patient 1, 2, 3–7, and 11) (Fig. 2 and Fig. S-2). Because clinical pictures were not available it has not been possible to evaluate whether or not the remaining patients shared this feature.
Clinical pictures of patient 1 at 3½ years (a), patient 2 at 11 months (b), patient 3 at 2½ years (c), patient 4 at 7 years (d), and patient 5 at 19 months (e). Sparse scalp hair, bitemporal narrowing, a high forehead, a long philtrum with closely placed philtral pillars, a tented mouth, full cheeks, a depressed nasal bridge, and a short nose in all patients is notable. Eyebrows were either straight (a–c) or arched (d, e). f Principal component analysis of gestalt scores from DeepGestalt of cases with disease-causing variant in PIGT, PIGN, PIGA, PIGV, and PGAP3. Facial gestalt of six of the novel patients (P-1 to P-5, and P-11) share a similar gestalt as previously published cases with variants in genes of the same subgroup PIGT, PIGA, and PIGN (MCAHS). 1- and 2-SD ellipses are drawn around the centroids of the clusters. HPMRS hyperphosphatasia with mental retardation syndrome, MCAHS multiple congenital anomalies–hypotonia–seizures syndrome.
MRI brain scan was available in all 13 patients and was abnormal in all but two patients (patients 5 and 13). MRI abnormalities included prominent cortical and subcortical volume loss with brainstem atrophy (patients 1, 2, 3, 8, and 10–11), white matter immaturity (patients 1, 2, and 12), hypoplastic cerebellum (patients 3–5, 8, and 11) and an abnormal corpus callosum (patients 1 and 2) (Supplementary Table 1).
A detailed clinical description of all 13 patients is available in the Supplementary File S-2.
Automated facial gestalt analysis
Clustering based on facial gestalt analysis by application of Face2Gene’s DeepGestalt algorithm shows a nonrandom distribution of published and novel PIGT cases in the glycosylphosphatidylinositol biosynthesis defect (GPIBD) syndrome space. Patients of the two phenotypic series MCAHS and hyperphosphatasia with mental retardation syndrome (HPMRS) form two clusters (Fig. 2e), in which the MCAHS cluster consists of patients with pathogenic variants in PIGT, PIGN, and PIGA, while the HPMRS cluster takes up cases with PIGV and PGAP3 variant. Some of the overlap in the visualization of the MCAHS and HPMRS cluster (Fig. 2f) is due to a reduction of information to the two principal components. A better separability between PIGT, PIGA, and PIGN cohorts is achieved in a three-dimensional PCA (Supplementary Figure S-1E). The special distribution of the five novel patients in the MCAHS cluster is similar to the published PIGT patients (Supplementary Figure S-1E). Calculation of centroid-based Euclidian distances between patients from the HPMRS and MCAHS phenotypic series showed a hierarchical delineation (Supplementary Figure S-1F). Confounder effects such as sex, age, and ethnicity were analyzed by distance consistency measure and showed no confinement (data not shown).
Analysis of variants
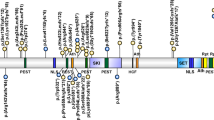
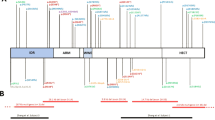
We identified nine different disease-causing variants including three novel variants (one missense and two splice-site variants). In all families the variants were biallelic. All variants were predicted to be damaging by different prediction tools (Supplementary File S-1). The protein positions of the different PIGT variants are shown in Fig. 3. Several recurrent pathogenic variants were identified. The c.550G>A variant was recently identified in homozygous state in a Chinese patient,29 which is shared in patient 11 from Pakistan. The c.709G>C variant was found in two Bangladeshi siblings (patients 12 and 13) and had previously been published in an Afghan male27, indicating an Asian origin of the variant. The c.709G>C was detected in a heterozygous state in three patients (patients 12, 13, and the patient 2 reported by Pagnamenta32,33). All three patients presented with epileptic seizures within the first two weeks of life suggesting that c.709G>C could be associated with a neonatal onset epileptic encephalopathy. The potentially European originating transversion c.1582G>A was found in four Polish patients (patients 3–5 and 10) with severe developmental delay with a treatable epilepsy and the same variant has been identified in compound heterozygous state with a severe loss-of-function variant (c.1730dupC) in a Caucasian patient.27 This patient had a global developmental delay with seizures; she became seizure-free with a combination of antiepileptic drugs (personal communication). This suggests that the missense variant c.1582G>A could cause a milder phenotype. The limited number of patients does not, however, allow a definitive conclusion regarding this genotype–phenotype correlation.
a Locations of variants in the PIGT gene. Novel variants described in this study are represented in red with the patient ID in superscript. Known pathogenic variants are shown with corresponding reference numbers. b Conservation analysis. Amino acid comparison of PIGT of different species. Variants affect highly conserved amino acids (black boxes).
The novel identified transition c.494-2A>G in patient 3 leads to a loss of a consensus acceptor site and is predicted to affect splicing. The c.1472T>A transversion leads to an exchange of a highly conserved leucine at position 491 to a histidine, which may alter the luminal portion of PIGT protein. The c.1484+2T>A transversion in patients 1 and 2 leads to loss of a consensus donor site and is predicted to affect splicing.
Overall landscape of disease-causing variants
From the complete data set of 26 patients (13 novel + 13 previously published) with pathogenic PIGT variants, 13 different variant sites emerged, of which three are novel (bold): c.250G>T (n = 2)28,34; c.494-2A>G (n = 1); c.547A>C (n = 2) (ref. 21); c.550G>A (n = 2) (ref. 29); c.709G>C (n = 3) (ref. 27); c.918dupC (n = 1) (ref. 35); c.1079G>T (n = 5) (ref. 26); c.1096G>T (n = 1) (ref. 28); c.1342C>T (n = 2) (refs. 34,35); c.1472T>A (n = 2); c.1484 + 2T>A (n = 2); c.1582G>A (n = 4) (ref. 27); and c.1730dupC (n = 2) (ref. 27). Variants were spread across the gene and there was no clustering in a specific domain making it difficult to establish a genotype–phenotype correlation based on the landscape of pathogenic variants.
DISCUSSION
So far 13 patients with a GPI anchor deficiency due to recessive PIGT variants have been described21,27,28,29,34,35 and all but 2 patients27,29 presented with an epileptic encephalopathy. We describe an additional 13 patients from eight unrelated families. The patients show a broad clinical spectrum and share several common features (Supplementary Table 1). The neurological findings include severe/profound ID, epilepsy with variable onset and severity, and a severe congenital hypotonia. The symptoms varied from profound ID and severe drug resistant epilepsy with neonatal–infantile onset epilepsy (8/13 patients), to severe ID and treatable epilepsy with later age at onset (8–24 months) (4/13 patients). Based on previously reported cases and our novel findings we conclude that the missense variant c.1582G>A seems to be associated with a milder phenotype, i.e., treatable epilepsy. Also our data suggests that the c.709G>C could be associated with a neonatal onset epileptic encephalopathy. The limited number of patients, however, does not allow a definitive conclusion regarding this genotype–phenotype correlation. Ophthalmological features including nystagmus and/or strabismus and a cortical visual impairment were also observed in 9 of 13 patients (Supplementary Table 1).
The skeletal and endocrine features in our patients differed from the initial five patients reported by Nakashima et al.34 and Kvarnung et al.;21 however, their features overlapped with the four patients presented by Lam et al.35 and Skauli et al.26 Our patients exhibited normal alkaline phosphatase, plasma calcium, plasma phosphate, and parathyroid hormone values. Bone age was only available in patient 1 and was normal. Apart from the brothers presented by Skauli et al.27 all previously described patients had reduced bone mineralization and scoliosis, features that could arise secondary to their neurologic complications. X-rays were only available in three of our patients (patient 1 and 2) and only two patients (patient 1 and 12) underwent a systematic skeletal survey. Scoliosis was found in one of our patients (patient 4), undermineralized long bones were found in three patients (patients 1, 2, and 12), and in patient 12 the reduced mineralization was present already at birth (pictures not shown). Dysplastic distal phalanges were only found in patient 1 (pictures not shown). Congenital fractures located at the right humerus, the right femoral diaphyses, and along the midshaft of the left humerus and also subtle contour abnormality at the anterior ends of the left seventh and eighth ribs were only found in one patient (patient 12), which is a feature not described previously (pictures not shown). So far craniosynostosis has only been described in two patients by Kvarnung et al.21—only a metopic and a sagittal ridge were found in one of our patients (patient 8) and did not require surgical intervention.
One of the affected sib pairs had abnormal dentition (patients 1 and 2), similar to the patients described by Kvarnung et al.21 This finding could be unrelated to PIGT, could reflect a genotype–phenotype effect, or could arise from differences in genetic background. Five patients (patients 1–2, 4–5, and 11) also exhibited significant joint hypermobility, which was also reported by Lam et al.35
Eleven of the published PIGT deficient patients had neuroimaging (MRI) performed and in all patients cortical and cerebellar atrophy was evident. Kvarnung et al.21 presented two patients in whom MRI at the age of 2.7 years demonstrated global atrophy with predominant vermis and cerebellar atrophy.
The patient described by Nakashima et al.34 demonstrated progressive atrophy of the cerebral hemispheres, cerebellum, and brainstem at the age of three years. The two brothers published by Skauli et al.26 were examined respectively at 14 months and 2.8 years of age. Both demonstrated cortical atrophy and cerebellar atrophy, primarily affecting the vermis. Lam et al.35 presented MRI observations supporting that atrophy in the cerebellum starts earlier and proceeds more rapidly than atrophy elsewhere in the brain. Finally, Yang et al.29 described in their patient cortical hypoplasia and cerebellar vermis dysplasia. So far published data suggest that atrophy in patients with pathogenic PIGT variants preferentially affects the cerebellum. In contrast to previous reports, eight of our patients (patients 1–2, 5–7, 9, 12, and 13) showed no signs of cerebellar atrophy (Fig. 4, pictures are only available for patient 1 and 2). Patients 2, 5, 12, and 13 had the MRI done at a very early age, which could perhaps explain why the cerebellar atrophy was not yet evident, however patient 1 had an MRI done at three days (Fig. 4a, b), three months (Fig. 4c, d), nine months (Fig. 4e, f), and four years (Fig. 4g, h) of age. Based on the previously published MRI findings in PIGT patients we would have expected a cerebellar hypoplasia to be detectable at four years of age. Although cortical atrophy was evident, we detected no signs of cerebellar atrophy in this patient. This further expands the clinical spectrum of PIGT-CDG.
Magnetic resonance image (MRI) of patient 1 and patient 2 showing sagittal T1W and axial T1W. MRI exams of patient 1 were performed at three days (a, b), three months (c, d), nine months (e, f) and four years of age (g, h). MRI of patient 2 was performed only once at 13 days of age (i, j). MRI at three days (a, b) shows delayed myelination, normal sulcations, and no atrophy; at three months (c,d) atrophy of the splenium of the corpus callosum, supratentorial atrophy of white matter, delayed myelination, including unmyelinated anterior limb of the internal capsule (ALIC) and only posterior limb internal capsule (PLIC) fully myelinated; at nine months (e, f) somewhat progressed myelination, still delayed, resembling a three-month-old on T1W with a little, but still insufficient growth of corpus callosum and supratentorial atrophy of white matter; at four years (g, h) clearly progressed myelination, but still not completed, supratentorial atrophy of white matter and corpus callosum. No atrophy of the cerebellum. Magnetic resonance spectroscopy (TE:135 ms) at 3T performed in patient 1, nine months old, from right parietal white matter and bioccipital gray matter showed near-normal N-acetylaspartate/choline, choline/creatine ratios, no lactate or other unusual metabolites. T1W sagittal (i) and axial (j) brain MRI exam of patient 2 taken at 13 days of age revealed global delayed myelination, including insufficient myelinated PLIC (shown), normal sulcation, and no atrophy.
Congenital heart defects were identified in only two individuals and both resolved spontaneously: a persistent foramen ovale in one (patient 8) and a combined persistent foramen ovale and an atrial septum aneurysm in another (patient 6).
The majority of the published patients had onset of febrile-induced seizures between four and six months of age followed by unprovoked and poorly controlled seizures. In the report by Kvarnung et al.21 the age at onset of seizures was not specified. We now know that the onset of febrile-induced seizures in their patients was around 12–18 months of age (personal communication). In our cohort the onset of epileptic seizures did also include the neonatal period, leading us to conclude that PIGT-CDG should be considered in patients with hypotonia, severe global developmental delay, and neonatal seizures. EEG recordings were severely abnormal in patients with neonatal onset epilepsy but might also be normal or almost normal in the other patients early in life. At follow up, the most common EEG features were the slowing of the background activity, often associated with focal slowing and multifocal epileptiform abnormalities, predominant in the frontotemporal regions. The severity of the EEG abnormalities seemed to correlate with the severity of the phenotypes.
Our novel cases shared similar facial features with previous patients leading us to conclude that there is a common facial gestalt that includes a high forehead with bitemporal narrowing, a depressed nasal bridge, a short anteverted nose, a long philtrum with closely placed philtral pillars, distinct and prominant philtrum, full cheeks, and an open mouth consistent with general hypotonia. Scalp hair, eyebrows, and eyelashes were sparse in 8 of 13 patients (patient 1, 2, 3–7, and 11). After reviewing the pictures of the previously described patients, we believe that the patients V-1, V-2, and V-4 described by Kvarnung et al.21 and the siblings reported by Lam et al.35 all show signs of sparse scalp hair including a high and thin anterior hairline. Abnormal distribution of hair on the body has not been described as a feature of any of the GPI anchor disorders36 and the hypotrichosis as a clinical feature has not been previously described in patients with PIGT-CDGs. We furthermore identified hypertrichosis in patient 1, which is a possible novel feature in this disease. Therefore, abnormal body hair distribution including hypotrichosis might be a recurrent but overlooked feature of PIGT-CDGs. The automated analysis of the facial gestalt confirmed the presence of a characteristic gestalt in published and novel PIGT cases and a high similarity to other patients from the MCAHS phenotypic series. By using the suggested syndromes from Face2Gene to represent a patient with a characteristic facial gestalt in a high-dimensional GPIBD gestalt space, we were able to show a hierarchical delineation between HPMRS and MCAHS. The GPIBD gestalt space may be further expanded by addition of novel patients into existing cohorts, as well as addition of novel patient cohorts. Thereby, formation of additional clusters may lead to delineation of novel disease entities. Evaluation of confounding factors (such as sex, age, and ethnicity) demonstrates the robustness of the applied methods. Thus, the combination of deep phenotyping, automated facial image analysis, and NGS is sufficient for correct molecular diagnosis.
There are no current FDA-approved therapies for GPI anchor disorders, but the progressive nature of PIGT-CDG is attractive for therapies that slow or halt the neurologic deterioration.36 Identification of additional PIGT-CDG patients should further define the clinical spectrum and assist in developing diagnostic criteria.
Data availability
Anonymized data will be shared by request with any qualified investigator for the sole purpose of replicating procedures and results presented in the article and as long as data transfer is in agreement with EU legislation on the general data protection regulation.
References
Kinoshita T, Fujita M, Maeda Y. Biosynthesis, remodelling and functions of mammalian GPI-anchored proteins: recent progress. J Biochem. 2008;144:287–294.
Fujita M, Kinoshita T. GPI-anchor remodeling: potential functions of GPI-anchors in intracellular trafficking and membrane dynamics. Biochim Biophys Acta. 2012;1821:1050–1058.
Kinoshita T, Fujita M. Biosynthesis of GPI-anchored proteins: special emphasis on GPI lipid remodeling. J Lipid Res. 2016;57:6–24.
Park S, Lee C, Sabharwal P, et al. GDE2 promotes neurogenesis by glycosylphosphatidylinositol-anchor cleavage of RECK. Science. 2013;339:324–328.
Nozaki M, Ohishi K, Yamada N, et al. Developmental abnormalities of glycosylphosphatidylinositol-anchor-deficient embryos revealed by Cre/loxP system. Lab Invest. 1999;79:293–299.
McKean DM, Niswander L. Defects in GPI biosynthesis perturb Cripto signaling during forebrain development in two new mouse models of holoprosencephaly. Biol Open. 2012;1:874–883.
Bessler M, Mason PJ, Hillmen P, et al. Paroxysmal nocturnal haemoglobinuria (PNH) is caused by somatic mutations in the PIG-A gene. EMBO J. 1994;13:110–117.
Almeida AM, Murakami Y, Layton DM, et al. Hypomorphic promoter mutation in PIGM causes inherited glycosylphosphatidylinositol deficiency. Nat Med. 2006;12:846–851.
Krawitz PM, Schweiger MR, Rödelsperger C, et al. Identity-by-descent filtering of exome sequence data identifies PIGV mutations in hyperphosphatasia mental retardation syndrome. Nat Genet. 2010;42:827–829.
Murakami Y, Tawamie H, Maeda Y, et al. Null mutation in PGAP1 impairing Gpi-anchor maturation in patients with intellectual disability and encephalopathy. PLoS Genet. 2014;10:e1004320.
Martin HC, Kim GE, Pagnamenta AT, et al. Clinical whole-genome sequencing in severe early-onset epilepsy reveals new genes and improves molecular diagnosis. Hum Mol Genet. 2014;23:3200–3211.
Ilkovski B, Pagnamenta AT, O’Grady GL, et al. Mutations in PIGY: expanding the phenotype of inherited glycosylphosphatidylinositol deficiencies. Hum Mol Genet. 2015;24:6146–6159.
Makrythanasis P, Kato M, Zaki MS, et al. Pathogenic variants in PIGG cause intellectual disability with seizures and hypotonia. Am J Hum Genet. 2015;98:1–12.
Johnstone DL, Nguyen T-T-M, Murakami Y, et al. Compound heterozygous mutations in the gene PIGP are associated with early infantile epileptic encephalopathy. Hum Mol Genet. 2017;26:1706–1715.
Edvardson S, Murakami Y, Nguyen TTM, et al. Mutations in the phosphatidylinositol glycan C (PIGC) gene are associated with epilepsy and intellectual disability. J Med Genet. 2017;54:196–201.
Maydan G, Noyman I, Har-Zahav A, et al. Multiple congenital anomalies-hypotonia-seizures syndrome is caused by a mutation in PIGN. J Med Genet. 2011;48:383–389.
Ng BG, Hackmann K, Jones MA, et al. Mutations in the glycosylphosphatidylinositol gene PIGL cause CHIME syndrome. Am J Hum Genet. 2012;90:685–688.
Krawitz PM, Murakami Y, Hecht J, et al. Mutations in PIGO, a member of the GPI-anchor-synthesis pathway, cause hyperphosphatasia with mental retardation. Am J Hum Genet. 2012;91:146–151.
Johnston JJJ, Gropman ALL, Sapp JCC, et al. The phenotype of a germline mutation in PIGA: the gene somatically mutated in paroxysmal nocturnal hemoglobinuria. Am J Hum Genet. 2012;10:295–300.
Hansen L, Tawamie H, Murakami Y, et al. Hypomorphic mutations in PGAP2, encoding a GPI-anchor-remodeling protein, cause autosomal-recessive intellectual disability. Am J Hum Genet. 2013;92:575–583.
Kvarnung M, Nilsson D, Lindstrand A, et al. A novel intellectual disability syndrome caused by GPI anchor deficiency due to homozygous mutations in PIGT. J Med Genet. 2013;50:521–528.
Chiyonobu T, Inoue N, Morimoto M, et al. Glycosylphosphatidylinositol (GPI) anchor deficiency caused by mutations in PIGW is associated with West syndrome and hyperphosphatasia with mental retardation syndrome. J Med Genet. 2014;51:203–207.
Howard MF, Murakami Y, Pagnamenta AT, et al. Mutations in PGAP3 impair GPI-anchor maturation, causing a subtype of hyperphosphatasia with mental retardation. Am J Hum Genet. 2014;94:278–287.
Ohishi K, Inoue N, Kinoshita T. PIG-S and PIG-T, essential for GPI anchor attachment to proteins, form a complex with GAA1 and GPI8. EMBO J. 2001;20:4088–4098.
Ohishi K, Nagamune K, Maeda Y, et al. Two subunits of glycosylphosphatidylinositol transamidase, GPI8 and PIG-T, form a functionally important intermolecular disulfide bridge. J Biol Chem. 2003;278:13959–13967.
Skauli N, Wallace S, Chiang S, et al. Novel PIGT variant in two brothers: expansion of the multiple congenital anomalies-hypotonia seizures syndrome 3 phenotype. Genes (Basel). 2016;7:108.
Pagnamenta AT, Murakami Y, Taylor JM, et al. Analysis of exome data for 4293 trios suggests GPI-anchor biogenesis defects are a rare cause of developmental disorders. Eur J Hum Genet. 2017;22:1–11.
Kohashi K, Ishiyama A, Yuasa S, et al. Epileptic apnea in a patient with inherited glycosylphosphatidylinositol anchor deficiency and pigt mutations. Brain Dev. 2017;40:53–57.
Yang L, Peng J, Yin X-M, et al. Homozygous PIGT mutation lead to multiple congenital anomalies-hypotonia seizures syndrome 3. Front Genet. 2018;9:153.
Gurovich Y, Hanani Y, Bar O, et al. DeepGestalt - Identifying Rare Genetic Syndromes Using Deep Learning. Nat Med. 2019;25:60–64.
Knaus A, Pantel JT, Pendziwiat M, et al. Characterization of glycosylphosphatidylinositol biosynthesis defects by clinical features, flow cytometry, and automated image analysis. Genome Med. 2018;10:3.
Pagnamenta AT, Murakami Y, Taylor JM, et al. Analysis of exome data for 4293 trios suggests GPI-anchor biogenesis defects are a rare cause of developmental disorders. Eur J Hum Genet. 2017;25:669–679.
Firth HV, Wright CF. The Deciphering Developmental Disorders (DDD) study. Dev Med Child Neurol. 2011;53:702–703.
Nakashima M, Kashii H, Murakami Y, et al. Novel compound heterozygous PIGT mutations caused multiple congenital anomalies-hypotonia-seizures syndrome 3. Neurogenetics. 2014;15:193–200.
Lam C, Golas GA, Davids M, et al. Expanding the clinical and molecular characteristics of PIGT-CDG, a disorder of glycosylphosphatidylinositol anchors. Mol Genet Metab. 2015;115:128–140.
Ng BG, Freeze HH. Human genetic disorders involving glycosylphosphatidylinositol (GPI) anchors and glycosphingolipids (GSL). J Inherit Metab Dis. 2015;38:171–178.
Acknowledgements
We thank the families for participating in this study. We thank Malin Kvarnung for kindly sharing previously unpublished data regarding the onset of epileptic seizures in their patients. I.H. was supported by intramural funds of the University of Kiel, and by a grant from the German Research Foundation (DFG) (HE5415/6-1). Y.W. was supported by a grant from the German Research Foundation (DFG) (WE4896/3-1 and WE4896/4-1). A.C. was sponsored by Polish National Science Centre grant number 2014/15/D/NZ5/03426. The DDD Study presents independent research commissioned by the Health Innovation Challenge Fund (HICF-1009-003), a parallel funding partnership between the Wellcome Trust and the Department of Health, and the Wellcome Trust Sanger Institute (WT098051). The research team acknowledges the support of the National Institute for Health Research, through the Comprehensive Clinical Research Network. The views expressed in this publication are those of the authors and not necessarily those of the Wellcome Trust or the Department of Health.
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Shared first authors: Allan Bayat, Alexej Knaus, Annika Wollenberg Juul.
Rights and permissions
About this article
Cite this article
Bayat, A., Knaus, A., Juul, A.W. et al. PIGT-CDG, a disorder of the glycosylphosphatidylinositol anchor: description of 13 novel patients and expansion of the clinical characteristics. Genet Med 21, 2216–2223 (2019). https://doi.org/10.1038/s41436-019-0512-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0512-3
Keywords
This article is cited by
-
Epidemiology of congenital disorders of glycosylation (CDG)—overview and perspectives
Journal of Rare Diseases (2022)
-
Analyzing clinical and genetic characteristics of a cohort with multiple congenital anomalies-hypotonia-seizures syndrome (MCAHS)
Orphanet Journal of Rare Diseases (2020)
-
EEG Monitoring of the Epileptic Newborn
Current Neurology and Neuroscience Reports (2020)