Abstract
Purpose
We evaluated genome sequencing (GS) as an alternative to multigene panel sequencing (PS) for genetic testing in dilated cardiomyopathy (DCM).
Methods
Forty-two patients with familial DCM underwent PS and GS, and detection rates of rare single-nucleotide variants and small insertions/deletions in panel genes were compared. Loss-of-function variants in 406 cardiac-enriched genes were evaluated, and an assessment of structural variation was performed.
Results
GS provided broader and more uniform coverage than PS, with high concordance for rare variant detection in panel genes. GS identified all PS-identified pathogenic or likely pathogenic variants as well as two additional likely pathogenic variants: one was missed by PS due to low coverage, the other was a known disease-causing variant in a gene not included on the panel. No loss-of-function variants in the extended gene set met clinical criteria for pathogenicity. One BAG3 structural variant was classified as pathogenic.
Conclusion
Our data support the use of GS for genetic testing in DCM, with high variant detection accuracy and a capacity to identify structural variants. GS provides an opportunity to go beyond suites of established disease genes, but the incremental yield of clinically actionable variants is limited by a paucity of genetic and functional evidence for DCM association.
Similar content being viewed by others
Introduction
Dilated cardiomyopathy (DCM) is a common heritable heart muscle disorder that frequently has a genetic etiology.1 Although long lists of disease-associated genes have been compiled,1 genetic testing yields positive results in relatively few individuals and has not been recommended as part of routine patient care.2 Knowing genotype status has enormous potential benefit for families, permitting tailored surveillance strategies and early detection of individuals at risk.3 The lack of results for most DCM families represents an unmet clinical need and a major roadblock for implementation of personalized therapy.
In recent years, next-generation sequencing has facilitated genetic testing by enabling high-throughput evaluation of multiple genes, including underinvestigated large genes with hundreds of coding exons. Available methods include multigene panel sequencing (PS), exome sequencing (ES), and genome sequencing (GS). PS uses libraries enriched for protein-coding regions of disease-associated genes. There is generally high sequence coverage and comparative studies with Sanger sequencing have found excellent reproducibility for variant detection.4 PS is widely used by clinical diagnostic laboratories with cardiomyopathy panel sizes increasing over time from <20 genes to >100 genes.5, 6 The finite number of genes is a major limitation, necessitating redesign of panels as new disease genes are discovered and additional costs if a second test is required in PS-negative cases. In contrast to PS, ES is not limited to specific genes and looks at all protein-coding sequences.7 Variability in breadth of coverage is a potential limiting factor for ES-based clinical testing and may necessitate extensive follow-up Sanger sequencing to fill in gaps, particularly for high-probability disease genes. In addition to protein-coding sequences, GS uniquely provides information about the vast tracts of noncoding sequences that are increasingly implicated in human disease and enables high-resolution structural variant (SV) detection.8, 9 However, because GS typically has a lower overall sequencing depth than PS and ES, its potential sensitivity for pathogenic variant (PV) screening has been questioned.10 Golbus and colleagues11 have recently reported promising results for GS in a pilot study of 11 DCM patients. A detailed appraisal of the role of GS for DCM genetic testing is now timely and warranted.
Here we compare PS and GS in a cohort of patients with familial DCM. We determined the concordance of PS and GS for rare variant detection, evaluated loss-of-function (LOF) variants in an extended gene panel, and performed the first comprehensive evaluation of SVs in DCM. Our data show that GS is a reliable method for screening established DCM disease genes as well as providing a wealth of sequence information for ongoing data mining in “unsolved” cases.
Materials and methods
Study subjects
Forty-two patients (19 [45%] males), aged 18 to 82 (mean 50) years with familial DCM were recruited from St Vincent’s Hospital and by referral from collaborating physicians. The clinical characteristics of study probands are provided in Supplementary Table S1. Familial DCM was defined by the presence of DCM and/or early (<35 years) sudden unexplained death in two or more individuals in the absence of another heritable cardiac or systemic cause. Probands and participating first-degree relatives provided informed written consent and were evaluated by history and physical examination, electrocardiogram (ECG), and transthoracic echocardiography. All study subjects were of self-reported European ancestry. Protocols were approved by St Vincent’s Hospital Human Research Ethics Committee.
DNA sequencing and variant calling
See Supplementary Methods for expanded sequencing methods. Briefly, 42 patient DNA samples were newly sequenced by GS, or previously sequenced using a custom capture panel for 67 or 69 DCM genes.12 ES data were generated in-house from the NA12878 cell line (Coriell Institute for Medical Research, Camden, NJ) using the SureSelectXT Human AllExon V5 ([SSv5], Agilent Technologies, Santa Clara, CA, n = 1) and the Clinical Research Exome V2 ([CREv2], Agilent, n = 13) capture kits. We also reanalyzed published data from samples that used SureSelectXT Human AllExon V6 ([SSv6], Agilent, n = 6) capture kit.13 All genomic data, including previously published data, were analyzed using a GATK best practices analysis pipeline.14 Short variants were annotated, filtered and prioritized using Seave.15 Structural variants (SVs) including copy-number variants were identified using ClinSV (Minoche et al., manuscript in preparation) which uses a combination of discordantly mapping read pairs, split-mapping reads, and depth of coverage changes. A genomic position was defined as “covered” if the sequencing depth had ≥15 high-quality reads.16 Selected variants were confirmed in probands and evaluated in family members using Sanger sequencing and/or polymerase chain reaction (PCR).
Variant concordance analysis
Variants that passed filters and were located within the genomic regions targeted in PS were included in this analysis. Variant concordance was assessed using bcftools (v1.2) and vcfeval from RTG-Core (Real Time Genomics, v3.4.4). In individual patients, sites at which the frequency of the two predominant alleles was <95% (allowing for sequencing errors) were considered nonbiallelic. The concordance analysis was performed for all single-nucleotide variants (SNVs) and indels, then separately for the subset that were annotated as high or medium impact.
Variant filtration and prioritization
Rare stop gain, splice donor or acceptor site loss, frameshift indels (defined as LOF variants), and missense variants were included if the maximal minor allele frequency (MAF) in the 1000 Genomes Project, Exome Sequencing Project, or Exome Aggregation Consortium (ExAC) databases was <1%. Missense variants were then excluded if they were predicted to be benign by both SIFT and PolyPhen2, or annotated as benign in ClinVar. SVs were included if they had population allele frequencies <1% and overlapped with exonic regions from genes of interest (Supplementary Methods). Variants were classified into one of five categories: pathogenic, likely pathogenic, uncertain significance (VUS), likely benign, and benign, according to recommendations for clinical reporting from the American College of Medical Genetics and Genomics (ACMG).17
Gene sets analyzed
Three sets of genes were evaluated (Supplementary Table S2). The first set included 67 “panel genes” evaluated by PS. An “extended gene set” was comprised of 406 genes, including reported DCM-associated genes that were not represented on the PS panel, genes with presumptive links to cardiac and skeletal myopathies, and cardiac-enriched genes from the human protein atlas.18 The third set included a list of 57 genes compiled by the ACMG in which secondary findings are deemed clinically reportable.19
Results
PS-identified SNV and indels
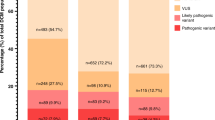
PS was performed in 42 probands with familial DCM. Seventy-eight rare (MAF <1%) LOF or potentially damaging missense variants were identified in 37 probands (Table 1). All 78 PS-identified variants were confirmed to be present in probands, and were also investigated in family members, using Sanger sequencing (Supplementary Methods). Twenty-one variants in 21 families (50%) were subsequently deemed pathogenic or likely pathogenic based on ACMG criteria,17 including 14 LOF TTN variants that we have reported previously.12
GS sequencing depth and coverage
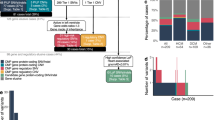
GS had an average depth of 34×, covering 97% of the genome, 98% of all exons, and 99% of PS gene exons (Fig. 1a). In comparison, PS had a much higher average read depth (486×) but covered only 91% of its targeted regions (Fig. 1a). GS coverage was compared with a ES dataset obtained in the human NA12878 cell line using the SSv5 capture kit. In this dataset, ES had an average read depth of 150× but covered only 69% of the PS targets (Fig. 1a). Similar results were found in ES datasets for which CREv2 and SSv6 capture kits had been used, with coverage of panel targets only 64 and 56%, respectively (Fig. 1a, Supplementary Table S3). The poor coverage of ES relative to PS highlights the benefit of a comprehensive disease-focused panel, compared with a generic ES design. The remaining causes of the lower coverage of panel genes with PS and ES, when compared with GS, were incompletely or mistargeted exons due to biases in probe design, synthesis, and hybridization (Fig. 1c,d).
a Bar graphs showing breadth of coverage, that is, the median (±1st and 3rd quartiles) percentage of positions covered by ≥15 sequence reads with genome sequencing (GS), panel sequencing (PS), and exome sequencing (ES) (SSv5 and CREv2) for targeted regions on the PS panel (“panel targets”) and b exons in Ensembl isoforms of these genes (“Ensembl exons”). c Histograms comparing GS depth of read coverage across representative gene exons with PS and d ES (SSv5). Note lack of uniform coverage due to capture bias in c and discrepancy between actual exons and WES target regions in d. e Overlap of protein-coding nucleotides contained within Ensembl isoforms with regions targeted on PS panel and f WES (SSv5), i.e., regions targeted by ES (“ES targets”) and targeted protein-coding exons (“ES exons”). g Breadth of coverage across panel genes with varying ES input in Gigabases (Gb) showing ES SSv5 exons (red) and Ensembl exons (orange); GS breadth of coverage across the same Ensembl exons is indicated by dashed line (teal), assuming the ~34× genome-wide average coverage used in this study
We further interrogated the reference sets of gene isoforms used in the design and analysis of target regions. GS data were analyzed with respect to protein-coding transcripts from the comprehensive Ensembl database, reporting 1–31 isoforms per panel gene. In comparison, PS targets included exons and conserved flanking sequences of isoforms from University of California–Santa Cruz (UCSC) knownGene and RefSeq, while ES SureSelect targets were based on subsets of isoforms from UCSC, RefSeq, GENCODE, and CCDS databases. Coverage of Ensembl isoforms of panel genes was high (99%) with GS, but was 86% with PS and ranged from 90 to 79% with ES (Fig. 1b, Supplementary Table S3). We found that this was mainly due to an extra 67 kb of Ensembl isoform sequences that were not included on the PS panel and 59 kb not included in ES SSv5 SureSelect targets/exons (Fig. 1e,f). Even with ES at >300× average depth of coverage, sequencing breadth plateaued at 90% for Ensembl exons and 95% for SureSelect targeted exons, which was less than the 99% achieved by GS (Fig. 1g).
Concordance between GS and PS
GS identified on average 3.8 × 106 SNVs and 1.2 × 106 small indels per proband genome-wide, of which 24,000 were coding and 42 were LOF and rare (MAF < 1%). Extensive Sanger sequencing of a subset of 115 SNV and indels in probands and family members showed that GS had an overall very low false-positive rate (0.81%, Supplementary Results).
For the variant concordance analysis between GS and PS, we used the same analytical pipeline and investigated the detection rates of all SNVs and indels in PS-targeted regions irrespective of MAF and read depth. A median of 483 variants per proband were concordant between GS and PS, with 131 variants identified exclusively by GS and 104 variants exclusively by PS (Supplementary Table S4). Seventy-seven percent of GS-specific variants were in positions that had <10 reads on PS, with the majority of these occurring at positions with no reads. In contrast, only 3% variants were missed by GS for the same reason. Variants listed in dbSNP are more likely to be true positives and this was the case for 87% of GS-specific variants but only 31% of the PS-specific variants, suggesting that two-thirds of PS-specific variants were either novel or false positives. Of these PS-specific variants, 22% were multiallelic (vs. 5% in GS) and 53% were located at homopolymers and short tandem repeats, likely resulting from replication slippage. This can occur naturally but more often arises during PCR-mediated DNA replication.20
To further investigate discordant variants, we looked at annotated LOF and missense variants, and manually inspected the read alignments. There was an average of 82 concordant variants per proband, with 11 variants exclusive to GS and 23 variants exclusive to PS (Supplementary Table S5). The most frequent cause of GS-specific variants was inadequate PS read coverage (Supplementary Fig. S1A), followed by missed variants occurring in Ensembl exons that were not represented on the PS panel. An additional 16% of GS-specific variants were present in the PS reads but missed by the variant caller. In contrast, most (84%) of the PS-specific variants were sequencing artifacts due to phasing (Supplementary Fig. S1B). This can occur during cycle-based sequencing when some of the DNA strands fall behind or jump ahead of the cycle.21 Other sources of discordance seen with both GS and PS included amplification of sequencing errors, incorrect variant calls (especially for indels), and ambiguous regions of low mapping quality (Supplementary Fig. S1C). In total, there was an average of 10 likely real LOF and missense variants per person that were missed by PS and 0.3 missed by GS.
GS detection of pathogenic/likely pathogenic variants in panel genes
GS data underwent further filtering to identify rare LOF or potentially damaging missense variants in panel genes. GS successfully detected the 78 prioritized PS variants, including all 21 of the pathogenic and likely pathogenic variants. A frameshift variant, SGCB p.M1Gfs, in family AF (Table 1), identified by GS and confirmed by Sanger sequencing, was missed by PS due to lack of coverage.
LOF variants in an extended gene set
To further explore the GS data, we compiled an extended set of 406 genes with proven and putative links to cardiomyopathy (Supplementary Table S2). GS successfully identified a reported likely pathogenic missense variant, p.I184M, in the NKX2-5 gene22 (Table 1). An additional 17 LOF variants in 14 families were evaluated (Table 2), all of which were validated by Sanger sequencing, but none met ACMG criteria for pathogenicity.
Five variants, in the FLNC, ANO5, ACADVL, TRIM63, and PDE4DIP genes, were present in all (two or more) family members with DCM in the respective kindreds (Supplementary Fig. S2). In family DF (negative after PS testing), a FLNC p.C2369* variant was identified. Variants in FLNC, which encodes the actin-binding protein filamin C, have been associated with cardiac and skeletal myopathies.23 Filamin C deficiency causes cardiac developmental defects in zebrafish and neonatal death in homozygous mice,23, 24 while heterozygous mice carrying a human FLNC p.W2710X PV show skeletal myopathy.25 To date, there is no definitive animal model evidence that heterozygous filamin C loss-of-function results in DCM. Although the nonsense FLNC variant was present in both affected individuals in family DF (II-1, II-2), it was also present in two unaffected siblings aged 52 (II-4, II-5) and 45 years, respectively, and it was classified as a VUS. In family CS, all affected individuals carried a likely pathogenic LOF TTN variant and an ANO5 p.N64fs variant. ANO5 encodes anoctamin 5, a transmembrane protein with putative calcium-activated chloride channel activity. Homozygosity for LOF ANO5 variants in human subjects has been associated with limb girdle and Miyoshi muscular dystrophies.26 This phenotype is not present in Ano5 knockout mice27 and none of the CS family members had overt skeletal myopathy. Despite good family cosegregation, the ANO5 variant was called a VUS. In family MO, individuals with a pathogenic LOF TTN variant also carried a splice acceptor site variant, ACADVL c.1077_+1G>T. The latter has been associated with a 36% reduction in very long chain acyl-CoA dehydrogenase activity but this level is predicted to be tolerated.28 Affected individuals in family BK had a likely pathogenic missense MYH7 variant as well as a TRIM63 p.Q274* variant, both of which have proposed associations with hypertrophic cardiomyopathy.29, 30 The two variant carriers (II-3, II-4) clearly had DCM with one also showing left ventricular hypertrophy. This nonsense TRIM63 variant has been associated with reduced autoubiquitination in transfected cells, and increased left ventricular mass in transgenic mice.30 The four affected individuals tested in family CZ all carried a PDE4DIP p.C18* variant in addition to a pathogenic missense MYH7 variant. PDE4DIP encodes an A-kinase anchoring protein that is involved in phosphorylation of the sarcomeric proteins, cMyBPC and cTNNI.31 This variant could plausibly modulate sarcomere kinetics but was classified as a VUS due to a lack of compelling evidence supporting PDE4DIP loss of function as a DCM mechanism.
Structural variants
Because GS sequence coverage extends beyond exonic regions, comprehensive genome-wide evaluation of SVs is possible. Using our in-house pipeline, ClinSV, we detected an average of 5379 SVs, including 4470 copy-number variants per proband, of which 232 were rare (MAF < 1%), and 23 that were rare and overlapped genic regions. When the 67 panel genes were evaluated, one rare SV was identified. This was a complex BAG3 deletion/duplication that included the BAG domain and is likely to have a loss-of-function effect (Fig. 2a, Table 3). This BAG3 SV was confirmed to be present in the proband using Sanger sequencing, segregated with DCM in family AA (Supplementary Fig. S2), and was deemed to be pathogenic. Numerous truncating BAG3 variants have been reported in DCM patients, with many cosegregating with disease in families.32 Eight SVs were found in the extended gene set, all of which were validated by independent sequencing methods (Supplementary Methods and Results), but none met ACMG criteria for pathogenicity or were likely primary causes of DCM (Fig. 2b, Table 3). For example, in the proband from family BG we identified a rare whole-gene duplication of triadin (TRDN; Fig. 2b), a developmentally regulated core member of the ryanodine receptor complex, whose copy number has been conserved in mammals.33 Although present in 4 of the 6 other affected family members (Supplementary Fig. S2), this variant was also seen in 4 unaffected family members, and was classified as a VUS. The remaining SVs mostly showed incomplete segregation with disease, occurred in families with other identified pathogenic/likely pathogenic variants, overlapped with SVs reported in population databases, or were in genes with unknown relevance to DCM, and were all classified as VUS.
Value of GS in additional family members
In two large kindreds that remained unsolved after PS and GS testing of the proband, we undertook GS in additional family members. In family BP (Supplementary Fig. S2), a MYBPHL p.R255* variant was identified in three family members but not in the proband. One of these, I-1, had been diagnosed with DCM and first-degree atrioventricular block at 84 years of age. The other two variant carriers included an asymptomatic 56-year-old male (II-1), and a 42-year-old female with ventricular ectopy (II-6), neither of whom had DCM. MYBPHL has been recently described as a DCM disease gene, with homozygous and heterozygous knockout mice showing DCM and conduction-system abnormalities.34 The same MYBPHL p.R255* variant was seen in a family with early-onset DCM and in an unrelated individual with left ventricular dilation.34 The mutant MyBP-HL protein was function-altering, with reduced expression in human cardiomyocytes and abnormal myofilament localization in transfected neonatal mouse cardiomyocytes.34 In family BG (Supplementary Fig. S2), GS in two of the proband’s aunts (II-2, II-4) identified a novel LOF TTN variant. Sanger sequencing of this variant in all family members confirmed its absence from the proband and presence in three affected siblings.
Overall yield of GS and reportable secondary findings
The yield of pathogenic/likely pathogenic variants was increased from 21 families (50%) with PS, to 24 families (57%) following GS analysis of panel genes and preliminary mining of the extended gene panel. Interrogation of the ACMG 57-gene list17 (Supplementary Table S2) yielded three significant secondary findings. In the family BG proband, we found a common function-altering GLA p.D313T variant associated with Fabry disease35 that did not segregate with DCM in the family (Supplementary Fig. S2). Two other probands were heterozygous carriers of common missense MUTYH variants that are annotated as pathogenic in ClinVar and associated with increased cancer risk.
Discussion
Our data support GS as a viable method for genetic testing in familial DCM with high detection accuracy for rare SNVs and indels in disease-associated genes. These findings concur with recently reported results for GS-based testing in hypertrophic cardiomyopathy36 and extend pilot data for GS in DCM.11 A compelling argument in favor of GS as a first-line testing method is its potential for ongoing data mining in unsolved cases. However, in an initial analysis we found that surprisingly few variants in an extended gene set data met clinical criteria for pathogenicity.
Despite a lower overall sequencing depth than PS (and ES), GS gave superior SNV and indel detection with less risk of variants being missed due to sequence gaps. GS’s broad and uniform sequence coverage, even across exons targeted specifically by PS, results from sample preparation methods that avoid capture bias37 and a lack of constraint to predefined sets of transcript isoforms and target regions. Some GS-identified variants in panel genes were missed by PS because different gene reference sets were used. This limitation of PS could potentially be reduced by optimized probe design and comprehensive representation of tissue-specific isoforms.6 The poor ES coverage observed over PS regions was recapitulated using three different exome capture kits, in data generated from two laboratories. Even at 300× depth, ES failed to achieve the same breadth of coverage as 34× GS. Importantly we show that on target ES performance can be good, but often the target does not correspond to the nearby exon, resulting in potentially missed pathogenic variants.
Coverage uniformity gives GS a distinctive advantage for identification of SVs, which account for 0.5–1% of heritable interindividual sequence differences (compared with 0.1% for SNVs).8 Although SVs are increasingly implicated in human disorders,8 we found relatively few pathogenic SVs in cardiomyopathy-associated genes. These results may be skewed by ascertainment bias because DCM associations of panel genes have predominantly been established by profiling SNVs and indels. SVs often involve pairs of flanking low copy repeats.8 Known DCM-associated genes may lack these regions and hence be less susceptible to structural rearrangements. Extending our analysis to 406 cardiac-enriched genes resulted in the discovery of only 8 additional SVs, none of which were clearly causative of DCM. One of the challenges in evaluating SVs is assessment of their functional effects, especially for those variants that involve complex or partial duplications and deletions. Intuitively, changes in gene copy number would impact on gene “dose,” but this is not necessarily the case and a number of adaptive mechanisms may result in dosage compensation.8 Even full-gene or balanced duplications and deletions can have unpredictable effects if there is disruption of local or long-range genomic architecture that involves gene-regulatory sequences.
To improve GS yield, more genetic information is needed to identify patterns of differential variant prevalence in cases and controls, recurrent variant types, and PV hotspots. Equally, more experimental data are required to identify function-altering variants and to show that these functional effects have plausible links to disease causation. In the ACMG criteria,17 segregation of variants in affected family members provides support for DCM association, but the level of evidence can increase to moderate or strong with larger family sizes and/or multiple families.
Segregation analysis in familial DCM relies on the assumption of single driver PV but this is increasingly open to question.38 As genetic evaluation has shifted from gene candidate screening studies to genome-wide analyses, it is not uncommon to find multiple rare variants in family probands39 and, as seen with families BG and BP, the number of potential function-altering variants in any one family can increase as more individuals are studied. These combinations of variants may have additive, synergistic, or neutralizing effects, and each person’s total burden of rare and common variants may determine threshold levels for myocardial dysfunction.
Should GS be used as a first-line genetic test for DCM or reserved for PS-negative cases? The answer to this question is currently unknown and involves a dynamic interplay between cost and yield (Supplementary Fig. S3). Currently, the cost of GS-based clinical testing is more than double that of PS. With increasing customer demand and technical efficiencies, the operational costs of both GS and PS are likely to decrease over time. For PS, these reductions will be offset by the ongoing need to design and optimize new probe sets as new disease genes are discovered. The manpower costs of expert clinical reporting are essentially equivalent for PS and GS because the same suites of disease genes are generally evaluated, but these costs may rise as the spectrum of reportable genes and variants expands. The turnaround time for GS is equivalent or faster than routine PS (or ES), because of the time required for targeted sequence capture, together with delays incurred by waiting for samples to be pooled. Recent data have shown that fast-tracked ES-based genetic testing in acutely ill pediatric patients is however feasible and cost-saving, due to expedited diagnosis and management.40 GS is a storehouse of medically relevant information that extends beyond identification of rare disease-causing variants, much of which cannot be obtained from PS or ES. For example, polygenic risk scores derived from suites of common variants (often in noncoding regions) may predict an individual’s risk of DCM complications, and pharmacogenomic associations may guide drug selection and doses.
Comprehensive economic models need to be developed that consider variables such as cost per test, number of tests ordered, yield of tests, requirement for secondary testing of probands, cascade testing of family members, and the impact of genotype information on requirements for clinical surveillance of family members. Genotype-based early interventions may also impact more broadly on long-term health outcomes and costs, potentially impacting hospitalization rates, device implantation, drug administration, workforce productivity, and use of social services. Compelling health economics data would provide a powerful argument to insurance companies or governments for the overall benefits of genetic testing and the preferred testing modality.
References
Fatkin D, Seidman CE, Seidman JG. Genetics and disease of ventricular muscle. Cold Spring Harb Perspect Med. 2014;1:a021063.
Ackerman MJ, Priori SG, Willems S, et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies. Europace. 2011;13:1077–1109.
Fatkin D, Johnson R, McGaughran J, et al. Position statement on the diagnosis and management of familial dilated cardiomyopathy. Heart Lung Circ. 2017;26:1127–1132.
Sikkema-Raddatz B, Johansson LF, de Boer EN, et al. Targeted next-generation sequencing can replace Sanger sequencing in clinical diagnostics. Hum Mutat. 2013;34:1035–1042.
Pugh TJ, Kelly MA, Gowrisankar S, et al. The landscape of genetic variation in dilated cardiomyopathy as surveyed by clinical DNA sequencing. Genet Med. 2014;16:601–608.
Pua CJ, Bhalshankar J, Miao K, et al. Development of a comprehensive sequencing assay for inherited cardiac condition genes. J Cardiovasc Transl Res. 2016;9:3–11.
Bamshad MJ, Ng SB, Bigham AW, et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet. 2011;12:745–755.
Weischenfeldt J, Symmons O, Spitz F, Korbel JO. Phenotypic impact of genomic structural variation: insights from and for human disease. Nat Rev Genet. 2013;14:125–138.
Li X, Montgomery SB. Detection and impact of rare regulatory variants in human disease. Front Genet. 2013;4:67.
Dewey FE, Grove ME, Pan C, et al. Clinical interpretation and implications of whole-genome sequencing. JAMA. 2014;311:1035–1045.
Golbus JR, Puckelwartz MJ, Dellefave-Castillo L, et al. Targeted analysis of whole genome sequence data to diagnose genetic cardiomyopathy. Circ Cardiovasc Genet. 2014;7:751–759.
Roberts AM, Ware JS, Herman DS, et al. Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations in health and disease. Sci Transl Med. 2015;7:270ra6.
Shi L, Zhang Y, Feng L, et al. Multi-omics study revealing the complexity and spatial heterogeneity of tumor-infiltrating lymphocytes in primary liver carcinoma. Oncotarget. 2017;8:34844–34857.
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–1303.
Gayevskiy V, Roscioli T, Dinger ME, Cowley MJ Seave: a comprehensive web platform for storing and interrogating human genomic variation. BioRxiv https://doi.org/10.1101/258061 (unpublished data in text).
Rehm HL, Bale SJ, Bayrak-Toydemir P, et al. ACMG clinical laboratory standards for next-generation sequencing. Genet Med. 2013;15:733–747.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Uhlén M, Fagerberg L, Hallström BM, et al. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Green RC, Berg JS, Grody WW, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15:565–574.
Viguera E, Canceill D, Ehrlich SD. In vitro replication slippage by DNA polymerases from thermophilic organisms. J Mol Biol. 2001;312:323–333.
Kircher M, Stenzel U, Kelso J. Improved base calling for the Illumina Genome Analyzer using machine learning strategies. Genome Biol. 2009;10:R83.
Costa MW, Guo G, Wolstein O, et al. Functional characterization of a novel mutation in NKX2-5 associated with congenital heart disease and dilated cardiomyopathy. Circ Cardiovasc Genet. 2013;6:238–247.
Begay RL, Tharp CA, Martin A, et al. FLNC gene splice mutations cause dilated cardiomyopathy. JACC: Basic Transl Sci. 2016;1:344–359.
Dalkilic I, Schienda J, Thompson TG, et al. Loss of filamin C (FLNC) results in severe defects in myogenesis and myotube structure. Mol Cell Biol. 2006;26:6522–6534.
Chevessier F, Schuld J, Orfanos Z, et al. Myofibrillar instability exacerbated by acute exercise in filaminopathy. Hum Mol Genet. 2015;24:7207–7220.
Bolduc V, Marlow G, Boycott KM, et al. Recessive mutations in the putative calcium-activated chloride channel Anoctamin 5 cause proximal LGMD2L and distal MMD3 muscular dystrophies. Am J Hum Genet. 2010;86:213–221.
Xu J, El Refaey M, Xu L, et al. Genetic disruption of Ano5 in mice does not recapitulate human Ano5-deficient muscular dystrophy. Skelet Muscle. 2015;5:43.
Hoffmann L, Haussmann U, Mueller M, et al. VLCAD enzyme activity determinations in newborns identified by screening: a valuable tool for risk assessment. J Inherit Metab Dis. 2012;35:269–277.
Erdmann J, Daehmlow S, Wischke S, et al. Mutation spectrum in a large cohort of unrelated consecutive patients with hypertrophic cardiomyopathy. Clin Genet. 2003;64:339–349.
Chen SN, Czernusczewicz G, Tan Y, et al. Human molecular genetic and functional studies identify TRIM63, encoding Muscle RING Finger Protein 1, as a novel gene for human hypertrophic cardiomyopathy. Circ Res. 2012;111:907–919.
Uys GM, Ramburan A, Loos B, et al. Myomegalin is a novel A-kinase anchoring protein involved in the phosphorylation of cardiac myosin binding protein C. BMC Cell Biol. 2011;12:18.
Norton N, Li D, Rieder MJ, et al. Genome-wide studies of copy number variation and exome sequencing identify rare variants in BAG3 as a cause of dilated cardiomyopathy. Am J Hum Genet. 2011;88:273–282.
Rice AM, McLysaght A. Dosage sensitivity is a major determinant of human copy number variant pathogenicity. Nat Commun. 2017;8:14366.
Barefield DY, Puckelwartz MJ, Kim EY, et al. Experimental modelling supports a role for MyBP-HL as a novel myofilament component in arrhythmia and dilated cardiomyopathy. Circulation. 2017;136:1477–1491.
Froissart R, Guffon N, Vanier MT, et al. Fabry disease: D313Y is an α-galactosidase A sequence variant that causes pesudodeficient activity in plasma. Mol Genet Metab. 2003;80:307–314.
Cirino AL, Lakdawala NK, McDonough B, et al. A comparison of whole genome sequencing to multigene panel testing in hypertrophic cardiomyopathy patients. Circ Cardiovasc Genet. 2017;10:e001768.
Bodi K, Perera AG, Adams PS, et al. Comparison of commercially available target enrichment methods for next-generation sequencing. J Biomol Tech. 2013;24:73–86.
Hershberger RE, Hedges DJ, Morales A. Dilated cardiomyopathy: the complexity of a diverse genetic architecture. Nat Rev Cardiol. 2013;10:531–547.
Haas J, Frese KS, Peil B, et al. Atlas of the clinical genetics of human dilated cardiomyopathy. Eur Heart J. 2015;36:1123–1135.
Stark Z, Lunke S, Brett GR, et al. Meeting the challenges of implementing rapid genomic testing in acute pediatric care [published online ahead of print March 15, 2018]. Genet Med 2018. https://doi.org/10.1038/gim.2018.37.
Acknowledgements
We thank the Kinghorn Centre for Clinical Genomics for assistance with production and processing of genome sequencing data. We acknowledge Robert Graham, Matthew Grounds, John Mattick, and John Schubert, as founders of the Cardiogenomics Project. This research was undertaken with the assistance of resources and services from the National Computational Infrastructure, which is supported by the Australian Government.
This work was undertaken as part of the Cardiogenomics Project, and would not have been possible without the support of John Schubert, the Kinghorn Foundation, Garvan Foundation, and the Victor Chang Cardiac Research Institute. Funding support was also received from the NSW Office of Health and Medical Research Collaborative Grants Program, National Health and Medical Research Council (D.F., C.S.), Estate of the Late RT Hall (D.F.), Simon Lee Foundation (D.F.), Howard Hughes Medical Institute (C.E.S.), National Institutes of Health (J.G.S.), Leducq Foundation (J.G.S., C.E.S.), Cancer Institute NSW (M.J.C.), and NSW Department of Health (M.J.C). J.I. is the recipient of a National Heart Foundation of Australia Future Leader Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
M.E.D., K.W., A.L.S., and B.L. are employed by Genome.One, a clinically accredited genetic testing provider that uses genome sequencing. The other authors declare no conflict of interest.
Additional information
A. Minoche and C. Horvat are joint first authors.
M. Dinger, M. Cowley, and D. Fatkin are joint senior authors.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Minoche, A.E., Horvat, C., Johnson, R. et al. Genome sequencing as a first-line genetic test in familial dilated cardiomyopathy. Genet Med 21, 650–662 (2019). https://doi.org/10.1038/s41436-018-0084-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0084-7
Keywords
This article is cited by
-
Exploring TTN variants as genetic insights into cardiomyopathy pathogenesis and potential emerging clues to molecular mechanisms in cardiomyopathies
Scientific Reports (2024)
-
Genetic testing and family screening in idiopathic pediatric cardiomyopathy: a prospective observational study from a tertiary care center in North India
Egyptian Journal of Medical Human Genetics (2023)
-
Cardiac splicing as a diagnostic and therapeutic target
Nature Reviews Cardiology (2023)
-
The genetic basis for adult-onset idiopathic dilated cardiomyopathy in people of African descent
Heart Failure Reviews (2023)
-
Whole genome sequencing delineates regulatory, copy number, and cryptic splice variants in early onset cardiomyopathy
npj Genomic Medicine (2022)