Abstract
Silver–Russell syndrome (SRS) is a rare imprinting disorder associated with prenatal and postnatal growth retardation. Loss of methylation (LOM) on chromosome 11p15 is observed in 40 to 60% of patients and maternal uniparental disomy (mUPD) for chromosome 7 (upd(7)mat) in ~5 to 10%. Patients with LOM or mUPD 14q32 can present clinically as SRS. Delta like non-canonical Notch ligand 1 (DLK1) is one of the imprinted genes expressed from chromosome 14q32. Dlk1-null mice display fetal growth restriction (FGR) but no genetic defects of DLK1 have been described in human patients born small for gestational age (SGA). We screened a cohort of SGA patients with a SRS phenotype for DLK1 variants using a next-generation sequencing (NGS) approach to search for new molecular defects responsible for SRS. Patients born SGA with a clinical suspicion of SRS and normal methylation by molecular testing at the 11p15 or 14q32 loci and upd(7)mat were screened for DLK1 variants using targeted NGS. Among 132 patients, only two rare variants of DLK1 were identified (NM_003836.6:c.103 G > C (p.(Gly35Arg) and NM_003836.6: c.194 A > G p.(His65Arg)). Both variants were inherited from the mother of the patients, which does not favor a role in pathogenicity, as the mono-allelic expression of DLK1 is from the paternal-inherited allele. We did not identify any pathogenic variants in DLK1 in a large cohort of SGA patients with a SRS phenotype. DLK1 variants are not a common cause of SGA.
Similar content being viewed by others
Introduction
Fetal growth restriction (FGR), defined as the failure of the fetus to reach its genetically determined growth potential, is one of the most common causes of perinatal mortality and morbidity [1]. It results from multiple causes, such as genetic and epigenetic alterations, the environment, hormonal dysregulation, or placental vascular dysfunction. More than 150 genetic disorders have been associated with FGR [2].
Silver–Russell syndrome (SRS, OMIM #180860) is a rare but well known imprinting disorder [3]. Clinical diagnosis of SRS is considered if a patient shows at least four of the six criteria of the Netchine–Harbison clinical scoring system (NH-CSS) [3, 4], which includes pre- and postnatal growth retardation, relative macrocephaly at birth, body asymmetry, protruding forehead, and early feeding difficulties. An underlying molecular cause is identified in ~60% of patients with SRS [3, 5]. Among them, loss of methylation (LOM) at H19/IGF2:IG-DMR (also called ICR1) on chromosome 11p15 (11p15 LOM) is observed in 40 to 60% and maternal uniparental disomy (mUPD) for chromosome 7 (upd(7)mat) in ~5 to 10% [3, 5,6,7]. The recent international consensus of SRS recommends additional molecular testing in cases of normal methylation on chromosomes 11 and 7, including screening of cyclin D kinase inhibitor 1c (CDKN1C) and insulin-like growth factor 2 (IGF2) genes [8, 9]. Since the first consensus on SRS, new molecular defects have been identified in high mobility group AT-hook 2 (HMGA2) and pleiomorphic adenoma gene 1 (PLAG1) in patients with a clinical presentation of SRS [10, 11]. The recent use of next-generation sequencing (NGS) for SRS patients has improved the molecular diagnosis [12,13,14,15,16], but more than 30% of patients with SRS remain without an identified molecular cause.
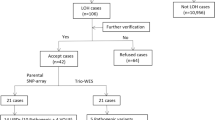
Temple Syndrome (TS) is another rare cause of prenatal and postnatal growth restriction caused by disruption of the 14q32 imprinted region. In this region, MEG3/DLK1:IG-DMR is normally methylated on the paternal allele [17], resulting in Delta like non-canonical Notch ligand 1 (DLK1), retrotransposon Gag like 1 (RTL1), and Iodothyronine Deiodinase 3 (DIO3) expression from the paternal allele [18]. In contrast, long noncoding RNAs (maternally expressed 3 (MEG3) and maternally expressed 8 (MEG8)), microRNAs, and small nucleolar RNAs are expressed by the unmethylated maternal allele (Fig. 1). mUPD of chromosome 14 (upd(14)mat), hypomethylation of MEG3/DLK1:IG-DMR, and paternal deletion of this region all lead to the phenotype of TS. Clinical overlap between SRS and TS has been described and Geoffron et al. reported 73% of patients with 14q32 disruption scoring positively for SRS, with a NHCSS ≥4/6 [19,20,21,22]. According to Geoffron et al., 14q32 disruption may be considered to be an alternative molecular cause of SRS and MEG3/DLK1:IG-DMR methylation should be tested in cases of negative results for other molecular testing of SRS patients [3].
The black line “IG-DMR” indicates the differentially methylated region (the imprinting control center of 14q32, named IG-DMR methylated on the paternal allele). The star represents methylated DMR. Black boxes indicate genes expressed from the paternal (pat) allele (DLK1, RTL1, and DIO3). White boxes indicate genes expressed from the maternal (mat) allele (the non-coding genes MEG3 and MEG8, and a cluster of snoRNAs and miRNAs).
DLK1 is widely expressed during fetal development. It encodes a transmembrane glycoprotein with six epidermal growth factor (EGF)-like motifs in its extracellular domain, a juxtamembrane region with a TACE-mediated cleavage site, a single transmembrane domain, and a short cytoplasmic tail [18]. The exact function of DLK1 is uncertain but it is involved in adipogenesis and appears to play an important role in preserving the pool of various progenitor cells until they differentiate [18]. With the generation of Dlk1-knockout mice, Moon et al. demonstrated overlapping phenotypes between Dlk1-null mice and human upd(14)mat, including growth retardation. They hypothesized that loss of Dlk1 expression may be responsible for most of the symptoms observed in human upd(14)mat [23]. Paternally inherited DLK1 variants have been recently identified in patients with central precocious puberty (CPP) but no growth retardation [24, 25]. Genetic defects of DLK1 have never been described in patients with FGR.
We screened DLK1 variants in a cohort of patients born SGA with a SRS phenotype using a NGS approach to search for new molecular defects responsible for SRS and assess the role of DLK1 in fetal growth.
Methods
Population studied
Patients included were referred to our molecular laboratory because they were born SGA (birth length and/or weight with a standard deviation score (SDS) < −2 [26]) with a clinical suspicion of SRS. Patients born SGA with a NH-CSS ≥4/6 or a NH-CSS = 3/6 with a strong clinical suspicion of SRS (relative macrocephaly and/or protruding forehead), negative molecular testing for H19/IGF2:IG-DMR (11p15.5) and DLK1/MEG3:IG-DMR (14q32.2) LOM and upd(7)mat, and negative molecular testing for CDKN1C, IGF2, HMGA2, and PLAG1 variants were included in this study. Written informed consent for participation was received from all patients or parents, in accordance with national ethics rules (Assistance Publique–Hôpitaux de Paris authorization no. 681). Patients were either followed at Armand Trousseau Children’s Hospital or referred by other clinical centers for molecular analysis. Postnatal growth parameters are expressed as SDS according to charts by Sempé and Pedron [27]. Blood samples were collected during routine biological follow-up at clinical visits. DNA was extracted in our laboratory from peripheral blood samples using an in-house protocol after cell lysis by a salting-out procedure, as previously described [28, 29].
Next-generation sequencing
DLK1 was sequenced as a SRS candidate gene using targeted sequencing. Library preparation, gene enrichment, sequencing, and data analysis were performed by IntegraGen SA (Evry, France) or by our laboratory with a pipeline designed by SOPHiA GENETICS (Lausanne, Switzerland).
Sanger sequencing
Variations of DLK1 identified through NGS were verified for the probands and their parents by Sanger sequencing using the ABI PRISM Big Dye Terminator v3.0 Cycle Sequencing Kit and an ABI 3100 Genetic Analyzer (Life Technologies, Courtaboeuf, France).
In silico analysis
The allele frequency was checked in the GnomAD database (online https://gnomad.broadinstitute.org/) to predict the functional consequences of any identified DLK1 variants. Interspecies alignment of DLK1 was performed using Clustal Omega (online tool from The European Bioinformatics Institute (EMBL-EBI), http://www.ebi.ac.uk/Tools/msa/clustalo/) and damage prediction scores were obtained using the Polyphen-2 bioinformatic tool [30]. Variants were also classified as benign or likely benign, pathogenic or likely pathogenic, or of uncertain significance following the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG/AMP) classification of variants [31]. Six main categories are evaluated according to these guidelines: population data (prevalence of the variant in control populations), computational in silico predictive data, functional characterization, segregation, de novo data, and allelic data.
Statistical analysis
Characteristics of the population are described as percentages for qualitative variables or as the SDS and mean (range) for quantitative variables.
Results
Clinical characteristics of patients screened for DLK1 variants
Samples from 132 patients referred for molecular genetic testing for SRS and without disturbances of 11p15 and 14q32 methylation or upd(7)mat were analyzed using a targeted NGS-based approach (93 by Integragen and 39 using the pipeline of SOPHiA GENETICS). Clinical data for at least four NH-CSS criteria were available for all patients and was complete (all six criteria available) for 61 (46%). All patients were born SGA. Patient characteristics are presented in Tables 1 and 2.
Screening for DLK1 variants
Among the 132 patients born SGA with a clinical suspicion of SRS, only two rare heterozygous variants of DLK1 were identified in two independent patients (NM_003836.6:c.103 G > C, p.(Gly35Arg) and NM_003836.6: c.194 A > G, p.(His65Arg)).
Phenotype of patients with identified DLK1 variants
Patient 1 was the third female child of two non-consanguineous parents. The proband’s parents were healthy. The final height of the father was 176 cm (0.2 SDS) and that of the mother 152 cm (−2.0 SDS). Proband’s mother was not born SGA and had no clinical features of SRS. The proband’s sisters were healthy, without prenatal or postnatal growth restriction. Patient 1 was born at 36 + 2 weeks of amenorrhea (WA). Her birth weight was 1930 g (−1.9 SDS), birth length 41 cm (−3.6 SDS), and head circumference 31 cm (−1.6 SDS). She did not experience catch up growth, with a height of 98.5 cm (−3.3 SDS), weight of 13.5 kg (−3.4 SDS), and head circumference of 49 cm (−1.5 SDS) at 6 years of age. She had no other remarkable features. She was not treated by growth-hormone (GH) therapy. She had feeding difficulties and a protruding forehead and fulfilled five of the six criteria of the NH-CSS. NGS sequencing of DLK1 revealed the heterozygous NM_003836.6: c.103 G > C variant located in exon 2, predicting an amino acid substitution at codon 35 (p.(Gly35Arg)). Gly35 is located within the first EGF-like motif in the extracellular domain of DLK1. This variant was inherited from her healthy mother, who carried the same heterozygous variant (Fig. 2).
Patient 2 was the first female child of two non-consanguineous parents. The mother’s final height was 169 cm (1.0 SDS). The proband’s father was born SGA at 35 WA (birth weight and height were 1290 g (−3.4 SDS) and 38 cm (−4.7 SDS)), but his head circumference at birth was uknown and we do not know if he had protruding forehead between 1 and 3 years. The proband’s father had no feeding difficulties during childhood and initially experienced catch-up growth with a height at −1 SDS between 10 and 14 years of age, but his final height was only 163 cm (−2.0 SDS). He was not treated with GH therapy. NH-CSS of proband’s father was 1/4. Patient 2 was a 29 WA-preterm girl with a birth weight of 920 g (−1.9 SDS), birth length of 34 cm (−3.0 SDS), and head circumference of 25 cm (−1.2 SDS). SGA was diagnosed in the second trimester of gestation. She did not develop feeding difficulties but had a protruding forehead. By 16 months of age, she had not experienced catch-up growth, with a height of 68.5 cm (−3.0 SDS) and a head circumference of 44.5 cm (−1.0 SDS). She was suspected of having SRS, with a NH-CSS = 4/6 and fifth finger clinodactyly. At 5 years of age, she experienced premature adrenarche without precocious puberty, responsible for catch-up growth, with a height of 105.4 cm (−0.8 SDS). NGS sequencing of DLK1 revealed that she carried a heterozygous NM_003836.6:c.194 A > G variation in exon 3 of DLK1, predicting an amino acid substitution at codon 65 (p.(His65Arg)). His65 is located within the second EGF-like motif of the extracellular domain of DLK1. This variant was inherited from her healthy mother who carried the same heterozygous variant (Fig. 2).
In silico analysis of the two DLK1 variations
The two variants NM_003836.6:c.103 G > C p.(Gly35Arg) and NM_003836.6: c.194 A > G p.(His65Arg) are described in GnomAD and dbSNP (rs762558665 and rs147224004) with an allele frequency in the general population of 7.0 × 10−6 and 4.7 × 10−4, respectively. Interspecies alignment of the amino acid sequences of DLK1 showed that residue Gly35 is invariant in vertebrates. The variation NM_003836.6:c.103 G > C p.(Gly35Arg) is predicted to be probably damaging, with a score of 1.000 by the Polyphen-2 bioinformatic tools of variation damage prediction. This variant is classified as a variant of uncertain significance (class 3) according to the ACMP/AMP classification (PM2-PP3).
Residue His65 is conserved only within Pan Troglodytes. Polyphen-2 predicted the variation p.His65Arg to be benign, with a score of 0.215. Variant NM_003836.6: c.194 A > G p.(His65Arg) is classified as likely benign (class 1) according to the ACMP/AMP classification (PM2-BP4-BS4).
The two variants NM_003836.6:c.103 G > C p.(Gly35Arg) and NM_003836.6: c.194 A > G p.(His65Arg) were not described in ClinVar. We submitted it (submission SUB9482112 and SUB9433818).
Discussion
We found two rare heterozygous variants of DLK1 in a cohort of 132 SGA patients with clinical suspicion of SRS and no identified molecular defects. The variants have already been reported in databases but with a low frequency. However, caution should be paid about variants frequencies regarding imprinted genes, as such a variant might have a different clinical impact depending on the maternal or paternal inheritance. Segregation analysis did not favor a pathogenic effect of these two variants, as they were both on the maternal allele, which is silent due to the maternal imprint of this gene. Thus, we did not identify any pathogenic DLK1 variants in our cohort.
No variants of DLK1 have been reported in SGA patients. Dauber et al. identified a complex defect of DLK1 (14-kb deletion and 269-bp duplication) in four patients with familial CPP. The four patients did not show prenatal or postnatal growth failure, and other classical clinical features of Temple or SRS, such as feeding difficulties, facial dysmorphia, precocious obesity, and relative macrocephaly, were excluded [24]. Gomes et al. identified three frameshift variants of DLK1 (NM_003836:c.594_594delC p.(Gly199Alafs*11), NM_003836:c.810_810delT p.(Val271Cysfs*14), and NM_003836:c.479_479delC p.(Pro160Leufs*50)) in five women from three families with CPP. Among them, three experienced postnatal growth failure, but no data were available about birth weight or length [25]. Montenegro et al. described a deletion (c.401_404 + 8del) in the splice-site junction of DLK1 in a girl with sporadic CPP without postnatal growth failure. No data about prenatal growth were available for this patient [32].
The role of DLK1 in fetal growth is not well established. Murine models have suggested a role for Dlk1 in fetal and postnatal growth. Indeed, Dlk1-null mice and heterozygous mice with paternal inheritance of the Dlk1-knockout allele showed prenatal and postnatal growth restriction [23, 33]. By contrast, heterozygous mice with maternal inheritance of the Dlk1-knockout allele did not experience growth restriction. Moreover, it has been demonstrated that Dlk1 promotes Gh expression. Dlk1-null mice showed reduced pituitary GH content and mice overexpressing Dlk1 had excessive pituitary and circulating levels of GH [33, 34]. The modulation of GH levels could explain, at least in part, the postnatal growth failure of mice lacking Dlk1, but not the prenatal growth failure. During fetal life, Dlk1 is expressed in the placenta in the endothelial cells of the placental labyrinth but is not required for its development [18]. Mice with a conditional deletion of Dlk1 in placental endothelial cells did not show FGR [35]. Reduced DLK1 levels in maternal blood samples have been shown in the second and third trimester of gestation with FGR [36, 37]. However, a causal relationship between low DLK1 levels and FGR has not been demonstrated or whether DLK1 levels simply reflect fetal weight.
To date, less is known about the contribution of individual genes of the 14q32 domain in the TS phenotype and the overlapping features with SRS. FGR could, for example, be explained by the action of several genes of the 14q32 domain in concert with genes in other imprinted domains [38]. Indeed, Abi Habib et al. demonstrated that overexpression of MEG3 and MEG8 in TS patients with 14q32 hypomethylation is associated with downregulation of IGF2 transcription from the 11p15 imprinting region [28].
In conclusion, we did not identify any variants in DLK1 in a cohort of 132 patients with suspected SRS. Although we screened a large cohort of patients for DLK1 variants, we cannot rule out the possibility of a role of DLK1 in fetal growth and the SRS phenotype. However, a frequent contribution of DLK1 variants among the molecular causes of SRS is unlikely. We did not identify a new molecular cause of SRS by the targeted NGS approach. Whole exome and genome sequencing and characterization of the entire methylome offer promising perspectives for the identification of new molecular causes of SRS.
References
Gaccioli F, ILMH Aye, Sovio U, Charnock-Jones DS, GCS Smith. Screening for fetal growth restriction using fetal biometry combined with maternal biomarkers. Am J Obstet Gynecol. 2018;218:S725–37.
Giabicani E, Pham A, Brioude F, Mitanchez D, Netchine I. Diagnosis and management of postnatal fetal growth restriction. Best Pr Res Clin Endocrinol Metab. 2018;32(Aug):523–34.
Wakeling EL, Brioude F, Lokulo-Sodipe O, O’Connell SM, Salem J, Bliek J, et al. Diagnosis and management of Silver-Russell syndrome: first international consensus statement. Nat Rev Endocrinol. 2017;13:105–24.
Azzi S, Salem J, Thibaud N, Chantot-Bastaraud S, Lieber E, Netchine I, et al. A prospective study validating a clinical scoring system and demonstrating phenotypical-genotypical correlations in Silver-Russell syndrome. J Med Genet. 2015;52(Jul):446–53.
Netchine I, Rossignol S, Dufourg M-N, Azzi S, Rousseau A, Perin L, et al. 11p15 imprinting center region 1 loss of methylation is a common and specific cause of typical Russell-Silver syndrome: clinical scoring system and epigenetic-phenotypic correlations. J Clin Endocrinol Metab. 2007;92:3148–54.
Gicquel C, Rossignol S, Cabrol S, Houang M, Steunou V, Barbu V, et al. Epimutation of the telomeric imprinting center region on chromosome 11p15 in Silver-Russell syndrome. Nat Genet. 2005;37:1003–7.
Kotzot D, Schmitt S, Bernasconi F, Robinson WP, Lurie IW, Ilyina H, et al. Uniparental disomy 7 in Silver-Russell syndrome and primordial growth retardation. Hum Mol Genet. 1995;4:583–7.
Begemann M, Zirn B, Santen G, Wirthgen E, Soellner L, Büttel H-M, et al. Paternally inherited IGF2 mutation and growth restriction. N Engl J Med. 2015;373:349–56.
Brioude F, Oliver-Petit I, Blaise A, Praz F, Rossignol S, Le Jule M, et al. CDKN1C mutation affecting the PCNA-binding domain as a cause of familial Russell Silver syndrome. J Med Genet. 2013;50:823–30.
Abi Habib W, Brioude F, Edouard T, Bennett JT, Lienhardt-Roussie A, Tixier F, et al. Genetic disruption of the oncogenic HMGA2-PLAG1-IGF2 pathway causes fetal growth restriction. Genet Med. 2018;20:250–8.
De Crescenzo A, Citro V, Freschi A, Sparago A, Palumbo O, Cubellis MV, et al. A splicing mutation of the HMGA2 gene is associated with Silver-Russell syndrome phenotype. J Hum Genet. 2015;60:287–93.
Akawi NA, Ali BR, Hamamy H, Al-Hadidy A, Al-Gazali L. Is autosomal recessive Silver-Russel syndrome a separate entity or is it part of the 3-M syndrome spectrum? Am J Med Genet A. 2011;155A:1236–45.
Inoue T, Nakamura A, Iwahashi-Odano M, Tanase-Nakao K, Matsubara K, Nishioka J, et al. Contribution of gene mutations to Silver-Russell syndrome phenotype: multigene sequencing analysis in 92 etiology-unknown patients. Clin Epigenetics. 2020;12:86.
Meyer R, Begemann M, Hübner CT, Dey D, Kuechler A, Elgizouli M, et al. One test for all: whole exome sequencing significantly improves the diagnostic yield in growth retarded patients referred for molecular testing for Silver-Russell syndrome. Orphanet J Rare Dis. 2021;16:42.
Meyer R, Soellner L, Begemann M, Dicks S, Fekete G, Rahner N, et al. Targeted next generation sequencing approach in patients referred for Silver-Russell syndrome testing increases the mutation detection rate and provides decisive information for clinical management. J Pediatr. 2017;187:206–212.e1.
Neuheuser L, Meyer R, Begemann M, Elbracht M, Eggermann T. Next generation sequencing and imprinting disorders: current applications and future perspectives: lessons from Silver-Russell syndrome. Mol Cell Probes. 2019;44:1–7.
Temple IK, Cockwell A, Hassold T, Pettay D, Jacobs P. Maternal uniparental disomy for chromosome 14. J Med Genet. 1991;28:511–4.
Traustadóttir GÁ, Lagoni LV, Ankerstjerne LBS, Bisgaard HC, Jensen CH, Andersen DC. The imprinted gene Delta like non-canonical Notch ligand 1 (Dlk1) is conserved in mammals, and serves a growth modulatory role during tissue development and regeneration through Notch dependent and independent mechanisms. Cytokine Growth Factor Rev. 2019;46:17–27.
Geoffron S, Abi Habib W, Chantot-Bastaraud S, Dubern B, Steunou V, Azzi S, et al. Chromosome 14q32.2 imprinted region disruption as an alternative molecular diagnosis of Silver-Russell syndrome. J Clin Endocrinol Metab. 2018;103:2436–46.
Kagami M, Nagasaki K, Kosaki R, Horikawa R, Naiki Y, Saitoh S, et al. Temple syndrome: comprehensive molecular and clinical findings in 32 Japanese patients. Genet Med. 2017;19:1356–66.
Kagami M, Mizuno S, Matsubara K, Nakabayashi K, Sano S, Fuke T, et al. Epimutations of the IG-DMR and the MEG3-DMR at the 14q32.2 imprinted region in two patients with Silver-Russell Syndrome-compatible phenotype. Eur J Hum Genet. 2015;23:1062–7.
Poole RL, Docherty LE, Al Sayegh A, Caliebe A, Turner C, Baple E, et al. Targeted methylation testing of a patient cohort broadens the epigenetic and clinical description of imprinting disorders. Am J Med Genet A. 2013;161:2174–82.
Moon YS, Smas CM, Lee K, Villena JA, Kim K-H, Yun EJ, et al. Mice lacking paternally expressed Pref-1/Dlk1 display growth retardation and accelerated adiposity. Mol Cell Biol. 2002;22:5585–92.
Dauber A, Cunha-Silva M, Macedo DB, Brito VN, Abreu AP, Roberts SA, et al. Paternally inherited DLK1 deletion associated with familial central precocious puberty. J Clin Endocrinol Metab. 2017;102:1557–67.
Gomes LG, Cunha-Silva M, Crespo RP, Ramos CO, Montenegro LR, Canton A, et al. DLK1 is a novel link between reproduction and metabolism. J Clin Endocrinol Metab. 2019;104:2112–20.
Usher R, McLean F. Intrauterine growth of live-born Caucasian infants at sea level: standards obtained from measurements in 7 dimensions of infants born between 25 and 44 weeks of gestation. J Pediatr. 1969;74:901–10.
Sempé (M). — Auxologie, méthode et séquences. Bulletins et Mémoires de la Société d’Anthropologie de Paris. 1980;7:77–77.
Abi Habib W, Brioude F, Azzi S, Rossignol S, Linglart A, Sobrier M-L, et al. Transcriptional profiling at the DLK1/MEG3 domain explains clinical overlap between imprinting disorders. Sci Adv. 2019;5:eaau9425.
Miller SA, Dykes DD, Polesky HF. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988;16:1215.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Montenegro L, Labarta JI, Piovesan M, Canton APM, Corripio R, Soriano-Guillén L, et al. Novel genetic and biochemical findings of DLK1 in children with central precocious puberty: a Brazilian-Spanish study. J Clin Endocrinol Metab. 2020;105:dgaa461.
Cheung LYM, Rizzoti K, Lovell-Badge R, Le Tissier PR. Pituitary phenotypes of mice lacking the notch signalling ligand delta-like 1 homologue. J Neuroendocrinol. 2013;25:391–401.
Charalambous M, Da Rocha ST, Radford EJ, Medina-Gomez G, Curran S, Pinnock SB, et al. DLK1/PREF1 regulates nutrient metabolism and protects from steatosis. Proc Natl Acad Sci USA. 2014;111:16088–93.
Appelbe OK, Yevtodiyenko A, Muniz-Talavera H, Schmidt JV. Conditional deletions refine the embryonic requirement for Dlk1. Mech Dev. 2013;130:143–59.
Cleaton MAM, Dent CL, Howard M, Corish JA, Gutteridge I, Sovio U, et al. Fetus-derived DLK1 is required for maternal metabolic adaptations to pregnancy and is associated with fetal growth restriction. Nat Genet. 2016;48:1473–80.
MacDonald TM, Walker SP, Hiscock R, Cannon P, Harper A, Murray E, et al. Circulating delta-like homolog 1 (DLK1) at 36 weeks is correlated with birthweight and is of placental origin. Placenta. 2020;91:24–30.
Howard M, Charalambous M. Molecular basis of imprinting disorders affecting chromosome 14: lessons from murine models. Reproduction. 2015;149:R237–249.
Acknowledgements
We thank the patients, their families and physicians, and the « Association Française des Familles ayant un enfant atteint du Syndrome Silver-Russell ou ne´ Petit pour l’âge Gestationnel (AFIF/PAG) ». We thank Cristina DAS NEVES and Nathalie THIBAUD for their contribution of this work.
Author contributions
A.P.: conception of the work, analysis and interpretation of the data, drafting of the manuscript, and final approval of the published version. M.-L.S., D.M., E.G., F.B., and I.N.: conception of the work, analysis and interpretation of the data, critical revision of the work for important intellectual content, and final approval of the published version. M.L.J.F.: Acquisition of the data and final approval of the published version.
Funding
This study received collaborative grant funding from the Agence Nationale de la Recherche (project “IMP-REGULOME”, ANR-18-CE12-0022-02).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Written informed consent for participation was received from all patients or parents, in accordance with national ethics rules (Assistance Publique–Hôpitaux de Paris authorization no. 681).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pham, A., Sobrier, ML., Giabicani, E. et al. Screening of patients born small for gestational age with the Silver-Russell syndrome phenotype for DLK1 variants. Eur J Hum Genet 29, 1756–1761 (2021). https://doi.org/10.1038/s41431-021-00927-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-021-00927-5
This article is cited by
-
Genomics elucidates both common and rare disease aetiology
European Journal of Human Genetics (2021)