Abstract
Purpose
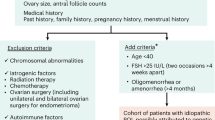
Primary ovarian insufficiency (POI) is a frequent disorder that affects ~1% of women under 40 years of age. POI, which is characterized by the premature depletion of ovarian follicles and elevated plasma levels of follicle-stimulating hormone (FSH), leads to infertility. Although various etiological factors have been described, including chromosomal abnormalities and gene variants, most cases remain idiopathic. The aim of the present study was to identify and validate functionally new sequence variants in ATG (autophagy-related genes) leading to POI.
Methods
We have reanalyzed, in silico, the exome sequencing data from a previously reported work performed in 69 unrelated POI women. Functional experiments using a classical hallmark of autophagy, the microtubule-associated protein 1 light chain 3β (LC3), were then used to link these genes to this lysosomal degradation pathway.
Results
We venture a functional link between ATG7 and ATG9A variants and POI. We demonstrated that variant ATG7 and ATG9A led to a decrease in autophagosome biosynthesis and consequently to an impairment of autophagy, a key biological process implicated in the preservation of the primordial follicles forming the ovarian reserve.
Conclusion
Our results unveil that impaired autophagy is a novel pathophysiological mechanism involved in human POI.
Similar content being viewed by others
Introduction
Primary ovarian insufficiency (POI) is a clinical syndrome characterized by the premature depletion of ovarian follicles, which clinically results in amenorrhea and elevated plasma levels of gonadotrophins.1 POI affects 1% of women before the age of 40 and 0.1% before the age of 30. POI generally leads to an irreversible infertility condition due to the early extinction of the ovarian reserve.2 POI etiology is highly heterogeneous and includes autoimmune diseases, metabolic disorders, viral infections, and iatrogenic causes.3 Genetic causes, such as chromosomal abnormalities and point mutations, have been described in syndromic (e.g., Turner; fragile X syndrome; and blepharophimosis, ptosis, and epicanthus inversus [BPES] syndrome) and nonsyndromic presentations of the disease.4 Although hundreds of proteins have been related to key developmental and physiological processes of the ovary, variants in fewer than 20 genes have been definitely validated, by functional tests, with POI etiology.5,6 Recently, we have reported a large genomic screening, via exome sequencing, performed in 69 Caucasian women affected by POI and described 55 candidate variants located in 49 genes potentially related to the phenotype.7 These variants can be considered strong etiogical POI candidates because they have been selected using stringent filtering, which took into account genetic, population, evolutionary, and functional variables. One patient displayed the c.1209T>A (p.Phe403Leu) drastic missense variant in the ATG7 gene, which encodes an ubiquitin E1-like activating enzyme involved in the two ubiquitin-like systems required for the expansion of autophagosomal membranes during macroautophagy.8 Autophagy-related genes (ATG) are the main regulators and implementers of the macroautophagy (hereafter referred to simply as autophagy).
Autophagy is a lysosomal degradative process of cytoplasmic components and damaged organelles that occurs in all eukaryotic cells from yeast to mammals. It is active in cells in a basal state, but can also be induced in response to nutritional deprivation or cellular stress.9 Autophagy starts with the formation of a double membrane, the phagophore, that expands into a vesicle known as the autophagosome, which engulfs part of the cytoplasm. The autophagosome then fuses with a lysosome to form an autolysosome within the sequestered material being degraded.10 This process is under the control of multiple ATG proteins. Some are engaged in two ubiquitin-like conjugation systems, which act sequentially during the elongation of the preautophagosomal membrane and autophagosome completion.11 However, only LC3 and other members of the ATG8 family remain anchored in the autophagosomal membrane.12 Basal autophagy is important for controlling the quality of proteins and organelles, to maintain cell function. Under stressful conditions, the principal role of autophagy is to supply nutrients for survival. Autophagy is involved in many physiological processes such as development, differentiation, and aging, and also contributes to both innate and adaptive immunity.13,14,15 Polymorphisms and variants in some of these genes have been associated with susceptibility to human diseases (e.g., Crohn disease, bacterial infections, cancer, Parkinson disease, asthma).16
In the ovary, autophagy is involved in maintaining the primordial oocyte pool in murine newborns.17 Moreover, it has been demonstrated that it is related to oocyte elimination during atresia.18 Interestingly, germ cell–specific knockout of Atg7 in mice led to POI, as the number of the follicles/oocytes was significantly reduced in the adult mutant animals.19 These results strongly suggested that ATG variants might also originate POI phenotype in humans.
Herein, we reanalyzed in silico exome sequencing data from the previous reported work7 and identified in a POI woman a novel variant in ATG9A (p.Arg758Cys). We performed functional assays for the previously described ATG7 p.Phe403Leu variant and for ATG9A p.Arg758Cys by studying their effect on autophagy. The ATG9A gene encodes a multipass transmembrane protein contained in vesicles that contribute at least in part to the membranes for the nucleation and the extension of the phagophore.20 The functional analysis revealed that ATG7 and ATG9A variants impair autophagy suggesting that they contribute to POI etiology.
Materials and methods
Selection of ATG variants and in silico analysis
The ATG7 p.Phe403Leu variant was previously identified by filtering exome sequencing data from 69 POI unrelated patients from French origin.7 The new ATG9A p.Arg758Cys variant was found by reanalyzing the data from Patiño's work. Details on these methodologies have been included in Supplementary Materials and Methods.
Cell culture and transfection
COS-7 cells grown in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal calf serum (FCS) in 5% CO2 at 37 °C were transiently transfected with expression vectors encoding ATG7-Myc or ATG9-GFP wild-type or mutated, using FuGENE 6 reagent (Roche Diagnostics) according to the manufacturer’s protocol. Transfected cells were used 24 or 48 h later. KGN cells21 were maintained in DMEM/F12 containing 10% FCS, in 5% CO2 at 37 °C. Cells were transfected with vectors expressing either an ATG7 short hairpin RNA (shRNA), an ATG9A shRNA, or a control shRNA and a gene imparting resistance to puromycin using FuGENE 6 reagent. Among puromycin-resistant clones, those expressing half of ATG7 (after 1 µg/mL doxycycline treatment for 48 h) and ATG9 compared with control cells were selected by western blot using anti-ATG7 and anti-ATG9 antibodies. Clonality was verified by immunostaining with the appropriated antibodies. Starvation-induced autophagy was carried out by culturing the cells in Earle’s Balanced Salt Solution (EBSS) (Gibco, Life Technologies) for 6 h. Chloroquine and bafilomycin A1 (Sigma) were used at 15–20 µM and 100 nM, respectively, to block autophagic degradation.
Indirect immunofluorescence analysis
Cells cultured in chamber slides (Nunc™ Lab-Tek™ II Chamber Slide™ System; Thermo Fisher Scientific) were washed with phosphate-buffered saline (PBS), fixed with ice-cold methanol for 2 min followed by ice-cold acetone for 2 min. They were incubated for 1 h in a blocking solution (DAKO® Protein Block Serum-Free) and then with appropriate primary antibodies diluted in DAKO® Antibody Diluent overnight at 4 °C. The cells were washed, incubated with appropriate secondary antibodies diluted in the same buffer, and stained with DAPI. The cells were mounted in Fluorescent Mounting Medium (DAKO). Imaging was carried out on a SP8 Leica confocal microscope (Leica Microsystems, Deerfield, IL, USA) or a Thermo Cellomics ArrayScan VTI HCS Reader (Thermo Fisher, Pittsburgh, PA, USA).
Automated quantification of LC3 dots by High Throughput Microscopy (HTM)
COS-7 transfected with ATG7 and ATG9 wild-type or mutated were processed as described above for LC3, ATG7, or ATG9 immunofluorescence staining. Sequential images (40× −0.75 NA lens) were acquired for each channel with the automated ArrayScan VTI imaging and analyzing platform (Thermo Fisher Scientific, Villebon-sur-Yvette, France). The SpotDetector Algorithm (v.4) was used to detect and quantify the LC3 dots as following. DAPI fluorescence was used to find the focus and was sequentially acquired with ATG (7/9A) and LC3 fluorescence to give separate image files. The DAPI staining (channel 1) identifying the nuclear region was used to define a binary nuclear mask. Nuclei clusters, mitotic cells, and apoptotic cells were gated out from the total cell population (total primary object) by using several progressive morphological filters. To quantify the amount of ATG7 and ATG9A fluorescence (channel 2) present in the cytoplasm of the primary selected object, this nuclear mask was dilated to cover the cytoplasmic region. Subtraction of the nuclear from the dilated mask created a binary cytoplasmic new mask covering the cytoplasmic region. An automatic cut-off threshold was used to specifically select the cells expressing ectopic ATG7 or ATG9A (secondary selected object, i.e., transfected cells). The cytoplasmic mask was lastly used to detect, select, and quantify the number, size, and fluorescence of dots/cell (LC3 channel) in the final population of transfected cells (n > 400) as described previously in Mouna et al.22 The images and masks were systematically visually inspected for accuracy. Note that the contributions from nuclear dots were totally excluded thereby restricting the analysis to perinuclear and cytoplasmic LC3 dots.
Statistical analysis
All data were analyzed using Prism 6 (GraphPad), and the results are presented as mean ± SEM using at least three independent experiments. An unpaired Student’s t test was performed when comparing two groups and a one-way analysis of variance (ANOVA) with Kruskal–Wallis’s post hoc test was performed when comparing more than two groups. A P value of 0.05 or less was considered statistically significant (*P < 0.05, **P < 0.01, ***P < 0.001).
Ethical approval
All clinical and experimental steps of this study were approved by Institutional Review Board (reference PHRC No. A0R03 052) and by the Bicêtre Ethical committee (CPP #PP 16–024 Ile-de-France VII). The clinical investigation was performed according to Helsinki Declaration guidelines (1975, as revised in 1996). All of the women had given their informed consent to participate.
Results
Patients and hormonal evaluation
Patients (Pt-7 and Pt-66) were of Caucasian origin and attended for the clinical evaluation of hypergonadotropic amenorrhea.7 Turner syndrome, X chromosome structural abnormalities, and FRMR1 premutations were ruled out by standard genetic analysis.
Pt-7 displayed normal pubertal development, with a menarche at age 14 then an idiopathic hypergonadotropic secondary amenorrhea at 34 but she was diagnosed at 37. Hormonal plasma values showed elevated follicle-stimulating hormone (FSH) and luteinizing hormone (LH) levels (FSH: 83 IU/L, LH: 17 IU/L; normal range of basal levels of FSH and LH are 3–9 IU/L and 1–5 IU/L, respectively), and low estradiol (<5 pg/mL; normal range is 20–350 pg/mL) and anti-Müllerian hormone (AMH) levels (<1 pmol/L; normal range is 15.7–48.5 pmol/L). Inhibin B was 12 ng/L (normal ranges are 60–200 ng/L). The ultrasound examination found a normal uterus but small ovaries lacking follicles (2.9 mL, normal ovarian volume ranges between 5 and 8 mL). Menopause of her mother and grandmother occurred before the age of 40, but their genetic evaluation was refused.
Pt-66 was 22 at first clinical evaluation. She presented primary amenorrhea, delayed puberty (Tanner stage B1) with an infantile uterus and strip gonads at ultrasound examination. Hormonal values showed elevated FSH and LH levels (FSH: 74 IU/L, LH: 21 IU/L), and low estradiol (6 pg/mL), AMH (<1 pmol/L), and inhibin B levels (<15 ng/L). She did not display family history of POI.
Detection of ATG variants and in silico analysis
Exome sequencing experiments and bioinformatics filtering of variants led to the identification in Pt-7 of the ATG7 (MIM 608760) c.1209T>A (p.Phe403Leu) variant.7 Sanger sequencing experiments performed in the work by Patiño et al.7 confirmed this missense variant. The ATG9A (MIM 612204) c.2272C>T (p.Arg758Cys) variant was identified in Pt-66 by reanalyzing our previously published data.7 Sanger sequencing confirmed the presence of the variant in this patient. Various databases displayed that the ATG7 p.Phe403Leu and ATG9 p.Arg758Cys variants are rare, which argued in favor of a potential pathogenic effect (Table S1). Notably, they were absent from the French Exome Project database (http://med-laennec.univ-brest.fr/FrExAC/), which can be used as a repository of Caucasian controls for association studies. In addition, various bioinformatic tools revealed scores associated with potential pathogenic effects for both variants (Table S1). Protein alignments revealed that the ATG7 p.Phe403Leu variant, located in the E1-like domain of the protein, affected an amino acid highly conserved among species (Fig. 1a). The ATG9A p.Arg758Cys variant, which was found in the C-terminal region of the protein outside the six transmembrane segments, also affected an amino acid highly conserved during evolution (Fig. 1b). To evaluate the potential effect of the ATG7 p. Phe403Leu substitution on the ATG7 homodimer conformation, we generated a three-dimensional homology model of a dimer of the human ATG7 by taking as guides the X-ray crystal structure of the dimeric yeast ATG7 and the homology model of the monomeric human ATG7 (Figure S1A).23
ATG7 and ATG9A variants identified in premature ovarian insufficiency (POI) patients. Schematic representation of (a) ATG7 and b ATG9A proteins and interspecific sequence alignments of genomic regions carrying the interchanged amino acids. E1-like domain E1 activating enzyme-like, TM transmembrane domain
This 3D model showed that the Phe403 residue within the wild-type (WT) ATG7 is located at the dimerization interface (Figure S1A). Phe403 from one monomer established stabilizing van der Waals contacts with residues from the same monomer (Tyr392 and Leu407) and with numerous ones from the second monomer (Pro325, Lys326, Ala329, Phe425, and Pro426) (Figure S1B). This network of van der Waals contacts was likely involved in the dimer stabilization. However, the replacement of the Phe by a Leu at the 403 position led to a drastic reduction of the number of van der Waals contacts between both monomers (Figure S1C).
Effect of ATG7 and ATG9A haploinsufficiency on autophagy
Our two POI patients carried the heterozygous ATG7 p.Phe403Leu and ATG9A p.Arg758Cys variants, which are predicted deleterious and affecting autophagy. A dominant negative effect was excluded for both variants because a biallelic loss was lethal in Atg7 and Atg9 deficient mouse models.24,25
To evaluate the potential effect of ATG7 and ATG9A haploinsufficiency on autophagy, we generated KGN cell lines stably transfected with vectors expressing either ATG7 shRNA or ATG9A shRNA. Then, we selected individual clones expressing half levels of ATG7 or ATG9A to mimic the consequences of the potential deleterious effect of heterozygous variants (Fig. 2). These cells were used to study their ability to produce autophagosomes compared with that from control cells. A classical hallmark of autophagy activation is the conversion of microtubule-associated protein 1 light chain 3β-I (LC3-I) to lipidated LC3-II form; the latter correlates with the autophagosome number. To evaluate autophagosomes biosynthesis, LC3-II levels must be analyzed in the presence of inhibitors of the lysosomal function, such as bafilomycin A1 (Baf A1) or chloroquine (CQ). In basal (nonstarvation) conditions, we identified that LC3-II levels after CQ treatment were not significantly decreased in KGN underexpressing ATG7 or ATG9A (data not shown).
Reduced expression of ATG7 and ATG9A decreases autophagy. a KGN cells carrying a Dox-inducible vector expressing an ATG7 short hairpin RNA (shRNA) or shRNA control (Ctl) were treated with Dox to reduce the expression of ATG7 by 50%. b KGN cells were stably transfected with a vector expressing a control shRNA (Ctl) or an ATG9A shRNA. Clones expressing half of ATG9A compared with control were used for the experiment. Control, shATG7-, or shATG9A-KGN were incubated in starved medium for 6 h in the absence or presence of chloroquine (CQ) during the last 2 h or 4 h. Expression of ATG7, ATG9A, LC3-I, LC3-II, and actin were detected by protein gel blotting. The graphs represent the LC3-II/actin ratio at the different time points. Data represent the mean ± SEM and are representative of four independent experiments. *P < 0.05 and **P < 0.01, by Mann–Whitney test
Our experiments showed that under starvation conditions, LC3-II level in cells expressing half ATG7 or ATG9A dose was similar to that from nonsilenced (control) cells. After 2 or 4 h of CQ treatment, an increased amount of LC3-II was observed in control cells. However, this effect was significantly less in cells underexpressing ATG7 (Fig. 2a) (a decrease of 33% and 39% compared with control after 2 h and 4 h CQ treatment with P < 0.05 and P< 0.01, respectively) or ATG9A (Fig. 2b) (a decrease of 35% and 52% compared with control after 2 h and 4 h CQ treatment with P < 0.05 and P < 0.01, respectively). These results indicated that ATG7 or ATG9A downexpression led to a decreased autophagosome biosynthesis and consequently to a decreased autophagy.
Study of ATG7 p.Phe403Leu and ATG9A p.Arg758Cys subcellular localization in COS-7 transfected cells
To perform functional analysis of variants found in POI patients, expression plasmids including WT and mutant versions of ATG7 and ATG9A were used to transfect COS-7 cells. Expression levels of WT and mutant proteins were evaluated by western blot.
We found that ATG7 or ATG9A variant from two independent clones were expressed similarly to the WT counterpart (Fig. 3a, b). The subcellular distribution of ATG7 and ATG9 proteins was studied by immunofluorescence. As shown in Fig. 3c (left panel), ATG7 WT displayed a diffuse cytoplasmic distribution, which was conserved in cells expressing the ATG7 p.Phe403Leu version. The distribution of ATG9A was not as diffuse in the cytoplasm and it was stored in perinuclear clusters as previously described by others (Fig. 3d left panel).20 Likewise, ATG9A p.Arg758Cys displayed an identical localization indicating that the cellular distribution was not affected by this variant.
Expression and localization of ATG7 and ATG9A variants. Results of western blot using (a) an anti-Myc tag antibody to detect ATG7 and b an anti-GFP antibody to detect ATG9A in cells transiently transfected with the corresponding vectors. The overall expression levels of ATG7 F403L and ATG9A R758C variants from two independent clones (1 and 2) were similar to that of their wild-type (WT) counterpart. Actin was used as loading control, and the empty vector was used as a negative control (Ctl) for protein expression. Ratio of ATG/actin is indicated below. c Immunofluorescence results from COS-7 cells expressing ATG7 WT or F403L stained with antibodies against Myc-Tag (ATG7, green) and LC3 (red). Nuclei were stained with DAPI (blue). White arrows point to LC3 dots in transfected cells (green). d Immunofluorescence results from COS-7 cells expressing ATG9A WT or R758C stained with antibodies against GFP-Tag (ATG9A, green) and LC3 (red). Nuclei were stained with DAPI (blue). White arrows point to LC3 dots in transfected cells (green). Scale bar, 10 µm
ATG7 p.Arg403Leu and ATG9A p.Arg758Cys variants alter autophagy
To investigate the consequence of both variants on the autophagy, we first examined the ability of COS-7 cells expressing WT or mutant ATGs proteins to produce autophagosomes (Fig. 3c, d right panel). Upon autophagy induction, LC3-II associates with autophagosomal membranes, resulting in the formation of punctuate structures, which can be visualized by immunofluorescent labeling of endogenous LC3. To better visualize LC3 dots, the cells were grown under nutrient-deficient conditions. We observed a reduced amount of dots in COS-7 cells expressing mutated ATG7 and ATG9A proteins compared with their WT counterparts. This observation suggested that both variants contributed to impair autophagy.
To better quantify this alteration, we monitored by western blot LC3-II expression in COS-7 cells transfected with WT and mutated ATG7 or ATG9A forms, under complete medium as well as after starvation. In basal conditions (complete medium) we did not observe any significant change of LC3-II between cells expressing control and mutated proteins, even in the presence of CQ (data not shown). Under starvation environment (Fig. 4a), the amount of LC3-II was stable after 2 and 4 h of CQ treatment in control cells. In transfected cells with ATG7 WT, the amount of LC3-II was not significantly different from control cells in the absence of CQ. However, it was strongly increased after 2 h of CQ treatment and even more after 4 h of treatment, suggesting that the overexpression of ATG7 was able to increase the autophagic capacity of the cells. On the contrary, cells expressing ATG7 p.Phe403Leu exhibited similar autophagic capacity to that of control cells, displaying similar levels of LC3-II after 2 h and 4 h of CQ treatment. These results confirmed that the ATG7 p.Phe403Leu variant impaired autophagy due to a decreased ability to produce autophagosomes. Cells expressing ATG9A WT also accumulated significantly more LC3-II after 2 and 4 h of CQ treatment than the control cells. However, this LC3-II accumulation under CQ treatment was less important in cells expressing the ATG9A p.Arg758Cys form suggesting that this variant also impaired autophagy (Fig. 4b).
ATG7 F403L and ATG9A R758C variants alter the autophagy. a Cells expressing ATG7 wild-type (WT) or F403L variant were incubated in starved medium for 6 h in the absence or presence of chloroquine (CQ) during the last 2 h or 4 h. Expression of ATG7, LC3-I, LC3-II, and actin was detected by protein gel blotting. The graph represents the LC3-II/actin ratio at the different time points. b Cells expressing ATG9A WT or R758C were incubated in starved medium for 6 h in the absence or presence of chloroquine (CQ) during the last 2 h or 4 h. Expression of ATG9A, LC3-I, LC3-II, and actin were detected by protein gel blotting. Control cells (Ctl) were transfected by an empty vector. Data represent the mean ± SEM and are representative of four independent experiments. *P < 0.05, **P < 0.01, and ****P < 0.0001 by one-way analysis of variance (ANOVA) test
To assess this alteration more precisely, we used an automated testing of LC3 dots, which allows the specific counting of LC3 dots in cells transfected uniquely with WT or mutant ATG forms (Fig. 5). First, we verified that the fluorescence from proteins was the same in cells expressing WT and mutated ATG7 p.Phe403Leu or ATG9A p.Arg758Cys (Figure S2). Then, we measured several parameters including the number of LC3 dots per cell (Fig. 5 b, c), the average area of individual LC3 dots and the average fluorescence intensity per cell (Figure S3). In starvation conditions (EBSS), compared with cells expressing WT proteins, the cells transfected with the ATG7 p.Phe403Leu or ATG9A p.Arg758Cys constructs displayed 36% and 42% decrease in the number of autophagosomes, respectively. The size of autophagosomes (LC3 spot average area) and average fluorescent intensity per cell were also slightly but significantly reduced (Figure S3). In the presence of bafilomycin A1, a lysosomal inhibitor that impairs autophagosomes degradation, the number of dots was reduced by 42% in COS-7 cells expressing the ATG7 p.Phe403Leu variant, compared with cells expressing the ATG7 WT form (Fig. 5b, right graph). A reduction in the number of dots by 20% was also observed in cells expressing the ATG9A p.Arg758Cys construct, compared with those expressing the ATG9A WT version (Fig. 5c, right graph). Again, these results demonstrated that in starvation conditions both variants impaired the formation of autophagosomes contributing to a deleterious autophagic process.
HTM quantification of autophagosome formation. a Representative pictures generated by HTM showing accurate dots identification through binary masks of transfected cells. Cells were transfected with the ATG7 wild-type (WT) vector and then starved in EBBS medium for 6 h in presence of bafilomycin A1. Immunocytochemistry was performed with specific antibodies to visualize ATG7 and LC3. Automated HTM acquisition and cellular dots analysis were achieved with an ArrayScan VTI microscope. Left panels show examples of sequential acquisition of one field that was acquired for each channel as indicated in the pictures (ATG7, LC3, and merge). Arrows show ATG7 transfected cells. Central panel shows binary intracellular mask generated by HCS Studio software for the ATG7 expressing cells (cell object in white), the DAPI labeling (nucleus in blue), and the autophagosome present in these cells (dots, in green). b Cells expressing ATG7 WT or F403L were starved in EBBS for 6 h in absence or presence of bafilomycin A1 (Baf A1). Cells were fixed and analyzed by automated HTM as described in (a). Binary mask was generated by HCS Studio software to detect, delineate, and quantify the dots corresponding to autophagosomes. Histograms show the quantification of the average number of dots found in each transfected cells expressing ATG7 WT or F403L mutant., n > 400 cells for each set of conditions. Error bars represent SEM. ****P < 0.0001 by Student’s t test. (c) Cells expressing ATG9A WT or R758C were starved in EBBS for 6 h in absence or presence of bafilomycin A1 (Baf A1), as indicated in the figure. Cells were fixed and analyzed by automated HTM as described in (a). Histograms show the quantification of the average number of dots found in each transfected cell expressing ATG9A WT or R758C mutant. n > 400 cells for each set of conditions. Error bars represent SEM. ****P < 0.0001 by Student’s t test
Discussion
During oogenesis, female germ cells are produced but die with up to two-thirds of the oocytes being lost before or shortly after birth.26 The ovarian reserve containing primordial follicles is an important determinant of the length of the ovarian lifespan, and therefore the fertility of women.3,27 POI is one of the main causes of female infertility owing to a declined ovarian reserve. POI phenotypes are highly heterogeneous probably due to different etiological mechanisms leading to ovary dysfunction.3 However, in most cases POI remains idiopathic, which has encouraged research into further genetic causative factors. Advances in sequencing technologies have highlighted the involvement of new genes in POI etiology, such as STAG3, MCM9, MCM8, LHX8, SOHLH, MSH4, MSH5, and SYCE1 (ref. 6).
In the present study, we tested functionally two variants affecting ATG genes, which were identified in women affected by POI. One of them (ATG7 p.Phe403Leu), was previously identified by using an innovative bioinformatic pipeline performed on genomic data issued from exome sequencing, which was carried out in 69 unrelated nonsyndromic POI patients.7 At that time, to select a coherent number of variants underlying potential strong functional effects, the in silico analysis was performed by applying stringent filters. These included scores compatible with potential deleterious effects obtained from SIFT and PolyPhen software. These programs have been massively used to analyze next-generation sequencing data, but they have displayed variable or inconsistent rates of false positive/negative results.28 This feature led us in the present work to reanalyze our previous data.7 We undertake manual interspecific alignments of ATG molecules carrying candidate missense variants that lack positive predictive deleterious scores in both SIFT and PolyPhen2 software. This approach allowed us to identify the novel ATG9A p. Arg758Cys variant that affected a highly conserved residue during evolution. Due to their low frequency in different populations and the potential deleterious effects detected by various in silico programs, we considered that these variants were coherent candidates for functional downstream analysis (Table S1).
ATG7 is part of the two ubiquitin-like systems involved in autophagy and is essential for the production of LC3-phosphatidylethanolamine (PE), which in turn is necessary for the expansion of autophagosomal membranes.8 ATG9A is a multispanning transmembrane ATG protein localized in the Golgi apparatus and the endosomes. Although its role is not still completely understood, this protein is crucial for autophagosome formation, as a decrease in LC3 conjugation to PE and in autophagosome formation has been observed in fibroblasts from Atg9a-deficient mice.25 Because both proteins are crucial for autophagosome formation, an alteration in ATG7 and ATG9 function results in a decreased capacity of the cells to respond to starvation-induced autophagy.
Several studies that used genetically modified mouse models have suggested the involvement of autophagy in the regulation of the ovarian reserve of primordial follicles. First, Gawriluck et al.17 observed that total or partial inhibition of autophagy in Atg7-/- mouse or Becn1+/- (Beclin1) mouse, respectively, resulted in an overloss of germ cell populations at postnatal day 1. They suggested that autophagy was an important regulator of germ cell survival prior to formation of the primordial follicular pool.17 Another study reported the use of mice displaying Atg7 deletion only in germ cells of female mice.19 This study showed that the number of oocytes is nearly the same at 17.5 days p.c. compared to WT animals while oocytes overloss was observed 3 days after birth. This suggested that autophagy is necessary for germ cells' survival during the neonatal transition. Indeed, in mammals the supply of nutrients through the placenta is interrupted at birth and neonates face severe starvation until supply can be restored through milk nutrients. During this neonatal starvation period, autophagy is induced in most of the organs to maintain energy homeostasis.29 Of note, among all organ systems only ovaries exhibit severe cell loss at birth.26 Autophagy is a critical process seeming to be essential to protect germ cells for overloss. In mouse the ovarian reserve is completed 3 days after birth, while this reserve is established earlier during the first trimester of pregnancy in woman.27 Thus, the induction of autophagy at birth is a crucial step to preserve the stock of primordial follicles.
In the present study, we showed that the decreased expression (50%) of ATG7 and ATG9A representing haploinsufficiency reduced the capacity of the cell to respond to starvation-induced autophagy and that ATG7 p.Phe403Leu and ATG9A p.Arg758Cys variants were deleterious for autophagy. The role of autophagy in the ovary is complex because it is dependent on cellular type and developmental stage. For example, the activation of PI3kinase signaling pathway suppresses granulosa cell autophagy during follicular development and is involved in the regulation of follicular atresia.30 In both heterozygous patients, it is quite likely that the induction of autophagy at birth was low in ovaries. This might lead to an alteration of the ovarian reserve. In Becn1+/- mouse ovaries, the number of germ cells 1 day after birth is decreased by half compared with wild-type ovaries.17
The phenotype of our two patients carrying ATG variants was heterogeneous. Pt-66 displaying the ATG9A variant (p.Arg758Cys) exhibited primary amenorrhea, absence of puberty, and a total depletion of follicles into very small ovaries, whereas Pt-7, carrying the ATG7 variant (p.Phe403Leu), was affected by secondary amenorrhea. The functional defects of autophagy observed in vitro and the age of POI onset may support a model of complex pathogenesis as already described for the BMP15 gene31 and NOBOX.32 Our data do not exclude the possible involvement of a “second hit” explaining the phenotype, as described in hypogonadotropic hypogonadism33 as well as in our previous study on POI patients.34 Following the last recommendations of the American College of Medical Genetics and Genomics/Association for Molecular Pathology (ACMG/AMP) and the Sherloc interpretation tool for classifying genetic variants, we consider that the ATG7-Phe453Leu and ATG9A-Arg758Cys can be categorized as pathogenic; this classification can be established due to the variants' rarity and their functional impact observed in our in vitro tests.35,36
Our approach may lose further candidates contributing to the phenotype. Indeed it seems likely that mutational analysis of multiple genes in apparently monogenic disorders will become increasingly frequent.
In conclusion, these results reveal for the first time a role of autophagy in the pathogenesis of POI and describe a novel molecular biomarker for diagnosis/prognosis purposes. Genotyping of these genes in women affected by POI from different ethnic origins should allow identifying novel variants and establishing potential genotype–phenotype correlations.
References
Beck-Peccoz P, Persani L. Premature ovarian failure. Orphanet J Rare Dis. 2006;1:9.
Nelson LM. Clinical practice. Primary ovarian insufficiency. N Engl J Med. 2009;360:606–614.
De Vos M, Devroey P, Fauser BC. Primary ovarian insufficiency. Lancet. 2010;376:911–921.
Rossetti R, Ferrari I, Bonomi M, Persani L. Genetics of primary ovarian insufficiency. Clin Genet. 2017;91:183–198.
Laissue P. Aetiological coding sequence variants in non-syndromic premature ovarian failure: from genetic linkage analysis to next generation sequencing. Mol Cell Endocrinol. 2015;411:243–257.
Laissue P. The molecular complexity of primary ovarian insufficiency aetiology and the use of massively parallel sequencing. Mol Cell Endocrinol. 2018;460:170–180.
Patino LC, Beau I, Carlosama C, et al. New mutations in non-syndromic primary ovarian insufficiency patients identified via whole-exome sequencing. Hum Reprod. 2017;32:1512–1520.
Feng Y, Yao Z, Klionsky DJ. How to control self-digestion: transcriptional, post-transcriptional, and post-translational regulation of autophagy. Trends Cell Biol. 2015;25:354–363.
Kroemer G, Marino G, Levine B. Autophagy and the integrated stress response. Mol Cell. 2010;40:280–293.
Lamb CA, Yoshimori T, Tooze SA. The autophagosome: origins unknown, biogenesis complex. Nat Rev Mol Cell Biol. 2013;14:759–774.
Mercer TJ, Gubas A, Tooze SA. A molecular perspective of mammalian autophagosome biogenesis. J Biol Chem. 2018;293:5386–5395.
Nakatogawa H, Suzuki K, Kamada Y, Ohsumi Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat Rev Mol Cell Biol. 2009;10:458–467.
Mizushima N, Levine B. Autophagy in mammalian development and differentiation. Nat Cell Biol. 2010;12:823–830.
Mizushima N. Physiological functions of autophagy. Curr Top Microbiol Immunol. 2009;335:71–84.
Shibutani ST, Saitoh T, Nowag H, Munz C, Yoshimori T. Autophagy and autophagy-related proteins in the immune system. Nat Immunol. 2015;16:1014–1024.
Levine B, Packer M, Codogno P. Development of autophagy inducers in clinical medicine. J Clin Invest. 2015;125:14–24.
Gawriluk TR, Hale AN, Flaws JA, et al. Autophagy is a cell survival program for female germ cells in the murine ovary. Reproduction. 2011;141:759–765.
Hulas-Stasiak M, Gawron A. Follicular atresia in the prepubertal spiny mouse (Acomys cahirinus) ovary. Apoptosis. 2011;16:967–975.
Song ZH, Yu HY, Wang P, et al. Germ cell-specific Atg7 knockout results in primary ovarian insufficiency in female mice. Cell Death Dis. 2015;6:e1589.
Noda T. Autophagy in the context of the cellular membrane-trafficking system: the enigma of Atg9 vesicles. Biochem Soc Trans. 2017;45:1323–1331.
Nishi Y, Yanase T, Mu Y, et al. Establishment and characterization of a steroidogenic human granulosa-like tumor cell line, KGN, that expresses functional follicle-stimulating hormone receptor. Endocrinology. 2001;142:437–445.
Mouna L, Hernandez E, Bonte D, et al. Analysis of the role of autophagy inhibition by two complementary human cytomegalovirus BECN1/Beclin 1-binding proteins. Autophagy. 2016;12:327–342.
Taherbhoy AM, Tait SW, Kaiser SE, et al. Atg8 transfer from Atg7 to Atg3: a distinctive E1-E2 architecture and mechanism in the autophagy pathway. Mol Cell. 2011;44:451–461.
Komatsu M, Waguri S, Ueno T, et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol. 2005;169:425–434.
Saitoh T, Fujita N, Hayashi T, et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc Natl Acad Sci USA 2009;106:20842–20846.
Tilly JL. Commuting the death sentence: how oocytes strive to survive. Nat Rev Mol Cell Biol. 2001;2:838–848.
Findlay JK, Hutt KJ, Hickey M, Anderson RA. How is the number of primordial follicles in the ovarian reserve established? Biol Reprod. 2015;93:111.
Dong C, Wei P, Jian X, et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum Mol Genet. 2015;24:2125–2137.
Kuma A, Hatano M, Matsui M, et al. The role of autophagy during the early neonatal starvation period. Nature. 2004;432:1032–1036.
Choi J, Jo M, Lee E, Choi D. AKT is involved in granulosa cell autophagy regulation via mTOR signaling during rat follicular development and atresia. Reproduction. 2014;147:73–80.
Toniolo D. X-linked premature ovarian failure: a complex disease. Curr Opin Genet Dev. 2006;16:293–300.
Bouilly J, Bachelot A, Broutin I, Touraine P, Binart N. Novel NOBOX loss-of-function mutations account for 6.2% of cases in a large primary ovarian insufficiency cohort. Hum Mutat. 2011;32:1108–1113.
Pitteloud N, Quinton R, Pearce S, et al. Digenic mutations account for variable phenotypes in idiopathic hypogonadotropic hypogonadism. J Clin Invest. 2007;117:457–463.
Bouilly J, Beau I, Barraud S, et al. Identification of multiple gene mutations accounts for a new genetic architecture of primary ovarian insufficiency. J Clin Endocrinol Metab. 2016;101:4541–4550.
Nykamp K, Anderson M, Powers M, et al. Sherloc: a comprehensive refinement of the ACMG-AMP variant classification criteria. Genet Med. 2017;19:1105–1117.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Acknowledgements
We thank S. Elledge, X.M. Yin, and T. Finkel for providing us with shATG7, pEGFP-ATG9A, and pCMV-ATG7 constructs, respectively. We thank V. Nicolas for assistance in confocal microscopy (Plate-Forme d’imagerie cellulaire-MIPSIT, University Paris-Sud). This work was supported by the Institut National de la Santé et de la Recherche Médicale. Paul Laissue´s Lab is supported by the Universidad del Rosario (grant CS/CIGGUR/ABN062/2018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Delcour, C., Amazit, L., Patino, L.C. et al. ATG7 and ATG9A loss-of-function variants trigger autophagy impairment and ovarian failure. Genet Med 21, 930–938 (2019). https://doi.org/10.1038/s41436-018-0287-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0287-y
Keywords
This article is cited by
-
Endometrial stem cells alleviate cisplatin-induced ferroptosis of granulosa cells by regulating Nrf2 expression
Reproductive Biology and Endocrinology (2024)
-
A matter of new life and cell death: programmed cell death in the mammalian ovary
Journal of Biomedical Science (2024)
-
Advanced oxidation protein products attenuate the autophagy-lysosome pathway in ovarian granulosa cells by modulating the ROS-dependent mTOR-TFEB pathway
Cell Death & Disease (2024)
-
Mechanisms of ovarian aging in women: a review
Journal of Ovarian Research (2023)
-
BNIP3-mediated autophagy via the mTOR/ULK1 pathway induces primordial follicle loss after ovarian tissue transplantation
Journal of Assisted Reproduction and Genetics (2023)