Abstract
Purpose
To characterize new molecular factors implicated in a hereditary congenital facial paresis (HCFP) family and otosclerosis.
Methods
We performed exome sequencing in a four-generation family presenting nonprogressive HCFP and mixed hearing loss (HL). MEPE was analyzed using either Sanger sequencing or molecular inversion probes combined with massive parallel sequencing in 89 otosclerosis families, 1604 unrelated affected subjects, and 1538 unscreened controls.
Results
Exome sequencing in the HCFP family led to the identification of a rare segregating heterozygous frameshift variant p.(Gln425Lysfs*38) in MEPE. As the HL phenotype in this family resembled otosclerosis, we performed variant burden and variance components analyses in a large otosclerosis cohort and demonstrated that nonsense and frameshift MEPE variants were significantly enriched in affected subjects (p = 0.0006–0.0060).
Conclusion
MEPE exerts its function in bone homeostasis by two domains, an RGD and an acidic serine aspartate-rich MEPE-associated (ASARM) motif inhibiting respectively bone resorption and mineralization. All variants associated with otosclerosis are predicted to result in nonsense mediated decay or an ASARM-and-RGD-truncated MEPE. The HCFP variant is predicted to produce an ASARM-truncated MEPE with an intact RGD motif. This difference in effect on the protein corresponds with the presumed pathophysiology of both diseases, and provides a plausible molecular explanation for the distinct phenotypic outcome.
Similar content being viewed by others
Introduction
Craniofacial abnormalities, characterized by abnormal growth and development of the head and facial bones, can be distinguished from other skeletal disorders because the mechanism of bone formation is notably different. Craniofacial bones are mainly formed through intramembranous ossification, whereas the development of the body skeleton requires endochondral ossification. Affected individuals with craniofacial bone development/growth disorders often have hearing loss (HL) due to bone abnormalities of the outer, middle, and/or inner ear.1
Hereditary congenital facial paresis (HCFP [MIM 601471]), a rare disorder belonging to the group of craniofacial abnormalities, is caused by uni- or bilateral maldevelopment of the VIIth cranial nerve (facialis) and its nucleus. Affected individuals have an inappropriate innervation of the facial muscles that leads to facial weakness or paresis and abnormal facial expression. Two loci have been identified by linkage analysis, HCFP1 on chromosome 3q21.2-q22.1 (ref. 2,3) (MIM 601471) and HCFP2 on chromosome 10q21.3-q22.1 (ref. 4) (MIM 604185), with no causative genes described yet. The only reported causative variants for HCFP (HCFP3 [MIM 614744]) are homozygous loss-of-function or missense variants altering the homeobox domain in HOXB1 (MIM 142968), which have been identified in a total of 12 HCFP subjects,5,6,7,8 and a single de novo nonsense variant in the REV3L gene (MIM 602776) (ref. 9).
Otosclerosis (MIM 166800) is a bone remodeling disorder of the human otic capsule,10 with a prevalence of 0.3% to 0.4% in the white population, and usually has an onset within the third decade of life.11 The otic capsule is a unique bony structure in the human body, because it retains small regions of immature cartilage called globuli interossei, and because it exhibits limited bone remodeling compared with other bones.10 Abnormal bone remodeling of the otic capsule is seen in otosclerosis, and can involve the stapes footplate, leading to conductive HL by reducing the motion of the stapes.12 Several studies suggest that otosclerosis is a complex disorder caused by a combination of several elements, such as environmental, immune, and endocrine factors; viral infections; and genetic factors.12,13 Major genetic risk factors for sporadic otosclerosis have been identified through association studies.13 Otosclerosis can also be present in a familial form, but these families are rare and often show autosomal dominant inheritance with reduced penetrance. The reduced penetrance has been estimated to be on average 40%, although this can vary greatly between families.1 Studies in families with autosomal dominant otosclerosis have led to the identification of eight loci,12 although the responsible genes in these loci have not yet been identified. Recently, genetic variation in SERPINF1 (MIM 172860) has been reported to be associated with familial otosclerosis.14
In this study, we investigated a Turkish family with nonprogressive HCFP and mixed HL due to diploic thickening and sclerosis of the skull. Exome sequencing revealed a single heterozygous frameshift variant in MEPE (MIM 605912) that cosegregates with the phenotype. Furthermore, we hypothesized that MEPE loss-of-function variants might be implicated in otosclerosis. Under this hypothesis, a large-scale MEPE screening of unrelated otosclerosis families, sporadic affected otosclerosis individuals and ethnically matched unscreened controls was performed, showing a significant enrichment of heterozygous loss-of-function MEPE variants in affected subjects.
Material and methods
Subjects
DNA samples of individuals from unrelated families with a HCFP diagnosis of Dutch, Spanish, Turkish, and Norwegian origin were collected by the Department of Human Genetics of the Radboud University Medical Center, Nijmegen (The Netherlands). In addition, 89 families with at least two affected sibs with otosclerosis from the Center of Medical Genetics (Antwerp, Belgium) and the Department of Human Genetics (Nijmegen, The Netherlands) were studied as well, including families of Belgian, Dutch, Finnish, Tunisian, French, Italian, Lebanese, Romanian, and Greek ethnicity. Finally, 1604 unrelated individuals with otosclerosis and ethnically matched control individuals of unknown affection status for otosclerosis (N = 1538) of Belgian, Dutch, French, Italian, Swiss, and Romanian origin were recruited by Departments of Otolaryngology in several hospitals: University Hospital of Antwerp (Belgium); the Center of Medical Genetics of the University Hospital of Antwerp (Belgium); GZA Hospital campus Sint-Augustinus (Antwerp, Belgium); University Hospital of Ghent (Belgium); the Departments of Otorhinolaryngology of the University Hospital in Zurich (Switzerland); and the Radboud University Medical Center (Nijmegen, The Netherlands); the Department of Clinical Sciences and Community Health, University of Milan (Italy); the Jean Causse Ear Clinic (Colombiers, France); and the ENT Department of Iuliu Hatieganu University of Medicine and Pharmacy (Cluj-Napoca, Romania). Subject evaluation and diagnostic criteria are described in the Supplemental Data (Supplemental Material and Methods).
Informed consent was obtained from all involved subjects and the local ethics committees, in accordance with the ethics of the World Medical Association Declaration of Helsinki, approved all procedures. Genomic DNA was isolated from either fresh or frozen peripheral blood using standard techniques. Informed consent for publication of identifiable photos from individuals from the Turkish HCFP family was received.
Exome sequencing and data analysis
Exome sequencing was performed using the SureSelect Human All Exon 50 Mb Kit (Agilent, Santa Clara, CA, USA) followed by multiplexed analysis on a SOLiD 4 System sequencing slide (Life Technologies, Carlsbad, CA, USA) as reported by Tomas-Roca et al.9 Color space reads were mapped to the hg19 reference genome with the SOLiD BioScope version 1.3 software, which utilizes an iterative mapping approach. Single-nucleotide variants were subsequently called by the Di–Bayes algorithm with high call stringency. Variant filtering was done in steps described in Supplementary Table S4. In short, only variants shared by subjects II:3 and IV:2 of the HCFP family were selected, and indels (frameshift and in-frame), nonsynonymous, nonsense, start/stop altering, and splice-site variants with a minor allele frequency <0.0005 in various databases (including gnomAD) were prioritized. Confirmation and cosegregation analysis of 21 candidate variants was carried out by Sanger sequencing.
MEPE screening in additional familial HCFP and otosclerosis subjects
HCFP-affected individuals of nine families and otosclerosis-affected individuals of 27 families were checked for variants in the complete coding region of MEPE using Sanger sequencing. Primers surrounding the exons of MEPE were designed using Primer3 (ref. 15). Polymerase chain reaction (PCR) amplification was carried out under standard conditions. Direct sequencing of PCR products was performed on an ABI3130XL sequencer (Applied Biosystems Inc., Foster City, CA, USA) or ABI PRISM 3730 DNA Analyzer (Applera Corp, Foster City, CA, USA). Analysis was performed using the Vector NTI® software (Life Technologies) or CodonCode Aligner (CodonCode Corporation, Centerville, MA, USA).
MEPE screening in a large set of individuals affected with otosclerosis and controls
Single-molecule molecular inversion probes (smMIPs) were designed for the complete coding regions, 5′ and 3′ untranslated regions (UTRs), and at least 10 bp overhang over the exon–intron boundaries of MEPE using MIPgen.16,17 MEPE was enriched using smMIPs (Integrated DNA Technologies, Coralville, IA, USA) in otosclerosis-affected individuals of 62 families, as well as in 1604 sporadic affected individuals and 1538 controls. The smMIP-enrichment was performed according to the protocol described elsewhere16,18,19 with some minor modifications (Supplemental Material and Methods). Sequencing of the smMIPs library was performed on the NextSeq 500 (Illumina, San Diego, CA, USA). Data was analyzed using an in-house developed analysis pipeline. Quality control parameters were established based on a subset of 111 Sanger sequenced variants to reduce the number of false positive variant calls and to exclude bad-performing and poorly covered samples (Supplemental Material and Methods).
For the case–control population, the case group consisted of unrelated otosclerosis subjects and one index individual per family. All frameshift and nonsense variants in MEPE in these groups were validated using Sanger sequencing. Statistical analyses of the identified variants was performed on a variant level using the Fisher’s exact test, and on gene level using the three implementations of variant burden tests (combined and multivariate collapsing test [CMC], kernel-based adaptive cluster [KBAC] test, and variable thresholds methods tests (VT]) and two variance components tests (SNP-set [sequence] kernel association test method and cAlpha test) using variant tools (vtools) and variant association tools (VAT).20,21
Three-dimensional protein modeling
The amino acid sequence of MEPE was retrieved from the UniProt database (identifier: Q9NQ76–1). Isoform A was chosen as it is the primary isoform expressed in many tissues (Genotype-Tissue Expression database; GTEx), and the only transcript expressed in a human bone tissue cell line (U2OS; Gene Expression Omnibus database [GEO] GSM2563988). The three-dimensional structure of wild-type (WT) and mutant MEPEs were predicted using the I-TASSER server.22 The predicted protein models were visualized using University of California–San Francisco (UCSF) Chimera.23
Results
Exome sequencing reveals frameshift variants in MEPE in a HCFP family
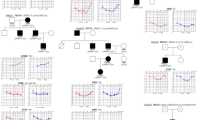
We first studied a four-generation Turkish HCFP family (Fig. 1a, b; Figure S2). In this family there are eight affected individuals diagnosed with HCFP. All had congenital, bilateral, nonprogressive facial paresis. The severity of the facial paresis in the patients was graded between 25 and 50 (Sunnybrook Facial Grading System; normal function = 100). Four of the affected individuals are also diagnosed with mixed HL (III:7, III:15, IV:1, and IV:2). According to family history and photographs, the facial paresis and HL manifested shortly after birth and did not progress over the years. Clinical monitoring in the period 2009–2015, including longitudinal audiometric analysis of one affected individual (IV.2), confirmed the nonprogressive nature of the clinical features.
Clinical features of hereditary congenital facial paresis (HCFP) affected individuals and pedigree of families in whom MEPE variants were identified DNA from individuals with underlined numbers. a Pedigree of the HCFP family showing that variant M1 [c.1273delC; p.(Gln425Lysfs*38)] fully segregates with the facial paresis phenotype. The affected status of individual III:16 is based on reporting by family members. b Clinical features of the affected individuals carrying the variants in MEPE. Affected individuals III:7, III:15, IV:1, and IV:2 show bilateral facial paresis; they are not able to smile. c Pedigrees of the families with otosclerosis (OTOS1 and OTOS2) in which the variant M2 [c.199_202delGAAA; p.(Lys70Ilefs*26)] was identified.
Individual III:16 was considered to have HL, based on subjective information provided by her family members. In audiometry, part of the hearing-impaired subjects displayed a Carhart notch at 2000 Hz and absence of stapedial reflexes (Figure S1, Table S1), suggestive for fixation of the ossicular chain. Although clinical symptoms with regard to hearing resembled otosclerosis, high-resolution computed tomography (HRCT) showed no radiological signs of otosclerosis. All affected subjects showed thickening of the skull, mainly at the base, due to an increased amount of diploic bone with varying degrees of sclerosis (Fig. 2 and Table S1). The bone expansion and appositional bone growth at the walls of the middle ear caused severe narrowing of the middle ear cavity with entrapment of the ossicles. The stapes appeared thickened and sclerotic when visible. In cases with extensive bone formation, the round window was also obliterated. These findings are compatible with the clinical presentation of conductive HL. All affected subjects showed narrowing of the internal auditory canal to some degree, potentially leading to sensorineural HL. The facial nerve canal was also narrowed. There was no or minimal pneumatization of the mastoid. The skull base foraminae were of normal size, indicating that other cranial nerves were not entrapped (Fig. 2). There were no longitudinal HRCT imaging data to assess progression of the bone abnormalities. The presence of extracranial bone abnormalities could not be assessed, because extracranial images were not available. The symptoms and clinical evaluations were consistent with the diagnosis of HCFP and mixed HL, as a result of diploic thickening and sclerosis of the skull.
a Axial skull base CT of individual III:15. The size of the external auditory canal is within normal limits, but due to bone apposition along the promontory (white arrow) and the epitympanic walls. (b/c) Detailed images of the right temporal bone show an increased amount and density of bone marrow in the thickened skull. The tympanic cavity is narrowed with entrapment of ossicles (b: black arrow). The stapes is thickened and sclerotic (b: black arrowhead). The internal auditory canal and facial canal are narrowed (c: white arrowheads). (d) Schematic overview of the three MEPE protein isoforms. The positions of the identified variants and their effects on the different protein isoforms are indicated. The frameshift variant identified in the HCFP family is highlighted in pink, the variants found enriched in otosclerosis patients are presented in black. For a schematic overview of the exon composition and variants see Figure S3. e Three-dimensional structures of wild-type (WT) MEPE (left) and the HCFP variant [p.(Gln425Lysfs*38)] (MT1; middle), and the superposition of WT and MT1. The conformational shape of MT1, including the mutant sequence, mimics WT, only missing its final ASARM motif (see also Figure S4). Isoform A osteoregulin domain: AA 97–256 (blue); RGD motif: AA 247–249 (yellow); ASARM motif: AA 507–525 (green); mutant sequence added due to MT1: AA 425–461 (pink). HL hearing loss, Mmutation, NMD nonsense mediated decay. (colour figs online only)
Exome sequencing was performed in two affected subjects of this family (II:3 and IV:2) (Figs. 1a, b and Table S1). Seventy-two percent of bases mapped on target, resulting in an average target coverage of 66.93× fold and 85% of the exons were covered at least 10 times (Table S3). Data analysis and variant filtering resulted in a list of 26 variants (Table S4). Confirmation and cosegregation analysis of 21 candidate variants in the rest of the family revealed only one variant cosegregating with the disease phenotype, namely a heterozygous c.1273delC (M1) variant in MEPE (Figure 1 and Table 1) located at chromosome 4q22.1. This heterozygous deletion creates a frameshift starting at codon Gln425 [p.(Gln425Lysfs*38)]. This variant is present in all HCFP patients available for testing (7/8), and absent in nonaffected individuals from the Turkish family. The variant was not present in the Exome Aggregation Consortium (ExAC) nor in the Genome Aggregation Database (GnomAD).24
MEPE variant screening using Sanger sequencing in familial otosclerosis and HCFP
The clinical symptoms of the Turkish HCFP family suggest a role of MEPE in facial paresis and mixed HL. Moreover, the nonradiological symptoms of the Turkish HCFP family resembled otosclerosis. Therefore, we screened 9 HCFP families and 27 families with otosclerosis for MEPE variants using Sanger sequencing (Figure S2). No rare variants were identified in the HCFP families. However, Sanger sequencing revealed a heterozygous frameshift variant (M2: c.199_202delGAAA [p.(Lys70Ilefs*26)]) (Fig. 2 and Table 1) in one otosclerosis family (OTOS1) (Fig. 1c). The variant was identified in three affected members of the family (II:1, III:11, and III:12) and seven unaffected members, possibly reflecting reduced penetrance of the otosclerosis phenotype.
smMIP-based MEPE variant screening in a large group of otosclerosis subjects and controls
To establish the contribution of variants in MEPE to the otosclerosis phenotype, a statistical case–control study was performed, using a targeted smMIP-based gene sequencing in a large population of otosclerosis families, cases, and controls.
Establishment of the sample quality control (QC) parameters on the data of the analyzed samples resulted in the inclusion of a total of 123 affected individuals in 62 families, as well as 1604 cases and 1538 controls. For these samples, on average between 94.9% and 97.9% of the target region reached at least 10× coverage. Furthermore, the quality-based filtering using the established cut-off values and parameters resulted in a validated set of variants with a sensitivity of 0.984 and specificity of 0.994 (Supplemental Material and Methods).
Sanger validation of the frameshift and nonsense variants was performed resulting in an overall variant detection in one familial affected subject (Fig. 1c and Table 1) and 18 unrelated affected subjects (Table 2 and Table S2) as well as three controls.
In the variant burden and variance components tests, nonsense and frameshift variants with MAF below 1% and 0.1% (calculated in the included controls) reached significance (p value below 0.006) in all variant burden and variance components tests (MAF < 0.01; CMC = 0.0010; KBAC = 0.0006; VT = 0.0006; SKAT = 0.0060; cAlpha = 0.0034; Table S5). The following loss-of-function variants were identified (Table 2 and Figure S3): (1) c.199_202delGAAA, p.(Lys70Ilefs*26) (M2) in one familial proband, nine unrelated affected individuals and one control; (2) c.49_54+17delinsCA, p.(Ala17Glnfs*37) (M3) in one affected subject; (3) M4: c.184G>T, p.(Glu62*) (M4) in one affected individual; (4) c.496dup, p.(Glu166Glyfs*9) (M5) in one affected subject; (5) c.617delG, p.(Ser206Ilefs*3) (M6) in six affected individuals and one control; and (6) c.679dup, p.(Ser227Phefs*9) (M7) in one control (complementary DNA [cDNA] positions based on NM_00184694). Variant burden and variance components tests of missense variants did not reveal significant results (Table S5).
Among these 19 affected subjects with a frameshift and nonsense variant, 1 was a proband of the OTOS2 family (Fig. 1). In this affected individual, the same variant was identified that was also present in family OTOS1, namely M2:c.199_202delGAAA [p.(Lys70Ilefs*26)]. This second family includes two male affected siblings and in the affected brother of the proband, the same frameshift variant was present in a heterozygous state (OTOS2) (Fig. 1c and Table 1). Both affected family members had bilateral, progressive mixed HL (Figure S1) and underwent bilateral stapedotomy, confirming the diagnosis of otosclerosis. HRCT imaging in subject II:1 demonstrated bilateral fenestral otosclerosis and a normal appearance of the skull. In none of the other families was a potential causative variant identified.
Three-dimensional protein modeling
Ab initio modeling techniques were utilized to construct the three-dimensional structures of WT and mutant MEPE proteins. Wild-type MEPE isoform A consists of 525 amino acids (AA) and includes an osteoregulin domain (blue) with RDG motif (yellow) and acidic serine aspartate-rich MEPE-associated (ASARM) motif (green) (Fig. 2d). We modeled HCFP mutant MEPE [MT1: p.(Gln425Lysfs*38)] in addition to all frameshift and nonsense variants present in gnomAD in >1 individual with intact osteoregulin domain and RGD motif, but missing the C-terminal ASARM motif (3 variants, 8 individuals; Fig. 2e, Figure S4). We found that MT1 (461 AA) is much longer than the three gnomAD frameshift variants of MEPE (303–410 AA). The superposition of WT and HCFP mutant MEPE (MT1) reveals a similar overall conformation, with MT1 only missing the final ASARM domain (green). In contrast, the conformational shape of the three variants observed in gnomAD is different from WT and MT1 (Figure S4).
Discussion
MEPE encodes a matrix extracellular phosphoglycoprotein that belongs to the family of small integrin-binding ligand proteins, N-linked glycoproteins (SIBLINGs).25 This protein family is characterized by an ASARM motif.26,27 After proteolytic cleavage, the ASARM peptide inhibits mineralization by binding to hydroxyapatite crystals present in bone.26 The ASARM motif of MEPE is expressed in osteocytes, osteoblasts, odontoblasts, renal proximal tubules, and salivary glands.28,29,30,31 Furthermore, MEPE has been shown to play a key role in the inhibition of bone mineralization and resorption, renal calcification suppression, and serum phosphate regulation. In addition, in vivo studies demonstrated that both homozygous (Mepe-/-) and heterozygous (Mepe+/-) knockout mice exhibit an increase of bone mass production and trabecular density due to a higher number of osteoblasts and their increased activity.32 These phenotypes become more severe with age in mice33 and they mainly affect the craniofacial bones, whereas the gross body skeletal structure remains unaffected. Conversely, administration of human MEPE in vitro and in vivo give rise to a mineralization defect and altered phosphate homeostasis.34 Besides the ASARM motif, MEPE contains another functional domain, namely the RGD motif, which plays an important role in bone homeostasis and osteogenesis.35
In this study, we used exome sequencing and targeted massive parallel sequencing to show that rare frameshift and nonsense variations in the MEPE gene are associated with familial HCFP and familial otosclerosis, and are significantly enriched in unrelated isolated otosclerosis subjects. As mentioned previously, two important functional domains in MEPE have been identified, an ASARM motif and an osteoregulin domain containing an RGD motif (Fig. 2d),35 both required in bone homeostasis (Fig. 3a).26,35,36 The rare variant identified in the HCFP family, M1: c.1273delC [p.(Gln425Lysfs*38)], is predicted to escape nonsense mediated decay (NMD)37 and predicted to result in an ASARM-truncated MEPE protein, still containing the RGD motif (Fig. 3b). In addition, p.(Gln425Lysfs*38) exhibits a similar three-dimensional shape to WT MEPE, only missing the ASARM motif (Fig. 2e; Figure S4). On the contrary, one otosclerosis-associated variant is expected to result in NMD (variant M3), and all other otosclerosis-associated variants are expected to lead to a truncated protein lacking both the RGD motif and the ASARM motif (Fig. 2d and Fig. 3c). All otosclerosis MEPE variants affect isoform A (Fig. 2d), which is the only isoform expressed in a human bone tissue cell line (GEO GSM2563988), and the main isoform expressed in other human tissues as well (GTEx).
Hypothesis on the mode of action of physiological MEPE and the truncated MEPE proteins in hereditary congenital facial paresis (HCFP) and otosclerosis. a The MEPE protein is proteolytically cleaved in an ASARM peptide and a larger peptide containing the small RGD motif. In normal physiological conditions, the ASARM peptide binds to hydroxyapatite crystals, thereby inhibiting mineralization. Furthermore, the RGD motif in the secreted MEPE peptide serves as a decoy receptor for the binding of pre-osteoclasts, thereby inhibiting the differentiation of these pre-osteoclasts to mature osteoclasts and hence controlling bone homeostasis. b In the HCFP family, MEPE is truncated and the ASARM motif is no longer present, leading to an increased mineralization because of the loss of the negative feedback loop. The RGD motif is still intact maintaining its inhibitory effect on pre-osteoclast differentiation and involvement in the control of bone homeostasis. c The variants identified in the otosclerosis study population, result in a truncated MEPE protein without ASARM and RGD motif or even nonsense mediated decay (NMD). As such, both mineralization and differentiation of the pre-osteoclast to the mature osteoclast is no longer inhibited, leading to an increased bone remodeling. Figure not to scale. Blue peptide: osteoregulin domain; yellow peptide: RGD motif; green peptide: ASARM motif; orange triangle: RGD motif; purple Y-shaped symbol: integrins; red stars: hydroxyapatite crystals. WT wild-type. (colour figs online only)
Furthermore, based on various in vitro and in vivo experiments, including several assessing truncated constructs of MEPE,34,35,38,39 we were able to postulate the underlying molecular mechanisms leading to the distinct bone phenotypes in otosclerosis and HCFP due to the location of the different variants (Fig. 2d; Fig. 3). We hypothesize that the RGD motif in MEPE serves as a decoy receptor for the pre-osteoclasts in normal circumstances, thereby inhibiting differentiation of this pre-osteoclast to a mature osteoclast. This is supported by the in vitro observation that RGD motif containing peptides inhibit the formation of multinucleated osteoclasts and consequently bone resorption.38 MEPE thus plays a role in the control of bone turnover. This is also supported by the observations that in normal bone homeostasis, osteoclasts anchor to the RGD motif in mineralized bone through integrin molecules. The importance of the RGD motif is further highlighted by the observation that a synthetic peptide (AC-100) containing the RGD motif and glycosaminoglycan attachment (GAG) sites shows the same functional effect as the ASARM-truncated MEPE, namely leading to increased osteogenesis.35 Additionally, subcutaneous injections of AC-100 onto the calvariae in mice lead to increased bone thickness and new bone formation whereas synthetic MEPE fragments with disturbed RGD motif or GAG attachment site, did not increase osteogenesis in vivo.39 On the other hand, the ASARM motif of MEPE inhibits mineralization after the proteolytic release of a 2.2-kDa C-terminal ASARM peptide by β-cathepsin.26,36 This cleaved ASARM peptide inhibits mineralization by binding to hydroxyapatite crystals.26 As such, ASARM-truncated MEPE leads to a normal bone turnover and increased mineralization. Based on the in vitro observation of Sprowson et al. related to the ASARM-truncated MEPE,35 we hypothesize that the HCFP phenotype is caused by a combination of increased mineralization and osteogenesis, while the inhibition of the differentiation of pre-osteoclasts into osteoclasts is preserved (Fig. 3b).
In otosclerosis, the identified MEPE variants either lead to NMD, with likely no protein or limited protein expression, or an RGD-and-ASARM-truncated protein. Here, the function of MEPE as decoy receptor is reduced, leading to increased differentiation of pre-osteoclasts into mature osteoclasts and, subsequently, increased bone turnover, combined with increased mineralization (Fig. 3c). It is plausible that this may specifically affect the otic capsule as this structure is characterized by extremely low bone remodeling in physiological circumstances. Therefore, the otic capsule may not possess compensating backup mechanisms for this abnormal bone remodeling, leading to otosclerotic foci in the otic capsule. In contrast, alternative pathways may be present in other bone tissues, leading to their capability to compensate for the increased osteoclastic activity and mineralization as a result of the loss-of-function MEPE variant.
The identification of genetic variation in MEPE in the HCFP family adds a new dimension to the pathogenesis of HCFP. Variants in previously identified genes implicated in congenital facial nerve paresis in HCFP and Möbius syndrome affect various stages of facial nerve development. Among these genes, HOXB1 is critical for the formation of rhombomere 4 (r4) and its derivatives, REV3L seems to be important for the maintenance of neuronal precursors,9 and PLXND1 is required for the proper migration of motoneurons from r4 to the facial branchiomotor nucleus in r6 (ref. 9). Most likely, HCFP caused by variants in MEPE results from an entirely different underlying pathophysiological mechanism. We hypothesize that there is obstruction of the outgrowth of the facial nerve due to diploic thickening of the bony facial canal.
The presence of some of the nonsense and frameshift RGD-and-ASARM affecting variants in unscreened controls and in unaffected family members from otosclerosis patients is probably due to reduced penetrance, as this is a common observation in otosclerosis.1 This could be attributable to several reasons. First, otosclerosis has a varying age of onset, which is usually between 16 and 50 years, but development at later ages is also possible, therefore we cannot exclude the development of otosclerosis in the unaffected individuals in control databases or families. On the other hand, abnormal bone remodeling in the temporal bone might not always lead to stapes fixation and conductive hearing loss. In fact, 2.5% of the Caucasian population has a histological form of otosclerosis, which is the presence of otosclerotic foci at the otic capsule without any clinical symptoms.11 This hypothesis can unfortunately not be confirmed because the diagnosis of histologic otosclerosis can only be made based on postmortem pathological examination of the otic capsule. Last, environmental factors and/or modifier genes could also play a role in the clinical presentation of otosclerosis, such as viral factors,1 that could exacerbate bone remodeling and lead to a clinical presentation of the disease.
In conclusion, our study shows that rare variations in MEPE contribute to the etiology of HCFP and otosclerosis. Because these variants were identified in a subset of affected individuals and controls, we think these represent a rare cause for HCFP and a rare high effect size risk factor for otosclerosis.
References
Menger DJ, Tange RA. The aetiology of otosclerosis: a review of the literature. Clin Otolaryngol Allied Sci. 2003;28:112–120.
Kremer H, Kuyt LP, van den Helm B, et al. Localization of a gene for Mobius syndrome to chromosome 3q by linkage analysis in a Dutch family. Hum Mol Genet. 1996;5:1367–1371.
Michielse CB, Bhat M, Brady A, et al. Refinement of the locus for hereditary congenital facial palsy on chromosome 3q21 in two unrelated families and screening of positional candidate genes. Eur J Hum Genet. 2006;14:1306–1312.
Verzijl HT, van den Helm B, Veldman B, et al. A second gene for autosomal dominant Mobius syndrome is localized to chromosome 10q, in a Dutch family. Am J Hum Genet. 1999;65:752–756.
Webb BD, Shaaban S, Gaspar H, et al. HOXB1 founder mutation in humans recapitulates the phenotype of Hoxb1-/- mice. Am J Hum Genet. 2012;91:171–179.
Uyguner ZO, Toksoy G, Altunoglu U, Ozgur H, Basaran S, Kayserili H. A new hereditary congenital facial palsy case supports arg5 in HOX-DNA binding domain as possible hot spot for mutations. Eur J Med Genet. 2015;58:358–363.
Sahin Y, Gungor O, Ayaz A, et al. A novel homozygous HOXB1 mutation in a Turkish family with hereditary congenital facial paresis. Brain Dev. 2017;39:166–170.
Vogel M, Velleuer E, Schmidt-Jimenez LF, et al. Homozygous HOXB1 loss-of-function mutation in a large family with hereditary congenital facial paresis. Am J Med Genet A. 2016;170:1813–1819.
Tomas-Roca L, Tsaalbi-Shtylik A, Jansen JG, et al. De novo mutations in PLXND1 and REV3L cause Mobius syndrome. Nat Commun. 2015;6:7199.
Milroy CM, Michaels L. Pathology of the otic capsule. J Laryngol Otol. 1990;104:83–90.
Declau F, Van Spaendonck M, Timmermans JP, et al. Prevalence of otosclerosis in an unselected series of temporal bones. Otol Neurotol. 2001;22:596–602.
Moumoulidis I, Axon P, Baguley D, Reid E. A review on the genetics of otosclerosis. Clin Otolaryngol. 2007;32:239–247.
Schrauwen I, Van Camp G. The etiology of otosclerosis: a combination of genes and environment. Laryngoscope. 2010;120:1195–1202.
Ziff JL, Crompton M, Powell HR, et al. Mutations and altered expression of SERPINF1 in patients with familial otosclerosis. Hum Mol Genet. 2016;25:2393–2403.
Untergasser A, Cutcutache I, Koressaar T, et al. Primer3—new capabilities and interfaces. Nucleic Acids Res. 2012;40:e115.
Ockeloen CW, Khandelwal KD, Dreesen K, et al. Novel mutations in LRP6 highlight the role of WNT signaling in tooth agenesis. Genet Med. 2016;18:1158–1162.
Boyle EA, O’Roak BJ, Martin BK, Kumar A, Shendure J. MIPgen: optimized modeling and design of molecular inversion probes for targeted resequencing. Bioinformatics. 2014;30:2670–2672.
O’Roak BJ, Vives L, Fu W, et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science. 2012;338:1619–1622.
Hiatt JB, Pritchard CC, Salipante SJ, O’Roak BJ, Shendure J. Single molecule molecular inversion probes for targeted, high-accuracy detection of low-frequency variation. Genome Res. 2013;23:843–854.
San Lucas FA, Wang G, Scheet P, Peng B. Integrated annotation and analysis of genetic variants from next-generation sequencing studies with variant tools. Bioinformatics. 2012;28:421–422.
Wang GT, Peng B, Leal SM. Variant association tools for quality control and analysis of large-scale sequence and genotyping array data. Am J Hum Genet. 2014;94:770–783.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2015;12:7–8.
Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–1612.
Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291.
Fisher LW, Fedarko NS. Six genes expressed in bones and teeth encode the current members of the SIBLING family of proteins. Connect Tissue Res. 2003;44 Suppl 1:33–40.
Martin A, David V, Laurence JS, et al. Degradation of MEPE, DMP1, and release of SIBLING ASARM-peptides (minhibins): ASARM-peptide(s) are directly responsible for defective mineralization in HYP. Endocrinology. 2008;149:1757–1772.
David V, Martin A, Hedge AM, Rowe PS. Matrix extracellular phosphoglycoprotein (MEPE) is a new bone renal hormone and vascularization modulator. Endocrinology. 2009;150:4012–4023.
Nampei A, Hashimoto J, Hayashida K, et al. Matrix extracellular phosphoglycoprotein (MEPE) is highly expressed in osteocytes in human bone. J Bone Miner Metab. 2004;22:176–184.
Rowe PS, de Zoysa PA, Dong R, et al. MEPE, a new gene expressed in bone marrow and tumors causing osteomalacia. Genomics. 2000;67:54–68.
Ogbureke KU, Fisher LW. Expression of SIBLINGs and their partner MMPs in salivary glands. J Dent Res. 2004;83:664–670.
Ogbureke KU, Fisher LW. Renal expression of SIBLING proteins and their partner matrix metalloproteinases (MMPs). Kidney Int. 2005;68:155–166.
Gowen LC, Petersen DN, Mansolf AL, et al. Targeted disruption of the osteoblast/osteocyte factor 45 gene (OF45) results in increased bone formation and bone mass. J Biol Chem. 2003;278:1998–2007.
Zelenchuk LV, Hedge AM, Rowe PS. Age dependent regulation of bone-mass and renal function by the MEPE ASARM-motif. Bone. 2015;79:131–142.
Rowe PS, Kumagai Y, Gutierrez G, et al. MEPE has the properties of an osteoblastic phosphatonin and minhibin. Bone. 2004;34:303–319.
Sprowson AP, McCaskie AW, Birch MA. ASARM-truncated MEPE and AC-100 enhance osteogenesis by promoting osteoprogenitor adhesion. J Orthop Res. 2008;26:1256–1262.
Estrada K, Styrkarsdottir U, Evangelou E, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet. 2012;44:491–501.
Neu-Yilik G, Amthor B, Gehring NH, et al. Mechanism of escape from nonsense-mediated mRNA decay of human beta-globin transcripts with nonsense mutations in the first exon. RNA. 2011;17:843–854.
van der Pluijm G, Mouthaan H, Baas C, de Groot H, Papapoulos S, Lowik C. Integrins and osteoclastic resorption in three bone organ cultures: differential sensitivity to synthetic Arg-Gly-Asp peptides during osteoclast formation. J Bone Miner Res. 1994;9:1021–1028.
Hayashibara T, Hiraga T, Yi B, et al. A synthetic peptide fragment of human MEPE stimulates new bone formation in vitro and in vivo. J Bone Miner Res. 2004;19:455–462.
Acknowledgements
This study was supported by funding from the Belgian Science Policy Office Interuniversity Attraction Poles (BELSPO-IAP) program (project IAP P7/43-BeMGI to G.V.C.) and grant 1505013N (Flemish National Fund for Scientific Research [FWO] to I.S.). L.T.-R. is a recipient of grants from the Fundación Séneca fellowship (04548/germ/06), EMBO short-term fellowship 200–2011, the IBRO Project In Europe grants program, and the Fundación Cultural Privada Esteban-Romero. H.V. is supported by a PhD grant from the Agency for Innovation by Science and Technology (IWT). Research in the group of H.v.B. is supported by the EU FP7 Large-Scale Integrating Project Genetic and Epigenetic Networks in Cognitive Dysfunction (241995). M.W., H. Kremer, and R.J.E.P. are supported by the Heinsius Houbolt Foundation, and H. Kayserili by the Scientific and Technological Research Council of Turkey (TUBITAK), grants 108S418 and 112S398 and CRANIRARE consortia of the European Research Area Network (E-RARE). Last, we would like to thank and acknowledge Robert Vincent (Causse Ear Clinic, France) and Marcel Cosgarea (Iuliu Hatieganu University of Medicine and Pharmacy, Romania) for their prior contributions to our otosclerosis research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Schrauwen, I., Valgaeren, H., Tomas-Roca, L. et al. Variants affecting diverse domains of MEPE are associated with two distinct bone disorders, a craniofacial bone defect and otosclerosis. Genet Med 21, 1199–1208 (2019). https://doi.org/10.1038/s41436-018-0300-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0300-5
Keywords
This article is cited by
-
Genome-wide screen of otosclerosis in population biobanks: 27 loci and shared associations with skeletal structure
Nature Communications (2023)
-
Genetics of otosclerosis: finally catching up with other complex traits?
Human Genetics (2022)
-
The functional extracellular matrix on the regulation of odontogenic differentiation of stem cells
Current Medicine (2022)
-
A wide range of protective and predisposing variants in aggrecan influence the susceptibility for otosclerosis
Human Genetics (2022)
-
Osteoprotegerin gene polymorphisms and otosclerosis: an additional genetic association study, multilocus interaction and meta-analysis
BMC Medical Genetics (2020)