Abstract
To elucidate whether Bronze Age population dispersals from the Eurasian Steppe to South Asia contributed to the gene pool of Indo-Iranian-speaking groups, we analyzed 19,568 mitochondrial DNA (mtDNA) sequences from northern Pakistani and surrounding populations, including 213 newly generated mitochondrial genomes (mitogenomes) from Iranian and Dardic groups, both speakers from the ancient Indo-Iranian branch in northern Pakistan. Our results showed that 23% of mtDNA lineages with west Eurasian origin arose in situ in northern Pakistan since ~5000 years ago (kya), a time depth very close to the documented Indo-European dispersals into South Asia during the Bronze Age. Together with ancient mitogenomes from western Eurasia since the Neolithic, we identified five haplogroups (~8.4% of maternal gene pool) with roots in the Steppe region and subbranches arising (age ~5–2 kya old) in northern Pakistan as genetic legacies of Indo-Iranian speakers. Some of these haplogroups, such as W3a1b that have been found in the ancient samples from the late Bronze Age to the Iron Age period individuals of Swat Valley northern Pakistan, even have sub-lineages (age ~4 kya old) in the southern subcontinent, consistent with the southward spread of Indo-Iranian languages. By showing that substantial genetic components of Indo-Iranian speakers in northern Pakistan can be traced to Bronze Age in the Steppe region, our study suggests a demographic link with the spread of Indo-Iranian languages, and further highlights the corridor role of northern Pakistan in the southward dispersal of Indo-Iranian-speaking groups.
Similar content being viewed by others
Introduction
As the world’s most widely spoken language family (3.2 billion speakers) [1], the Indo-European (IE) languages recorded to predominate the Eurasian Steppe and beyond, extending some 8000 km across Eurasia from East Asia (northeast China) to as far as Central Europe. One of the most widely accepted hypotheses posits that this language family originated in the Pontic Steppe in association with the formation and extension of the Yamnaya cultural complex during the Bronze Age (3000–1000 bc) [2, 3]. An alternative popular view suggests relatively earlier diffusion of IE languages from Anatolia with the expansion of farming (7000–6000 bc) [4], which also has received support from ancient genome data [5]. In Europe, the expansion of some branches of IE languages from their eastern periphery has been suggested with the dispersal of Yamnaya-related populations, largely consistent with Bronze Age migrations from the Steppe [3, 6]. However, controversy still exists regarding how the largest extant branch of IE languages, i.e., the Indo-Iranian family, expanded southward to South Asia. Certain studies have indicated substantial genetic contributions from western Eurasia introduced with the expansion of IE languages into South Asia (especially northern India) [7,8,9,10], whereas others have suggested that this genetic introgression is rather limited [11,12,13,14]. It is also widely argued that the early proto-Indo-Aryan speakers began to appear at the Bactria–Margiana Archaeological Complex (BMAC) in Southern Central Asia ~2000 bc before they expanded to the South Asian borderlands [2]; however, recent report of genome-wide ancient DNA data from Central and South Asia suggests no substantial genetic contribution from BMAC to South Asians [10]. Therefore, it is still controversial about whether there are substantial genetic contributions from the Steppe into the South Asia along with the expansion of IE speakers. Moreover, due to the different extent of genetic admixtures between Indo-Iranian speakers and local populations in South Asia, such as Dravidian speakers [14], the genetic contributions from early Indo-Iranian speakers to the modern South Asians would be underestimated. Therefore, to address these issues, genetic studies on populations that are likely to be the first descendants from west Eurasia into the Indian subcontinent are critical.
Located in the northernmost outpost of South Asia, at the brink of the Inner Asian Mountain Corridor, the Hindu Kush range of northern Pakistan has long been considered as a conduit for the migration of early Indo-Iranian speakers (often referred to as pastoralist groups from the Steppe) [2, 15] who were likely pioneers for the introduction of IE languages into South Asia [2, 16]. Distinct Indo-Aryan dialect complexes, natively spoken in this region and known as “Dardic,” were broadly assumed to be the independent surviving remnants of the ancestors of Indo-Iranian speakers [17]. Anthropological and genetic studies have suggested that settlers in northern Pakistan, such as the Kalash and Kho, were historically and culturally isolated from their urbanized surroundings in South Asia and other extant Eurasian populations [18, 19], thus probably representing early offshoots of the Vedic Aryans [20]. Moreover, the other major languages in this range, i.e., the eastern Iranians, can be traced back to Avestan scriptures [21], implying antiquity and long-term occupation of these IE language bearers along the northwestern region of South Asia. Therefore, genetic investigation into Dardic and Iranian speakers residing in northern Pakistan could help to elucidate the initial expansion of IE languages into South Asia if it can be linked to the Bronze Age demographic dispersals.
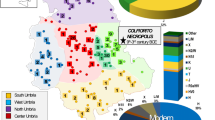
In this study, we sequenced maternally transmitted mitochondrial genomes (mitogenomes) from 213 Indo-Iranian language speakers (53 Iranian and 160 Dardic speakers) from the Hindu Kush mountain valleys along the northwestern periphery of Pakistan (Fig. 1). Together with published ancient and modern mitochondrial DNA (mtDNA) data and radiocarbon-dated human bones and archeological artifacts from this region, we explored whether there are significant genetic contributions from the Eurasian Steppe in northern Pakistanis along with the expansion of IE languages.
A Present grouping of Indo-Iranian languages is shown with a black solid line; red dotted line indicates classical independent position of Dardic within the Indo-Iranian domain proposed in previous study [17]. Unique three letter language identifiers are enclosed in brackets according to the ISO 639-3 standard. Numbers in open brackets show number of language(s) in respective language family, four linguistic groups sampled in present study are highlighted in green, with numbered circles corresponding to their locations on the map (B) inset. Figure is optimized according to the language catalog Ethnologue global data set by [1]. B Geographic locations of populations and archeological sites in northern Pakistan analyzed in this study. Different shapes represent different geographic areas and shape colors represent language families to which populations belong, as defined in the legend. Three main Steppe regions (from Mongolia passing through Central Asia occupying most of Europe) are approximately labeled on the map. Inset shows location of populations and archeaological sites in northern Pakistan, and numbers in circles are sampling locations in present study, corresponding to their linguistic affiliations in (A). Base map was selected from online map server in Surfer. Close coordinates are slightly jittered on map to minimize overlapping between locations.
Materials and methods
Data collection
Blood samples from 213 IE speakers (193 new and 20 reported in our previous study [22]) that covered major Indo-Iranian-speaking regions in northern Pakistan were collected (Fig. 1). All experimental protocols were approved by the Ethics Committee at the Kunming Institute of Zoology, Chinese Academy of Sciences, China. Written and signed informed consent was obtained from each participant before the study. Furthermore, a total of 19,568 mtDNA sequences covering native IE speakers from west Eurasia and Asia were obtained, including 6339 complete mitogenomes (Table S1) and 13,229 partial mtDNA (with control and partial coding region information) (Table S2). Published archeological data from northern Pakistan (N = 116, Table S3) starting from the Bronze Age (5102 ± 60 to 310 ± 30 bp), which mainly include radiocarbon-dated human bones, were also included for comparisons. Sample locations were plotted on a map using the Kriging algorithm in Surfer 17.1.288 (Golden Software Inc. Golden, Colorado, USA).
Sequencing of complete mitogenomes
Total genomic DNA was isolated using the standard phenol/chloroform method [23]. DNA yield and purity were measured via UV spectroscopy. Complete mitogenomes of samples were enriched by capture-based strategies using a Human Mitochondria Capture Kit (MyGenostics Inc., Beijing, China) followed by whole mtDNA sequencing. Sequencing was carried out using an Illumina HiSeq X Ten platform at MyGenostics with a sequencing depth of >1000 X.
Quality control and analysis of mtDNA genome sequences
All sequenced data were visually inspected to correct any potential phantom mutations and artifacts. We obtained the variants for each sequence relative to the revised Cambridge Reference Sequence (NC_012920.1) [24] and assigned them to specific haplogroups using PhyloTree Build 17 (http://phylotree.org/) [25]. The haplogroup results were further confirmed using HaploGrep [26]. We followed the general criteria of PhyloTree Build 17 for assigning new haplogroup and sub-haplogroup statuses [25], and searched for diagnostic motifs for those new branches using our in-house database (Tables S1 and S2) (19,781 mtDNA sequences) to further assess genetic affinity between geographic regions (Table S4). The variable lengths in homopolymeric C-stretches in regions 16,180–16,193 and 303–315 and AC indels at 515–522 were disregarded in phylogenetic tree reconstruction. The DnaSP software package was used to assess the haplotype (H) and nucleotide (π) diversities, including Tajima’s D statistics.
Principal component analysis (PCA) and admixture estimates
PCA was performed based on the haplogroup frequency matrix of all obtained population mtDNA data (Table S5) using the FactoMineR v1.00 package in R (http://www.rproject.org). We excluded populations with a limited number of samples (<5) when calculating haplogroup frequencies. For the mtDNA data sequences obtained from the 1000 Genomes Project (1KGP) [27], the kinship of individuals in their respective families was checked to ensure that the samples represented a random population survey. SPSS v20 (Chicago, IL, USA) was used for regression (REGR), correspondence analysis, and admixture estimation with the weighted least squares method [28].
Phylogenetic reconstructions and haplogroup age estimates
Phylogenetic trees of the mitochondrial genome sequences were reconstructed based on contemporary and ancient mitogenome sequences by mt-Phyl software (https://sites.google.com/site/mtphyl/home) and checked manually. Coalescent ages of haplogroups were calculated by different methods, including the rho (ρ) and standard error (σ) methods [29, 30], which provide unbiased and overlapping estimates of clade age [31], as well as the Bayesian estimate by BEAST. Branches with ≥3 sequences were considered for age estimations. Substitution rates at complete coding regions and synonymous positions of coding regions from previous study [32] were adopted. The protein coding regions with synonymous positions were selected for coalescent time estimations utilizing the Bayesian Markov chain Monte Carlo (MCMC) approach implemented in BEAST v1.5.2 [33] with the best-fit model. For west Eurasian haplogroups, we obtained all contemporary (Table S6) and ancient mtDNA sequences (Table S7) from the Mitomap (https://www.mitomap.org) [34], European Nucleotide Archives (https://www.ebi.ac.uk/ena), ancient human mitochondrial genome database (AmtDB) [35], and available literature for phylogeographic reconstructions.
Bayesian skyline plot analysis
Bayesian skyline plots (BSP) which showing the changes of effective population size (Ne) through time were reconstructed using BEAST v1.5.2 [33]. To circumvent the confounding effects of common homoplasy in the hypervariable control region and to focus on neutrally evolving sites, 13 protein coding regions (spanning nucleotides 576–16,023 of mitogenomes with the reverse complemented ND6 gene, thus ensuring the same read direction for all genes in the sequences) were utilized for BSP analysis, as described in previous studies [36]. The best-fit model (HKY + G) was selected with Model test (v3.7) [37] for all BEAST-related analyses. A strict molecular clock with the fixed rate of 1.691E−8 substitutions per site per year was chosen [38] and was further evaluated with an uncorrelated relaxed clock model. Each MCMC simulation was run for 40,000,000 generations and sampled every 4000 generations, with the first 40,000 generations discarded as burn-in. After each cycle, the runs were checked with Tracer v1.5 (http://tree.bio.ed.ac.uk/software/tracer/) to confirm convergence and sufficient effective population size (ESS ≥ 200) was considered for all relevant parameters of traces. The BEAST trees were reconstructed using DensiTree [39].
Results and discussion
Northern Pakistani maternal gene pool experienced substantial contributions from western Eurasians
Based on the obtained mitogenome information, all 213 IE speakers in northern Pakistan could be allocated into haplogroups derived from non-African lineages M, N, and R. Most individuals in northern Pakistan (66.19%, 141/213) were affiliated to haplogroups HV, JT, and U (Table S8), which are reported to be prevalent in west Eurasians [40,41,42], at a much higher rate than that of other South Asian populations (24.9%) based on the 1KGP [27], Human Genome Diversity Project (HGDP) [43], and Bengali population data [13]. The distribution of unique diagnostic motifs of the haplogroups defined in our study is shared across Eurasia; however, some substitutions were more frequent in South Asians (Table S4), implying the consistent presence and de novo origination of these lineages in north Pakistan after differentiation from western Eurasian counterparts.
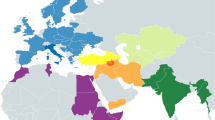
To further investigate the genetic relationship between the northern Pakistani populations and surroundings, a total of 19,757 samples from 294 population studies were analyzed covering IE and other language family-speaking populations from northern Pakistan and its surroundings, including Europe, the Middle East and Caucasus, Central Asia, South Asia, and East Asia (Fig. 1). The PCA plot based on haplogroup frequencies showed a general differentiation in IE-speaking populations between South Asia and Europe, with South Asian IE speakers distributed together with the Dravidians (Figs. 2A and S1), consistent with the genetic admixture of these two linguistic groups observed in previous studies [14]. Haplogroups ubiquitous in present-day west Eurasians (e.g., mainly the descendants of haplogroups HV and JT) contributed most to PC1, whereas haplogroups prevalent in South Asia contributed to PC2 (Fig. 2B). The association of the western Eurasian components with present-day South Asians is further supported by the correspondence analysis (Fig. S2). Moreover, some components, e.g., U5a sub-haplogroups that were prevalent in the Steppe during the Bronze Age [10], also contributed to PC2, implying potential genetic contributions from the Steppe into South Asia, including northern Pakistan.
A Different colors represent different language groups and different shapes represent different geographic locations. Languages labeled as “Others” include Mongolic, Andamanese, Ongan, Tai-Kadai, Uralic, Japonic, Niger-Congo, Tungusic, Caucasian, Northwest Caucasian, dialects, and unclassified. Geographic regions labeled as “Others” include regions out of Europe and Asia. Each population is indicated by a corresponding number (Table S1). Populations in this study are labeled in bold and their closest Tajik populations from the north are labeled in italics. Percentage of variation explained by each axis is shown in parentheses. B Plot of haplogroup contribution to first and second PCs calculated as factor scores for PC1 and PC2 using REGR in SPSS.
In comparison to IE speakers in India, populations residing in the northwestern regions of the subcontinent, including the Kho and Pashtuns in northern Pakistan, were more akin to populations from the west, especially those from the Central Asian Steppe, e.g., the Pamiris and Tajiks (Fig. 2). This could be attributed to less deeply rooted Asian-specific lineages, such as macrohaplogroup “M” and its derivatives, in our northern Pakistani samples (18.3%) than in the southern subcontinent populations [27, 43] (47%), as well as substantial haplogroup sharing (e.g., HV and J) between northern Pakistanis and Central Asians (Fig. S3). These affinities would have been established since ancient times, as evident from archaeological studies documenting ancient population movements between west Eurasia and the Indus peripheral region [44]. Further evidence comes from the gradual decrease in west Eurasian component from north Pakistan to the southeastern Indian subcontinent based on admixture estimation (Table S9). Moreover, the diversity indices of those lineages in northern Pakistan, which are prevalent in western Eurasians, show higher nucleotide diversities in northern Pakistan than in the southern subcontinent, implying longer differentiation of these lineages in the northwestern subcontinent (Table S10). These results together suggest that the Indo-Iranian populations in northern Pakistan retained more genetic components from western Eurasia in comparison with the IE-speaking populations in the southern Indian subcontinent.
Eurasian Steppe legacy in South Asia is largely retained in Indo-Iranian-speaking northern Pakistani groups
Based on phylogeographic analysis of all available mitogenome data, we updated the phylogeographic status of the western Eurasian lineages in South Asia (Table S4) and identified 29 haplogroups which are prevalent in South Asia, and would have differentiated independently with derived branches in this region (Fig. S4). Five of these haplogroups, which had been reported in our previous work [22] (i.e., H2a1a3, J2b1a7a, R0a5a, U2e1h1, and U4a1f) were also reassessed together with the new haplogroup branches. Age estimations utilizing different mtDNA mutation rates and different methods indicated that the ages of these 29 haplogroups ranged from 10.14 to 0.5 kya (Table S11), with most (20/29) <5 kya old in South Asia (Table S11 and Fig. S4), a time subsequent to the major Bronze Age processes occurring in the Eurasian Steppe, e.g., eastern extensions of IE and related cultures advancing from Yamnaya about 3000 bc [2]. Specifically, among the 20 haplogroups that can be traced back to the Bronze Age period, 12 were distributed exclusively in northern Pakistan and 4 were ubiquitous in both the north and southern mainland (Fig. S4), accounting for ~23% of our northern Pakistani samples. Considering the restrictive distribution of these haplogroups in north Pakistan and the surroundings, it is possible that these haplogroups differentiated independently in this area. Therefore, the ages of these haplogroups would reflect the upper bound of their arrival times into north Pakistan. These results thus suggest substantial genetic contributions to the northern Pakistani gene pool since the beginning of the Bronze Age. This finding was further supported by Bayesian-based analysis, which illustrated that the majority of these west Eurasian lineages coalesced 5 kya in north Pakistan (Fig. S5). Interestingly, this extensive genetic contributions from west Eurasia during the Bronze Age is in agreement with the substantial settlements evidenced by the increasing number of archaeological remains in northern Pakistan with 14C dates from the beginning of the Bronze Age (Table S3 and Fig. S6), which are broadly interpreted as remnants of Indo-Iranian speakers from the region [45].
Further analysis, together with ancient mitogenome data, was conducted to assess past genetic contact between northern Pakistan and western Eurasia. Some haplogroups were found to have ancestors or sister lineages predominantly in the Middle East and the Caucasus (i.e., R0a5a, T1a2c, R2e, U7a3b1, and J1b1b1d) (Table S7 and Fig. S4), reflecting a likely genetic connection between northern Pakistan and the Middle East [46]. Similarly, several lineages (i.e., W3b2, I4b1, J1d7, U7a6, U7a2c, U7a3a1, U7b3, T2o, and T2d1b3) showed even more complex branching, with siblings found in the Middle East as well as further north in the Steppe, but mostly lacking upstream branches from the southern subcontinent (Fig. S4). Given the existence of genetic introgression from Anatolian farmers into Yamnaya ancestries [5], and the lack of Anatolian ancestry in South Asians after ~6000 bc [10], it is likely that these lineages would have been introduced into north Pakistanis by Steppe pastoralists, who accepted genetic contributions from Anatolian farmers. However, the possibility that the bearers of those haplogroups were early Iranian farmers [46] who further mixed with pastoralists in the BMAC [10] cannot be ruled out. Moreover, the absence of haplogroup K1 in our data, which is often associated with Neolithic dispersals from Anatolia [47], further supports the above admixture scenarios. Other haplogroups that would have been transported with these past genetic contacts, e.g., H5a1r and especially J2b1a7a, with ancestor types and sibling lineages in Europe, would have moved from the Europe and reached the Hindu Kush region, probably during the Bronze Age onward [22].
We further focused on haplogroups displaying direct connections between northern Pakistan and the Steppe region. Results indicated that haplogroup U2e1h1 (age 7.06–9.55 kya), which had ancestor types distributed in the Steppe region, likely originated therein. One possible explanation for the prevalence in northern Pakistan with coalescent ages >7 kya could be its presence in this region for more than 7000 years, and advent to northern Pakistan before the early Bronze Age Yamnaya expansion [12]. However, whether this kind of haplogroups are part of the common Eurasian-shared ancestry and remain visible in north Pakistan through genetic drift or originated elsewhere ~7–9 kya and dispersed into this region during the Bronze Age cannot be ruled out. Given the uncertainty of their expansion time into northern Pakistan, it is difficult to speculate whether U2e1h1 expanded with the spread of IE languages during the Bronze Age.
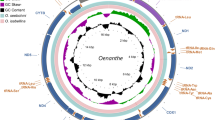
Differently, other haplogroups, with coalescent ages estimated since about 5 kya, such as J1b1a1f (age 2.63–3.53 kya, one outlier 12.07 kya was disregarded), U4a1f (age 2.52–5.00 kya), and H2b1 (2.35–5.25 kya) (Table S11), have ancestor and sister lineages found in Europe and the Steppe region (Fig. S4). These haplogroups, especially J1b1a1f and U4a1f which showed independent differentiations in north Pakistan, thus would have been introduced into the northern Pakistani Indo-Iranian-speaker gene pool after 5 kya, consistent with the arrival of Indo-Iranian-speaking groups in northern South Asia during the Bronze Age (~2300–1200 bc) [12]. Thus, these haplogroups could be considered as genetic legacies of IEs from the Steppe region. Additional support comes from haplogroup U4a1f, whose ancestor type (U4a1) was found in Neolithic samples from Europe and, importantly, in Yamnaya samples from the western Steppe (Fig. S4). Similarly, haplogroup W3a1a4 (age 2.35–5.25 kya) and sub-haplogroups of W3a1b (e.g., W3a1b1, age 2.63–6 kya, distributed in both northern Pakistan and the southern subcontinent), with the sister type W3a1a found in ancient Yamnaya samples, could also have Yamnaya-related origination. Other haplogroups, e.g., H2a1a3 (age 0.88–3.00 kya) with ancestor node in the eastern and central Steppe, likely suggest dispersals into northern Pakistan along with the expansion of Bronze Age cultures such as the Andronovo (a late Bronze Age culture also associated with early Indo-Iranian groups) (Fig. 3A). These haplogroups, viz., J1b1a1f, U4a1f, H2a1a3, W3a1a4, and W3a1b subbranches, can therefore trace their ancestry to the Steppe region and would have descended from populations therein during the Bronze Age (Fig. 3A, B). As their coalescent ages in northern Pakistan and the radiocarbon dates of their ancient Eurasian counterparts (Fig. 3B) fit well with the timing of the spread of IE languages into South Asia, the most probable explanation would be that these lineages were introduced into the gene pool of northern Pakistani Indo-Iranian speakers from the Steppe region from the beginning of the Bronze Age, at least according to currently available evidence.
A Distribution of individuals belonging to haplogroups H2a1a, J1b1a1, U4a1, and W3a1. Numbers in circles represent number of present-day mitogenomes belonging to haplogroup(s) plotted on map with different colors for different haplogroups. Sizes of circles are proportional to total number of modern mitogenomes from corresponding regions. Numbers of ancient mitogenomes used in phylogeographic reconstructions are also shown on their respective locations on the map in bold italics (without any punctuation). B Five Steppe-driven lineages coalescing since ~5 kya in northern Pakistan. Each haplogroup is represented by a triangle with its base proportional to the number of mitogenomes and height to age of clades (kya) in northern Pakistan. Different colors represent different geographic regions. Diamond shapes on branches are ancient mitogenomes positioned either correspond to calibrated radiocarbon ages (italic) or average of lower and upper limit of sample age (95.4% probability interval for 14C samples). C Phylogeographic reconstruction of haplogroup W3a1. Geographic origins of samples are represented by different colors. Each square represents a mitogenome unless otherwise indicated by the number on the square. Samples from aDNA specimens are encircled in red. Each dot on branch represents a mutation and color of dot corresponds to type of mutation. Branches with incomplete mitogenome sequence alignments with Gap(s) are colored in red, their position in the tree can be dubious, and had been excluded in age estimations.
Southward dispersal of haplogroup W3a1 to Indian subcontinent
Interestingly, the IE genetic legacy from the Steppe region, i.e., H2a1a3, J1b1a1f, U4a1f, and subbranch of W3a1, accounted for 8.4% of our northern Pakistani samples, which is much more pronounced than that in the southern subcontinent populations (~1.7%) in the 1KGP [27] and HGDP [43] data sets. This implies a greater genetic contribution from the Steppe region to the northern Pakistanis than to the southern South Asians. One explanation could be the influence of demographic processes (e.g., genetic drift or founder effect) resulting in the retention of a greater genetic legacy from IE speakers in northern Pakistanis. However, considering the closer genetic affinity between the southern subcontinent Indo-Iranian and Dravidian speakers (Fig. S1), it is more likely that this difference is due to the southward migration of IE speakers via northern Pakistan, during which they assimilated maternal genetic components from contemporary non-IE-speaking groups, consistent with substantial ancestry from the Steppe in present-day South Asians [10]. This scenario finds further support from the southward dispersals of Steppe legacy of IEs in northern Pakistan in our analysis. For example, W3a1b, with root types found in both modern individuals and individuals of prehistorical and early historical periods in the Swat Valley, northern Pakistan [10], had subbranches W3a1b1, W3a1b2, and W3a1b3 that arose in situ in the southern subcontinent, especially India (Table 1 and Fig. 3A, C). The estimated ages of sub-haplogroups of W3a1b (W3a1b1: 3.53 kya, W3a1b2: 4.14 kya) were further confirmed with BEAST estimations (Table 1) and echoed the arrival time of IE speakers in India in the late Bronze Age (~1500 bc). Therefore, the genetic legacy of IE speakers in northern Pakistan also contributed to the gene pools of the southern subcontinent populations along with IE language expansion, indicating that the southward IE language expansion was not simply mediated by cultural diffusion. These results highlight the corridor role of the Hindu Kush region of northern Pakistan in genetic introgression from the Steppe to South Asia during the Bronze Age, which would have been mediated by the southward expansion of IE speakers, consistent with views from archaeological and linguistic studies [17, 19, 44, 45].
Impact of Bronze Age Eurasian Steppe expansion on population dynamics of South Asia
BSP analysis was carried out to elucidate the impact of Bronze Age Eurasian migrations on the population dynamics of South Asia. Results indicated an increase in the effective population size (Ne) for northern Pakistani populations from ~18 kya (Fig. 4A), although relatively flat, likely facilitated by the climatic improvements after the last glacial maximum. The Ne of northern Pakistanis remained stable during the postglacial period and even during the Neolithic, in agreement with recent reports on modern and ancient genomes from South Asia [9, 48]. In brief, our results indicated that the Neolithic expansion into South Asia from 8 kya did not strongly affect the population size in northern Pakistan, thus suggesting any major genetic influx from the Iranian plateau during the Neolithic into the region to be unlikely. Surprisingly, a population decrease was observed in the northern Pakistanis after 5 kya, which lasted until 1.02 kya (Fig. 4A). This differs from the signal observed from Bronze Age lineages and the genetic legacy of IE speakers from the Steppe, which displayed intensive growth from 5 kya (Fig. 4B, C) and likely reflected a rapid population expansion of pastoralist groups from the Steppe after their migration into northern Pakistan. These BSP trends were reevaluated utilizing a relaxed clock model using the same data sets, with highly similar results, but with less intense population decline trend in northern Pakistan since 5 kya (Fig. S7). To further avoid any potential artifacts arising in BSP methodology [49], we calculated Tajima’s D and obtained a positive value (0.63550), indicative of population shrinkage in this region (Table S10) and lending support to the BSP results. One explanation for this decline may be the depletion of natural resources or habitat in the region after the arrival of Steppe pastoralists ~5.5 kya, which may have led to a population decrease [44, 50]. However, the influence of other factors, such as social structure changes, population size fluctuations, natural selection, and quaternary climatic oscillations between the northern hemisphere and South Asian monsoon [44], should not be ignored. More studies should be carried out to confirm this population decrease.
A BSP for all northern Pakistanis analyzed in our study (N = 213). B BSP indicating effective population size trends for all lineage ages estimated to Bronze Age in South Asia (N = 88). C BSP for H2a1a, J1b1a1, U4a1, and derivatives of W3a1 (W3a1a and W3a1b) in South Asia (N = 46). Solid lines represent mean population size estimate from Bayesian posterior distribution, dashed lines show 95% highest posterior density limits, visualized using Tracer v1.5.
In summary, based on available archeological and high-resolution mitogenome data from northwestern Pakistan, especially from Iranian and Dardic populations, who are suggested to be the surviving traces of early Indo-Iranian groups, we identified the genetic contributions of different dispersals from west Eurasia into northern Pakistan during the Bronze Age onward. Importantly, we identified five haplogroups as the genetic legacy of IE speakers from the Eurasian Steppe, likely dispersed along with the migration of IE-speaking populations during the Bronze Age into northern Pakistan, thus implying that IE language expansion into South Asia was not simply mediated by cultural diffusion. This migration contributed 8.4% of the gene pool of northern Pakistani IE speakers, suggesting this demographic connection, which is a possible source of IE language diffusion, could be one part of the complex demographic history of the region. Our results also provide implications on the two main hypotheses of IE language origination, viz. Anatolia and Steppe hypotheses. Considering that Steppe components were observed in all Indo-Iranian groups in northern Pakistan in our study, as well as in other regions in South Asia [10], while lineages possibly representing the genetic legacy of Neolithic farmers, e.g., R2e, K1, were either absent or not found in all of the IE-speaking groups in northern Pakistan, our results lend more support to the Steppe hypothesis, at least from a matrilineal perspective. Furthermore, these IE speakers, as evidenced by the genetic legacy identified here, also moved southward and contributed genetically, though to a rather limited extent, to the Indian subcontinent, suggesting northern Pakistan as a corridor in the spread of IE languages during the Bronze Age dispersals into South Asia. Since our study is only based on mtDNA data, which only reflect maternal histories of populations, more investigations based on genome-wide data are also needed to intensively dissect the expansion of IE speakers into South Asia.
Data availability
The 213 complete mitochondrial DNA sequences were deposited in GenBank under accession numbers MN595684–MN595896.
References
Eberhard DM, Simons GF, Fennig CD. Ethnologue: languages of the world. 23rd ed. Dallas: SIL International; 2020.
Parpola A. The roots of Hinduism: the early Aryans and the Indus Civilization. New York, Oxford University Press; 2015.
Anthony DW. The horse, the wheel, and language: how Bronze-Age Riders from the Eurasian Steppes shaped the modern world. New Jersy, Princeton University Press; 2007.
Renfrew C. Archaeology and language: the puzzle of Indo-European origins. London: Jonathan Cape; 1987.
Wang CC, Reinhold S, Kalmykov A, Wissgott A, Brandt G, Jeong C, et al. Ancient human genome-wide data from a 3000-year interval in the Caucasus corresponds with eco-geographic regions. Nat Commun. 2019;10:590.
Haak W, Lazaridis I, Patterson N, Rohland N, Mallick S, Llamas B, et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature. 2015;522:207–11.
Basu A, Mukherjee N, Roy S, Sengupta S, Banerjee S, Chakraborty M, et al. Ethnic India: a genomic view, with special reference to peopling and structure. Genome Res. 2003;13:2277–90.
Cordaux R, Saha N, Bentley GR, Aunger R, Sirajuddin SM, Stoneking M. Mitochondrial DNA analysis reveals diverse histories of tribal populations from India. Eur J Hum Genet. 2003;11:253–64.
Silva M, Oliveira M, Vieira D, Brandao A, Rito T, Pereira JB, et al. A genetic chronology for the Indian Subcontinent points to heavily sex-biased dispersals. BMC Evol Biol. 2017;17:88.
Narasimhan VM, Patterson N, Moorjani P, Rohland N, Bernardos R, Mallick S, et al. The formation of human populations in South and Central Asia. Science. 2019;365:eaat7487.
Kivisild T, Bamshad MJ, Kaldma K, Metspalu M, Metspalu E, Reidla M, et al. Deep common ancestry of Indian and western-Eurasian mitochondrial DNA lineages. Curr Biol. 1999;9:1331–4.
de Barros Damgaard P, Martiniano R, Kamm J, Moreno-Mayar JV, Kroonen G, Peyrot M, et al. The first horse herders and the impact of early Bronze Age steppe expansions into Asia. Science. 2018;360:eaar7711.
Li Y-C, Wang H-W, Tian J-Y, Li R-L, Rahman ZU, Kong Q-P. Cultural diffusion of Indo-Aryan languages into Bangladesh: a perspective from mitochondrial DNA. Mitochondrion. 2018;38:23–30.
Moorjani P, Thangaraj K, Patterson N, Lipson M, Loh PR, Govindaraj P, et al. Genetic evidence for recent population mixture in India. Am J Hum Genet. 2013;93:422–38.
Cavalli-Sforza LL, Cavalli-Sforza L, Menozzi P, Piazza A. The history and geography of human genes. New Jersy, Princeton University Press; 1994.
Jettmar K. Ethnological research in Dardistan 1958: preliminary report. Proc Am Philos Soc. 1961;105:79–97.
Grierson AG. Linguistic survey of India: Vol. VIII. India: Office of the Superintendent of Government Printing; 1919.
Ayub Q, Mezzavilla M, Pagani L, Haber M, Mohyuddin A, Khaliq S, et al. The Kalash genetic isolate: ancient divergence, drift, and selection. Am J Hum Genet. 2015;96:775–83.
Masica CP. The Indo-Aryan languages. Cambridge: Cambridge University Press; 1991.
Jettmar K. Urgent tasks of research among the Dardic peoples of Eastern Afghanistan and Northern Pakistan. Anthropol Ethnol Res. 1959;2:85–96.
Michael MTH. Four varieties of Pashto. J Am Orient Soc. 1983;103:595–7.
Rahman ZU, Li Y-C, Tian J-Y, Kong Q-P. Exploring European ancestry among the Kalash population: a mitogenomic perspective. Zool Res. 2020;41:552–6.
Sambrook J, Fritsch EF, Maniatis T. Extraction with phenol: chloroform. Molecular cloning: a laboratory manual. New York, Cold Spring Harbor Laboratory Press; vol 3. 1989.
Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, Howell N. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nat Genet. 1999;23:147.
van Oven M, Kayser M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum Mutat. 2009;30:E386–94.
Weissensteiner H, Pacher D, Kloss-Brandstatter A, Forer L, Specht G, Bandelt HJ, et al. HaploGrep 2: era of high-throughput sequencing. Nucleic Acids Res. 2016;44:W58–63.
Sudmant PH, Rausch T, Gardner EJ, Handsaker RE, Abyzov A, Huddleston J, et al. An integrated map of structural variation in 2,504 human genomes. Nature. 2015;526:75–81.
Long JC, Williams RC, McAuley JE, Medis R, Partel R, Tregellas WM, et al. Genetic variation in Arizona Mexican Americans: estimation and interpretation of admixture proportions. Am J Phys Anthropol. 1991;84:141–57.
Saillard J, Forster P, Lynnerup N, Bandelt HJ, Nørby S. mtDNA variation among Greenland Eskimos: the edge of the Beringian expansion. Am J Hum Genet. 2000;67:718–26.
Forster P, Harding R, Torroni A, Bandelt HJ. Origin and evolution of Native American mtDNA variation: a reappraisal. Am J Hum Genet. 1996;59:935–45.
Macaulay V, Soares P, Richards MB. Rectifying long-standing misconceptions about the ρ statistic for molecular dating. PLoS ONE. 2019;14:e0212311.
Soares P, Ermini L, Thomson N, Mormina M, Rito T, Rohl A, et al. Correcting for purifying selection: an improved human mitochondrial molecular clock. Am J Hum Genet. 2009;84:740–59.
Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29:1969–73.
Kogelnik AM, Lott MT, Brown MD, Navathe SB, Wallace DC. MITOMAP: an update on the status of the human mitochondrial genome database. Nucleic Acids Res. 1997;25:196–9.
Ehler E, Novotny J, Juras A, Chylenski M, Moravcik O, Paces J. AmtDB: a database of ancient human mitochondrial genomes. Nucleic Acids Res. 2018;47:D29–D32.
Atkinson QD, Gray RD, Drummond AJ. mtDNA variation predicts population size in humans and reveals a major Southern Asian chapter in human prehistory. Mol Biol Evol. 2008;25:468–74.
Posada D, Buckley TR. Model selection and model averaging in phylogenetics: advantages of Akaike information criterion and bayesian approaches over likelihood ratio tests. Syst Biol. 2004;53:793–808.
Ho SY, Endicott P. The crucial role of calibration in molecular date estimates for the peopling of the Americas. Am J Hum Genet. 2008;83:142–6.
Bouckaert RR. DensiTree: making sense of sets of phylogenetic trees. Bioinformatics. 2010;26:1372–3.
Torroni A, Lott MT, Cabell MF, Chen YS, Lavergne L, Wallace DC. mtDNA and the origin of Caucasians: identification of ancient Caucasian-specific haplogroups, one of which is prone to a recurrent somatic duplication in the D-loop region. Am J Hum Genet. 1994;55:760–76.
Richards M, Corte-Real H, Forster P, Macaulay V, Wilkinson-Herbots H, Demaine A, et al. Paleolithic and neolithic lineages in the European mitochondrial gene pool. Am J Hum Genet. 1996;59:185–203.
Macaulay V, Richards M, Hickey E, Vega E, Cruciani F, Guida V, et al. The emerging tree of West Eurasian mtDNAs: a synthesis of control-region sequences and RFLPs. Am J Hum Genet. 1999;64:232–249.
Cann HM, de Toma C, Cazes L, Legrand MF, Morel V, Piouffre L, et al. A human genome diversity cell line panel. Science. 2002;296:261–2.
Dambricourt Malassé A, Gaillard C. Relations between climatic changes and prehistoric human migrations during Holocene between Gissar Range, Pamir, Hindu Kush and Kashmir: the archaeological and ecological data. Quat Int. 2011;229:123–31.
Dani AH. Gandhara grave culture and the Aryan problem. J Cent Asia. 1978;1:42–55.
Lazaridis I, Nadel D, Rollefson G, Merrett DC, Rohland N, Mallick S, et al. Genomic insights into the origin of farming in the ancient Near East. Nature. 2016;536:419–24.
Hofmanová Z, Kreutzer S, Hellenthal G, Sell C, Diekmann Y, Díez-del-Molino D, et al. Early farmers from across Europe directly descended from Neolithic Aegeans. Proc Natl Am Sci. 2016;113:6886–91.
Shinde V, Narasimhan VM, Rohland N, Mallick S, Mah M, Lipson M, et al. An Ancient Harappan genome lacks ancestry from Steppe Pastoralists or Iranian farmers. Cell. 2019;179:729–35.
Heller R, Chikhi L, Siegismund HR. The confounding effect of population structure on Bayesian skyline plot inferences of demographic history. PLoS ONE. 2013;8:e62992.
Hewitt GM. Genetic consequences of climatic oscillations in the Quaternary. Philos Trans R Soc Lond B Biol Sci. 2004;359:183–95.
Acknowledgements
We would like to thank Dr. Christine Watts for help in refining the paper. We also extend our gratitude to all sample donors for making this study possible. Z-UR is grateful to Dr. Jan Alam and Abdul Hameed from Hazara University, Pakistan, for discussions on the anthropological/archeological perspectives of the study. Z-UR is also grateful to former colleagues from Dr. Obaid lab at University of Karachi, Pakistan, for their support. We are also thankful to the four reviewers for providing helpful comments and suggestions.
Funding
This work was supported by the Strategic Priority Research Program (Grant No. XDA20040102), Second Tibetan Plateau Scientific Expedition, Research (STEP) (Grant No. 2019QZKK0607), National Natural Science Foundation of China (31620103907, 31601017), Chinese Academy of Sciences (QYZDB-SSW-SMC020), and Yunnan Applied Basic Research Project (2017FB044).
Author information
Authors and Affiliations
Contributions
Q-PK and Y-CL designed the research; Z-UR collected samples; Z-UR and J-YT collected the data; Z-UR, J-YT, B-YY, and L-QY performed the experiments; Z-UR, Y-CL, and J-YT analyzed the data; Z-LG, H-TW, and W-XX assisted in data analysis and discussed the results; Z-UR, Y-CL, and Q-PK wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Rahman, Z.U., Tian, JY., Gao, ZL. et al. Complete mitogenomes document substantial genetic contribution from the Eurasian Steppe into northern Pakistani Indo-Iranian speakers. Eur J Hum Genet 29, 1008–1018 (2021). https://doi.org/10.1038/s41431-021-00829-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-021-00829-6
This article is cited by
-
Mitogenome analysis reveals predominantly ancient Yellow River origin of population inhabiting Datong agro-pastoral ecotone along Great Wall
Molecular Genetics and Genomics (2023)