Abstract
Purpose
With the advent of gene therapies for inherited retinal degenerations (IRDs), genetic diagnostics will have an increasing role in clinical decision-making. Yet the genetic cause of disease cannot be identified using exon-based sequencing for a significant portion of patients. We hypothesized that noncoding pathogenic variants contribute significantly to the genetic causality of IRDs and evaluated patients with single coding pathogenic variants in RPGRIP1 to test this hypothesis.
Methods
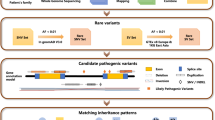
IRD families underwent targeted panel sequencing. Unsolved cases were explored by exome and genome sequencing looking for additional pathogenic variants. Candidate pathogenic variants were then validated by Sanger sequencing, quantitative polymerase chain reaction, and in vitro splicing assays in two cell lines analyzed through amplicon sequencing.
Results
Among 1722 families, 3 had biallelic loss-of-function pathogenic variants in RPGRIP1 while 7 had a single disruptive coding pathogenic variants. Exome and genome sequencing revealed potential noncoding pathogenic variants in these 7 families. In 6, the noncoding pathogenic variants were shown to lead to loss of function in vitro.
Conclusion
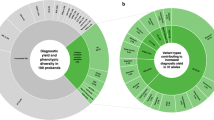
Noncoding pathogenic variants were identified in 6 of 7 families with single coding pathogenic variants in RPGRIP1. The results suggest that noncoding pathogenic variants contribute significantly to the genetic causality of IRDs and RPGRIP1-mediated IRDs are more common than previously thought.
Similar content being viewed by others
Introduction
Inherited retinal degenerations (IRDs) are a group of monogenic diseases that are the most common cause of blindness in the working age population.1 About 260 genes have been associated with IRDs with functions spanning almost every aspect of cellular function, from splicing machinery, to microtubular transport and phototransduction.1 State-of-the-art clinical diagnostics using next-generation sequencing (NGS) of known IRD genes successfully identifies the causal pathogenic variant (PV) in only 50 to 70% of cases.2,3 Although copy-number changes4 and intronic PV5 contribute to disease, they are not routinely assessed and likely contribute to the genetic causality in a significant portion of currently unsolved cases. With the advent of successful gene therapies for IRDs,6 understanding such noncoding PV and developing assays to evaluate them is of increasing importance. One example is the autosomal recessive RPGRIP1-associated disease, which is an attractive candidate for gene therapy with already established success in murine7 and canine8 models. Yet, almost all of the PV in RPGRIP1 have been described in the coding region.9
RPGRIP1 plays a critical role in opsin trafficking, outer-segment disc organization and photoreceptor survival.10,11 While it primarily localizes to the transition zone of rods and cones, various of its isoforms can be found in the outer segment, along the microtubules as well as in the amacrine cells of the inner plexiform layer.12,13 Its largest transcript variant, NM_020366, is composed of 3861 coding base pairs distributed over 24 exons.14 This encodes a 1287–amino acid protein that interacts with a variety of other IRD proteins such as retinitis pigmentosa GTPase regulator (RPGR), SPATA7, and NPHP415. The expression of RPGRIP1 is limited to the retina and testis.16
Confident genetic diagnosis with RPGRIP1 as the causal gene will be crucial for effective clinical trials of potential therapies. In our analysis of IRD families with targeted panel sequencing of coding regions of IRD-associated genes,17 we repeatedly noted identification of single likely pathogenic variants in RPGRIP1 in families without PV in other IRD disease genes. To test the hypothesis that PV in RPGRIP1 are the likely cause of disease in these families, we performed exome (ES) and genome sequencing (GS) to search for noncoding PV and structural variations accounting for the loss of function (LoF) of the second allele.
Materials and methods
Ethical guidelines
The study was approved by the institutional review board at the Massachusetts Eye and Ear (Human Studies Committee MEE in USA) and adhered to the Declaration of Helsinki. Informed consent was obtained from all individuals on whom genetic testing and further molecular evaluations were performed.
Clinical evaluation
All the patients in this study underwent clinical assessment by ophthalmologists subspecializing in inherited retinal degenerations. The clinical characteristics are outlined in Table 1.
Sequencing
DNA was extracted from venous blood using the DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany). All samples underwent GEDi sequencing as described previously.17 Exome and genome sequencing were done at the Center for Mendelian Genomics at the Broad Institute of MIT and Harvard using methodology described previously.18 Sanger sequencing was performed on ABI 3730xl using BigDye Terminator v3.1 kits (Life Technologies, Carlsbad, CA) and using polymerase chain reaction (PCR) primers indicated in Supplementary Information. When PCR products were sequenced, they were purified prior to sequencing (ExoSap-IT, Affymetrix, Santa Clara, CA). Gel bands that were Sanger sequenced had DNA extracted via the ZymocleanTM Gel DNA Recovery Kit (Zymo Research, Irvine, CA).
Bioinformatics
Analyses of DNA sequence data were performed as described previously.17,19 Briefly, Burrows–Wheeler Aligner (BWA) was used for alignment. SAMtools and custom programs were used for single-nucleotide polymorphism and insertion/deletion calls.19 Variants of interest were limited to polymorphisms with less than 0.005 allelic frequency in the gnomAD (http://gnomad.broadinstitute.org/) and ExAC (http://exac.broadinstitute.org/) databases.18 Genome copy-number analysis, with consideration of structural changes, was done using Genome STRiP 2.0 (ref. 20). For the analysis of splicing patterns from amplicon sequencing, STAR (version 2.5.3a) aligner21 was used to generate an index of the human genome (GRCh37.75.dna.primary_assembly.fa) and to align the reads. Integrative Genomics Viewer (IGV)22 was used to load the aligned sequences (BAM files) and for data visualization with Sashimi plots.
PCR, cloning, and site-directed mutagenesis
PCR was performed using PfuUltra II Fusion polymerase (Agilent Technologies, Santa Clara, CA) on genomic DNA of patients harboring the PV of interest (primers are listed in the Supplementary Material). The PCR products were cloned into pENTR Directional TOPO vector (Thermo Fisher, Waltham, MA) and used to transform chemically competent Escherichia coli (One Shot TOP10, Thermo Fisher, Waltham, MA). Plasmid DNA from single colonies was extracted with miniprep kits (ZymoPURE, Zymo Research) and analyzed by restriction enzyme digestion with BsrGI (NE Biolabs, Ipswich, MA) and Sanger sequencing. Essential splice-site PV were introduced by site-directed mutagenesis (QuickChange II Site Directed mutagenesis kit, Agilent Technologies) and verified by Sanger sequencing. Colonies with the correct sequence and restriction enzyme pattern were then subcloned into the pCS2 + GW vector (kind gift from Erica Davis) via Gateway LR clonase II (Thermo Fisher) and similar analyses as before was done to isolate vectors with the appropriate inserts for transfection experiments. The final vector included RPGRIP1 exons 11–16 including extensions into introns 10 and 16 on the 5′ and 3′ ends, which was cloned into pCS2 + GW and used for splicing assays.
Quantitative polymerase chain reactions (qPCR)
Five nanograms (ng) of genomic DNA, 200 nM of each primer, and 10 µl of Fast SYBR Green Master Mix (Life Technologies, Grand Island, NY) were used for qPCR reactions, which were performed on a Stratagene Mx3000P instrument (Agilent Technologies) using the standard thermocycling program (95 °C for 3 min, 40 cycles of 95 °C for 20 s, and 60 °C for 1 min, followed by a melting curve). The ddCT method was used for the analysis of results where ZNF80 was used as a reference gene and an in-house DNA sample with wild-type RPGRIP1 (OGI081-200) used for normalization. Each sample was tested in triplicate and the average value was used. Standard deviation with error propagation was used to calculate up and down errors.
Cell culture and transfections
Human embryonic kidney (HEK293T) and retinoblastoma (WERI-Rb1) cells purchased from American Type Culture Collection (ATCC, Manassas, VA) and maintained in RPMI medium supplemented with 10% fetal bovine serum 1640 (Thermo Fisher). Then, 2 ml of 5 × 105 cells/ml were plated into each well of a 6-well plate (Corning Inc., Corning, NY) 12 h prior to transfections; 1–5 μg of vector DNA per well was used for transfections using a commercial reagent (Lipofectamine 2000, Invitrogen, Carlsbad, CA). Cells were harvested for RNA extraction 48 h after transfection.
RNA isolation and complementary DNA synthesis
Cells were lysed with TRIzol (Thermo Fisher). After 1-Bromo-3-chloropropane or chloroform (Sigma-Aldrich, St. Louis, MO) treatment, the aqueous phase was transferred to mRNeasy columns with DNase I digestion performed on-column (Qiagen). Quantification was performed via NanoDrop (Thermo Fisher) and 500 ng of RNA was converted to complementary DNA (cDNA) using oligo(dT) primers and SuperScript II (Thermo Fisher).
Splicing assay and amplicon sequencing
The mutant, control, and wild-type vectors described above were transfected into HEK293T and WERI-Rb1 cells. Two days posttransfection, cDNA was generated as described and reverse transcription polymerase chain reaction (RT-PCR) performed amplifying the flanking exons of the point PV of interest. The PCR products were purified by DNA-clean and concentrator kits (Zymo Research). Amplicon sequencing was then performed at the Massachusetts General Hospital Center for Computational and Integrative Biology, where the PCR products were fragmented using sonication and sequenced with the standard NGS pipeline. Visualization and analysis of the data were performed as described under bioinformatics.
Epigenetic features surrounding the RPGRIP1 locus
ATAC-Seq and ChIP-Seq were performed according to previously published methods.23,24 Briefly, human retinal tissue was obtained from donors 25–65 years old with a postmortem interval <8 h (Lions Vision Gift, Portland, OR). Approximately 20,000 nuclei were isolated for ATAC-Seq and the transposition reaction was performed for 60 min at 37 °C. ChIP-Seq was performed on approximately 25 million cells per reaction using the following antibodies: CTCF (Abcam AB70303); H3K4me2 (Abcam AB7766 lot GF160184-1; RRID AB_2560996); H3K27ac (Abcam AB4729, lot GF150367-1; RRID AB_2118291); CRX (Santa Cruz, B11X, lot E1409); OTX2 (Abcam, AB21990, lot GR242019-1); NRL (Abcam, AB137193, lot GR104520-2); RORB (Diagenode, pAb-001-100, lot HS-0010); MEF2D (Greenberg lab, 2373). All sequencing was performed on an Illumina NextSeq500 to a depth of >10 M reads. Mapping, alignment, and normalization of reads was performed as previously described.24 Genome tracks were displayed using the University of California–Santa Cruz Genome Browser.
Results
Genetic analysis of 1722 IRD probands who underwent targeted exon sequencing of known IRD genes17 revealed 3 patients with biallelic loss-of-function (LoF) PV in RPGRIP1 and 7 with only one LoF change in this gene (Table 1, Fig. S1). In the latter 7 families no other significant PV in RPGRIP1 or other IRD genes were identified. Because they were all diagnosed with an early-onset IRD, a characteristic presentation of RPGRIP1 disease,25 further testing was performed to search for noncoding and structural variants in RPGRIP1 or other IRD genes through exome and genome sequencing. Among these 7 families, three of the second-allele PV were predicted to be copy-number changes and three were intronic PV (Table 1). In one family, OGI-578, we did not validate second pathogenic variants in RPGRIP1 (Fig. S1).
ES and GS studies showed that three families have structural variants as the second PV in RPGRIP1. Analysis of ES data showed that patient OGI-1797-3128 had a predicted deletion of exon 19 (Table 1), which was confirmed by qPCR (Fig. S2). Exon 19 has 139 nucleotides, and thus its deletion is predicted to lead to a frameshift resulting in a premature stop codon and likely subsequent nonsense-mediated decay (NMD).26 In OGI-281-608, the gain of a copy of exon 2 was detected through structural analysis20 in addition to coverage-based predictions of GS data (Fig. 1). The structural change led to misalignment of paired-end reads (Fig. 1c, d), which indicated a tandem duplication. This PV was validated by qPCR (Fig. 1b), and the predicted breakpoint confirmed through PCR and Sanger sequencing (Fig. 1e, f). Sanger sequencing also identified 135 bp of missing DNA upstream of the breakpoint, suggesting a possible complex rearrangement as the causal event.27 Given that exon 2 is 133 bp, its tandem duplication would lead to a LoF allele.
Exon 2 duplication in OGI-281. (a) Pedigree of the family showing deceased parents and the three siblings all of whom were analyzed. (b) Quantitative polymerase chain reaction (qPCR)-based copy-number results along the first three exons of RPGRIP1. All three siblings have a duplication of exon 2 in an RPGRIP1 allele. Exons 1 and 3 are not affected. The bottom panel shows the locations of RPGRIP1 exons based on the NM_020366 transcript. (c) Integrative Genomics Viewer22 (IGV) view of the sequenced genome sequencing (GS) reads where the duplication was discovered for OGI-281-608. The bottom of the figure shows the location of exon 2 of RPGRIP1. The gray thick arrows correspond to expected paired-end reads. The green thick arrows are mapped reads that have aligned abnormally and hint to a PV. (d) Schematic explanation of the how genomic duplication would lead to the abnormal paired-end reads seen in (c). The gray region is the area of hypothetical duplication, while the green thick arrows are the paired-end reads that will align abnormally. The top of the figure shows what is actually sequenced in the mutant sample, while the bottom shows how such sequenced reads would map to a reference wild-type (WT) model. The aligned paired-end reads of the mutant will have a greater distance between them and will point away from one another as seen in (c). The dark and light green hash lines correspond to the aligning sequenced of the paired-end reads. The primers used for (e) are shown as blue and red arrows. Their directionality is indicated relative to the mutant (top) and WT models (bottom). (e) Polymerase chain reaction (PCR) across the predicted duplication breakpoint using primers represented in (d). Presence of a tandem duplication would yield a product while its absence would lead to no amplification as the primers would be pointing away from each other. The predicted duplication is present in all OGI-281 family members while it is absent in HEK293T cells. The control (Cntrl) PCR on the bottom was done to ensure that larger products could be amplified from all samples thus ensuring DNA fragmentation or quality was not a confounding factor. (f) Sanger sequencing identifying the exact breakpoint (black arrowhead) using OGI-281-608 PCR product from (e). There is a 135-bp deletion, upstream of the breakpoint.
Patient OGI-237-523 similarly had a tandem copy-number gain but in both exons 1 and 2 detected by ES (Fig. 2a, b). The 5′ breakpoint was mapped 2 Kb upstream of exon 1 (Figs. 2c, S3). However, given that the second copy would maintain 2 Kb upstream of the exon 1, which would include a proximal presumed promoter (Fig. 2c), we questioned whether a second transcriptional start site (TSS) within the duplicated 5′ upstream region would exclude the mutant exons 1′ and 2′ thus leading to normal transcription. Therefore, we hypothesized that perhaps a critical RPGRIP1 regulatory domain exists outside of the 2-Kb region upstream of exon 1. Review of transcriptome data of normal human retina28 revealed an additional exon upstream of the annotated transcript, which was identified by numerous split reads as well as high sequence coverage approximately 8 Kb upstream of the annotated RPGRIP1 exon 1 (Fig. 2d). We further assessed this using ATAC-seq and ChIP-seq data, which showed open chromatin and transcription factor binding to this region and not to the previously annotated RPGRIP1 TSS (Fig. 2e). These results suggest that the retinal TSS of RPGRIP1 and its regulatory regions are 8 Kb upstream of the currently annotated exon 1. This also explains the likely pathogenic effect of the structural PV seen in OGI-237-523. The tandem exon 1 and 2 duplication excludes a TSS and will lead to a premature stop codon 29 codons past the start of the duplicated 5′UTR of exon 1.
(a) Pedigree of OGI-237 showing the segregation of the RPGRIP1 PV in the family. (b) Quantitative polymerase chain reaction (qPCR) of OGI-237 family members, showing the presence of duplication of both exons 1 and 2 in the mother (1776) and the proband (523) while the father (1777) has a normal copy number across these exons. The bottom panel shows the locations of RPGRIP1 exons based on the NM_020366 transcript. (c) Representation of the predicted wild-type (WT) and mutant alleles with the duplication of exons 1 and 2 (M4). Arrowed lines are introns, tall bars are exons, and short bars indicate untranslated regions (UTR). Further analysis (Fig. S3), shows that M4 is a result of tandem duplication with a breakpoint at ~2 Kb upstream of exon 1 (black arrowhead). If the transcriptional start site (TSS) is within this 2-Kb region, then M4 could lead to normal transcripts given uninterrupted exon 1 and its upstream region. (d) Exploration of the retina transcriptome28 shows, however, that there is an additional novel exon (1n) upstream of the currently annotated exon 1 in the canonical NM_020366 transcript model. Thus the actual TSS is expected to be upstream of exon 1n rather than exon 1. The red bars are indicative of read depth of the transcriptome data. The green and blue arrowed lines at the top indicate split reads between exons. The green are across unannotated exons, while the blue corresponds to annotated exons. There are 550 split reads between exons 1n and 1, further confirming the presence of this novel exon. The NM_020366 canonical transcript model is shown at the bottom. The gray and light blue highlighted areas corresponded to the gray and light blue areas of (e) respectively. (e) ATAC-Seq and ChIP-Seq from adult human retina of histone modifications and transcription factor binding at the RPGRIP1 locus as in (d). The light blue shading represents the area directly upstream of the annotated RPGRIP1 transcript. The light gray shading represents a promoter region and newly discovered retina-specific exon suggested by RNA-Seq, ATAC-Seq, and ChIP-Seq. The transcript models of RPGRIP1 are shown on top
Three families with single PV identified in RPGRIP1 by panel-based genetic testing had deep intronic PV that segregated in the families and had low frequencies in the gnomAD database18 (Table 1). We hypothesized that these PV could potentially lead to aberrant splicing creating LoF alleles as well. To assess the effect of these changes on splicing, midi-gene assays were performed in HEK293T cells with mutant and wild-type (WT) constructs, as well as essential splice-site changes serving as positive controls. In OGI-827-1591, the c. 2367+23delG change led to a significant retention of intron 15 in mutant versus wild type comparable to the findings in the c. 2367+1G>A positive control (Fig. 3a). While the ratio of splicing between exons 15 and 16 to that between exons 14 and 15 was 6090/6207 or 0.98 in wild type, this proportion was altered to 2209/3982 or 0.55 in mutant. This suggests a relative reduction of 43% of the appropriate splicing at this locus. The aberrant splice isoform leads to a premature stop codon 19 codons into intron 15.
Sashimi plots of the amplicon sequencing of splicing assays. The Sashimi plots are generated using amplicon sequencing on reverse transcription polymerase chain reaction (RT-PCR) products across the indicated regions in HEK293T cells that were transfected with wild type (WT) (red), positive control (blue), or mutant (green) RPGRIP1 vectors. The annotated NM_020366 exons are indicated on the bottom of each plot in dark blue. (a) In OGI-827, the intronic PV caused increased intron retention (green), similar to the essential splice-site PV control (blue). (b) In OGI-601, as described in the text, the intronic mutation shifts splice to extend exon 12 (green), as does the essential splice-site PV control (blue). Interestingly, the transcript variant XM_005267881.3 has been reported to include this extension in its exon 1, which is entirely untranslated. Untranslated and translated regions of the transcript models are indicated as thin and thick dark blue lines respectively. (c) In OGI-949, the deep intronic PV leads to inclusion of a cryptic exon 11′ in the mutant transcript (green). The dashed lines highlight the immediate upstream sequence of the exon 11′ showing the start of the cryptic exon (green) 13 bp downstream of the PV (red). The fourth codon of 11′ is a stop codon (underlined). M11 was a de novo PV in patient OGI-949-1907, confirmed via Sanger sequencing
In OGI-601-1236, the c.1611+27G>A PV led to a splicing pattern that became apparent only through amplicon sequencing (Figs. 3b, S4, S5). In the WT allele, three splicing signals of various strengths were detected between exons 12 and 13 (Fig. 3b). The two prominent splice variants have been annotated for RPGRIP1, the main event (9598 reads) being present in the NM_020366 transcript and indicating the canonical splice junction between exons 12 and 13. The second splice donor occurring 104 bp into intron 12 (8499 reads) is part of the 5′UTR of predicted transcript model XM_005267881.3 (Fig. 3b). The third splice donor site occurs 227 bp into intron 12 and has not been reported. In both the positive control, c.1611+1G>A, and the PV under study, c.1611+27G>A, the primary splicing event that would lead to full-length protein failed to occur whereas the alternate transcript donor site, 104 bp past the NM_020366 exon 12, was preferred (Fig. 3b). This variant would lead to a premature stop codon 12 codons after the exon 12 of the full-length transcript (i.e., NM_020366).
In OGI-949-1907, the intronic c.1468-263G>C variant resulted in an activation of a cryptic splice acceptor site 13 bp past the PV (Fig. 3c). The resultant cryptic exon 11′, although 117 bp and in-frame, has a premature stop codon four codons past the cryptic acceptor site. Thus the aberrant transcript will lead to NMD. The second PV in OGI-949-1907 was a de novo deletion of an essential splice site, c.3618-1_3621del5 (M11), confirmed by Sanger sequencing of the proband and other family members.
To account for potential differences in splicing between the neuronal and nonneuronal cell types,29 all the splicing experiments were repeated with a retinoblastoma cell line WERI-Rb1, derived from photoreceptor cells.30 The splicing changes resulting from the PV assessed led to similar patterns in both HEK293T as well as WERI-Rb1 lines (Fig. S5). Overall, we found four novel noncoding PV, four unreported coding PV in RPGRIP1 (Fig. 4), and we have corrected the transcript model for RPGRIP1 in the retina with a new TSS and 5′ exon 8 kb upstream of the annotated exon 1 (named exon 1n in this paper).
Among the seven families with one LoF RPGRIP1 allele, OGI-578 did not reveal a second RPGRIP1 PV after GS and copy-number analyses. None of the RPGRIP1 calls showed proper segregation or were of poor quality and failed Sanger validation. GS, however, revealed another IRD candidate gene, the most prominent of which was CNGA1, with biallelic high-quality calls.
Discussion
In our analysis of nine families with biallelic RPGRIP1 PV, six had a second noncoding PV leading to a LoF allele. This not only highlights the importance of noncoding PV in pathogenesis of recessive IRDs, but also implies a greater prevalence of RPGRIP1-mediated disease than previously thought. Three among the 1722 IRD families had bilallelic coding PV in RPGRIP1, which corresponds roughly to the previously reported rates in the literature (0.17 vs. 0.25% [ref. 14,31]). However, when taking into account noncoding PV, our rate increased from 0.17 to 0.51%. This significant increase in frequency of RPGRIP1-mediated disease is still likely an underestimate given the bias of choosing samples with one LoF coding sequence PV. Considering the possibility of biallelic noncoding LoF alleles, the rate of RPGRIP1-mediated disease is likely to be much greater than what was previously thought.
In the six families solved with the addition of structural and intronic PV analyses, we validated the second LoF PV in vitro showing aberrations in the reading frame caused by splicing or copy-number changes. While deletions are simpler to interpret given clear interruption of reading frame, copy-number gains can be just as disruptive. In both cases of copy-number gain, OGI-237 and OGI-281, the PV were tandem duplications leading to premature stop codons. While investigating the potentially pathogenic effect of the duplication in the first two annotated RPGRIP1 exons in OGI-237, we discovered a novel exon and a retinal TSS 8 Kb upstream of the previously annotated transcriptional start site. This is important as PV in regulatory regions and 5′UTR have been shown to be disease-causing, such as in the case of PRPF31 and OPN1MW.9 We propose the inclusion of this RPGRIP1 exon, chr14:21748265-21748318 (hg19), in future IRD diagnostic panels.
We also detected three PV causing intron retentions or inclusion of cryptic exons in the resulting transcripts, two of which were in the flanking 30 bp of annotated exons. The traditional way of assessing for the effect of intronic PV includes use of exon-trapping vectors with a small (mini-gene) or medium (midi) sized insert harboring the PV under study.32 Subsequent transfection into HEK293T cells, and analysis of the processed RNA via RT-PCR, allows one to assess the alterations via gel electrophoresis. However, as seen in the case of OGI-601 (Fig. 3b, S4b), there can be a multitude of splicing signals, some of which are not annotated for the genes of interest thus making the interpretation of the electrophoresis results difficult. We found that amplicon sequencing of the RT-PCR product can increase the sensitivity of detection (OGI-601, Figs. 3b, S4b) and also add a quantitative value to assessment of splicing changes. Additionally, the PV c.2367+23delG, had been noted previously3 as a potential disease-causing and splicing altering PV. Yet its precise effect proved challenging to interpret, while the authors suggested a possible exon 15 and 16 skipping in blood messenger RNA (mRNA). Through amplicon sequencing (Figs. 3a, S5) we found evidence of intron 15 retention in both HEK293T and the tissue-relevant WERI-Rb1 line (Fig. S5). Amplicon sequencing analysis clarifies the exact splicing events and proportions thus offering an advantage in interpreting results. It should be noted, however, that we cannot rule out exon 15–16 skipping as suggested by Riera et al.3 because exon 17 was not part of the cloned RPGRIP1 used in the splicing assay. Weakening of splicing between exons 15 and 16 with this PV is supportive of both models.
Splicing is a tissue-specific phenomenon and neuronal tissues have a unique splicing machinery that can lead to unique exonic retentions.29 Thus the splicing patterns we identified in HEK293T cells were confirmed in a retina-relevant line. We chose WERI-Rb1, a retinoblastoma line shown to possess the neuronal specific splicing machinery.29 Although we did not detect a significant difference between the two lines (Fig. S5), we foresee a benefit in using a tissue-relevant cell line, such as WERI-Rb1 for IRDs, when assessing splicing PV.
Despite the decreasing costs of genome sequencing, it is still not feasible for routine clinical assessment. Hence, assessing for noncoding PV in RPGRIP1 and other recessive IRD genes still demands a step-wise approach to reduce screening costs. Our recommendation is to include and analyze at least 30 bp of the flanking exonic regions and to include copy-number analysis4 in targeted panel sequencing of recessive IRD genes. In this study, we noted only one significant PV outside of these criteria (c.1468-263G>C). The remaining unsolved cases can then be submitted for genome sequencing to identify novel genes and PV.
The effect of deep intronic PV on splicing and their contribution to disease has been noted by others.32,33,34 For example, noncanonical splice-site PV in ABCA4 have been thoroughly studied by midi-gene splicing assays and showed pathogenicity of multiple variants of previously unknown significance.32,34 Another approach of proving pathogenicity was used by Cumming et al. in muscular dystrophies, where genome sequencing coupled with RNA sequencing of the affected tissue was used to detect rare variants leading to splicing aberrations.33 This method increased the diagnostic rate in rare muscular diseases by 25%33.
As demonstrated above, the empirical validation of second-mutant alleles in recessive diseases when one mutant allele is identified can be time consuming and expensive. One might argue that such stringency may not be necessary if the gene identified with a single PV matches the presenting phenotype. Statistical models predict a small false positive diagnosis rate when a single PV is identified in a disease-specific gene such as MYO7A in Usher syndrome type I.35 Yet when a large number of potential genes can lead to the phenotype of interest, such as in early-onset IRD, the calculations will not yield as small a false positive rate.35 Thus we strongly believe that to make the correct diagnosis, it is important to identify and characterize both mutant alleles.
Accurate genetic diagnostics for inherited retinal degenerations is increasingly critical with the emergence of gene therapy. Clinical trials of gene augmentation therapy for RPE65-associated retinal degeneration have been completed, and the treatment was recently approved by the US Food and Drug Administration (FDA).6 Further, clinical trials of gene therapies for eight other genetic forms of retinal degeneration are in progress.36,37,38,39,40 Successful preclinical studies of gene therapies for multiple other genetic forms of IRD have been reported, including for RPGRIP1-associated IRD.7,8 Evidence of biallelic gene PV are critical for inclusion of patients in trials when dealing with autosomal recessive diseases. We believe the results reported here are also relevant for other genetic forms of IRD, and that genetic testing that incorporates detection of noncoding and structural PV will increase the diagnostic sensitivity for all IRDs.
References
Farrar GJ, Carrigan M, Dockery A, et al. Toward an elucidation of the molecular genetics of inherited retinal degenerations. Hum Mol Genet. 2017;26:R2–R11. R1
Ellingford JM, Barton S, Bhaskar S, et al. Molecular findings from 537 individuals with inherited retinal disease. J Med Genet. 2016;53:761–767.
Riera M, Navarro R, Ruiz-Nogales S, et al. Whole exome sequencing using ion proton system enables reliable genetic diagnosis of inherited retinal dystrophies. Sci Rep. 2017;7:42078.
Bujakowska KM, Fernandez-Godino R, Place E, et al. Copy-number variation is an important contributor to the genetic causality of inherited retinal degenerations. Genet Med. 2017;19:643–651.
Liquori A, Vache C, Baux D, et al. Whole USH2A gene sequencing identifies several new deep intronic mutations. Hum Mutat . 2016;37:184–193.
Russell S, Bennett J, Wellman JA, et al. Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet. 2017;390:849–860.
Pawlyk BS, Bulgakov OV, Liu X, et al. Replacement gene therapy with a human RPGRIP1 sequence slows photoreceptor degeneration in a murine model of Leber congenital amaurosis. Hum Gene Ther. 2010;21:993–1004.
Lhériteau E, Petit L, Weber M, et al. Successful gene therapy in the RPGRIP1-deficient dog: a large model of cone–rod dystrophy. Mol Ther. 2014;22:265–277.
Institute of Medical Genetics. Human Gene Mutation Database (HGMD) [database online]. Cardiff, UK: Cardiff University; 2017. http://www.hgmd.cf.ac.uk/ Accessed 23 July 2018.
Shu X, Fry A, Tulloch B, et al. RPGR ORF15 isoform co-localizes with RPGRIP1 at centrioles and basal bodies and interacts with nucleophosmin. Hum Mol Genet. 2005;14:1183–1197.
Zhao Y, Hong D-H, Pawlyk B, et al. The retinitis pigmentosa GTPase regulator (RPGR)-interacting protein: subserving RPGR function and participating in disk morphogenesis. Proc Natl Acad Sci U S A. 2003;100:3965–3970.
Mavlyutov TA, Zhao H, Ferreira PA. Species-specific subcellular localization of RPGR and RPGRIP isoforms: implications for the phenotypic variability of congenital retinopathies among species. Hum Mol Genet. 2002;11:1899–1907.
Castagnet P, Mavlyutov T, Cai Y, et al. RPGRIP1s with distinct neuronal localization and biochemical properties associate selectively with RanBP2 in amacrine neurons. Hum Mol Genet. 2003;12:1847–1863.
Gerber S, Perrault I, Hanein S, et al. Complete exon-intron structure of the RPGR-interacting protein (RPGRIP1) gene allows the identification of mutations underlying Leber congenital amaurosis. Eur J Hum Genet. 2001;9:561.
Patnaik SR, Raghupathy RK, Zhang X, et al. The role of RPGR and its interacting proteins in ciliopathies. J Ophthalmol . 2015;2015:414781.
Boylan JP, Wright AF. Identification of a novel protein interacting with RPGR. Hum Mol Genet. 2000;9:2085–2093.
Consugar MB, Navarro-Gomez D, Place EM, et al. Panel-based genetic diagnostic testing for inherited eye diseases is highly accurate and reproducible and more sensitive for variant detection than exome sequencing. Genet Med. 2015;17:253.
Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291.
Falk MJ, Zhang Q, Nakamaru-Ogiso E, et al. NMNAT1 mutations cause Leber congenital amaurosis. Nat Genet. 2012;44:1040–1045.
Handsaker RE, Van Doren V, Berman JR, et al. Large multiallelic copy number variations in humans. Nat Genet . 2015;47:296–303.
Dobin A, Davis CA, Schlesinger F, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013;29:15–21.
Robinson JT, Thorvaldsdóttir H, Winckler W, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–26.
Buenrostro JD, Giresi PG, Zaba LC, et al. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods. 2013;10:1213–1218.
Andzelm MM, Cherry TJ, Harmin DA, et al. MEF2D drives photoreceptor development through a genome-wide competition for tissue-specific enhancers. Neuron. 2015;86:247–263.
Li T. Leber congenital amaurosis caused by mutations in RPGRIP1. Cold Spring Harb Perspect Med. 2015;5:a017384.
Baker KE, Parker R. Nonsense-mediated mRNA decay: terminating erroneous gene expression. Curr Opin Cell Biol . 2004;16:293–299.
Usher CL, McCarroll SA. Complex and multi-allelic copy number variation in human disease. Brief Funct Genomics. 2015;14:329–338.
Farkas MH, Grant GR, White JA, et al. Transcriptome analyses of the human retina identify unprecedented transcript diversity and 3.5 Mb of novel transcribed sequence via significant alternative splicing and novel genes. BMC Genomics. 2013;14:486.
Calarco JA, Superina S, O’Hanlon D, et al. Regulation of vertebrate nervous system alternative splicing and development by an SR-related protein. Cell. 2009;138:898–910.
McFall RC, Sery TW, Makadon M. Characterization of a new continuous cell line derived from a human retinoblastoma. Cancer Res. 1977;37:1003–1010.
Dryja TP, Adams SM, Grimsby JL, et al. Null RPGRIP1 alleles in patients with Leber congenital amaurosis. Am J Hum Genet. 2001;68:1295–1298.
Sangermano R, Bax NM, Bauwens M, et al. Photoreceptor progenitor mRNA analysis reveals exon skipping resulting from the ABCA4 c.5461-10T-->C mutation in Stargardt disease. Ophthalmology. 2016;123:1375–1385.
Cummings BB, Marshall JL, Tukiainen T, et al. Improving genetic diagnosis in Mendelian disease with transcriptome sequencing. Sci Transl Med. 2017;9:eaal5209.
Albert S, Garanto A, Sangermano R, et al. Identification and rescue of splice defects caused by two neighboring deep-intronic ABCA4 mutations underlying Stargardt disease. Am J Hum Genet . 2018;102:517–527.
Kimberling WJ. Estimation of the frequency of occult mutations for an autosomal recessive disease in the presence of genetic heterogeneity: application to genetic hearing loss disorders. Hum Mutat . 2005;26:462–470.
Zallocchi M, Binley K, Lad Y, et al. EIAV-based retinal gene therapy in the shaker1 mouse model for Usher syndrome type 1B: development of UshStat. PLoS One. 2014;9:e94272.
Banin E, Gootwine E, Obolensky A, et al. Gene augmentation therapy restores retinal function and visual behavior in a sheep model of CNGA3 achromatopsia. Mol Ther. 2015;23:1423–1433.
Komáromy AM, Alexander JJ, Rowlan JS, et al. Gene therapy rescues cone function in congenital achromatopsia. Hum Mol Genet. 2010;19:2581–2593.
Bainbridge JW, Smith AJ, Barker SS, et al. Effect of gene therapy on visual function in Leber’s congenital amaurosis. N Engl J Med. 2008;358:2231–2239.
Maguire AM, Simonelli F, Pierce EA, et al. Safety and efficacy of gene transfer for Leber’s congenital amaurosis. N Engl J Med. 2008;358:2240–2248.
Acknowledgements
This work was supported by grants from the National Eye Institute (RO1EY012910 [EAP], R01EY026904 [KMB/EAP], and P30EY014104 [MEEI core support]), and the Foundation Fighting Blindness (USA, EAP). Sequencing and analysis was provided by the Center for Mendelian Genomics at the Broad Institute of MIT and Harvard and was funded by the National Human Genome Research Institute, the National Eye Institute, and the National Heart, Lung, and Blood Institute grant UM1 HG008900 to Daniel MacArthur and Heidi Rehm. The authors would like to thank the patients and their family members for their participation in this study and the Ocular Genomics Institute Genomics Core members for their experimental assistance. The authors would like to thank the Exome Aggregation Consortium, the Genome Aggregation Database (gnomAD), and the groups that provided exome variant data for comparison. A full list of contributing groups can be found at http://exac.broadinstitute.org/about and http://gnomad.broadinstitute.org/about.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Jamshidi, F., Place, E., Mehrotra, S. et al. Contribution of noncoding pathogenic variants to RPGRIP1-mediated inherited retinal degeneration. Genet Med 21, 694–704 (2019). https://doi.org/10.1038/s41436-018-0104-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0104-7