Abstract
Some of the most widely studied variants in psychiatric genetics include variable number tandem repeat variants (VNTRs) in SLC6A3, DRD4, SLC6A4, and MAOA. While initial findings suggested large effects, their importance with respect to psychiatric phenotypes is the subject of much debate with broadly conflicting results. Despite broad interest, these loci remain absent from the largest available samples, such as the UK Biobank, limiting researchers’ ability to test these contentious hypotheses rigorously in large samples. Here, using two independent reference datasets, we report out-of-sample imputation accuracy estimates of >0.96 for all four VNTR variants and one modifying SNP, depending on the reference and target dataset. We describe the imputation procedures of these candidate variants in 486,551 UK Biobank individuals, and have made the imputed variant data available to UK Biobank researchers. This resource, provided to the scientific community, will allow the most rigorous tests to-date of the roles of these variants in behavioral and psychiatric phenotypes.
Similar content being viewed by others
Introduction
Early genetic association studies of psychiatric traits were predicated on optimism regarding the existence of common variants with substantial effects on disease liability [1]. A collection of common variable number tandem repeat variants (VNTRs), located in SLC6A3, DRD4, SLC6A4, and MAOA were central to these early investigations and continue to receive considerable attention, each sharing two common qualities: plausible biological relevance to psychiatric traits and established assay methods [2,3,4,5]. As a prominent example, the 5HTTLPR variant in SLC6A4 was hypothesized to contribute to liability for affective disorders due to its functional role in serotonin uptake [2] and soon became a popular research target across a variety of psychiatric and behavioral traits, including anxiety [6], schizophrenia [7], and personality [8]. A highly-cited (>8000 citations as of May, 2018) gene-by-environment study in 2003 [9] further fueled interest in the 5HTTLPR variant, which has yet to decline; at least 15 meta-analyses of the effects of 5HTTLPR on behavioral phenotypes were published between 2015 and 2017 (see Supplement).
Despite broad and continued interest in contributions of these variants to psychiatric outcomes, the validity of much of the research supporting their relevance remains controversial. Specifically, critics have pointed to replication failures at the variant- and whole-gene levels [10, 11], evidence for systematic publication bias [12], and inadequate statistical power [13]. Further, results from modern genome-wide association studies (GWAS), derived from samples of hundreds of thousands of individuals, do not implicate the great majority of previous candidate variants comprised of (or in high linkage disequilibrium with) single nucleotide polymorphisms (SNPs) [14, 15]. However, the failure to examine the role of many candidate repeat variants in GWAS has been a long-standing complaint of GWAS critics [16], and the absence of these variants within large GWAS datasets has prevented direct replication attempts of several prominent candidate VNTRs using GWAS data. While several studies have attempted to leverage GWAS data to infer candidate gene VNTRs (for instance, SLC6A4 5HTTLPR [17, 18]), these variants are absent from the largest datasets. Given these limitations, as well as the continued controversy surrounding past candidate variant results [19, 20], the current research sought to impute highly-studied candidate VNTRs in SLC6A3 (estimated position hg19 chr5:g.(1393863_1393862)ins(3_13)), DRD4 (hg19 chr11:g.(639989_640194)ins(3_10)), SLC6A4 (hg19 chr17:g.(28564296_28564497)ins(14_16)), and MAOA (hg19 chrX:g.(43514349_43514453)ins(2_5)), and the modifying SNP (rs25531; hg19 chr17:g.28564346 A > G) in SLC6A4, using genome-wide SNP data in 486,551 individuals in the widely-used UK Biobank (UKBB) sample [21]. In addition to imputed genotypes, which are available to qualified researchers through the UKBB, we provide validation data and describe an approach generally applicable to the imputation of variants previously unavailable in GWAS data. Our results aim to provide resources for the reconciliation of candidate variant studies and GWAS findings, with the broader goal of identifying the lines of inquiry most likely to provide insight into the genetic architecture of psychiatric traits.
Materials and methods
Reference datasets
The Family Transitions Project (FTP) initiated in 1989, was developed to examine factors influencing family economics in rural Iowa and is largely of European ancestry [22]. We used previously published VNTR and genome-wide SNP array data. Individuals were genotyped for VNTRs in the four target genes at the CU IBG Genotyping Core Facility as previously described [23,24,25]. SNP array genotypes were obtained from FTP participants using the Illumina HumanOmni-1 Quad and Illumina HumanOmniExpressExome platforms (Stallings et al. in preparation). We assigned the physical position of each SNP using the UCSC Genome Browser build hg19. The number of individuals with both SNP array data and candidate gene variant data varied among loci: 1982 individuals at the SLC6A3 VNTR, 1951 individuals at the DRD4 VNTR, 1963 individuals at SLC6A4 5HTTLPR, 1949 individuals at SLC6A4-rs25531, and 1936 individuals at the MAOA VNTR (895 males and 1041 females).
The Center for Antisocial Drug Dependence (CADD) and the Genetics of Antisocial Drug Dependence (GADD) studies were established to evaluate links among genetic variation and risk behaviors [26, 27]. The samples were collected from subjects in Colorado and California, and reflects more diverse European, Hispanic and African American ancestry, and were genotyped using the Affymetrix 6.0 SNP array [26]. VNTRs were genotyped at the CU IBG Genotyping Core Facility. The number of individuals with both SNP array data and candidate gene variant data varied among loci: 1,050 individuals at the SLC6A3 VNTR, 1,031 individuals at the DRD4 VNTR, 1052 individuals at SLC6A4 5HTTLPR, 658 individuals at SLC6A4 rs25531, and 838 individuals at the MAOA VNTR (565 males and 273 females). The numbers of individuals varied across VNTRs because of successful or failed PCR amplification during the genotyping. Such variability among loci is not uncommon for these VNTRs [23].
Population structure of reference panels with respect to the UK Biobank
We used principal components analysis (PCA) to compare the two reference panels to the UK Biobank. Due to the size of the UK Biobank, we randomly selected 50,000 individuals for this analysis. We combined the three datasets, retaining only SNPs that were present in all. We then filtered SNPs based on minor allele frequency and linkage disequilibrium (LD) with version 1.9 of plink2 [28]. (command: --maf 0.05 --geno 0.001 --hwe 0.0001 --indep-pairwise 50 5 0.2), and used this set of SNPs for PCA with flashpca2 [29]). A total of 40,037 biallelic SNPs were used in the final analysis.
Estimation of imputation accuracy by reciprocal reference imputation
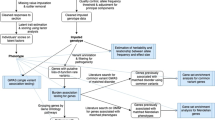
To estimate the accuracy of our imputation of the candidate gene variants, we used the two reference datasets (with both SNP array and directly-genotyped VNTR data) to reciprocally impute the VNTRs (Supplementary Figure S1). We chose to assess imputation accuracy via reciprocal imputation rather than combining the two reference panels and using a cross-validation strategy because such an estimate is more conservative, and incorporates inaccuracy induced by imputation into an independent sample, such as the UK Biobank. As the two samples were genotyped on different arrays, we first imputed both to the Haplotype Reference Consortium (HRC) [30]. To do this, we first extracted all array SNPs within 1.5 Mbp of the focal variant (physical positions listed in Tables S1-S6, size chosen to reflect a balance between computational efficiency and the number of markers in the analyses). We then phased the each of the 3 Mbp regions independently within each sample using shapeit2 [31] and imputed to the HRC using Minimac3 using default parameters [32]. For the MAOA region on chromosome X, we imputed males and females separately as recommended. We retained all imputed, biallelic SNPs with imputation INFO scores of ≥0.6. These were then used to reciprocally impute masked VNTR data within the CADD/GADD and FTP datasets with Minimac3, again using default parameters [32].
In all cases, VNTRs were treated as biallelic, using either short/long allele designation or based on the putative risk allele from published literature [10, 25, 33,34,35]. While the VNTRs contained multiple alleles, preliminary tests imputing multiallelic genotypes with Beagle v4.1 [36] had poor accuracy compared to biallelic imputation. Furthermore, candidate gene association studies often treat these VNTRs as biallelic, with risk or wildtype alleles used rather than the repeat number [10, 25, 33,34,35]. VNTR repeat numbers corresponding to the biallelic designations are reported in Supplemental Tables S1-S6.
We compared the imputed genotypes to directly genotyped candidate gene variants to assess accuracy. For each biallelic, imputed variant, we calculated the imputed risk variant frequency, the Minimac3 INFO score, the empirical squared correlation between the imputed and observed number of risk alleles, the overall proportion of genotypes correctly imputed, and the proportion of alleles correctly imputed. As these measures are in part impacted by minor allele frequency [32, 37], we also estimated the allelic match rate of the minor allele only. We estimated LD between candidate gene variants (as biallelic) and surrounding array SNPs using the --r2 plink2 command. We assessed these first using all imputed genotypes, and second restricting to those imputed calls with genotype probabilities ≥ 0.99.
Combined reference panel and imputation of the UK Biobank
To impute the candidate variants in the UK Biobank, we combined the CADD/GADD and FTP data to maximize reference panel size and diversity. We merged the independently phased CADD/GADD and FTP array and VNTR data, then imputed the combined reference to the HRC with Minimac3, retaining the target variants and all imputed SNPs with INFO scores of ≥0.6.
In the UK Biobank sample, which was imputed to the HRC by the UK Biobank [38], we retained all biallelic SNPs with imputation INFO scores of ≥0.6 within 3 Mbp of the target variants. For computational efficiency, we phased each of these candidate variant regions in four equally-sized, randomly-chosen batches (three of 121,642 and one of 121,439 individuals) using shapeit2. In none of the analyses did we remove related individuals; the presence of cryptic relatives should have no detriment to the imputation accuracy, and can improve accuracy as relatives will share longer stretches of identical-by-descent haplotypes [39]. We then imputed these batches to the combined CADD/GADD and FTP reference panel using Minimac3.
We used a one-way ANOVA to assess how self-reported ethnicity (field 21000.0.0 in the UK Biobank data) influenced imputed variant genotype probability.
Results
Population structure of reference panels with respect to the UK Biobank
We used two independent reference datasets with both directly genotyped VNTR and genome-wide SNP array data to assess the accuracy of VNTR imputation. We first compared these two reference datasets to the UK Biobank using PCA to assess ancestry of the samples, as reference and target panel diversity and ancestry can impact imputation accuracy [40]. Samples from the Family Transitions Project (FTP) [22, 25], the CADD and the GADD [23, 24, 26] have directly genotyped candidate variants and genome-wide array data. The FTP dataset was collected from participants in rural Iowa and is of largely European ancestry, while the CADD/GADD dataset, collected from subjects in Colorado and California, is more diverse, including a substantial proportion of Hispanic ancestry participants (Fig. 1). There are few individuals of South Asian ancestry in either CADD/GADD or the FTP sample (Fig. 1, negative PC3 axis); therefore, genotypes of South Asian ancestry individuals in the UK Biobank were likely imputed with lower accuracy. However, as we did not have an independent sample with VNTR genotypes reflecting this population, we were unable to directly test this hypothesis. Still, the combined FTP and CADD/GADD dataset comprised a reasonable reference panel for the majority of the UK Biobank.
Estimation of VNTR imputation accuracy with reference datasets
For each candidate variant, sample sizes of the two independent reference datasets with both directly genotyped VNTR and genome-wide SNP data were, for FTP and CADD/GADD, respectively: SLC6A3 VNTR: 1982 and 1050; DRD4 VNTR: 1951 and 1031; SLC6A4 5HTTLPR: 1963 and 1052; SLC6A4 rs25531: 1949 and 658; and MAOA VNTR: 1936 and 838. We reciprocally imputed the target variants (see Methods) in each sample using the other as the reference panel. Initial attempts to impute the exact number of repeats of the VNTRs (using Beagle v4.1 [36]) had poor accuracy compared to treating the VNTRs as biallelic. As the vast majority of candidate gene association studies (e.g., [10, 25, 33,34,35]) treat these as biallelic long/short or risk/wild-type, we used Minimac3 [32] to impute them as biallelic variants, which greatly improved accuracy. Imputation quality of biallelic variants using Minimac3 or Beagle v4.1 is likely to be similar [32, 36].
Overall, imputation accuracies, as measured by the proportion of correctly imputed biallelic genotypes, ranged from 0.81–0.99 (Table 1, Supplementary Tables S1-S6). VNTR imputation accuracy was greater when using CADD/GADD as a reference panel and FTP as the target, with genotypic match rates > 0.9, as expected because CADD/GADD is more diverse than FTP, and perhaps due to array differences in tagging the focal variants (Supplementary Figure S2). Minor allele match rates were similar to overall allelic match rates, perhaps because all imputed biallelic variants were relatively common (MAF > 0.05).
Restricting the comparisons to high-quality imputed genotypes with genotype probabilities ≥ 0.99 increased genotypic and allelic match rates (Table 1, Supplementary Tables S1-S6). While genotypic match rates in the CADD/GADD dataset improved, all match rates were >0.96 in the FTP dataset when CADD/GADD was used as a reference panel, reflecting the better performance of the more diverse reference panel. For SLC6A4 5HTTLPR, the genotype accuracies of >0.93 were higher than those obtained from a previously-published vertex discriminant analysis (0.89–0.92) [18], and the allelic match rate of >0.96 (Table 1) was higher than that suggested by a two-SNP haplotype-based method (~0.94) [17].
Empirical squared correlations showed similar patterns and increased when restricted to high-quality imputed genotypes with genotype probabilities ≥ 0.99. Imputation INFO scores from Minimac3 across all target/reference panel combinations and across all variants were over 0.92 (Supplementary Tables S1-S6).
Imputed VNTR risk variant frequencies were similar to the true risk variant frequencies. Restricting to high-quality imputed genotypes with genotype probabilities ≥ 0.99 did not alter frequencies greatly (Supplementary Tables S1-S6). Furthermore, they were also similar to estimates from other populations [23, 41].
Imputation INFO scores in the UK Biobank
We used the FTP and CADD/GADD datasets as a combined reference panel to impute the VNTRs and one moderating SNP (rs25531 in SLC6A4) to the UKBB. In the UKBB, Minimac3 INFO scores across the target variants were >0.88 and four of the five variants had INFO > 0.9 (Table 2, Supplementary Table S7), similar to the reciprocally-imputed reference panel estimates. The imputed variant frequencies were also very similar to previously published estimates [23] and those in the CADD/GADD and FTP datasets (Table 1). While we did not have a way to independently assess the imputation accuracy in the UK Biobank, genotypic match rates are likely to be >0.9 and even higher if restricted to high-quality imputed genotypes (genotype probability ≥ 0.99), given estimates from reciprocally imputing the two reference panels and the fact that the combined CADD/GADD and FTP reference panel was larger and more diverse than either individually. Of the 486,551 individuals, the imputed genotype probability was ≥0.99 for 347,916 (DRD4), 254,998 (MAOA), 326,546 (SLC6A3), 228,274 (SLC6A4 5HTTLPR), and 419,411 (SLC6A4 rs25531). Imputation accuracy, as measured by genotype probability of the imputed variants, was highest in individuals of self-reported European ancestry, as expected because the combined CADD/GADD and FTP reference panel was primarily of European and Hispanic ancestry (Supplementary Figure S3 and Fig. 1).
All imputed UK Biobank genotypes are available through the UK Biobank Data Showcase (http://www.ukbiobank.ac.uk/).
Discussion
The present work successfully imputed four highly studied candidate VNTRs and one moderating SNP in a sample of 486,551 individuals in the UKBB sample, the largest sample to-date for which these candidate variants are available. Additionally, we provide estimates of out-of-sample misclassification probabilities for each variant, as well as outline a general approach for the imputation of common repeat variants currently absent from GWAS reference panels. To the extent that imputation is imperfect as measured by an information score α ≤ 1, it will reduce the effective sample size, within a sample of size N, to approximately αN [40]. Given the large size of the UK Biobank and the INFO scores of Table 2, this is unlikely to reduce power substantially, except for subsamples for whom the reference panel used was not a good ancestry match (Supplementary Figure S3), as ancestry differences can impact imputation quality [42]. As reference panels become larger and more diverse, we anticipate future improvement. Limitations included a modest reference panel size and the lack of an independent test of accuracy for the UK Biobank sample itself when using the combined CADD/GADD and FTP reference panel. Furthermore, we imputed the VNTRs as biallelic risk/wild-type or short/long alleles, rather than the actual number of repeats. While this is the standard approach to association testing and functional characterization with these loci [10, 25, 33,34,35], it does not reflect their total allelic diversity. The rich variety of phenotypes available through the UK Biobank will permit future interrogation of several widely-studied hypotheses previously inaccessible in the context of GWAS data (e.g., stressful life event × 5HTTLPR effects on liability for depression), and in doing so will provide the most robust tests of these highly debated candidate variant hypotheses.
Data access
All imputed UK Biobank candidate variants have been returned to the UK Biobank (http://www.ukbiobank.ac.uk/).
References
McInnes LA, Freimer NB. Mapping genes for psychiatric disorders and behavioral traits. Curr Opin Genet Dev. 1995;5:376–81.
Ramamoorthy S, Bauman AL, Moore KR, et al. Antidepressant and cocaine-sensitive human serotonin transporter: molecular cloning, expression, and chromosomal localization. Proc Natl Acad Sci USA. 1993;90:2542–6.
Sabol SZ, Hu S, Hamer D. A functional polymorphism in the monoamine oxidase A gene promoter. Hum Genet. 1998;103:273–9.
Tol HHM Van, Wu CM, Guan H-C, et al. Multiple dopamine D4 receptor variants in the human population. Nature. 1992;358:149–52.
Vandenbergh DJ, Persico AM, Hawkins AL, et al. Human dopamine transporter gene (DAT1) maps to chromosome 5p15.3 and displays a VNTR. Genomics. 1992;14:1104–6.
Lesch KP, Bengel D, Heils A, et al. Association of anxiety-related traits with a polymorphism in the serotonin transporter gene regulatory region. Science. 1996;274:1527–31.
Collier DA, Arranz MJ, Sham P. The serotonin transporter gene is a potential susceptibility factor for biplor affective disorder. Neuroreport. 1996;7:1675–9.
Hamer DH, Greenberg BD, Sabol SZ, Murphy DL. Role of the serotonin transporter gene in temperament and character. J Pers Disord. 1999;13:312–27.
Caspi A, Sugden K, Moffitt TE, et al. Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science. 2003;301:386–9.
Culverhouse RC, Saccone NL, Horton AC, et al. Collaborative meta-Analysis finds no evidence of a strong interaction between stress and 5-HTTLPR genotype contributing to the development of depression. Mol Psychiatry. 2018;23:133–42.
Johnson EC, Border R, Melroy-Greif WE, de Leeuw CA, Ehringer MA, Keller MC. No evidence that schizophrenia candidate genes are more associated with schizophrenia than noncandidate genes. Biol Psychiatry. 2017;82:702–8.
Duncan LE, Keller MC. A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am J Psychiatry. 2011;168:1041–9.
Burton PR, Hansell AL, Fortier I, et al. Size matters: just how big is BIG?: quantifying realistic sample size requirements for human genome epidemiology. Int J Epidemiol. 2009;38:263–73.
Bosker FJ, Hartman CA, Nolte IM, et al. Poor replication of candidate genes for major depressive disorder using genome-wide association data. Mol Psychiatry. 2011;16:516–32.
Farrell MS, Werge T, Sklar P, et al. Evaluating historical candidate genes for schizophrenia. Mol Psychiatry. 2015;20:555–62.
Brookes KJ. The VNTR in complex disorders: The forgotten polymorphisms? A functional way forward? Genomics. 2013;101:273–81.
Vinkhuyzen AAE, Dumenil T, Ryan L, et al. Identification of tag haplotypes for 5HTTLPR for different genome-wide SNP platforms. Mol Psychiatry. 2011;16:1073–5.
Lu AT-H, Bakker S, Janson E, Cichon S, Cantor RM, Ophoff RA. Prediction of serotonin transporter promoter polymorphism genotypes from single nucleotide polymorphism arrays using machine learning methods. Psychiatr Genet. 2012;22:182–8.
Assary E, Vincent JP, Keers R, Pluess M. Gene-environment interaction and psychiatric disorders: review and future directions. Semin Cell Dev Biol. 2018;77:133–43.
Duncan LE, Pollastri AR, Smoller JW. Mind the gap: why many geneticists and psychological scientists have discrepant views about gene-environment interaction (G × E) research. Am Psychol. 2014;69:249–68.
Sudlow C, Gallacher J, Allen N, et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12:1–10.
Conger RD, Schofield TJ, Neppl TK. Intergenerational continuity and discontinuity in harsh parenting. Parenting. 2012;12:222–31.
Haberstick BC, Smolen A, Stetler GL, et al. Simple sequence repeats in the national longitudinal study of adolescent health: an ethnically diverse resource for genetic analysis of health and behavior. Behav Genet. 2014;44:487–97.
Haberstick BC, Smolen A, Williams RB, et al. Population frequencies of the triallelic 5HTTLPR in six ethnicially diverse samples from North America, Southeast Asia, and Africa. Behav Genet. 2015;96:255–61.
Masarik AS, Conger RD, Brent Donnellan M, et al. For better and for worse: genes and parenting interact to predict future behavior in romantic relationships. J Fam Psychol. 2014;28:357–67.
Derringer J, Corley RP, Haberstick BC, et al. Genome-wide association study of behavioral disinhibition in a selected adolescent sample. Behav Genet. 2015;45:375–81.
Young SE, Stallings MC, Corley RP, Krauter KS, Hewitt JK. Genetic and environmental influences on behavioral disinhibition. Am J Med Genet Part B Neuropsychiatr Genet. 2000;695:684–95.
Chang CC, Chow CC, Tellier LC, et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:7.
Abraham G, Inouye M. Fast principal component analysis of large-scale genome-wide data. PLoS One. 2014;9:e92766.
McCarthy S, Das S, Kretzschmar W, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279–83.
Delaneau O, Zagury J-F, Marchini J. Improved whole-chromosome phasing for disease and population genetic studies. Nat Methods. 2013;10:5–6.
Das S, Forer L, Schönherr S, et al. Next-generation genotype imputation service and methods. Nat Genet. 2016;48:1287–1287.
Drury SS, Theall KP, KB JB, Scheeringa M. The role of the dopamine transporter (DAT) in the development of preschool children. J Trauma Stress. 2009;22:534–9.
Yu YWY, Tsai SJ, Hong CJ, Chen TJ, Chen MC, Yang CW. Association study of a Monoamine oxidase A gene promoter polymorphism with major depressive disorder and antidepressant response. Neuropsychopharmacology. 2005;30:1719–23.
Hutchison KE, McGeary J, Smolen A, Bryan A, Swift RM. The DRD4 VNTR polymorphism moderates craving after alcohol consumption. Heal Psychol. 2002;21:139–46.
Browning BL, Browning SR. Genotype imputation with millions of reference samples. Am J Hum Genet. 2016;98:116–26.
Mitt M, Kals M, Pärn K, et al. Improved imputation accuracy of rare and low-frequency variants using population-specific high-coverage WGS-based imputation reference panel. European Journal of Human Genetics. 2017;25:869–76.
Bycroft C, Freeman C, Petkova D et al. The UK Biobank resource with deep phenotyping and genomic data. Nature. 2018;562:203–9.
Li Y, Willer C, Sanna S, Abecasis GR. Genotype imputation. Annu Rev Genom Hum Genet. 2009;10:387–406.
Marchini J, Howie B. Genotype imputation for genome-wide association studies. Nat Rev Genet. 2010;11:499–511.
Chang FM, Kidd JR, Livak KJ, Pakstis AJ, Kidd KK. The world-wide distribution of allele frequencies at the human dopamine D4 receptor locus. Hum Genet. 1996;98:91–101.
Deelen P, Menelaou A, Leeuwen EM Van et al. Improved imputation quality of low-frequency and rare variants in European samples using the “Genome of The Netherlands”. European Journal of Human Genetics. 2014;1321–6.
Acknowledgements
We thank the participants of the FTP, CADD/GADD and UK Biobank studies. This work was supported by NIH R01MH100141 to MCK and the Institute for Behavioral Genetics. RB is supported by NIH T32MH016880. The FTP was supported by NICHD HD064687. CADD was supported by NIDA DA011015 and DA035804. GADD was supported by DA012845, DA035804, and DA021692. This work utilized the RMACC Summit supercomputer, which is supported by the National Science Foundation (awards ACI-1532235 and ACI-1532236), the University of Colorado Boulder, and Colorado State University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
BIORXIV PREPRINT: https://doi.org/10.1101/358267
Supplementary information
Rights and permissions
About this article
Cite this article
Border, R., Smolen, A., Corley, R.P. et al. Imputation of behavioral candidate gene repeat variants in 486,551 publicly-available UK Biobank individuals. Eur J Hum Genet 27, 963–969 (2019). https://doi.org/10.1038/s41431-019-0349-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-019-0349-x
This article is cited by
-
Cross-ancestry genetic architecture and prediction for cholesterol traits
Human Genetics (2024)
-
A method for an unbiased estimate of cross-ancestry genetic correlation using individual-level data
Nature Communications (2023)
-
Revisiting tandem repeats in psychiatric disorders from perspectives of genetics, physiology, and brain evolution
Molecular Psychiatry (2022)
-
A summary-statistics-based approach to examine the role of serotonin transporter promoter tandem repeat polymorphism in psychiatric phenotypes
European Journal of Human Genetics (2022)
-
A Re-evaluation of Candidate Gene Studies for Well-Being in Light of Genome-Wide Evidence
Journal of Happiness Studies (2022)