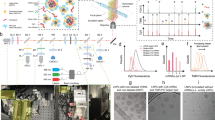
Abstract
Lipid nanoparticles (LNPs) are currently the most advanced non-viral clinically approved messenger ribonucleic acid (mRNA) delivery systems. The ability of a mRNA vaccine to have a therapeutic effect is related to the capacity of LNPs to deliver the nucleic acid intact into cells. The role of LNPs is to protect mRNA, especially from degradation by ribonucleases (RNases) and to allow it to access the cytoplasm of cells where it can be translated into the protein of interest. LNPs enter cells by endocytosis and their size is a critical parameter impacting their cellular internalization. In this work, we studied different formulation process parameters impacting LNPs size. Taylor dispersion analysis (TDA) was used to determine the LNPs size and size distribution and the results were compared with those obtained by Dynamic Light Scattering (DLS). TDA was also used to study both the degradation of mRNA in the presence of RNases and the percentage of mRNA encapsulation within LNPs.

Similar content being viewed by others
Introduction
Prophylactic vaccines stimulate the immune system to produce immunity against a specific pathogen, protecting the person from future infections. Thereby, during the last century, conventional vaccine approaches have drastically reduced the mortality and morbidity associated with infectious diseases [1, 2]. Despite this success, the number of untreated medical needs remains high and new infectious diseases are appearing periodically [3]. The development of new vaccine strategies is therefore required.
Messenger ribonucleic acid (mRNA) vaccines have emerged as promising and versatile alternatives to conventional vaccine approaches [4]. These vaccines do not carry a typical antigen but rather the information for producing it using the cell machinery and turning the body into its own antigen factory. This is close to the process of natural viral infections, where viral genetic material is used to produce viral proteins from within a host cell. The use of mRNA vaccine has several beneficial features over whole inactivated, live-attenuated, subunit or DNA-based vaccines. Indeed, mRNA is a noninfectious platform that does not need to enter into the cell nucleus to be effective [5] and has the potential for rapid development and manufacturing. mRNA vaccines are manufactured using defined processes, regardless of the antigen encoded by the mRNA, which allows to simplify vaccine production and quality control and therefore reduce development and validation timelines [6]. The first report of successful use of in vitro transcribed (IVT) mRNA was published in 1990, when mRNAs were injected into mice and protein production was detected [7]. However, until recently, use of mRNA vaccine has been hold back by mRNA instability, high innate immune activation, and inefficient intra cellular delivery [8]. Recent advances on IVT-mRNA structure and RNA delivery systems have allowed to overcome these issues, and mRNA vaccines show now considerable promises for both prophylactic and therapeutic vaccines [9]. Onpattro® (patisiran) was the first RNA-based drug approved by the Food and Drug Administration (FDA) and the European Medicines Agency in 2018 [10]. This drug uses small interfering RNA encapsulated into lipid nanoparticles (LNPs) for the treatment of hereditary transthyretin-mediated amyloidosis [11, 12]. Since then, other highly effective RNA-based vaccines using LNPs have been developed and approved in a record time during the COVID-19 epidemic [13]. This opens the way and the hope for the acceleration of the development of new vaccines based on mRNA vaccine technology [14]. Yet, some points still need to be optimized such as the stability of the formulations [15, 16], the endosomal escape efficiency [17, 18], the reduction of reactogenicity and adverse events after injection [19], and the precise targeting of certain organs [20,21,22].
LNPs are not simple carriers, they play an active role in the vaccine efficacy [23]. Among other physicochemical parameters, the size of the LNPs is a critical parameter that has an important effect on their in vivo behavior. Particle size is known to influence the pathway of cell internalization and intracellular processing [24,25,26,27]. Size also influences the pharmacokinetics of the nanoparticles [28, 29]. The size of LNPs can therefore be optimized to specifically target certain organs [30,31,32,33,34]. Moreover, size characterization is also of major importance during stability studies as it is necessary to verify that there is no aggregation, fusion or degradation of the LNPs and that their size remains stable [15]. Optimizing mRNA-based vaccine formulations requires the development of new analytical methods to better understand current limitations and to challenge existing analytical methods [35].
In this work, Taylor dispersion analysis (TDA) was used as a powerful tool for the optimization of LNPs formulations. LNPs of different sizes were formulated by modifying three parameters: (i) the PEG-lipid content, (ii) the total flow rate (TFR) which is the fluid volume which passes per unit of time and (iii) the flow rate ratio (FRR) between the aqueous and the organic phase. The sizes and size distribution obtained by TDA were then compared to those obtained by Dynamic Light Scattering (DLS) which is a method widely used for the monitoring of LNPs formulations [28, 36]. TDA was also used to follow the degradation of mRNA in the presence of ribonucleases and to determine the percentage of mRNA encapsulation within the LNPs.
Materials and methods
Chemicals and materials
FLuc (Cyanine 5 Firefly Luciferase) mRNA (1929 nucleotides) at 1 mg/mL in 1 mM sodium citrate pH 6.4 was purchased from TriLink BioTechnologies (San Diego, CA, USA). RNase-free water DEPC treated was purchased from Thermo Fisher Scientific (Illkirch-Graffenstaden, France). Trisodium citrate dihydrate HOC(COONa)(CH2COONa)2 ·2H2O, (Mw = 294.1 g/mol) and citric acid monohydrate HOC(COOH)(CH2COOH)2·H2O (Mw = 210.1 g/mol) were purchased from Merk (Darmstadt, Germany). DLin-MC3-DMA ((6Z,9Z,28Z,31Z)-heptatriacont-6,9,28,31-tetraene-19-yl 4-(dimethylamino)butanoate) was purchased from Sai Life Sciences Ltd. (Telangana, India). DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine), Chol (cholesterol) and DMG-PEG-2000 (1,2-dimyristoyl-rac-glycero-3-methoxypolyethylene glycol-2000) were purchased from Sigma-Aldrich (St. Quentin Fallavier, France). Absolute ethanol was purchased from Carlo Erba Reagents (Val de Reuil, France). PBS (phosphate-buffered saline) 10X buffer was purchased from Eurobio Scientific (Les Ulis, France). Slide-A-Lyzer™ dialysis cassettes, 10 K MWCO, 3 mL, were purchased from Thermo Fisher Scientific (Illkirch-Graffenstaden, France). µSIL-FC (fluorocarbon polymer) coated capillaries were purchased from Agilent Technologies (Santa Clara, CA, USA). Invitrogen™ Quant-iT™ RiboGreen™ RNA Assay Kit and Pierce™ White Opaque 96-Well Plates were purchased from Thermo Fisher Scientific (Illkirch-Graffenstaden, France). Ribonuclease A from bovine pancreas (RNase A, Mw = 14 kDa, pI = 9) and Triton™ X-100 were purchased from Merk (Darmstadt, Germany). Deionized water was further purified with a Milli-Q system from Millipore (Molsheim, France).
Formulation of LNPs
LNPs self-assemble by the rapid dilution of an organic phase containing lipids in an aqueous phase containing the mRNA. The organic phase was prepared by mixing the four lipids in absolute ethanol at the following molar ratio DLinMC3-DMA:DSPC:Chol:DMG-PEG-2000 (50:10:40-x:x). Stock solutions of lipids were previously prepared in absolute ethanol at the following concentrations: DLinMC3-DMA 100 mg/mL, DSPC 30 mg/mL, Chol 18 mg/mL, DMG-PEG-2000 30 mg/mL. The aqueous phase was prepared by diluting mRNA at 0.25 g/L in 50 mM RNase free citrate buffer, pH 4. The mixing step was performed using NanoAssemblr® IgniteTM instrument from Precision NanoSystems (Vancouver, Canada). One syringe was filled with the aqueous solution and another one with the ethanolic solution. LNPs are then produced setting up the NanoAssemblr® software with the following parameters: FRR and TFR. Both parameters varied according to each formulation, as summarized in Table 1. 2 mL solution were involved per formulation with 0.45 mL of start waste volume and 0.05 mL of end waste volume, resulting in 1.5 mL per LNP formulation. After formulation step, each LNP-mRNA sample was immediately dialyzed one night against 500 mL of 50 mM citrate buffer, pH 4 using Slide-A-Lyzer dialysis cassette 10 K MWCO. This first dialysis was performed to remove residual ethanol. After that, two successive dialysis of one day and one night long in total were performed against 500 mL PBS buffer at pH 7.4 to raise the pH to physiological value. The size of the different LNPs formulations was then determined by DLS and the formulations were filtered using 0.2 or 0.45 micrometer filters according to their size. The formulations were then stored at +5 °C.
Taylor Dispersion Analysis (TDA)
The equipment used for TDA experiments was a 7100 Capillary Electrophoresis Agilent system (Waldbronn, Germany). This system was equipped with a diode array detector and with a Zetalif LED induced fluorescence detector purchased from Adelis (Grabels, France), both connected in series. The UV measurements were performed at 200 nm. The fluorescent measurements were performed with excitation at 480 nm (for RiboGreen® RNA dye). A neutral μSIL-FC capillary of 61 cm total length (52.5 cm to the UV detector, 40 cm to the fluorescence detector) × 50 μm was used. The temperature of the capillary cartridge was set at 25 °C. The capillary was presaturated before use with the LNP formulations for 10 min at 60 mbar. The presaturation is performed to limit further LNP adsorption on the capillary surface, it allows to saturate the interaction sites and therefore creates a dynamic coating inside the capillary. The optimization of TDA parameters to ensure reliable results has been described in a previous article [37]. The LNP formulations were injected without prior dilution into the capillaries filled with PBS buffer. TDA experiments were performed using 60 mbar mobilization pressure of a sample plug injected at 20 mbar for 6 s. The elution peaks obtained were fitted using two Gaussians, depending on the number of objects in the solution, according to equation (1) using a home-developed Excel spreadsheet:
where S(t) is the absorbance signal, σi is the temporal variance, Ai is a constant that depended on the response factor and the injected quantity of solute and t0 is the average elution time. t0 is directly obtained from the position of the maximum of absorbance and σi, and Ai are adjusting parameters obtained by nonlinear least square regression using Excel solver.
The temporal variance σi allows to calculate the molecular diffusion coefficient Di according to Eq. (2):
where Rc is the capillary radius. Stokes–Einstein Eq. (3) allows then to determine the hydrodynamic diameter Dh,i:
where kB is the Boltzmann constant, T is the temperature, and η is the eluent viscosity. Taylor dispersion analysis is suitable for determining the hydrodynamic diameter of objects ranging from 0.1 nm to 300 nm [38].
To get the size distribution, the elution profile can be fitted using a second approach based on Regularized Linear Inversion (CRLI) algorithm [39] according to Eq. (4):
where C is an instrumental constant, M(D) and ρ(D) are the mass and the molar concentration of the objects with the diffusion coefficient D, respectively. The polydispersity of the sample can further be determined using Eqs. (5) and (6):
Dynamic Light Scattering (DLS)
The equipment used for DLS experiments was a Malvern Panalytical Zetasizer Nano ZS system (Palaiseau, France). LNPs formulations were diluted by adding 10 µL of the formulation into 1 mL PBS buffer. The temperature of the cuvettes was set at 25 °C. The measurement angle was 173°. Cumulant fit was used to fit the experimental data of the autocorrelation function and to get the intensity average hydrodynamic diameter.
Encapsulation efficiency by Ribogreen assay
To determine mRNA encapsulation efficiency, two calibration curves were first performed with unformulated mRNA. The first curve was determined with mRNA diluted to 0.04, 0.10, 0.40, 0.80, and 1.20 µg/mL in 1X RNase free TE buffer (10 mM Tris-HCl with 1 mM EDTA) and the second one in 1X RNase free TE buffer with 0.5% (v/v) Triton X-100 in a 96-well plate. Quant-iT RiboGreen fluorescent reagent was diluted by 200 and then added to each well in volume equal the RNA solution (dilution by 2). The fluorescence signal (excitation/emission: 480/520 nm) was quantified using SpectraMax® i3k from Molecular Devices (San Jose, CA, USA).
LNP-mRNA formulations were then diluted in 1X RNase free TE buffer and in 1X RNase free TE buffer with 0.5% (v/v) Triton X-100, followed by Quant-iT RiboGreen reagent addition. mRNA samples were diluted to target the middle of the calibration range (≈0.40 µg/mL). mRNA encapsulation was determined by comparing the fluorescence in the presence and in the absence of Triton X-100. In the absence of Triton X-100, the signal only comes from the free (unencapsulated) mRNA while, in the presence of Triton X-100, the signal comes from the total mRNA as the detergent allows to break the LNPs. The encapsulation efficiencies (ee%) were therefore calculated according to Eq. (7):
Results
Nine formulations of LNPs were produced by dilution of an ethanolic phase containing the lipids in an aqueous citrate buffer containing mRNA, as shown in Fig. 1. The first three formulations [F1–F3] have been formulated with mRNA in the citrate buffer and the next six [F4–F9] without mRNA but with the same other parameters of formulation. Three formulation parameters have been studied: (i) the PEG-lipid content, (ii) the total flow rate and (iii) the flow rate ratio between the aqueous and the organic phase, as summarized in Table 1.
At low pH, the ionizable lipids are positively charged and electrostatic interactions between the negatively charged mRNA and the ionizable lipids result in the formation of the LNPs nuclei. At the same time, hydrophobic interactions between the lipids and the aqueous phase cause the lipids to group, avoiding contact with the aqueous phase, and resulting in the formation of the LNPs envelope.
The formulations were analyzed in triplicates by TDA and the mean hydrodynamic diameters were calculated by fitting the elution peaks obtained with two gaussian curves using a home-developed Excel spreadsheet (see Fig. S1 to visualize the taylorgram fits). Two-Gaussian fitting is used for taking into account the LNP population and the presence of small molecules in the samples (represented by the small sharp peak at the top of the elution profile). Figure 2A shows that decreasing the PEG-lipid content from 5 to 0.5% led to an increase in the hydrodynamic diameters of the LNPs from 52 ± 2 to 216 ± 4 nm, respectively, leading to broader elution profiles. Figure 2B shows that a decrease in the total flow rate from 6 to 1 mL/min resulted in an increase in the hydrodynamic diameters from 83 ± 5 to 121 ± 5 nm, respectively. Figure 2C shows that a decrease in the flow rate ratio from 5:1 to 2:1 also resulted in an increase in the hydrodynamic diameters from 79 ± 4 to 100 ± 5 nm, respectively.
Impact of formulation parameters: PEG-lipid content (A), Total Flow Rate (TFR) (B), Flow Rate Ratio (FRR) (C). Experimental conditions: µSIL-FC coated capillaries of 61 cm total length (52.5 cm to the UV detector) × 50 μm i.d. Buffer: 10 mM PBS, pH 7.4, η = 0.9 × 10−3 Pa s. Capillary presaturation: LNPs for 10 min at 60 mbar. Capillary preconditioning: H2O for 2 min at 960 mbar followed by 2 min PBS at 960 mbar. Injection of LNPs: 20 mbar, 6 s. Mobilization pressure: 60 mbar. Sample: LNPs (DLin-MC3-DMA:DSPC:Chol:DMG-PEG-2000 at 50:10:40-x:x molar ratio) empty or encapsulating Firefly Luciferase (FLuc) mRNA. UV detection: 200 nm. Temperature: 25 °C. All results are presented in triplicates.
The mean hydrodynamic diameters and the polydispersity indices of all LNPs formulations were also measured by DLS for comparison (see Table 1). The size distributions obtained by DLS and by TDA using Regularized Linear Inversion (CRLI) algorithm [39] were given in Figs. S2 and S3, respectively, and the corresponding polydispersity indices (PDI) were gathered in Table 1. The results obtained by TDA and DLS were compared and the sizes and PDI obtained by TDA were systematically lower than those obtained by DLS for all the LNPs formulations, due to intrinsic differences in these two techniques [37].
Figure 3A displays the size distributions (in triplicates) obtained by TDA for formulation F1, F3, and for a 50/50 (v/v) F1–F3 mixture. Figure 3B displays the size distributions of the same samples obtained by DLS for comparison. In DLS, the size distributions of the two formulations analyzed separately were shown to overlap with [30–300] nm for F1 and [110–600] nm for F2, which was not the case in TDA, [35–70] nm for F1 and [160–280] nm for F2. When analyzing the F1–F3 mixture, a single population with a very broad peak of [30–600] nm and an average size of 211 ± 4 nm was observed in DLS, while two distinct populations of [40–70] nm and [160–240] nm could be distinguished in TDA.
Analysis of formulations F1, F3 and a 50/50 (v/v) mixture of F1 and F3. TDA experimental conditions as in Fig. 2. DLS experimental conditions: 10 mM PBS buffer, pH 7.4, η = 0.9 × 10−3 Pa s. Sample: LNPs (DLin-MC3-DMA:DSPC:Chol:DMG-PEG-2000 at 50:10:40-x:x molar ratio) empty or encapsulating Firefly Luciferase (FLuc) mRNA. Sample dilution: 10 µL of the formulation into 1 mL PBS buffer. Temperature: 25 °C. Measurement angle: 173°. All results are presented in triplicates.
Subsequently, TDA was used to study the degradation of mRNA in presence of ribonuclease A (RNase A). Figure 4A shows the analysis of naked mRNA (unencapsulated) at 0.5 g/L in PBS buffer (without RNase A, see blue trace) giving an average hydrodynamic diameter of 43 ± 2 nm (mRNA Length: 1929 nucleotides). Repeating the same operation after addition of RNase A at 5 mg/L (below RNase A UV detection limit in TDA) in the mRNA solution, led to much a thinner peak (see green trace), the hydrodynamic diameter being only 1.4 ± 0.2 nm. The same experiment was then carried out on a LNP formulation encapsulating mRNA. No significant difference in size was observed with or without the presence of RNase A, the hydrodynamic diameters being similar (82 ± 4 and 84 ± 4 nm, respectively). Finally, a 50/50 (v/v) mixture of a LNPs solution and a free mRNA solution was prepared. In the absence of RNase A, two populations were distinguished by TDA, one of 43 ± 1 nm corresponding to the free mRNA and one of 84 ± 1 nm corresponding to LNPs (see blue trace). In the presence of RNase A, a population of 1.4 ± 0.1 nm, corresponding to the naked mRNA degraded by RNase A, and another one of 86 ± 2 nm, corresponding to LNPs, were observed. The Gaussian fits of the elution profiles are presented in Fig. S4.
Analysis of free mRNA (A), mRNA loaded LNP formulation F2 (B) and a mixture of free mRNA and LNP formulation F2 (C). Experimental conditions: free mRNA is Firefly Luciferase (FLuc) mRNA. LNP formulation F2 is DLin-MC3-DMA:DSPC:Chol:DMG-PEG-2000 at 50:10:38.5:1.5 molar ratio encapsulating FLuc mRNA. RNase A concentration in the final samples (green traces): 2.5 mg/L. Other conditions as in Fig. 2. All results are presented in triplicates.
In order to access the integrity of the mRNA encapsulated within the LNPs and to calculate the percentage encapsulation, RiboGreen® Reagent was used with a fluorescence detector on the capillary electrophoresis equipment. This fluorescent reagent does not enter LNPs and allows to label the free mRNA (red traces in Fig. 5). TE (10 mM Tris-HCl, 1 mM EDTA, pH 7.5) was used as buffer instead of PBS for TDA measurement involving RiboGreen® Reagent as it is provided in the Invitrogen™ Quant-iT™ RiboGreen™ RNA Assay Kit. A first visual observation of the free mRNA peaks obtained for the two formulations F1 and F3 in Fig. 5, shows that F1 has a higher rate of free mRNA than F3. To access the amount of total mRNA, the LNPs were deformulated by adding a powerful surfactant Triton X-100 at 0.5% (v/v) (see purple traces in Fig. 5). The ratio between the two peak areas (free mRNA/total mRNA) directly gave access to the percentage of mRNA encapsulation. Encapsulations of 79 and 95% were obtained for F1 and F3, respectively. To confirm these results, the encapsulation efficiencies of the two formulations were also calculated by Ribogreen assay using SpectraMax® i3k, (see materials and methods). This orthogonal method gave similar percentage of mRNA encapsulation (75% and 94% for F1 and F3, respectively).
Use of RiboGreen® fluorescent Reagent to label the free (unencapsulated) mRNA (red traces) and the total mRNA after deformulation of the LNP with Triton X-100 (purple traces). Experimental conditions: µSIL-FC coated capillaries of 61 cm total length (40 cm to the fluorescence detector) × 50 μm i.d. Buffer: TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 7.5). Capillary presaturation: LNP for 10 min at 60 mbar. Capillary preconditioning between runs: H2O for 2 min at 960 mbar followed by 2 min TE buffer at 960 mbar. Injection: 20 mbar, 6 s. Mobilization pressure: 60 mbar. Sample: LNP formulations F1 and F3 with RiboGreen® fluorescent Reagent and with (purple trace) or without (red trace) Triton X-100 at 0.5% (v/v) in the sample. Fluorescence excitation at 480 nm, emission light wavelength ranges from 515 to 760 nm. All results are presented in triplicates.
Additionally, it was possible to access the size of the mRNA by TDA, giving information on its integrity, by fitting the elution profile before and after LNP deformulation using Triton X-100. In the case of formulation F1, the average hydrodynamic diameter was 28 ± 2 nm and 37 ± 2 nm for the free mRNA (before deformulation) and the total mRNA (after deformulation), respectively. In the case of formulation F2, the average hydrodynamic diameter was 10 ± 2 nm and 42 ± 2 nm for the free mRNA and the total mRNA, respectively.
Finally, using calibration curves performed with naked mRNA, as presented in Fig. S5, it was possible to access the concentration of the total mRNA and free mRNA present in one formulation. 12.3 µg/mL free mRNA and 60.7 µg/mL total mRNA were found for formulation F1 and 3.2 µg/mL free mRNA and 62.1 µg/mL total mRNA were found for formulation F3.
Discussion
The size of LNPs is a critical parameter for cellular uptake [33, 40]. In this work, three parameters impacting LNPs size were studied. First, the PEG-lipid content was shown to have a significant impact on the size of LNPs, as a decrease from 5% to 0.5% multiplies the hydrodynamic diameter by 4. The strong influence of the PEG-lipid content has already been reported in the literature and the results obtained are in agreement with those of Belliveau et al. [41] and Yanez Arteta et al. [42]. Furthermore, Bao et al. [43]. showed that LNPs with higher PEG-C-DMA amount altered gene silencing efficacy by potentially reducing endosomal disruption. In addition, several studies reported the presence of anti-PEG antibodies [44, 45], therefore the development of new strategies to optimize or replace the PEG lipid was explored [46,47,48,49,50]. But its replacement is tricky and other strategies must be considered for size optimization, especially trying to obtain small-sized LNPs with diameter <100 nm.
Two other parameters of the formulation that have also been reported to have an impact on LNP size [51] were studied, namely the TFR and the FRR. It appeared in this work that the higher the TFR and the FRR, the smaller the LNPs size. Indeed, the faster the dilution of the ethanol phase in the aqueous phase, the less time needed for lipids to group and stabilize, and the smaller the LNPs size. Therefore, the system used for mixing the organic and the aqueous phases is also a critical point for controlling the LNPs size. To obtain small LNPs of well-defined size, the system must allow a rapid and homogeneous dilution of the ethanolic phase into the aqueous phase by maximizing the contact interface between the two fluids [52, 53].
The sizes, size distributions, and polydispersity indices of the LNPs formulations measured by TDA using CRLI fit [39] were compared to the ones obtained by DLS using Cumulant fit. The size distributions obtained by TDA were all narrower and shifted towards smaller sizes than those obtained by DLS. Therefore, the mean hydrodynamic diameters and the polydispersity indices obtained by TDA were smaller than those one obtained by DLS. These differences can be explained by the fact that the two techniques do not determine the same size distributions. The size distribution obtained by TDA is related to the weight-average (or mass-average) distribution for a detection sensitive to the mass concentration [54], each LNP contributing to the distribution relative to their mass proportion in the mixture. In contrast, the size distribution obtained by DLS is related to the intensity-average distribution, that gives more weight to the larger entities, since the distribution is weighted by the scattered intensity scaling as the hydrodynamic radius to the power 6. As the size distributions obtained by DLS are wider than those obtained by TDA, it is thus more difficult to distinguish two populations in DLS than in TDA. This was confirmed by the analysis of a 50–50 mixture of two LNP formulations of different hydrodynamic diameters (52 and 216 nm). Indeed, DLS experiments showed a single mode distribution with an average size close to the one of the largest LNP in the mixture, while TDA was able to identify the presence of two LNP populations. These differences show the complementarity of the two methods for a better characterization of LNP size distributions. For instance, DLS can easily detect the presence of few quantities of larger or aggregated LNP in the sample, while TDA brings a distribution relative to the mass proportion of each nanoparticle in the mixture, and consequently a non-biased mass-average size. Moreover, the complementary use of these two methods comply with the US FDA recommendation about the use of orthogonal methods to address technique-related differences for critical drug product parameters [55].
Subsequently, TDA was used to study the degradation of mRNA in the presence of ribonucleases. Degradation of mRNA is one of the main reasons why mRNA should be encapsulated into LNPs [56, 57]. It was observed that the addition of small amounts of RNase A is able to cut mRNA into small pieces as its hydrodynamic diameter instantly drops from 43 to 1.4 nm. TDA thus allowed a clear observation of mRNA degradation. Concerning the mRNA loaded LNPs, no visible degradation was directly observed by TDA after RNase addition.
It was demonstrated that TDA can be also used to determine the percentage of mRNA encapsulation within LNPs and the size integrity of the encapsulated mRNA. The results obtained were consistent with those obtained by Ribogreen assay using a microplate reader. The degree of encapsulation is an important parameter, as unencapsulated mRNA can easily be degraded and hardly penetrate into the cytoplasm of cells due to its negative charge. For the two studied formulations, the size of the free mRNA obtained by TDA was smaller than the one of the encapsulated mRNA, which shows a potential degradation of the free mRNA when the formulations are stored in non-RNase free conditions. mRNA integrity is a critical parameter, as in case of degradation, it can no longer be translated into the protein of interest. Therefore, the vaccine efficacy highly depends on the rate of encapsulation and on mRNA integrity.
Finally, this study showed that a single capillary electrophoresis equipment is able to provide valuable and complementary information on mRNA-LNP formulations. As TDA is easy to automatize, inexpensive and allow to work under physiological conditions, it could be advantageously used for quality control and stability studies to control LNPs size and mRNA size integrity. As it requires small volumes (nL are injected, 20 µL typically handled), TDA can also be used for research purpose to optimize LNP formulations parameters.
Data availability
Additional data are available in the Supplementary Information file.
References
Rappuoli R, Pizza M, Del Giudice G, De Gregorio E. Vaccines, new opportunities for a new society. Proc Natl Acad Sci. 2014;111:12288–93.
Pollard AJ, Bijker EM. A guide to vaccinology: from basic principles to new developments. Nat Rev Immunol. 2021;21:83–100.
Morens DM, Fauci AS. Emerging pandemic diseases: how we got to COVID-19. Cell. 2020;182:1077–92.
Hajj KA, Whitehead KA. Tools for translation: non-viral materials for therapeutic mRNA delivery. Nat Rev Mater. 2017;2:17056.
Nanomedicine and the COVID-19 vaccines. Nat Nanotechnol. 2020;15, 963–963.
Gebre MS, Brito LA, Tostanoski LH, Edwards DK, Carfi A, Barouch DH. Novel approaches for vaccine development. Cell. 2021;184:1589–603.
Wolff JA, Malone RW, Williams P, Chong W, Acsadi G, Jani A, et al. Direct gene transfer into mouse muscle in vivo. Science. 1990;247:1465–8.
Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines—a new era in vaccinology. Nat Rev Drug Discov. 2018;17:261–79.
Guan S, Rosenecker J. Nanotechnologies in delivery of mRNA therapeutics using nonviral vector-based delivery systems. Gene Ther. 2017;24:133–43.
Zhang X, Goel V, Robbie GJ. Pharmacokinetics of patisiran, the first approved RNA interference therapy in patients with hereditary transthyretin-mediated amyloidosis. J Clin Pharmacol. 2020;60:573–85.
Urits I, Swanson D, Swett MC, Patel A, Berardino K, Amgalan A, et al. A review of patisiran (ONPATTRO®) for the treatment of polyneuropathy in people with hereditary transthyretin amyloidosis. Neurol Ther. 2020;9:301–15.
Akinc A, Maier MA, Manoharan M, Fitzgerald K, Jayaraman M, Barros S, et al. The Onpattro story and the clinical translation of nanomedicines containing nucleic acid-based drugs. Nat Nanotechnol. 2019;14:1084–7.
Kim YC, Dema B, Reyes-Sandoval A. COVID-19 vaccines: breaking record times to first-in-human trials. Npj Vaccines. 2020;5:34.
Friedrichs S, Bowman DM. COVID-19 may become nanomedicine’s finest hour yet. Nat Nanotechnol. 2021;16:362–4.
Schoenmaker L, Witzigmann D, Kulkarni JA, Verbeke R, Kersten G, Jiskoot W, et al. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int J Pharm. 2021;601:120586.
Crommelin DJA, Anchordoquy TJ, Volkin DB, Jiskoot W, Mastrobattista E. Addressing the cold reality of mRNA vaccine stability. J Pharm Sci. 2021;110:997–1001.
Gilleron J, Querbes W, Zeigerer A, Borodovsky A, Marsico G, Schubert U, et al. Image-based analysis of lipid nanoparticle–mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat Biotechnol. 2013;31:638–46.
Sabnis S, Kumarasinghe ES, Salerno T, Mihai C, Ketova T, Senn JJ, et al. A novel amino lipid series for mRNA delivery: improved endosomal escape and sustained pharmacology and safety in non-human primates. Mol Ther. 2018;26:1509–19.
Chapin-Bardales J, Gee J, Myers T. Reactogenicity following receipt of mRNA-based COVID-19 vaccines. JAMA. 2021;325:2201–2.
Cheng Q, Wei T, Farbiak L, Johnson LT, Dilliard SA, Siegwart DJ. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat Nanotechnol. 2020;15:313–20.
Dammes N, Goldsmith M, Ramishetti S, Dearling JLJ, Veiga N, Packard AB, et al. Conformation-sensitive targeting of lipid nanoparticles for RNA therapeutics. Nat Nanotechnol. 2021;16:1030–8.
Wilhelm S, Tavares A, Dai Q, Ohta S, Audet J, Dvorak HF, et al. Analysis of nanoparticle delivery to tumours. Nat Rev Mater. 2016;1:16014.
Jiang W, Kim BYS, Rutka JT, Chan WCW. Nanoparticle-mediated cellular response is size-dependent. Nat Nanotechnol. 2008;3:145–50.
Conner SD, Schmid SL. Regulated portals of entry into the cell. Nature. 2003;422:37–44.
Rejman J, Oberle V, Zuhorn IS, Hoekstra D. Size-dependent internalization of particles via the pathways of clathrin- and caveolae-mediated endocytosis. Biochem J. 2004;377:159–69.
Petros RA, DeSimone JM. Strategies in the design of nanoparticles for therapeutic applications. Nat Rev Drug Discov. 2010;9:615–27.
Andar AU, Hood RR, Vreeland WN, DeVoe DL, Swaan PW. Microfluidic preparation of liposomes to determine particle size influence on cellular uptake mechanisms. Pharm Res. 2014;31:401–13.
Evers MJW, Kulkarni JA, Van der Meel R, Cullis PR, Vader P, Schiffelers RM. State-of-the-art design and rapid-mixing production techniques of lipid nanoparticles for nucleic acid delivery. Small Methods. 2018;2:1700375.
Hoshyar N, Gray S, Han H, Bao G. The effect of nanoparticle size on in vivo pharmacokinetics and cellular interaction. Nanomed. 2016;11:673–92.
Chauhan VP, Jain RK. Strategies for advancing cancer nanomedicine. Nat Mater. 2013;12:958–62.
Le-Vinh B, Steinbring C, Wibel R, Friedl JD, Bernkop-Schnürch A. Size shifting of solid lipid nanoparticle system triggered by alkaline phosphatase for site specific mucosal drug delivery. Eur J Pharm Biopharm. 2021;163:109–19.
Trevaskis NL, Kaminskas LM, Porter CJH. From sewer to saviour—targeting the lymphatic system to promote drug exposure and activity. Nat Rev Drug Discov. 2015;14:781–803.
Mitchell MJ, Billingsley MM, Haley RM, Wechsler ME, Peppas NA, Langer R. Engineering precision nanoparticles for drug delivery. Nat Rev Drug Discov. 2021;20:101–24.
Blanco E, Shen H, Ferrari M. Principles of nanoparticle design for overcoming biological barriers to drug delivery. Nat Biotechnol. 2015;33:941–51.
Demelenne A, Servais AC, Crommen J, Fillet M. Analytical techniques currently used in the pharmaceutical industry for the quality control of RNA-based therapeutics and ongoing developments. J Chromatogr. A. 2021;1651:462283.
Fan Y, Marioli M, Zhang K. Analytical characterization of liposomes and other lipid nanoparticles for drug delivery. J Pharm Biomed Anal. 2021;192:113642.
Malburet C, Leclercq L, Cotte JF, Thiebaud J, Bazin E, Garinot M, et al. Size and charge characterization of lipid nanoparticles for mRNA vaccines. Anal Chem. 2022;94:4677–85.
Chamieh J, Leclercq L, Martin M, Slaoui S, Jensen H, Østergaard J, et al. Limits in size of taylor dispersion analysis: representation of the different hydrodynamic regimes and application to the size-characterization of cubosomes. Anal Chem. 2017;89:13487–93.
Cipelletti L, Biron JP, Martin M, Cottet H. Measuring arbitrary diffusion coefficient distributions of nano-objects by Taylor Dispersion Analysis. Anal Chem. 2015;87:8489–96.
Zhang S, Li J, Lykotrafitis G, Bao G, Suresh S. Size-dependent endocytosis of nanoparticles. Adv Mater. 2009;21:419–24.
Belliveau NM, Huft J, Lin PJ, Chen S, Leung AK, Leaver TJ, et al. Microfluidic synthesis of highly potent limit-size lipid nanoparticles for in vivo delivery of siRNA. Mol Ther—Nucleic Acids. 2012;1:e37.
Yanez Arteta M, Kjellman T, Bartesaghi S, Wallin S, Wu X, Kvist AJ, et al. Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc Natl Acad Sci. 2018;115:E3351.
Bao Y, Jin Y, Chivukula P, Zhang J, Liu Y, Liu J, et al. Effect of PEGylation on biodistribution and gene silencing of siRNA/lipid nanoparticle complexes. Pharm Res. 2013;30:342–51.
Kozma GT, Shimizu T, Ishida T, Szebeni J. Anti-PEG antibodies: properties, formation, testing and role in adverse immune reactions to PEGylated nano-biopharmaceuticals. Adv Liposome Res. 2020;154–155:163–75.
Hoang Thi TT, Pilkington EH, Nguyen DH, Lee JS, Park KD, Truong NP. The importance of poly(ethylene glycol) alternatives for overcoming PEG immunogenicity in drug delivery and bioconjugation. Polymers. 2020;12:298.
Nogueira SS, Schlegel A, Maxeiner K, Weber B, Barz M, Schroer MA, et al. Polysarcosine-functionalized lipid nanoparticles for therapeutic mRNA delivery. ACS Appl Nano Mater. 2020;3:10634–45.
Joh DY, Zimmers Z, Avlani M, Heggestad JT, Aydin HB, Ganson N, et al. Architectural modification of conformal PEG-bottlebrush coatings minimizes Anti-PEG antigenicity while preserving stealth properties. Adv Healthc Mater. 2019;8:1801177.
Ou H, Cheng T, Zhang Y, Liu J, Ding Y, Zhen J, et al. Surface-adaptive zwitterionic nanoparticles for prolonged blood circulation time and enhanced cellular uptake in tumor cells. Acta Biomater. 2018;65:339–48.
Pitek AS, Jameson SA, Veliz FA, Shukla S, Steinmetz NF. Serum albumin ‘camouflage’ of plant virus based nanoparticles prevents their antibody recognition and enhances pharmacokinetics. Biomaterials. 2016;89:89–97.
Gulati NM, Stewart PL, Steinmetz NF. Bioinspired shielding strategies for nanoparticle drug delivery applications. Mol Pharm. 2018;15:2900–9.
Roces CB, Lou G, Jain N, Abraham S, Thomas A, Halbert GW et al. Manufacturing considerations for the development of lipid nanoparticles using microfluidics. Pharmaceutics. 2020;12:1095.
Zhigaltsev IV, Belliveau N, Hafez I, Leung AK, Huft J, Hansen C, et al. Bottom-up design and synthesis of limit size lipid nanoparticle systems with aqueous and triglyceride cores using millisecond microfluidic mixing. Langmuir. 2012;28:3633–40.
Chiesa E, Greco A, Riva F, Tosca EM, Dorati R, Pisani S, et al. Staggered herringbone microfluid device for the manufacturing of chitosan/TPP nanoparticles: systematic optimization and preliminary biological evaluation. Int. J. Mol. Sci. 2019;20:6212.
Cottet H, Biron JP, Martin M. Taylor dispersion analysis of mixtures. Anal. Chem. 2007;79:9066–73.
U.S. FDA. Drug products, including biological products, that contain nanomaterials—guidance for Industry. 2017.
Let’s talk about lipid nanoparticles. Nat Rev Mater. 2021;6, 99–99.
Reichmuth AM, Oberli MA, Jaklenec A, Langer R, Blankschtein D. mRNA vaccine delivery using lipid nanoparticles. Ther Deliv. 2016;7:319–34.
Funding
This work was partly funded by Sanofi Pasteur under a Cooperative Research and Development Agreement with the University of Montpellier and the CNRS.
Author information
Authors and Affiliations
Contributions
CM, LL, J-FC, JT, and HC designed this study. CM and EB formulated the LNPs. CM performed the TDA and DLS experiments. LL, J-FC, JT, EB, MG, and HC reviewed the data. CM wrote the manuscript. LL, J-FC, JT, EB, MG, and HC reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
Camille Malburet, Jean-François Cotte, Jérôme Thiebaud, Emilie Bazin and Marie Garinot are Sanofi employees and may hold shares or stocks in the company.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Malburet, C., Leclercq, L., Cotte, JF. et al. Taylor Dispersion Analysis to support lipid-nanoparticle formulations for mRNA vaccines. Gene Ther 30, 421–428 (2023). https://doi.org/10.1038/s41434-022-00370-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41434-022-00370-1