Abstract
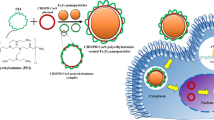
The metabolic instability of mRNA currently limits its utility for gene therapy. Compared to plasmid DNA, mRNA is significantly more susceptible to digestion by RNase in the circulation following systemic dosing. To increase mRNA metabolic stability, we hybridized a complementary reverse mRNA with forward mRNA to generate double-stranded mRNA (dsmRNA). RNase A digestion of dsmRNA established a 3000-fold improved metabolic stability compared to single-stranded mRNA (ssmRNA). Formulation of a dsmRNA polyplex using a PEG-peptide further improved the stability by 3000-fold. Hydrodynamic dosing and quantitative bioluminescence imaging of luciferase expression in the liver of mice established the potent transfection efficiency of dsmRNA and dsmRNA polyplexes. However, hybridization of the reverse mRNA against the 5′ and 3′ UTR of forward mRNA resulted in UTR denaturation and a tenfold loss in expression. Repeat dosing of dsmRNA polyplexes produced an equivalent transient expression, suggesting the lack of an immune response in mice. Co-administration of excess uncapped dsmRNA with a dsmRNA polyplex failed to knock down expression, suggesting that dsmRNA is not a Dicer substrate. Maximal circulatory stability was achieved using a fully complementary dsmRNA polyplex. The results established dsmRNA as a novel metabolically stable and transfection-competent form of mRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Wu GY, Wu CH. Receptor-mediated gene delivery and expression in vivo. J Biol Chem. 1988;263:14621–4.
Wu CH, Wilson JM, Wu GY. Targeting genes: delivery and persistent expression of a foreign gene driven by mammalian regulatory elements in vivo. J Biol Chem. 1989;264:16985–7.
Wu GY, Wilson JM, Shalaby F, Grossman M, Shafritz DA, Wu CH. Receptor-mediated gene delivery in vivo. Partial correction of genetic analbuminemia in nagase rats. J Biol Chem. 1991;266:14338–42.
Wilson JM, Grossman M, Wu CH, Chowdhury NR, Wu GY, Chowdhury JR. Hepatocyte-directed gene transfer in vivo leads to transient improvement of hypercholesterolemia in low density lipoprotein receptor-deficient rabbits. J Biol Chem. 1992;267:963–7.
Perales JC, Ferkol T, Beegen H, Ratnoff OD, Hanson RW. Gene transfer in vivo: sustained expression and regulation of genes introduced into the liver by receptor-targeted uptake. Proc Natl Acad Sci USA. 1994;91:4086–90.
Stankovics J, Crane AM, Andrews E, Wu CH, Wu GY, Ledley FD. Overexpression of human methylmalonyl CoA mutase in mice after in vivo gene transfer with asialoglycoprotein polylysine DNA complexes. Hum Gene Ther. 1994;5:1095–104.
Hara T, Tan Y, Huang L. In vivo gene delivery to the liver using reconstituted chylomicron remnants as a novel nonviral vector. Proc Natl Acad Sci USA. 1997;94:14547–52.
Nishikawa M, Takemura S, Takakura Y, Hashida M. Targeted delivery of plasmid DNA to hepatocytes in vivo: optimization of the pharmacokinetics of plasmid DNA/galactosylated Poly(L-lysine) complexes by controlling their physicochemical properties. JPET. 1998;287:408–15.
Kwoh DY, Coffin CC, Lollo CP, Jovenal J, Banaszczyk MG, Mullen P, et al. Stabilization of poly-L-lysine/DNA polyplexes for in vivo gene delivery to the liver. Biochim Biophys Acta. 1999;1444:171–90.
Kawakami S, Fumoto S, Nishikawa M, Yamashita F, Hashida M. In vivo gene delivery to the liver using novel galactosylated cationic liposomes. Pharm Res. 2000;17:306–13.
Nishikawa M, Yamauchi M, Morimoto K, Ishida E, Takakura Y, Hashida M. Heptocyte-targeted in vivo gene expression by intraveneous injection of plasmid DNA complexed with synthetic multi-functional gene delivery system. Gene Ther. 2000;7:548–55.
Al-Dosari MS, Knapp JE, Liu D. Hydrodynamic delivery. Adv Genet. 2005;54:65–82.
Phua KKL, Leong KW, Nair SK. Transfection efficiency and transgene expression kinetics of mRNA delivered in naked and nanoparticle format. J Control Release. 2013;166:227–33.
Wang Y, Su H-h, Yang Y, Hu Y, Zhang L, Blancafort P, et al. Systemic delivery of modified mRNA encoding herpes simplex virus 1 thymidine kinase for targeted cancer gene therapy. Mol Ther. 2013;21:358–67.
Li J, He Y, Wang W, Wu C, Hong C, Hammond PT. Polyamine-mediated stoichiometric assembly of ribonucleoproteins for enhanced mRNA delivery. Angew Chem Int Ed Engl. 2017;56:13709–12.
Li J, Wang W, He Y, Li Y, Yan EZ, Zhang K, et al. Structurally programmed assembly of translation initiation nanoplex for superior mRNA delivery. ACS Nano. 2017;11:2531–44.
Pardi N, Secreto AJ, Shan X, Debonera F, Glover J, Yi Y, et al. Administration of nucleoside-modified mRNA encoding broadly neutralizing antibody protects humanized mice from HIV-1 challenge. Nat Commun. 2017;8:14630.
Pardi N, Tuyishime S, Muramatsu H, Kariko K, Mui BL, Tam YK, et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J Control Release. 2015;217:345–51.
Thess A, Grund S, Mui BL, Hope MJ, Baumhof P, Fotin-Mleczek M, et al. Sequence-engineered mRNA without chemical nucleoside modifications enables an effective protein therapy in large animals. Mol Ther. 2015;23:1456–64.
Kauffman KJ, Mir FF, Jhunjhunwala S, Kaczmarek JC, Hurtado JE, Yang JH, et al. Efficacy and immunogenicity of unmodified and pseudouridine-modified mRNA delivered systemically with lipid nanoparticles in vivo. Biomaterials. 2016;109:78–87.
Ramaswamy S, Tonnu N, Tachikawa K, Limphong P, Vega JB, Karmali PP, et al. Systemic delivery of factor IX messenger RNA for protein replacement therapy. Proc Natl Acad Sci USA. 2017;114:E1941–E1950.
Jacobs F, Wisse E, De Geest B. The role of liver sinusoidal cells in hepatocyte-directed gene transfer. Am J Pathol. 2010;176:14–21.
Wisse E, Jacobs F, Topal B, Frederik P, De Geest B. The size of endothelial fenestrae in human liver sinusoids: implications for hepatocyte-directed gene transfer. Gene Ther. 2008;15:1193–9.
Hu Y, Haynes MT, Wang Y, Liu F, Huang L. A highly efficient synthetic vector: nonhydrodynamic delivery of DNA to hepatocyte nuclei in vivo. ACS Nano. 2013;7:5376–84.
Yao J, Fan Y, Li Y, Huang L. Strategies on the nuclear-targeted delivery of genes. J Drug Target. 2013;21:926–39.
Lam AP, Dean DA. Progress and prospects: nuclear import of nonviral vectors. Gene Ther. 2010;17:439–47.
Guan S, Rosenecker J. Nanotechnologies in delivery of mRNA therapeutics using nonviral vector-based delivery systems. Gene Ther. 2017;24:133–43.
DeRosa F, Guild B, Karve S, Smith L, Love K, Dorkin JR, et al. Therapeutic efficacy in a hemophilia B model using a biosynthetic mRNA liver depot system. Gene Ther. 2016;23:699–707.
Tsui NB, Ng EK, Lo YM. Stability of endogenous and added RNA in blood specimens, serum, and plasma. Clin Chem. 2002;48:1647–53.
Chen Q, Qi R, Chen X, Yang X, Wu S, Xiao H, et al. A targeted and stable polymeric nanoformulation enhances systemic delivery of mRNA to tumors. Mol Ther. 2017;25:92–101.
Kizzire K, Khargharia S, Rice KG. High-affinity PEGylated polyacridine peptide polyplexes mediate potent in vivo gene expression. Gene Ther. 2013;20:407–16.
Fernandez CA, Baumhover NJ, Duskey JT, Khargharia S, Kizzire K, Ericson MD, et al. Metabolically stabilized long-circulating PEGylated polyacridine peptide polyplexes mediate hydrodynamically stimulated gene expression in liver. Gene Ther. 2011;18:23–37.
Khargharia S, Kizzire K, Ericson MD, Baumhover NJ, Rice KG. PEG length and chemical linkage controls polyacridine peptide DNA polyplex pharmacokinetics, biodistribution, metabolic stability and in vivo gene expression. J Control Release. 2013;170:325–33.
Crowley ST, Poliskey JA, Baumhover NJ, Rice KG. Efficient expression of stabilized mRNA PEG-peptide polyplexes in liver. Gene Ther. 2015;22:993–9.
Nellimarla S, Mossman KL. Extracellular dsRNA: its function and mechanism of cellular uptake. J Interferon Cytokine Res. 2014;34:419–26.
Ueda T, Tohda H, Chikazumi N, Eckstein F, Watanabe K. Phosphorothioate-containing RNAs show mRNA activity in the prokaryotic translation systems in vitro. Nucleic Acids Res. 1991;19:547–52.
Hunter T, Hunt T, Jackson RJ, Robertson HD. The characteristics of inhibition of protein synthesis by double-stranded ribonucleic acid in reticulocyte lysates. J Biol Chem. 1975;250:409–17.
McAnuff MA, Rettig GR, Rice KG. Potency of siRNA versus shRNA mediated knockdown in vivo. J Pharm Sci. 2007;96:2922–30.
Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines - a new era in vaccinology. Nat Rev Drug Discov. 2018;17:261–279.
Schlake T, Thess A, Fotin-Mleczek M, Kallen K-J. Developing mRNA-vaccine technologies. RNA Biol. 2012;9:1319–30.
McCaffrey AP, Ohashi K, Meuse L, Shen S, Lancaster AM, Lukavsky PJ, et al. Determinants of hepatitis C translational initiation in vitro, in cultured cells and mice. Mol Ther. 2002;5:676–84.
Wilber A, Frandsen JL, Geurts JL, Largaespada DA, Hackett PB, McIvor RS. RNA as a source of transposase for sleeping beauty-mediated gene insertion and expression in somatic cells and tissues. Mol Ther. 2006;13:625–30.
Tam YYC, Chen S, Cullis PR. Advances in lipid nanoparticles for siRNA delivery. Pharmaceutics. 2013;5:498–507.
Nair JK, Willoughby JL, Chan A, Charisse K, Alam MR, Wang Q, et al. Multivalent N-acetylgalactosamine-conjugated siRNA localizes in hepatocytes and elicits robust RNAi-mediated gene silencing. J Am Chem Soc. 2014;136:16958–61.
Oupický D, Koňák Č, Dash PR, Seymour LW, Ulbrich K. Effect of albumin and polyanion on the structure of DNA complexes with polycation containing hydrophilic nonionic block. Bioconjug Chem. 1999;10:764–72.
Khargharia S, Baumhover NJ, Crowley ST, Duskey J, Rice KG. The uptake mechanism of PEGylated DNA polyplexes by the liver influences gene expression. Gene Ther. 2014;21:1021–8.
Baumhover NJ, Duskey JT, Khargharia S, White CW, Crowley ST, Allen RJ, et al. Structure–activity relationship of PEGylated polylysine peptides as scavenger receptor inhibitors for non-viral gene delivery. Mol Pharm. 2015;12:4321–28.
McCaffrey AP, Meuse L, Pham TT, Conklin DS, Hannon GJ, Kay MA. RNA interference in adult mice. Nature. 2002;418:38–9.
Qu X, Wen JD, Lancaster L, Noller HF, Bustamante C, Tinoco I Jr. The ribosome uses two active mechanisms to unwind messenger RNA during translation. Nature. 2011;475:118–21.
Ameres SL, Martinez J, Schroeder R. Molecular basis for target RNA recognition and cleavage by human RISC. Cell. 2007;130:101–12.
Karikó K, Muramatsu H, Welsh FA, Ludwig J, Kato H, Akira S, et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol Ther. 2008;16:1833–40.
Chattopadhyay A, Raghuraman H. Application of fluorescence spectroscopy to membrane protein structure and dynamics. Curr Sci. 2004;87:175–80. pp
Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD. How cells respond to interferons. Annu Rev Biochem. 1998;67:227–64.
Ramanathan A, Robb GB, Chan SH. mRNA capping: biological functions and applications. Nucleic Acids Res. 2016;44:7511–26.
Zust R, Cervantes-Barragan L, Habjan M, Maier R, Neuman BW, Ziebuhr J, et al. Ribose 2’-O-methylation provides a molecular signature for the distinction of self and non-self mRNA dependent on the RNA sensor Mda5. Nat Immunol. 2011;12:137–43.
Acknowledgements
The authors gratefully acknowledge support from NIH Grants GM117785 and T32 GM00865.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Poliskey, J.A., Crowley, S.T., Ramanathan, R. et al. Metabolically stabilized double-stranded mRNA polyplexes. Gene Ther 25, 473–484 (2018). https://doi.org/10.1038/s41434-018-0038-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41434-018-0038-3