Abstract
RNA binding motif protein X‐linked (RBMX) encodes the heterogeneous nuclear ribonucleoprotein G (hnRNP G) that regulates splicing, sister chromatid cohesion and genome stability. RBMX knock down experiments in various model organisms highlight the gene’s importance for brain development. Deletion of the RGG/RG motif in hnRNP G has previously been associated with Shashi syndrome, however involvement of other hnRNP G domains in intellectual disability remain unknown. In the current study, we present the underlying genetic and molecular cause of Gustavson syndrome. Gustavson syndrome was first reported in 1993 in a large Swedish five-generation family presented with profound X-linked intellectual disability and an early death. Extensive genomic analyses of the family revealed hemizygosity for a novel in-frame deletion in RBMX in affected individuals (NM_002139.4; c.484_486del, p.(Pro162del)). Carrier females were asymptomatic and presented with skewed X-chromosome inactivation, indicating silencing of the pathogenic allele. Affected individuals presented minor phenotypic overlap with Shashi syndrome, indicating a different disease-causing mechanism. Investigation of the variant effect in a neuronal cell line (SH-SY5Y) revealed differentially expressed genes enriched for transcription factors involved in RNA polymerase II transcription. Prediction tools and a fluorescence polarization assay imply a novel SH3-binding motif of hnRNP G, and potentially a reduced affinity to SH3 domains caused by the deletion. In conclusion, we present a novel in-frame deletion in RBMX segregating with Gustavson syndrome, leading to disturbed RNA polymerase II transcription, and potentially reduced SH3 binding. The results indicate that disruption of different protein domains affects the severity of RBMX-associated intellectual disabilities.
Similar content being viewed by others
Introduction
RNA binding proteins (RBPs) are ubiquitously expressed regulators of RNA processing, especially important for post-mitotic neurons to control temporal individual differences in axonal growth, plasticity, and function [1]. Heterogeneous nuclear ribonucleoproteins (hnRNPs) belong to the RBP family of structurally related proteins, recently highlighted in Gillentine et al. 2021, as not fully investigated candidate genes for neurodevelopmental disorders (NDDs). NDDs associated with hnRNPs have many shared phenotypes including severe structural brain abnormalities, intellectual disability, seizures, speech delay and hypotonia, and are suggested to share a common molecular pathogenesis [2]. RNA binding motif protein X‐linked (RBMX) encodes hnRNP G important for splicing regulation [3], sister chromatid cohesion regulation [4] and genome stability [5] specifically in neurons [6]. hnRNP G is part of the supraspliceosome, previously reported to be important for alternative splice site selection by protein binding (C-terminal) and RNA binding (RNA recognition motif;RRM) [3]. Overexpression of RBMX leads to exon skipping or inclusion where hnRNP G-dependent exons are significantly enriched in CCA/CCC motifs, suggesting a function in exon skipping/exon inclusion regulation [3]. RBMX morpholino knock down in zebrafish [7], African frog [8] and neuronal in-vitro studies [9, 10] highlight the gene’s importance for brain development and function. Moreover, a 23 bp frameshift deletion disrupting the RBMX RGG/RG motif in the RNA binding domain is associated with a mild to moderate X-linked intellectual disability (XLID) called Shashi syndrome (OMIM #300238) [10, 11]. However, the involvement of other hnRNP G domains in intellectual disability remains unknown.
Gustavson syndrome was first described in 1993 in six males and one female of a Swedish five-generation family (OMIM #309555) [12, 13]. The syndrome was characterized by profound intellectual disability, microcephaly, severe structural brain abnormalities, epileptic seizures, severe vision defect, hearing loss, congenital heart defects, psychomotor deficits, and an early death before 4 years of age due to pulmonary infections. Linkage analysis of 21 affected and unaffected family members indicated an association to the Xq26 region, including RBMX and 399 other genes, and an X-linked recessive inheritance pattern [13]. However, the genetic variant/s causing Gustavson syndrome was not established at that time.
In this study, we investigated the underlying genetic and molecular cause of Gustavson syndrome. Three additional affected family members have been identified since 1993 and presented striking phenotypic overlap with previously described patients. Genomic analyses of 36 family members revealed hemizygosity for an in-frame deletion in RBMX (NM_002139.4; c.484_486del, p.(Pro162del)) that segregated with the disease in the large family pedigree. Asymptomatic heterozygous females presented with a skewed X-chromosome inactivation (XCI) pattern, indicating silencing of the pathogenic allele. Transcriptomics and differential gene expression analysis of neuronal cells harboring the variant revealed that the top 100 differentially expressed genes (DEGs) were significantly enriched for transcription factors and genes involved in RNA polymerase II transcription, highlighting hnRNP Gs important role during transcription. Variant predictions and fluorescence polarization assay show that the variant is located in a highly conserved region which is likely to be a novel polyproline II helix/Src homology-3 (SH3) binding domain of hnRNP G. The fluorescence polarization assay indicates that the variant potentially lowers the binding affinity to proteins with SH3 domains.
Materials and methods
Patient samples
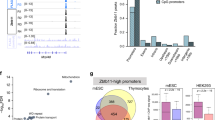
The investigated Swedish five-generation family consist of 91 individuals (Fig. 1A). Ten individuals (III:12, III:15, III:16, IV:3, IV:5, IV:16, IV:19, IV:23, V:3, V:7) were affected with Gustavson syndrome. Thirty-six individuals of the extended pedigree were included in the genetic analysis. DNA and RNA were extracted from blood by standard procedures (available upon request).
A Extended family pedigree of the family affected with Gustavson syndrome (91 individuals). Sanger sequencing of the RBMX variant in 36 individuals (blue squares) revealed that all investigated affected individuals were hemizygous (dark gray) and had inherited the allele from their carrier mothers. Patients with trisomy 18 is highlighted in light gray. B X-chromosome inactivation (XCI) analysis and variant detection on DNA and RNA confirms that asymptomatic carrier females (n = 11) are protected from disease by silencing the pathogenic allele (black arrow). XCI result from all individuals are presented in Supplementary Fig. 1. C Schematic of hnRNP G with known domains. Top: RBMX variants listed in the gnomAD and ClinVar databases. ClinVar variants are underlined, green: homozygous/hemizygous variants (all reported stop gained, start lost, splice acceptor, in-frame insertions/deletions or frameshift insertions/deletions), blue: heterozygous variants (all reported stop gained, start lost, splice acceptor, in-frame deletions or frameshift deletions), black: homozygous/hemizygous missense variants. Red lines: variants associated with Gustavson syndrome (p.(Pro162del)) and Shashi syndrome (p.Glu346fs). GnomaD variants obtained 22-07-28, including variants in the canonical transcript and excluding variants flagged as of poor quality or in low complexity regions. Bottom: Domains of hnRNP G are RNA recognition motif, RNA binding Arg-Gly-Gly-box, Nascent transcripts targeting domain, RNA binding domain (52, 54).
Genetic characterization of Gustavson syndrome
Genome sequencing by Illumina
Genome sequencing and analysis was performed on two family trios (III:19, III:20, IV:19 or IV:26, IV:27 and V:7) by two independent labs. The first trio (III:19, III:20, IV:19) were sequenced from 1 μg DNA using the TruSeq PCRfree DNA sample preparation kit (350 bp, cat.20015962/3, Illumina) and library prepped according to the manufacturers’ instructions (guide#1000000039279). Sequencing was performed on a NovaSeq S4 flowcell, with v1 sequencing chemistry, paired-ends and 150 bp read length. Alignment to reference genome (hg19) was performed using BWA-MEM and variant calling was performed using GATK 3.3.0. Manta Structural Variant Caller 1.0.3 was used to identify structural variants, and small insertions and deletions [14]. Variants were analyzed using MOON software (www.diploid.com/moon) with the filtering criteria: depth ≥8, genotype quality ≥40 and allele frequency ≤1.0% in gnomAD and Diploid database (www.diploid.com/moon). The other trio (IV:26, IV:27 and V:7) was sequenced and bioinformatically analyzed as previously reported [15]. All inheritance patterns were considered. All SNVs and SVs segregating with Gustavson syndrome in the extended pedigree were considered for further analysis.
Genome sequencing by 10X Genomics
Linked-read barcoded sequencing libraries were prepared from 1.25 ng DNA of a family trio (III:19, III:20, IV:19) using the Chromium Genome reagent kit v2 (cat.120257/58/61/62, 10X Genomics) according to the manufacturers’ protocol (#CG00043 Chromium Genome Reagent Kit v2 User Guide, 10X Genomics). Genome sequencing was performed on a NovaSeq 6000 S4 flowcell with v1 sequencing chemistry, paired-ends and a read length of 150 bp. The reads were mapped towards the reference genome hg38, and variants were called using LongRanger 2.2 (10X Genomics). Strict filtering was performed where large SVs (>30 kb) and deletions (50 bp-30 kb), shared in the mother and affected fetus were further investigated. Linked-read data were visualized using Loupe software 2.1.2 (10X Genomics). Variants were only considered if they segregated with Gustavson syndrome in the extended pedigree. Genome sequencing data from the family were used for segregation analysis. Variants were excluded if they were sequencing artefacts or present in our in-house database consisting of 1424 exomes from individuals with no signs of Gustavson syndrome [16].
In silico predictions of the p.(Pro162del) in RBMX
hnRNP G (P38159) 3D structure was predicted using AlphaFold2 [17]. The evolutionary conservation of the deleted bases of RBMX was investigated using phyloP and GERP scores. MutationTaster2 and PROVEAN were used to investigate the variant effect on protein function, SpliceAI and Human Splicing Finder were used to investigate the variant effect on alternative splicing. The genome databases gnomAD and SweGen [18] were used to investigate the variant landscape in RBMX present in the general population. Polyproline II secondary structures of hnRNP G (P38159) were predicted using the PPIIPRED software (http://bioware.ucd.ie/PPIIPRED) [19], and disordered regions were predicted using Database of Disordered Protein Predictions (D2P2) [20], and AlphaFold2 [17].
Segregation analysis
DNA was extracted from peripheral blood from 36 individuals in the extended pedigree (Fig. 1A and Supplementary Table 1). The region spanning the RBMX variant (NM_002139.4; c.484_486del, p.(Pro162del)) was amplified using 20–50 ng DNA, the RBMX_DNA primers and PCR protocol described in Supplementary material 1. Cycle sequencing was then performed using BigDye Terminator v3.1 Cycle Sequencing kit (Thermo Fisher Scientific, Carlsbad, CA, USA) according to manufacturer’s instructions with cycling parameters: 96°C 1 min, 30x (96°C 10 s, 50°C 5 s, 60°C 30 s) 4 °C hold. All samples were cleaned up using BigDye XTerminator Purification kit (Thermo Fisher Scientific, Bedford, MA, USA) and capillary electrophoresis was performed on 3130XL ABI Genetic Analyzer (Thermo Fisher Scientific, Foster City, CA, USA). The result was analyzed using CodonCode Aligner v9.0.1 (CodonCode Corporation, www.codoncode.com).
Molecular characterization of Gustavson syndrome
X-chromosome inactivation analysis of healthy carrier females
XCI patterns were investigated in eleven asymptomatic carrier females aged 20–60 years (III:20, III:18, III:22, II:3, III:4, IV:2, II:5, III:6, III:14, IV:26, IV:20), six asymptomatic non-carriers (III:9, III:2, IV:2, IV:9, II:10, II:12) and seven controls. XCI analysis of the AR and RP2 genes were performed using PCR and Fragment Length Analysis (FLA) as described before [21, 22]. Two hundred ng of genomic DNA were cut using HpaII FastDigest in a total volume of 20 µl following manufacturer’s instructions (Thermo Fisher Scientific, Baltics UAB, Vilnius, Lithuania). PCR was performed using 50 ng of DNA or 2 µl digested DNA as input, primers and PCR conditions as described in Supplementary material 1. FLA was performed on the 3130xl ABI Genetic Analyzer with ROX500 Size Standard (Thermo Fisher Scientific, Carlsbad, CA, USA) and the Amplified Fragment Length Polymorphism was determined using GeneMarker software v2.6.3 (SoftGenetics, State College, PA, USA). Polymorphic repeats in non-carrier females, parents and/or offspring from the family were used to indicate the pathogenic allele. The XCI result was confirmed with Sanger sequencing spanning the RBMX variant using cDNA synthesized from RNA blood (800 ng and 24 ng of RNA) of two carrier mothers (III:20, III:22) according to protocol (Maxima H Minus First Strand cDNA Synthesis Kit with dsDNase, Thermo Fisher Scientific, Waltham, MA, USA). PCR was performed as described in Supplementary material 1 using 50 ng cDNA and the primers RBMX_RNA.
RBMX splicing investigation
Due to the predicted variant effect on alternative splicing, a splicing analysis was performed on RNA extracted from peripheral blood of an affected individual (V:7). cDNA synthesis was performed on 2.2 µg RNA as described above. PCR was performed using 1 µl cDNA, primers spanning the whole gene (RBMX_Splicing_1 and RBMX_Splicing_2) and protocol described in Supplementary material 1. Sanger sequencing was performed using the Mix2seq kit (Eurofin Genomics, Ebersberg, Germany).
To confirm the result in other cell types, a mini-gene splicing assay was performed in HeLa cells and SH-SY5Y cells. The mini-gene construction, cell transfection, RNA extraction and cDNA synthesis were performed as described in Supplementary methods 1.
hnRNP G interaction study
To further investigate if the tri-proline region spanning the variant had the capacity to interact with SH3 domains, a fluorescence polarization assay with hnRNP G peptides and SH3 domains were performed. Wildtype and mutant (△P162) hnRNP G peptides of different lengths spanning the region of interest (P38159) were ordered from Genecast (Supplementary material 2) with or without Fluorescein isothiocyanate (FITC)-label (purity >95%, dissolved in 50 mM potassium phosphate pH 7.5). Two SH3 domains were expressed and purified: ASAP1-SH3 (Addgene: 91501), BIN1-SH3 (GenScript) according to standard protocol (Supplementary material 2).
To determine the affinity between the SH3 domains and the FITC-labeled hnRNP G peptides, saturation binding experiments were performed by preparing the 1:1 dilution series of increasing protein concentration (0.3–620 μM for ASAP1-SH3 and 0.425–870 μM for BIN1-SH3) and mixing it with a fixed concentration of FITC-labeled peptides (10 nM). Fluorescence polarization was measured with SpectraMax iD5 plate reader (Molecular Devices) at 25°C and excitation/emission wavelengths of 485/535 nm.
To determine the affinity between BIN-SH3 domain and the non-labeled hnRNP G peptides, displacement experiments were performed where the fixed concentration of FITC-labeled peptide and protein was mixed with an increasing concentration of displacer non-labeled peptide. The mP signal was investigated to determine the IC50 values, which were then converted to dissociation constants as previously described [23]. All results were analyzed with GraphPad Prism to determine the dissociation constant for labeled peptide and the IC50 values, and Microsoft Excel was used to calculate KD from IC50. All fluorescence polarization experiments were performed in 50 mM potassium phosphate pH 7.5, containing 1 mM tris(2-carboxyethyl)phosphine (TCEP).
Transcriptomics on neuronal cells expressing the p.(Pro162del) in RBMX
Transient transfection was performed in SH-SY5Y cells using pcDNA3.1( + )-C-eGFP plasmid constructs containing RBMX cDNA sequence with and without the variant (c.484_486del, p.(Pro162del)) (Supplementary material 3). To control for technical bias, two independent cell and sequencing experiments were performed, one in duplicates and the other in triplicates. For each experiment, SH-SY5Y cells (passage 65 or 67) were cultured as described previously and 3.4 × 106 cells were seeded to nine wells of 6-well plates. Transfection and RNA extraction was performed as previously described. Libraries were prepared using 260 ng of poly(A)-selected RNA with the TruSeq Stranded mRNA sample preparation kit according to the manufacturer’s protocol (Illumina Inc., San Diego, CA, USA). The quality of the libraries was evaluated using Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA, USA) using the RNA 6000 Nano reagent kit (Agilent, Hopkinton, MA, USA). RNA sequencing was performed on a SP flowcell using the NovaSeq 6000 system and v1.5 sequencing chemistry (Illumina Inc., San Diego, CA, USA) according to manufacturer’s instructions. Demultiplexing and conversion to FASTQ format was performed using bcl2fastq (v2.20.0.422). Trimmomatic (v0.39) was used for trimming adapter contamination [24], and quality was assessed using FastQC (v0.11.9) [25] and MultiQC (v1.12) [26]. STAR (v2.7.9a) was used for alignment to the reference genome (GRCh38.p13/hg38). Reads were counted to exons by using featureCounts (Rsubread v1.5.2) [27] followed by differential gene expression analysis with adjustment for batch effects using DESeq2 [28]. To identify enriched protein classes and pathways in the top 100 DEGs, PANTHER statistical overrepresentation test (v17.0) was used with correction for multiple testing using Bonferroni [29]. Data visualization was performed using Rstudio (v2021.09.1) and pheatmap with default settings (v1.0.12).
Protein expression and localization analysis
To investigate if the variant alters the location of hnRNP G a new transfection (SH-SY5Y cells, passage 69) was performed as previously described but using a µ-Slide 8 Well (ibidi, Gräfelfing, Germany). Images were taken 48 h post transfection using EVOS FL Auto 2 (Fisher Scientific, Willow Creek, Eugene, OR, USA) and protein expression was approximated using ImageJ (Rasband, W.S., ImageJ, U. S. National Institutes of Health, Bethesda, Maryland, USA, https://imagej.nih.gov/ij/, 1997–2018). A permutation test (N = 10,000) was performed to determine if the difference in mean values were significant (p < 0.05). Localization of hnRNP G was determined using a combination of GFP expression and DAPI staining. Imaging was performed using Zeiss LSM700 Confocal Microscope, at 40x magnification (Supplementary methods 3).
Results
Case reports of affected individuals
Six affected males (III:12, III:16, IV:3, IV:5, IV:16, IV:19) and one affected female (III:15) were first described in 1993 [19]. Since then, three additional affected males (IV:23, V:3, V:7) have been identified in this family, with striking phenotypic overlap to previously described affected individuals, including profound intellectual disability, seizures, microcephaly (Fig. 2; detailed case reports in Supplementary data 1).
A Magnetic resonance imaging showing lissencephaly with very thick cortex and dorsal dominance. As well as, some hypotrophy of vermis, while the frontal part of corpus callosum is well developed and the back part is thinner. B Patient V:7 showing dysmorphic features include bitemporal narrowing, wide mouth, puffy eyelids, broad mouth, high palate, two hemangiomas, microcephaly, short stature and overweight. For more detailed clinical description and phenotype comparison to other patients with Gustavson syndrome, see Supplementary data 1 and Supplementary Table 4B.
Genetic characterization of Gustavson syndrome
RBMX p.(Pro162del) segregates with Gustavson syndrome in the large family
Initially exome sequencing (Illumina) was performed on six family members (IV:5, III:6, IV:19, III:20, IV:26, V:7), but no candidate variants segregating with disease was identified. Then, genome sequencing was performed using Illumina and 10X Genomics sequencing. MOON software listed 60 variants following all inheritance patterns using Illumina genome sequencing. Loupe software detected 1262 deletions (50 bp-30 kb) and 41 large SVs (>30 kb) shared by the mother and child when using 10X Genomics sequencing. Both Illumina and Linked-read sequencing uncovered a variant of unknown significance (NM_002139.4; c.484_486del, p.(Pro162del)) in RBMX (RNA binding motif protein X-linked) located in the previously identified linked region Xq26. All variants except c.484_486del in RBMX were excluded as sequencing artefacts or because they did not segregate with disease (Supplementary Table 2). The result was confirmed by an independent second trio analysis in the same family. The variant is located in an evolutionary conserved locus (phyloP score 4.88, GERP 5.39). MutationTaster2 and PROVEAN suggest that the variant has a deleterious effect on protein function (PROVEAN score −8.666) and Human Splicing Finder predicted alternative splicing while SpliceAI did not. The c.484_486del variant is not present in public reference genomes (gnomAD, SweGen) and RBMX was reported to be depleted from pLoF variants (gnomAD pLI 0.83, o/e 0.14 CI 0.06–0.43) (Fig. 1C). Sanger sequencing of 36 samples from the family, spanning the identified RBMX variant confirmed that the variant segregated with Gustavson syndrome with an X-linked recessive inheritance pattern (Fig. 1A and Supplementary Table 1).
Molecular characterization of Gustavson syndrome
Skewed X-chromosome inactivation in healthy carrier females
XCI analysis [21, 22] was performed using PCR and FLA spanning the AR and RP2 genes in eleven carrier females, six non-carrier females and controls. Segregation of the polymorphic repeats in AR and RP2 indicated skewed XCI (100:0), silencing of the pathogenic allele in carrier females (Fig. 1B, Supplementary Fig. 1B). FLA analysis of non-carrier females showed normal XCI patterns, consistent with previously published data of the general population (Supplementary Fig. 1C) [30]. RNA sequencing spanning the RBMX variant confirmed that XCI leads to silencing of the pathogenic allele in two carrier females by only expressing the wildtype allele (Fig. 1B).
RBMX splicing assay reveals no variant effect on alternative splicing
Due to a predicted effect of the c.484_486del variant on alternative splicing, we performed a RBMX splicing analysis on blood from an affected hemizygous individual and in SH-SY5Y cells and HeLa cells overexpressing the mutant RBMX mini-gene. Sanger sequencing of cDNA showed no splicing effect within RBMX when the variant was present in the investigated tissues (Supplementary Fig. 2).
The p.(Pro162del) in hnRNP G is located in a novel SH3-binding motif, and potentially reduces the binding affinity to SH3 domains
The tri-proline region of hnRNP G (aa 160–162) was predicted to form a polyproline type-II helix structure (PPIIPRED aa 162;PPII score 0.66), located in a disordered region (aa 162;IUPred score 0.98, aa 160–162;AlphaFold2 pLDDT score 70, D2P2 consensus score) (Fig. 3A and Supplementary Fig. 3). Since polyproline II helices frequently bind to SH3 domains [31, 32], we further investigated if the tri-proline region had the capacity to interact with SH3 domains. Saturation binding experiments revealed binding to SH3 domains (BIN1 and ASAP1) with an affinity of 400 μM for a FITC-labeled peptide corresponding to residues 150–170 (Fig. 3B), and 40 μM for a non-labeled peptide (Fig. 3C), thus confirming SH3 domain binding capacity. To investigate if the putative SH3 recognition motif in the vicinity of the deletion is sufficient for the interaction [33], we measured the affinity for a shorter hnRNP G peptide (aa 156–169). The peptide showed weaker but comparable affinity for BIN1-SH3 domain (150 μM) (Fig. 3D), thus confirming the importance of the region. The adjacent N-terminal residues in hnRNP G (aa 150–170) contributed to the affinity.
A hnRNP G amino acid sequence (P38159) with the p.(Pro162del) variant highlighted with a star, predicted polyproline II helices highlighted in red (strong prediction) and blue (weaker prediction) using PPIIPRED prediction. B Saturation binding experiments between two SH3 domains and the labeled hnRNP G peptide (amino acids 150–170) revealed binding with moderate affinities to SH3 domains. C Displacement binding experiments between BIN1-SH3 domain and two longer hnRNP G peptides (wildtype and △P162 variant) showed no clear difference in SH3 binding affinity between the wildtype and the mutant peptide. D Displacement experiments between BIN1-SH3 and the shorter hnRNP G peptides (wildtype and △P162 variant), showing a reduced binding affinity to SH3 domains when the p.(Pro162del) variant was introduced.
The effect of the p.(Pro162del) on SH3-binding was tested both for the long and the short hnRNP G peptide described above. The short peptide (aa 156–169 △P162) displayed a threefold weaker affinity for BIN1-SH3 (KD = 400 μM), while the long peptide (aa 150–170 △P162) showed no clear difference in binding affinity compared to the wildtype peptide (KD = 30 μM) (Fig. 3).
Localization and expression of hnRNP G in neuronal cells
To investigate if the P162del variant affects the location and/or expression of hnRNP G, mutant and wildtype cDNA was transiently expressed in SH-SY5Y cells. Confocal fluorescent images showed no difference in location for hnRNP G between the cell-lines. Merged images of the GFP expression and DAPI show that hnRNP G is expressed in the nucleus (Supplementary Fig. 3A). Comparing the mean expression of the two populations resulted in a difference in intensity of 488.2 units. The permutation test resulted in a 95% confidence interval of [−2217.3, 2093.4], and indicate no significant difference in expression (p = 0.6641) (Supplementary Fig. 3B).
Transcriptomics of neuronal cells expressing the RBMX p.(Pro162del) variant
Transcriptomics and differential gene expression analysis of neuronal cells expressing the variant revealed seven significant DEGs, where mutant cells presented a significant upregulation (padj<0.05) of ZNF805, PCDHA10, LYSMD3 and downregulation of COL2A1, EVPL, TTN, AC010207.1. EVPL encodes for a protein with an SH3 domain (Supplementary Table 3). Hierarchical clustered heatmap visualization of the top 100 DEGs revealed a distinct difference in expression pattern when comparing the wildtype and mutant expression levels (Fig. 4). The top 100 DEGs were significantly enriched for transcription factors and genes in the RNA polymerase II transcription process (Supplementary Table 3B, C).
A Transcriptomics experimental setup with transient transfection of the wildtype vector (pink) or mutant vector (p.(Pro162del): turquoise) in SH-SY5Y cells. B The top 100 most differentially expressed genes between wildtype (pink) and mutant cells (turquoise). Each line represents the expression of one gene, where upregulated genes are red and downregulated genes are blue. The genes (rows) and samples (columns) of the top 100 most differentially expressed genes are hierarchical clustered according to expression patterns. Significantly upregulated genes (star) in the mutant cells (padj<0.05): ZNF805;ENSG00000204524.7, PCDHA10;ENSG00000250120.8, LYSMD3;ENSG00000176018.13. Significantly downregulated genes (star) in the mutant cells (padj<0.05): COL2A1;ENSG00000139219.19, EVPL;ENSG00000167880.8, TTN;ENSG00000120332.17, AC010207.1;ENSG00000260633.1.
Discussion
We have identified a novel in-frame deletion in RBMX (c.484_486del, p.(Pro162del)) leading to Gustavson syndrome (OMIM #309555) [12, 13] by disturbed RNA polymerase II transcription and potentially reduced SH3 binding. Linkage analysis previously mapped Gustavson syndrome to the Xq26 region, which includes RBMX and 399 other genes [13]. The RBMX variant was identified using genome sequencing, but missed during exome analyses due to poor alignment in a complex GC-rich region and presence of human retroposon-derived RBMX-like genes, emphasizing the usefulness of genome sequencing in RBMX-related disorders. XCI analysis of eleven carrier females showed skewed X-chromosome inactivation (100:0), indicating silencing of the pathogenic allele, and protection from disease. RNA analysis confirmed the XCI result by expression of only the wildtype allele. A crossover event in the Xq26 region, leading to exclusion of the RBMX variant, was observed in one non-carrier female (III:2) and her daughter (IV:1) (Supplementary Fig. 1) [13], strengthening the predicted pathogenic effect, since both were asymptomatic and the daughter (IV:1) had random X-inactivation pattern. Interestingly, a girl has been reported to suffer from Gustavson syndrome [12, 13]; however, due to lack of patient material further investigation of her is not possible, but could be caused by uniparental isodisomy, or an additional/alternative diagnosis such as Turner syndrome.
The variant p.(Pro162del) was predicted to have a deleterious effect on protein function and cause alternative splicing of RBMX. However, alternative splicing of RBMX caused by the variant was excluded as the mechanism of disease by no detected aberrant splicing events in RNA of an affected hemizygous individual, SH-SY5Y cells and HeLa cells overexpressing the mutant RBMX mini-gene. The variant is located in an evolutionary conserved locus (phyloP score 4.88, GERP 5.39) with an unknown function [34, 35]. The variant deletes a proline of a tri-proline structure, predicted to be a polyproline type-II helix [19]. Polyproline type-II helices are important for protein-protein interaction motifs especially for SH3 and Enabled/VASP Homology-1 domains [31, 32]. Polyproline type-II segments are often structurally flexible, with an important structure for protein folding [32], appearing within intrinsically disordered regions. The RBMX variant is located in a disordered region which has a flexible undefined structure depending on ligand binding and environment [17, 19, 20]. To our knowledge, the function of tri-prolines at amino acid residues 160–162 (hnRNP G; P38159) and protein interactions with this region, have not been published before.
To confirm the effect of the p.(Pro162del) variant on hnRNP G function, fluorescence polarization displacement experiments were performed. hnRNP G aa 156–169 was found to bind to BIN1-SH3 and ASAP1-SH3 domains with moderate affinities, thus confirming that this region of hnRNP G can bind to SH3 domains. However, residues 150–170 also contributed to the affinity. Published affinities of SH3 domains vary from low nanomolar to high micromolar [36, 37], and are affected by adjacent domains and motifs [33, 38]. Thus, the variant effect is context dependent and likely complex in vivo. The BIN1-SH3 and ASAP1-SH3 domains used in this experiment were chosen based on predicted specificities of recognition motifs [33]; however, they are likely not the natural cellular interaction partner/s of the hnRNP G SH3 binding motif. Further studies are needed to reveal the interaction partner/s in cells. In summary, our data suggest that residues 150–170 can interact with SH3 domains and the p.(Pro162del) variant may reduce the affinity.
Differential gene expression analysis on neuronal cells expressing the wildtype or mutant (c.484_486del;p.(Pro162del)) RBMX was performed. Transcriptomics analysis revealed seven significant DEGs with where mutant cells presented a significant downregulation of EVPL, COL2A1, TTN, AC010207.1 and upregulation of ZNF805, LYSMD3, PCDHA10. The low number of significant DEGs is likely a consequence of few replicates, indicated by a clustered heatmap of the top 100 DEGs showing a distinct expression pattern among the genes in the two groups (Fig. 4). Top 100 DEGs revealed an overrepresentation of genes involved in RNA polymerase II transcription, and specifically genes encoding transcription factors. One of the significantly downregulated genes encodes a protein with a SH3 domain (EVPL), further supporting the possibility that the variant reduces SH3-binding affinity. We also show that these differences are not due to a difference in expression or localization of hnRNP G. EVPL encodes envoplakin, and EVPL -/- mice and zebrafish indicate a function in the skin barrier development [39, 40]. The SH3 domain in EVPL is highly conserved in plakin family members, important for mechano-sensing [41] and have been suggested to not bind the canonical PXXP-binding groove [42], correlating with our suggested binding motif in hnRNP G (PPP) (Supplementary Fig. 4). COL2A is important for collagen production, bones, tissue, and sensory development [43]. ZNF805 is a transcription factor involved in RNA polymerase II transcription, consistent with the overrepresentation of transcription factors seen in our top 100 DEGs. LYSMD3 encodes a receptor on human airway epithelial cells, important for immune response to airway pathogens [44]. TTN is an important muscle protein in the heart, associated to a number of different disorders including neuromuscular disorders [45] and respiratory failure [46]. PCDHA10 encodes protocadherin α expressed in brain and eye [47], important in neurodevelopment [48].
RBMX has previously been associated with Shashi syndrome (OMIM #300238) [11] caused by a p.Glu346fs variant [11], leading to disruption of the RGG/RG motif and thus aberrant p53 activation and neuronal differentiation [10]. Patients with Gustavson syndrome have minor shared phenotypes with Shashi syndrome, severe phenotypes such as structural brain abnormalities, epilepsy, severe vision defects, hearing loss, and early death are not present in Shashi patients. This suggests that different variants disrupting different hnRNP G domains lead to distinct phenotypes. Furthermore, Gustavson syndrome has a larger phenotypic overlap with other hnRNP-associated neurodevelopmental disorders, including the most commonly reported symptoms such as intellectual disability, seizures, hypotonia and severe structural brain abnormalities [2] (Supplementary Table 4). To our knowledge, death has only been described in one patient with a hnRNP-associated neurodevelopmental disorder before (15 years old female), associated with a heterozygous predicted splice variant in hnRNPK (NM_031263.2;c.258-3 C > T) [2]. However, other likely gene disrupting variants in hnRNPK were not lethal, and missense variants in the same gene and/or domain result in the same severity as likely gene disrupting variants, indicating no clear genotype-phenotype correlation in this gene. Thus, Gustavson syndrome is the most severe intellectual disability syndrome described in this gene family so far. Early death is consistent with knock-down experiments performed in zebrafish [7] and depletion of pLoF variants in RBMX in the general human population (Fig. 1C), which may indicate that the p.(Pro162del) causes loss of hnRNP G function.
Human retroposon-derived RBMX Like 1 (RBMXL1) and RBMX Like 9 (RBMXL9) have 96% sequence similarity to RBMX and are expressed in various tissues including brain [49]. The gene retrocopies may potentially replace or compensate hnRNP G function; however, the tri-proline region of interest (aa 160–162; P38159) are disrupted in the retrocopies and therefore unlikely to compensate for the RBMX variant effect in our patients (Supplementary Fig. 5). Moreover, we did not identify any disease-modifying or additional potential disease-causing variants in RBMXL1 and RBMXL9 as likely contributions to Gustavson syndrome. However, more studies are needed to confirm the function of the retrocopies and we would like to highlight the possibility that phenotypic differences to Shashi syndrome may be due to variant rescue by gene retrocopy/ies.
In conclusion, an in-frame deletion in RBMX (c.484_486del, p.(Pro162del)) leads to Gustavson syndrome. Protein interaction analyses revealed the first indication that amino acid 156–169 of hnRNP G possibly is an SH3-binding motif and that the variant could reduce binding affinity to SH3-domains. Transcriptomics analyses indicated that the variant region has important functions in transcription by RNA polymerase II.
Data availability
The data generated and analyzed during the current study are not publicly available due to Swedish legislation. Data from transcriptomics analysis is available upon request.
References
Khalil B, Morderer D, Price PL, Liu F, Rossoll W. mRNP assembly, axonal transport, and local translation in neurodegenerative diseases. Brain Res. 2018;1693:75–91.
Gillentine MA, Wang T, Hoekzema K, Rosenfeld J, Liu P, Guo H, et al. Rare deleterious mutations of HNRNP genes result in shared neurodevelopmental disorders. Genome Med. 2021;13:63.
Heinrich B, Zhang Z, Raitskin O, Hiller M, Benderska N, Hartmann AM, et al. Heterogeneous nuclear ribonucleoprotein G regulates splice site selection by binding to CC(A/C)-rich regions in Pre-mRNA*. J Biol Chem. 2009;284:14303–15.
Matsunaga S, Takata H, Morimoto A, Hayashihara K, Higashi T, Akatsuchi K, et al. RBMX: a regulator for maintenance and centromeric protection of sister chromatid cohesion. Cell Rep. 2012;1:299–308.
Adamson B, Smogorzewska A, Sigoillot FD, King RW, Elledge SJ. A genome-wide homologous recombination screen identifies the RNA-binding protein RBMX as a component of the DNA-damage response. Nat Cell Biol. 2012;14:318–28.
Reid DA, Reed PJ, Schlachetzki JCM, Nitulescu II, Chou G, Tsui EC, et al. Incorporation of a nucleoside analog maps genome repair sites in postmitotic human neurons. Science. 2021 ;372:91–4.
Tsend-Ayush E, O’Sullivan LA, Grützner FS, Onnebo SMN, Lewis RS, Delbridge ML, et al. RBMX gene is essential for brain development in zebrafish. Dev Dyn. 2005;234:682–8.
Dichmann DS, Fletcher RB, Harland RM. Expression cloning in Xenopus identifies RNA-binding proteins as regulators of embryogenesis and Rbmx as necessary for neural and muscle development. Dev Dyn. 2008;237:1755–66.
Zhang G, Neubert TA, Jordan BA. RNA binding proteins accumulate at the postsynaptic density with synaptic activity. J Neurosci. 2012 ;32:599–609.
Cai T, Cinkornpumin JK, Yu Z, Villarreal OD, Pastor WA, Richard S. Deletion of RBMX RGG/RG motif in Shashi-XLID syndrome leads to aberrant p53 activation and neuronal differentiation defects. Cell Rep. 2021;36:109337.
Shashi V, Xie P, Schoch K, Goldstein DB, Howard TD, Berry MN, et al. The RBMX gene as a candidate for the Shashi X-linked intellectual disability syndrome. Clin Genet. 2015;88:386–90.
Gustavson KH, Annerén G, Malmgren H, Dahl N, Ljunggren CG, Bäckman H. New X-linked syndrome with severe mental retardation, severely impaired vision, severe hearing defect, epileptic seizures, spasticity, restricted joint mobility, and early death. Am J Med Genet. 1993;45:654–8.
Malmgren H, Sundvall M, Dahl N, Gustavson KH, Annerén G, Wadelius C, et al. Linkage mapping of a severe X-linked mental retardation syndrome. Am J Hum Genet. 1993;52:1046–52.
Chen X, Schulz-Trieglaff O, Shaw R, Barnes B, Schlesinger F, Källberg M, et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics. 2016;32:1220–2.
Stranneheim H, Lagerstedt-Robinson K, Magnusson M, Kvarnung M, Nilsson D, Lesko N, et al. Integration of whole genome sequencing into a healthcare setting: high diagnostic rates across multiple clinical entities in 3219 rare disease patients. Genome Med. 2021;13:40.
Ameur A, Bunikis I, Enroth S, Gyllensten U. CanvasDB: a local database infrastructure for analysis of targeted- and whole genome re-sequencing projects. Database (Oxford). 2014;2014:bau098.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Ameur A, Dahlberg J, Olason P, Vezzi F, Karlsson R, Martin M, et al. SweGen: a whole-genome data resource of genetic variability in a cross-section of the Swedish population. Eur J Hum Genet. 2017;25:1253–60.
O’Brien KT, Mooney C, Lopez C, Pollastri G, Shields DC. Prediction of polyproline II secondary structure propensity in proteins. R Soc Open Sci. 2020;7:191239.
Oates ME, Romero P, Ishida T, Ghalwash M, Mizianty MJ, Xue B, et al. D2P2: database of disordered protein predictions. Nucleic Acids Res. 2013;41:D508–516.
Allen RC, Zoghbi HY, Moseley AB, Rosenblatt HM, Belmont JW. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am J Hum Genet. 1992;51:1229–39.
Machado FB, Machado FB, Faria MA, Lovatel VL, Alves da Silva AF, Radic CP, et al. 5meCpG epigenetic marks neighboring a primate-conserved core promoter short tandem repeat indicate X-chromosome inactivation. PLoS One. 2014;9:e103714.
Nikolovska-Coleska Z, Wang R, Fang X, Pan H, Tomita Y, Li P, et al. Development and optimization of a binding assay for the XIAP BIR3 domain using fluorescence polarization. Anal Biochem. 2004;332:261–73.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Andrews S. FASTQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics, Babraham Institute, Cambridge, United Kingdom. 2010.
Ewels P, Magnusson M, Lundin S, Käller M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics. 2016;32:3047–8.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Mi H, Ebert D, Muruganujan A, Mills C, Albou LP, Mushayamaha T, et al. PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucleic Acids Res. 2021;49:D394–403.
Shvetsova E, Sofronova A, Monajemi R, Gagalova K, Draisma HHM, White SJ, et al. Skewed X-inactivation is common in the general female population. Eur J Hum Genet. 2019;27:455–65.
Gushchina LV, Gabdulkhakov AG, Nikonov SV, Filimonov VV. High-resolution crystal structure of spectrin SH3 domain fused with a proline-rich peptide. J Biomol Struct Dyn. 2011;29:485–95.
Rath A, Davidson AR, Deber CM. The structure of ‘unstructured’ regions in peptides and proteins: role of the polyproline II helix in protein folding and recognition. Biopolymers. 2005;80:179–85.
Teyra J, Huang H, Jain S, Guan X, Dong A, Liu Y, et al. Comprehensive analysis of the human SH3 domain family reveals a wide variety of non-canonical specificities. Structure. 2017;25:1598–1610.e3.
Elliott DJ, Dalgliesh C, Hysenaj G, Ehrmann I. RBMX family proteins connect the fields of nuclear RNA processing, disease and sex chromosome biology. Int J Biochem Cell Biol. 2019;108:1–6.
Soulard M, Della Valle V, Siomi MC, Piñol-Roma S, Codogno P, Bauvy C, et al. hnRNP G: sequence and characterization of a glycosylated RNA-binding protein. Nucleic Acids Res. 1993;21:4210–7.
Lewitzky M, Harkiolaki M, Domart MC, Jones EY, Feller SM. Mona/Gads SH3C binding to hematopoietic progenitor kinase 1 (HPK1) combines an atypical SH3 binding motif, R/KXXK, with a classical PXXP motif embedded in a polyproline type II (PPII) helix. J Biol Chem. 2004;279:28724–32.
Pisabarro MT, Serrano L, Wilmanns M. Crystal structure of the abl-SH3 domain complexed with a designed high-affinity peptide ligand: implications for SH3-ligand interactions. J Mol Biol. 1998;281:513–21.
Saksela K, Permi P. SH3 domain ligand binding: what’s the consensus and where’s the specificity? FEBS Lett. 2012;586:2609–14.
Määttä A, DiColandrea T, Groot K, Watt FM. Gene targeting of envoplakin, a cytoskeletal linker protein and precursor of the epidermal cornified envelope. Mol Cell Biol. 2001;21:7047–53.
Inaba Y, Chauhan V, van Loon AP, Choudhury LS, Sagasti A. Keratins and plakin family cytolinker proteins control the length of epithelial microridge protrusions. ELife. 2020;9:e58149.
Daday C, Kolšek K, Gräter F. The mechano-sensing role of the unique SH3 insertion in plakin domains revealed by Molecular Dynamics simulations. Sci Rep. 2017;7:11669.
Ortega E, Buey RM, Sonnenberg A, de Pereda JM. The structure of the plakin domain of plectin reveals a non-canonical SH3 domain interacting with its fourth spectrin repeat. J Biol Chem. 2011;286:12429–38.
Deng H, Huang X, Yuan L. Molecular genetics of the COL2A1-related disorders. Mutat Res Rev Mutat Res. 2016;768:1–13.
He X, Howard BA, Liu Y, Neumann AK, Li L, Menon N, et al. LYSMD3: a mammalian pattern recognition receptor for chitin. Cell Rep. 2021;36:109392.
Savarese M, Sarparanta J, Vihola A, Udd B, Hackman P. Increasing role of titin mutations in neuromuscular disorders. J Neuromuscul Dis. 2016;3:293–308.
Pfeffer G, Povitz M, Gibson GJ, Chinnery PF. Diagnosis of muscle diseases presenting with early respiratory failure. J Neurol. 2015;262:1101–14.
Kohmura N, Senzaki K, Hamada S, Kai N, Yasuda R, Watanabe M, et al. Diversity revealed by a novel family of cadherins expressed in neurons at a synaptic complex. Neuron. 1998;20:1137–51.
Wu Q, Maniatis T. A striking organization of a large family of human neural cadherin-like cell adhesion genes. Cell. 1999;97:779–90.
Lingenfelter PA, Delbridge ML, Thomas S, Hoekstra HE, Mitchell MJ, Graves JA, et al. Expression and conservation of processed copies of the RBMX gene. Mamm Genome. 2001;12:538–45.
Acknowledgements
We wish to acknowledge the family for participating in this long-term study. Thanks to Prof. Dr. Christel Depienne and Assistant Prof. Dr. Juliette Godin for great scientific discussions and collaboration. Exome sequencing, genome sequencing and transcriptome sequencing were performed by the SNP&SEQ Technology Platform in Uppsala, Sweden. The facility is part of the National Genomics Infrastructure (NGI) Sweden and Science for Life Laboratory. Thanks to BioVis, Uppsala University for helping with confocal microscopy and image analysis. This study was performed in memory of Prof. Karl-Henrik Gustavson who was very dedicated to find the underlying cause to disease in this family. The computations were enabled by resources in projects SNIC 2021/23-723 and SNIC 2021/22-897, provided by the Swedish National Infrastructure for Computing (SNIC) at UPPMAX, partially funded by the Swedish Research Council through grant agreement no. 2018-05973.
Funding
This work was supported by grants from Uppsala University, Faculty of Medicine, for psychiatric and neurological research, grants from Uppsala University Hospital, grants form Svenska Läkaresällskapet (SLS), grants from Jeansson foundations and grants from the Swedish Society for Medical Research (SSMF). The SNP&SEQ Platform is also supported by the Swedish Research Council and the Knut and Alice Wallenberg Foundation. S.G. was supported by grants from Knut and Alice Wallenberg Foundation. J.J. was supported by grants from the Sävstaholm Foundation. P.J. acknowledges a grant from the Swedish Research Council (2020-04395). Open access funding provided by Uppsala University.
Author information
Authors and Affiliations
Contributions
JJ, SL, FM, SG, SE, A-MM, MP, MV and MW performed the experiments. JJ, SL, FM, SG, SE, A-MM, MP, AA, MV, KL-R and MW analyzed the data. MV, AN, PJ, GA, MW and M-LB supervised research. CF, CG, KL-R, AN, GA and M-LB performed the clinical investigations.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study was approved by the local ethics committee for human research in Uppsala, Sweden (Dnr 2012/321, Dnr 2019/05635, Dnr 2020/06592) and Linköping, Sweden (Dnr 2015/129-31) prior to research. The genetic analyses were conducted in accordance with the guidelines of the Declaration of Helsinki. Written Informed consent for the study and all images was obtained from the participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Johansson, J., Lidéus, S., Frykholm, C. et al. Gustavson syndrome is caused by an in-frame deletion in RBMX associated with potentially disturbed SH3 domain interactions. Eur J Hum Genet 32, 333–341 (2024). https://doi.org/10.1038/s41431-023-01392-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-023-01392-y
This article is cited by
-
Solving medical mysteries with genomics
European Journal of Human Genetics (2024)
-
Comment on Gustavson syndrome is caused by an in-frame deletion in RBMX associated with potentially disturbed SH3 domain interactions
European Journal of Human Genetics (2023)