Abstract
Purpose
Genetic testing in pediatric cholestasis can be very informative but genetic causes have not been fully characterized.
Methods
Exome sequencing and positional mapping in seven families with cholestatic liver disease and negative clinical testing for known disease genes.
Results
KIF12, which encodes a microtubule motor protein with a tentative role in cell polarity, was found to harbor three homozygous likely deleterious variants in three families with sclerosing cholangitis. KIF12 expression is dependent on HNF-1β, deficiency which is known to cause bile duct dysmorphogenesis associated with loss of KIF12 expression. In another extended family, we mapped an apparently novel syndrome of sclerosing cholangitis, short stature, hypothyroidism, and abnormal tongue pigmentation in two cousins to a homozygous variant in PPM1F (POPX2), a regulator of kinesin-mediated ciliary transport. In the fifth family, a syndrome of normal gamma glutamyltransferase (GGT) cholestasis and hearing loss was found to segregate with a homozygous truncating variant in USP53, which encodes an interactor with TJP2. In the sixth family, we mapped a novel syndrome of transient neonatal cholestasis, intellectual disability, and short stature to a homozygous variant in LSR, an important regulator of liver development. In the last family of three affected siblings, a novel syndrome of intractable itching, hypercholanemia, short stature, and intellectual disability was mapped to a single locus that contains a homozygous truncating variant in WDR83OS (C19orf56), known to interact with ATP13A2 and BSEP.
Conclusion
Our results expand the genetic heterogeneity of pediatric cholestatic liver disease and highlight the vulnerability of bile homeostasis to a wide range of molecular perturbations.
Similar content being viewed by others
Introduction
Bile is a concentrated mixture of cholesterol-derived bile acids and conjugated bilirubin that facilitates, among other physiological roles, the emulsification and absorption of lipids in the intestine.1 Hepatocytes and cholangiocytes are the epithelial cells in the liver, and they both participate in the production of bile. Bile secretion depends on the function of membrane transport systems in hepatocytes and cholangiocytes and on the structural integrity of the biliary tree. Canalicular bile is produced by polarized hepatocytes that possess distinct transporters in their basolateral (sinusoidal) and apical (canalicular) plasma membrane.2 These transporters are trafficked accordingly from their sites of synthesis and storage, and undergo distinct posttranslational modifications.3
Defects in any of the above developmental, structural, or functional characteristics may impair bile homeostasis and result in cholestatic jaundice, a disease that affects approximately 1:2500 term infants.4 Apart from the detrimental effects impaired bile circulation can have on nutrition, detoxification, and cholesterol homeostasis, serious liver injury is a common complication of longstanding cholestasis owing to the toxic (detergent) nature of bile acids. In fact, severe cholestatic liver disease is a leading referral to pediatric liver transplant centers.5
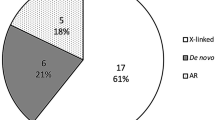
Despite attempts to subclassify cholestasis based on clinical characteristics, the resulting phenotypic categories remain broad and heterogeneous.6 Genetic profiling of cholestasis, on the other hand, offers a precise molecular classification that can have a meaningful impact on management and certainly on prevention.7 It is currently unknown what percentage of pediatric cholestasis is genetic in etiology, with estimates ranging from 25% to 50% (refs.8,9). In a recent study, we demonstrated the utility of a comprehensive gene panel that covers known cholestasis-related genes in patients with advanced cholestatic liver disease.7 A likely causal pathogenic variant was identified in the majority (n = 60, 61%) with pathogenic variants in ABCB11, ABCB4, and TJP2 being the most common. Other genetic diagnoses made include PFIC1 (progressive familial intrahepatic cholestasis caused by ATP8B1 pathogenic variants), Alagille syndrome, bile acid synthesis defect, tyrosinemia, Wilson disease, G6PC-related glycogen storage disease, SLC10A2-related bile acid malabsorption, MYO5B-related microvillus inclusion disease and VIPAS39-related cholestasis, renal impairment, and arthrogryposis (ARC) syndrome.7
In another recent study in which we aimed to characterize the landscape of Mendelian pathogenic variants in 1000 Saudi families with various phenotypes, we identified a potential genetic cause in 53% (10/19) of patients with severe cholestatic liver disease.10 This consistent diagnostic yield of screening known disease genes in children with advanced cholestatic liver disease may be interpreted as the limit of monogenic contribution to the etiology. However, several of the “negative” cases have a strong family history compatible with an autosomal recessive etiology, which suggests the possibility of novel genetic causes as the alternative explanation. Indeed, new genetic etiologies continue to be described in the literature, e.g., a novel UNC54A-related syndrome of cholestasis with diarrhea, deafness, and impaired bone strength was very recently published.11
In this study, we exploited the special characteristics of our population (consanguinity and large family size) to search for similarly novel disease genetic factors in pediatric cholestasis by employing genome sequencing and positional mapping.
Materials and methods
Human subjects
All the patients in this study were referred from two large pediatric and organ transplant centers in Riyadh, Saudi Arabia with abnormal bile metabolism and cholestatic liver disease following negative clinical sequencing of known cholestasis-related genes (targeted panels and exomes). Informed consent was obtained (a separate consent to publish photographs was also obtained), and venous blood was collected in EDTA tubes and sodium heparin tubes for genomic DNA extraction and establishment of lymphoblastoid cell lines from the affected and available family members per an institutional review board–approved protocol (KFSHRC RAC #2121053).
Positional mapping
Positional mapping was performed using autozygome analysis as described before.12,13 Genotyping was performed on the Axiom SNP chip platform and calling autozygous intervals was using AutoSNPa (http://dna.leeds.ac.uk/autosnpa/) and AgileMultiIdeogram (http://dna.leeds.ac.uk/autozygosity/), with runs of homozygosity (ROH) that are >2 MB in size considered as surrogates of autozygosity. Linkage analysis was performed with the easyLINKAGE package under a fully penetrant autosomal recessive model (http://nephrologie.uniklinikum-leipzig.de/nephrologie.site,postext,easylinkage,a_id,797.html).
Exome sequencing
The index individual from each of the recruited families was subjected to exome analysis. DNA samples were prepared using Agilent SureSelect Target Enrichment Kit, constructed libraries were sequenced on Illumina Hiseq2000 platform, and Genome Analysis Toolkit (GATK) was used for variant calling. Called variants were further filtered based on autozygosity and low frequency in ethnically matched exomes and gnomAD.14 Postfiltering variants were analyzed based on prioritizing truncating variants, and predicted pathogenicity in silico based on PolyPhen, SIFT, and combined annotation-dependent depletion (CADD) in case of missense variants.15 Segregation of candidate variants with the disease was confirmed by Sanger sequencing in all available family members.
RNA extraction, RT-PCR, and expression analysis
Total RNA was extracted from patient and control lymphoblast cells using the RNeasy Mini Kit (Qiagen Inc., MD, Cat #74106) and DNase treated by the RNase-Free DNase Set (Qiagen), as recommended by the manufacturer. Mouse Total RNA Master Panel was used to assess expression across mouse organs (Clontech Laboratories, Cat #636644). Complementary DNA (cDNA) was synthesized using the iScript™ cDNA synthesis kit (Applied Biosystems, CA) according to the manufacturer’s instructions. Reverse transcription–polymerase chain reaction (RT-PCR) was performed with gene-specific primers. Amplicons were analyzed on agarose gel (1.5%), and further sequence was confirmed by subjecting purified bands to Sanger sequencing. Primers used in these studies are available in Table S1.
Results
We recruited seven consanguineous families with one (in one family) or more (in six families) affected members who were diagnosed with infantile onset cholestasis or hypercholanemia. Patient characteristics (demographics, family history, and clinical features) are summarized in Table 1. Because negative screening for all known genes using panel testing or exome sequencing was a prerequisite, it was not surprising that our analysis did not reveal pathogenic variants in any of the known genes. Rather, in each of these families a likely novel candidate gene was identified as described below.
Pathogenic variants in KIF12 cause high GGT neonatal cholestasis/sclerosing cholangitis
We initially compared the candidate autozygome of the affected members across all seven families. Only three families showed an overlapping pattern. Specifically, family 1 (17DG0864), family 2 (17DG0877), and family 3 (17DG0404/17DG0551) showed an overlapping region of autozygosity on chr9 (112862200–117180500) but with different haplotypes, indicating that the causal founder recessive pathogenic variant may be different (Fig. 1). Linkage analysis confirmed this as the only candidate locus in these three families (Figure S1). Exome sequencing revealed a different novel homozygous variant in KIF12 within the critical locus in each of the three families. Specifically, we identified a homozygous truncating pathogenic variant in KIF12 (NM_138424.1:c.463C>T:p.[Arg155*]) in family 1, a homozygous missense variant (NM_138424.1:c.656G>A:p.[Arg219Gln]) in family 2, and a different homozygous missense variant (NM_138424.1:c.610G>A:p.[Val204Met]) in the same gene in family 3 (Fig. 1). The three variants are absent in 2379 ethnically matched exomes and all but one were also absent in gnomAD (123,136 exomes and 15,496 genomes) (NM_138424.1:c.610G>A:p.[Val204Met] was present at a very low frequency of 0.00002092 but never in homozygosity). Identified variants in KIF12 fully segregated with the disease in the respective families. The in silico prediction of both missense variants suggest pathogenicity (PolyPhen 0.953, SIFT 0 and CADD 34 for p.[Arg219Gln], and 0.989, 0.02, and 34 for p.[Val204Met]).
a Pedigrees of the families with KIF12-related high gamma glutamyltransferase (GGT) neonatal cholestasis and sclerosing cholangitis. b AgileMultiIdeogram showing a single locus on chr9 (112862200–117180500) spanning KIF12. c–e Histopathology at age of 14 months of patient 17DG0404. c Liver tissue showed biliary cirrhosis (H&E, 10×). d Trichrome stain is highlighting the degree of fibrosis, (TC, 10×). e Illustrates the degree of fibrosis surrounding the bile duct branches before complete destruction (H&E 40×). f~ Axial T2 weighted image from magnetic resonance cholangiopancreatography (MRCP) examination revealed intervening short segmental strictures and slight dilatation in the central intrahepatic bile ducts alternating with normal ducts resulted in beaded appearance (red arrow). g Sanger sequencing showing the three homozygous variants in KIF12. h Cartoon representing protein structure of KIF12 (513 amino acids in size, the kinesin motor domain [1–225] mediates intracellular transport of organelles and cell division) and the corresponding location of the three variants
Interestingly, patients from these three families share the same clinical phenotype characterized by high GTT neonatal cholestasis with rapid progression to liver fibrosis and portal hypertension. Two patients had living-related liver transplant at the age of 10 months (17DG0551; currently 17 years old) and 6 years (17DG0877; currently 20 years old). Both are alive and well. Liver biopsy for patient 17DG0551 at age 6 months was suggestive of biliary atresia despite normal gallbladder by ultrasound. He was transplanted at age 10 months because of liver failure. His cousin (17DG0404) presented with neonatal cholestasis. At age 14 months, he presented with itching and decreasing jaundice while GGT and alkaline phosphatase (ALP) were persistently elevated (Table 1), a typical biochemical pattern of childhood sclerosing cholangitis. Liver biopsy was suggestive of biliary cirrhosis (Figure S1). Magnetic resonance cholangiopancreatography (MRCP) at age 4 years showed beading of intrahepatic bile ducts indicating sclerosing cholangitis (Fig. 1). Currently, he is 11 years old and suffers from recurrent bleeding attacks because of portal hypertension and esophageal varices.
A novel locus for the syndrome of sclerosing cholangitis, short stature, hypothyroidism, and abnormal tongue pigmentation
In a fourth consanguineous family (family 4, two siblings and a cousin), we similarly applied positional mapping and exome sequencing and identified a novel homozygous variant in PPM1F (NM_014634.3:c.904C>T:p.[Arg302Cys]) in the critical locus (chr22:21917190–23055630) (Fig. 2). This variant is absent in gnomAD and local exome database, is predicted pathogenic by in silico tools (PolyPhen: 1, SIFT: 0, CADD: 31), and fully segregated with the disease in the family. One child died at age 13 years because of liver failure. The female sibling (17DG0439) presented with high GGT neonatal cholestasis and jaundice. Bilirubin normalized with time but GGT continued to be elevated. Liver biopsy at age 3 years showed paucity of bile ducts and liver fibrosis. MRCP at age of 19 years showed beading of major bile ducts indicating sclerosing cholangitis (Fig. 2). Currently, she is 21 years old and suffers from portal hypertension, short stature, hypothyroidism, and hyperpigmented tongue. Their cousin (17DG0930) was transplanted at age 2 years because of liver failure. He also suffers from hypothyroidism, hyperpigmented tongue, and short stature that is not corrected after successful living-related liver transplant (Fig. 2). Currently, he is 20 years old with normal liver function tests.
a Pedigree of family 4 with a proband and a cousin with sclerosing cholangitis, short stature, hypothyroidism, and abnormal tongue pigmentation. b AgileMultiIdeogram showing a single shared region of homozygosity (ROH) on chr22 (21917190–23055630) spanning PPM1F. (c) Pigmented tongue of IV:3 d, e Coronal magnetic resonance cholangiopancreatography (MRCP) image (3D Maximum-intensity projection (MIP) reformat) of patient 17DG0439 showed multifocal areas of narrowing, involving intrahepatic bile ducts of both lobes with intervening areas of dilatation producing beaded appearance, and multifocal short segmental strictures and dilatation in intrahepatic bile ducts (red arrow) alternating with normal ducts producing beaded appearance. f Sanger sequencing showing the change of C to T in the genomic DNA. g Graphical representation of PPM1F protein (PP2C domain [171–413 amino acids]). PP2C domain mediates the serine/threonine phosphatase activity
A novel locus for the syndrome of normal GGT cholestasis and deafness
In the fifth family, two sisters and a cousin presented with normal GGT cholestasis. The first sibling presented at age 4 months with jaundice, high alanine aminotransferase (ALT)/aspartate aminotransferase (AST), very high ALP, and normal GGT (Table 1). She had living-related liver transplantation at age 6 years because of intractable itching not responsive to medical treatment. She had persistent unexplained posttransplant hypocalcemia necessitating long term oral calcium supplements. At age 14 years, she had bilateral hearing loss. Currently she is 23 years old with normal liver function tests. Her sister presented at age 5 months with hypocalcemic seizure and very high ALP. She continued to have high ALP with mild elevation of ALT/AST, associated with intermittent episodes of intractable itching while on ursodeoxycholic acid. Itching was always responsive to the administration of rifampicin. She also had bilateral hearing loss at the age of 9 years. Currently, she is 15 years old with normal ALT/AST and no itching. A 3-year-old male cousin is affected by the same disease but has no deafness. He presented at age 1 year with intractable itching, very high bilirubin and ALP, moderately increased ALT/AST, and normal GGT (Table 1). He also had persistent hypocalcemia. He showed a dramatic response to rifampicin with normalization of liver enzymes, bilirubin, and disappearance of itching. We identified a single autozygous interval (chr4:118701500–127631100) that is exclusive to the affected siblings, i.e., not shared by the unaffected members, and the locus spans no known cholestasis-related genes. Exome sequencing revealed a single novel homozygous truncating variant therein in USP53 (NM_019050.2:c.951delT:p.[Phe317Leufs*6]) that fully segregated with the disease in the family (Fig. 3). Importantly, no shared candidate variant for hearing loss was identified in the two siblings.
a Pedigree of family 5 with two affected siblings and their affected cousin with the syndrome of normal gamma glutamyltransferase (GGT) cholestasis and deafness. b Sanger sequencing showing deletion of T in the genomic DNA. c AgileMultiIdeogram showing a single shared region of homozygosity (ROH) on chr4 (118701500–127631100) spanning USP53. d Graphical representation of USP53 protein (1073 amino acids, the UCH domain critically regulates protein turnover by modulating deubiquitinating process). Pathogenic variant (indicated in red) truncates the UCH domain. e Pedigree of family 6 with normal GGT cholestasis, intellectual disability, and short stature. f, g Liver biopsy images of patient 18DG04520 showing multiple portal areas with marked ductal proliferation associated with lymphocytic infiltrate. The hepatocytes show giant cell transformation with feathery degeneration associated with intrahepatic and canalicular cholestasis. h Sanger sequencing showing the identified variant in LSR and a cartoon showing the protein structure (Ig domain plays a role in protein folding and other biological pathways)
A novel candidate in a syndrome of normal GGT cholestasis, developmental delay, and short stature
In the sixth consanguineous family, we describe a 9-year-old boy presented at 4 months of age with hypocalcemic convulsion and severe itching with high total serum bile acids, total/direct bilirubin, ALT, AST, ALP, and normal GGT (Table 1). He has mild intellectual disability (IQ 70–77), speech delay but severe fine motor discoordination and short stature with normal skeletal survey. At age 1 year his liver biopsy showed a morphology consistent with paucity of the intrahepatic bile duct (Fig. 3). The hepatocytes showed giant cell transformation with feathery degeneration associated with intrahepatic and canalicular cholestasis (Fig. 3). Severe itching was difficult to control with ursodeoxycholic acid and rifampicin. Exome sequencing revealed a novel homozygous missense variant in LSR gene (NM_001260489.1:c.704A>G:p.[Glu235Gly]), which is pathogenic by all in silico (PolyPhen: 0.989/probably damaging; SIFT: 0.02/deleterious with CADD score 33), absent in our database and gnomAD (Fig. 3), and segregated within available family members.
A novel locus for the syndrome of intractable itching, hypercholanemia, dysmorphism, and intellectual disability
In the seventh family, three affected members presented with the consistent combination of hypercholanemia (high total serum bile acids and ALP with normal ALT, AST, and total bilirubin), intractable itching, intellectual disability (IQ 45), progressive microcephaly, dysmorphic facies, short stature, and genital abnormalities (very large clitoris in one affected girl, and undescended testes status postorchidopexy in the affected boy) (Fig. 4). We mapped this apparently novel syndrome to a single locus on chr19:8201051–16183420, which spans 351 genes, none of which is known to be mutated in cholestasis. Therefore, we proceeded with exome sequencing and only considered variants within the critical locus. The only novel homozygous coding/splicing variant that was identified therein was WDR83OS (NM_016145.3:c.156+1G>T). This variant abolishes a canonical splice donor site with resulting frameshift and premature truncation as confirmed by RT-PCR using RNA from patient lymphoblasts (r.51_156del:p.[Arg17Serfs*2]) (Fig. 4).
a Pedigree of family 7 with the syndrome of intractable itching, hypercholanemia, dysmorphism, and intellectual disability. b–d Facial dysmorphic features in the three affected siblings. e AgileMultiIdeogram showing a single region of homozygosity (ROH) on chr19:8201051–16183420. f Reverse transcription–polymerase chain reaction (RT-PCR) showing aberrant transcript with deletion of 105 bps (r.51_156del). g Sanger sequencing of WDR83OS showing the pathogenic variant at the DNA level. h Cartoon representation of WDR83OS (106 amino acids; the two transmembrane domains are highlighted). N.C. normal control, P patient
Expression profiling in mouse
We assessed the expression profiles of identified candidate genes KIF12, PPM1F, USP53, LSR, and WDR83OS in various mouse tissues using their murine orthologs as surrogates. Reassuringly, transcripts of all five genes were detected in the liver (Figure S2).
Discussion
Our understanding of cellular factors that are required for bile metabolism has greatly benefited from deciphering the genetics of hereditary forms of cholestatic liver diseases. In this study, we specifically focused on consanguineous cases of cholestasis with no identifiable etiology in known genes to enrich for novel presumably recessive gene discovery. While several novel loci were defined by this study, the one spanning KIF12 deserves a special emphasis because three independent likely deleterious variants were identified in this gene in three families with a strikingly similar phenotype. KIF12 encodes an orphan member of the kinesin family of molecular motors, which are involved in microtubule cytoskeleton organization and intracellular transport.16,17 Several lines of evidence, apart from the human genetics data presented in this report, support the candidacy of KIF12 in the context of sclerosing cholangitis. First, KIF12 is known to associate with the mitotic spindle and cleavage furrow and its deficiency has been shown to impair cytokinesis.18,19 This suggests a role in cellular polarity, a critical requirement for the proper arrangement of hepatocytes.20 Second, KIF12 expression is regulated by HNF1β, loss of which is known to cause bile duct morphogenesis defects and liver dysfunction.21 Third, KIF12 has been shown to modulate the phenotype of cpk mouse, which is characterized by severe cystic disease of the kidneys and liver.22 Finally, we note that pathogenic variants in DCDC2, encoding doublecortin domain containing 2 (DCDC2), which is known to interact with the Kinesin-2 complex, the KIF3A, are known to cause neonatal sclerosing cholangitis.23,24
Both DCDC2 and KIF3A localize to the primary cilium and centrosome and KIF3A is proposed to play a role in cell polarity.23 Interestingly, the phosphatase protein encoded by PPM1F gene, which we found to be mutated in a family with sclerosing cholangitis and hyperpigmented tongue, is also known to regulate centrosome orientation and kinesin-mediated transport and trafficking by dephosphorylating KIF3A on S690 (refs.25,26). It is tempting to speculate, therefore, that familial high GGT cholestasis/sclerosing cholangitis caused by primary defects in DCDC2, KIF12, and PPM1F represents a cholangiociliopathy caused by primary cilia dysfunction.27
In contrast to the presumed KIF12- and PPM1F-related cholangiociliopathy, USP53-related phenotype is probably related to defective tight junction complex. The syndrome of normal GGT cholestasis with bilateral hearing loss was found to fully segregate with a homozygous truncating variant in USP53. It is worth highlighting that Usp53 deficiency has been shown to cause deafness in mice, and that Usp53 colocalizes and interacts with the tight junction scaffolding proteins TJP1 and TJP2 in polarized epithelial cells, findings that suggest that USP53 is a component of the tight junction complex.28 Interestingly, our patients with USP53-related disease have normal GGT cholestasis similar to TJP2-related PFIC4 and this was indeed the primary differential diagnosis for the three children.29 Pathogenic variants in TJP2 and DCDC2 are also known to cause deafness in addition to liver involvement.30,31 In addition, primary loss of the P4-ATPase ATP8B1, known to cause PFIC1 with normal GGT cholestasis, is associated with deafness.32
In a family of three siblings with hypercholanemia, dysmorphism, and intellectual disability, we identified a homozygous truncating variant in WDR83OS (C19orf56). WDR83OS has been shown on yeast two-hybrid assays to interact with ATP13A2 (ref.33), deficiency of which is known to cause various neurodegenerative disorders including Parkinson disease and neuronal ceroid lipofuscinoses.34 WDR83OS also interacts with BSEP, deficiency of which is an established cause of neonatal cholestasis (PFIC2) (ref.35).
Lipolysis-stimulated lipoprotein receptor (LSR) has an established role in liver development in mouse and zebrafish. LSR deficient mouse exhibited small liver size with embryonic lethality at day 15.5 of gestation, and a similar phenotype was observed in zebrafish where morpholino-induced knockdown causes reduction in size of the liver.36,37 While cognitive/behavioral assessment was not possible in homozygous knockout mice due to embryonic lethality, it is interesting that a recent study has shown that heterozygous knockout mice are susceptible to Aβ42-induced cognitive changes.38 Taken together, the likely deleterious variant identified in LSR gene is an interesting candidate etiology of both the liver and cognitive phenotypes observed in the patient with a likely deleterious LSR variant.
It is rather remarkable that our small cohort of only seven families in whom known cholestasis-related genes had been excluded by clinical testing revealed such an array of novel candidate pathogenic variants . This suggests that the monogenic contribution to cholestasis etiology is much more diverse than previously thought and that the current estimate of 25–50% will likely be revised as more novel genetic etiologies such as the ones we report here are identified and reported.
With the exception of KIF12, which we found to be independently mutated in three families with an identical cholestasis phenotype, we note that our novel candidate genes, despite compelling positional mapping data and relevant biological functions, require confirmation through additional cases. We hope that our study will facilitate the genetic diagnosis of a larger proportion of pediatric cholestasis patients, because this has been shown to have important clinical utility in terms of management and recurrence risk determination.
References
Hofmann AF. Chemistry and enterohepatic circulation of bile acids. Hepatology. 1984;4 Suppl 5:4S–14S.
Keppler D, Arias IM. Hepatic canalicular membrane. Introduction: transport across the hepatocyte canalicular membrane. FASEB J. 1997;11:15–18.
Wakabayashi Y, Lippincott-Schwartz J, Arias IM. Intracellular trafficking of bile salt export pump (ABCB11) in polarized hepatic cells: constitutive cycling between the canalicular membrane and Rab11-positive endosomes. Mol Biol Cell. 2004;15:3485–3496.
McKiernan P. Neonatal cholestasis. Semin Neonatol. 2002;7:153–165.
Fayyad A, Shagrani M, AlGoufi T, et al. Progress and outcomes of the first high-volume pediatric liver transplantation program in Saudi Arabia. Clinical transplants. 2012:77–83.
Fawaz R, Baumann U, Ekong U, et al. Guideline for the evaluation of cholestatic jaundice in infants: joint recommendations of the North American Society for Pediatric Gastroenterology, Hepatology, and Nutrition and the European Society for Pediatric Gastroenterology, Hepatology, and Nutrition. J Pediatr Gastroenterol Nutr. 2017;64:154–168.
Shagrani M, Burkholder J, Broering D, et al. Genetic profiling of children with advanced cholestatic liver disease. Clin Genet. 2017;92:52–61.
Chen HL. Mining the idiopathic genetic cholestasis syndrome. J Gastroenterol Hepatol. 2013;28:389–391.
Feldman AG, Sokol RJ. Neonatal cholestasis. Neoreviews. 2013;14:e63–e73.
Monies D, Abouelhoda M, Al Sayed M, et al. The landscape of genetic diseases in Saudi Arabia based on the first 1000 diagnostic panels and exomes. Hum Genet. 2017;136:921–939.
Esteve C, Francescatto L, Tan PL, et al. Loss-of-function mutations in UNC45A cause a syndrome associating cholestasis, diarrhea, impaired hearing, and bone fragility. Am J Hum Genet. 2018;102:364–374.
Alkuraya FS. Autozygome decoded. Genet Med. 2010;12:765–771.
Alkuraya FS. Discovery of rare homozygous mutations from studies of consanguineous pedigrees. Curr Protoc Hum Genet. 2012 Oct;Chapter 6:.
Alkuraya FS. The application of next-generation sequencing in the autozygosity mapping of human recessive diseases. Hum Genet. 2013;132:1197–1211.
Alkuraya FS. Discovery of mutations for Mendelian disorders. Hum Genet. 2016;135:615–623.
Hirokawa N, Noda Y, Tanaka Y, Niwa S. Kinesin superfamily motor proteins and intracellular transport. Nat Rev Mol Cell Biol. 2009;10:682–696.
Nakagawa T, Tanaka Y, Matsuoka E, et al. Identification and classification of 16 new kinesin superfamily (KIF) proteins in mouse genome. Proc Natl Acad Sci USA 1997;94:9654–9659.
Lakshmikanth GS, Warrick HM, Spudich JA. A mitotic kinesin-like protein required for normal karyokinesis, myosin localization to the furrow, and cytokinesis in Dictyostelium. Proc Natl Acad Sci USA 2004;101:16519–16524.
Chen Q, Lakshmikanth GS, Spudich JA, De Lozanne A. The localization of inner centromeric protein (INCENP) at the cleavage furrow is dependent on Kif12 and involves interactions of the N terminus of INCENP with the actin cytoskeleton. Mol Biol Cell. 2007;18:3366–3374.
Wang L, Boyer JL. The maintenance and generation of membrane polarity in hepatocytes. Hepatology. 2004;39:892–899.
Coffinier C, Gresh L, Fiette L, et al. Bile system morphogenesis defects and liver dysfunction upon targeted deletion of HNF1β. Development. 2002;129:1829–1838.
Mrug M, Li RH, Cui XQ, Schoeb TR, Churchill GA, Guay-Woodford LM. Kinesin family member 12 is a candidate polycystic kidney disease modifier in the cpk mouse. J Am Soc Nephrol. 2005;16:905–916.
Massinen S, Hokkanen ME, Matsson H, et al. Increased expression of the dyslexia candidate gene DCDC2 affects length and signaling of primary cilia in neurons. PLoS ONE. 2011;6:e20580.
Grammatikopoulos T, Sambrotta M, Strautnieks S, et al. Mutations in DCDC2 (doublecortin domain containing protein 2) in neonatal sclerosing cholangitis. J Hepatol. 2016;65:1179–1187.
Hoon JL, Li HY, Koh CG. POPX2 phosphatase regulates cell polarity and centrosome placement. Cell Cycle. 2014;13:2459–2468.
Phang HQ, Hoon JL, Lai SK, et al. POPX2 phosphatase regulates the KIF3 kinesin motor complex. J Cell Sci. 2014;127 Pt 4:727–739.
Mansini AP, Peixoto E, Thelen KM, Gaspari C, Jin S, Gradilone SA. The cholangiocyte primary cilium in health and disease. Biochim Biophys Acta. 2018;1864(4 Pt B):1245–1253.
Kazmierczak M, Harris SL, Kazmierczak P, et al. Progressive hearing loss in mice carrying a mutation in Usp53. J Neurosci. 2015;35:15582–15598.
Sambrotta M, Strautnieks S, Papouli E, et al. Mutations in TJP2 cause progressive cholestatic liver disease. Nat Genet. 2014;46:326–328.
Kim MA, Kim YR, Sagong B, et al. Genetic analysis of genes related to tight junction function in the Korean population with non-syndromic hearing loss. PLoS ONE. 2014;9:e95646.
Grati M, Chakchouk I, Ma Q, et al. A missense mutation in DCDC2 causes human recessive deafness DFNB66, likely by interfering with sensory hair cell and supporting cell cilia length regulation. Hum Mol Genet. 2015;24:2482–2491.
Verhulst PM, van der Velden LM, Oorschot V, et al. A flippase‐independent function of ATP8B1, the protein affected in familial intrahepatic cholestasis type 1, is required for apical protein expression and microvillus formation in polarized epithelial cells. Hepatology. 2010;51:2049–2060.
Usenovic M, Knight AL, Ray A, et al. Identification of novel ATP13A2 interactors and their role in α-synuclein misfolding and toxicity. Hum Mol Genet. 2012;21:3785–3794.
de Tezanos Pinto F, Adamo HP. The strategic function of the P5-ATPase ATP13A2 in toxic waste disposal. Neurochem Int. 2018;112:108–113.
Przybylla S, Stindt J, Kleinschrodt D, et al. Analysis of the bile salt export pump (ABCB11) interactome employing complementary approaches. PLoS ONE. 2016;11:e0159778.
Mesli S, Javorschi S, Berard AM, et al. Distribution of the lipolysis stimulated receptor in adult and embryonic murine tissues and lethality of LSR-/- embryos at 12.5 to 14.5 days of gestation. Eur J Biochem. 2004;271:3103–3114.
Dokmanovic-Chouinard M, Chung WK, Chevre JC, et al. Positional cloning of “Lisch-Like”, a candidate modifier of susceptibility to type 2 diabetes in mice. PLoS Genet. 2008;4:e1000137.
Pincon A, Thomas MH, Huguet M, et al. Increased susceptibility of dyslipidemic LSR+/- mice to amyloid stress is associated with changes in cortical cholesterol levels. J Alzheimers Dis. 2015;45:195–204.
Acknowledgements
We thank the study families for their enthusiastic participation. We thank the Genotyping and Sequencing Core Facilities at King Faisal Specialist Hospital & Research Center (KFSHRC) for their technical help. This work was supported in part by King Salman Center for Disability Research (F.S.A.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Maddirevula, S., Alhebbi, H., Alqahtani, A. et al. Identification of novel loci for pediatric cholestatic liver disease defined by KIF12, PPM1F, USP53, LSR, and WDR83OS pathogenic variants. Genet Med 21, 1164–1172 (2019). https://doi.org/10.1038/s41436-018-0288-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0288-x
Keywords
This article is cited by
-
Diagnostic yield and novel candidate genes by next generation sequencing in 166 children with intrahepatic cholestasis
Hepatology International (2024)
-
Cholestatic Liver Disease in a Child with KIF12 Mutation
Indian Journal of Pediatrics (2023)
-
The diagnostic yield of exome sequencing in liver diseases from a curated gene panel
Scientific Reports (2023)
-
Computational identification of host genomic biomarkers highlighting their functions, pathways and regulators that influence SARS-CoV-2 infections and drug repurposing
Scientific Reports (2022)
-
A ZFYVE19 gene mutation associated with neonatal cholestasis and cilia dysfunction: case report with a novel pathogenic variant
Orphanet Journal of Rare Diseases (2021)