Abstract
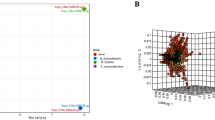
Identifying small compounds capable of inhibiting Mycobacterium tuberculosis polyketide synthase 13 (Pks13), in charge of final step of mycolic acid biosynthesis, could lead to the development of a novel antituberculosis drug. This study screened for lead compounds capable of targeting M. tuberculosis Pks13 from a chemical library comprising 154,118 compounds through multiple in silico docking simulations. The parallel compound screening (PCS), conducted via two genetic algorithm-based programs was applied in the screening strategy. Out of seven experimentally validated compounds, four compounds showed inhibitory effects on the growth of the model mycobacteria (Mycobacterium smegmatis). Subsequent docking simulation of analogs of the promising leads with the assistance of PCS resulted in the identification of three additional compounds with potent antimycobacterial effects (compounds A1, A2, and A5). Further, molecular dynamics simulation predicted stable interaction between M. tuberculosis Pks13 active site and compound A2, which showed potent antimycobacterial activity comparable to that of isoniazid. The present study demonstrated the efficacy of in silico structure-based drug screening through PCS in antituberculosis drug discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lönnroth K, Castro KG, Chakaya JM, Chauhan LS, Floyd K, Glaziou P, et al. Tuberculosis control and elimination 2010-50: cure, care, and social development. Lancet. 2010;375:1814–29.
Dye C, Williams BG. The population dynamics and control of tuberculosis. Science. 2010;328:856–61.
Torres JN, Paul LV, Rodwell TC, Victor TC, Amallraja AM, Elghraoui A, et al. Novel katG mutations causing isoniazid resistance in clinical M. tuberculosis isolates. Emerg Microbes Infect. 2015;4:e42
Shaku M, Ealand C, Kana BD. Cell surface biosynthesis and remodeling pathways in mycobacteria reveal new drug targets. Front Cell Infect Microbiol. 2020;10:603382.
Takayama K, Wang C, Besra GS. Pathway to synthesis and processing of mycolic acids in Mycobacterium tuberculosis. Clin Microbiol Rev. 2005;18:81–101.
Portevin D, De sousa-D'auria C, Houssin C, Grimaldi C, Chami M, Daffe M, et al. A polyketide synthase catalyzes the last condensation step of mycolic acid biosynthesis in mycobacteria and related organisms. Proc Natl Acad Sci USA. 2004;101:314–9.
Gavalda S, Leger M, van der Rest B, Stella A, Bardou F, Montrozier H, et al. The Pks13/FadD32 crosstalk for the biosynthesis of mycolic acids in Mycobacterium tuberculosis. J Biol Chem. 2009;284:19255–64.
Gavalda S, Bardou F, Laval F, Bon C, Malaga W, Chalut C, et al. The polyketide synthase Pks13 catalyzes a novel mechanism of lipid transfer in mycobacteria. Chem Biol. 2014;21:1660–9.
Lun S, Xiao S, Zhang W, Wang S, Gunosewoyo H, Yu LF, et al. Therapeutic potential of coumestan Pks13 inhibitors for tuberculosis. Antimicrob Agents Chemother. 2021;65:e02190–20.
Ioerger TR, O’Malley T, Liao R, Guinn KM, Hickey MJ, Mohaideen N, et al. Identification of new drug targets and resistance mechanisms in Mycobacterium tuberculosis. PLoS ONE. 2013;8:e75245
Wilson R, Kumar P, Parashar V, Vilcheze C, Veyron-Churlet R, Freundlich JS, et al. Antituberculosis thiophenes define a requirement for Pks13 in mycolic acid biosynthesis. Nat Chem Biol. 2013;9:499–506.
North EJ, Jackson M, Lee RE. New approaches to target the mycolic acid biosynthesis pathway for the development of tuberculosis therapeutics. Curr Pharm Des. 2014;20:4357–78.
Aggarwal A, Parai MK, Shetty N, Wallis D, Woolhiser L, Hastings C, et al. Development of a novel lead that targets M. tuberculosis polyketide synthase 13. Cell. 2017;170:249–59.e225.
Wilson C, Ray P, Zuccotto F, Hernandez J, Aggarwal A, Mackenzie C, et al. Optimization of TAM16, a benzofuran that inhibits the thioesterase activity of Pks13; evaluation toward a preclinical candidate for a novel antituberculosis clinical target. J Med Chem. 2022;65:409–23.
Koseki Y, Aoki S. Computational medicinal chemistry for rational drug design: Identification of novel chemical structures with potential anti-tuberculosis activity. Curr Top Med Chem. 2014;14:176–88.
Kuriki K, Taira J, Kuroki M, Sakamoto H, Aoki S. Computer-assisted screening of mycobacterial growth inhibitors: Exclusion of frequent hitters with the assistance of the multiple target screening method. Int J Mycobacteriol. 2021;10:307–11.
Izumizono Y, Arevalo S, Koseki Y, Kuroki M, Aoki S. Identification of novel potential antibiotics for tuberculosis by in silico structure-based drug screening. Eur J Med Chem. 2011;46:1849–56.
Kanetaka H, Koseki Y, Taira J, Umei T, Komatsu H, Sakamoto H, et al. Discovery of InhA inhibitors with anti-mycobacterial activity through a matched molecular pair approach. Eur J Med Chem. 2015;94:378–85.
Taira J, Umei T, Inoue K, Kitamura M, Berenger F, Sacchettini JC, et al. Improvement of the novel inhibitor for Mycobacterium enoyl-acyl carrier protein reductase (InhA): a structure-activity relationship study of KES4 assisted by in silico structure-based drug screening. J Antibiot. 2020;73:372–81.
Taira J, Ito T, Nakatani H, Umei T, Baba H, Kawashima S, et al. In silico structure-based drug screening of novel antimycobacterial pharmacophores by DOCK-GOLD tandem screening. Int J Mycobacteriol. 2017;6:142–8.
Taira J, Morita K, Kawashima S, Umei T, Baba H, Maruoka T, et al. Identification of a novel class of small compounds with anti-tuberculosis activity by in silico structure-based drug screening. J Antibiot. 2017;70:1057–64.
Nakashima J, Takeuchi M, Kawamoto S, Monobe K, Taira J, Aoki S. Establishing parallel compound screening and identification of novel antimicrobial compounds targeting Staphylococcus aureus dihydrofolate reductase. J Appl Pharm Sci. 2022, in press.
Ewing TJ, Makino S, Skillman AG, Kuntz ID. DOCK 4.0: search strategies for automated molecular docking of flexible molecule databases. J Comput Aided Mol Des. 2001;15:411–28.
Verdonk ML, Cole JC, Hartshorn MJ, Murray CW, Taylor RD. Improved protein-ligand docking using GOLD. Proteins. 2003;52:609–23.
Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. 2010;31:455–61.
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 2001;46:3–26.
Durant JL, Leland BA, Henry DR, Nourse JG. Reoptimization of MDL keys for use in drug discovery. J Chem Inf Comput Sci. 2002;42:1273–80.
T JAS, J R, Rajan A, Shankar V. Features of the biochemistry of Mycobacterium smegmatis, as a possible model for Mycobacterium tuberculosis. J Infect Public Health. 2020;13:1255–64.
Bloemberg GV, Keller PM, Stucki D, Trauner A, Borrell S, Latshang T, et al. Acquired resistance to bedaquiline and delamanid in therapy for tuberculosis. N Engl J Med. 2015;373:1986–8.
Olayanju O, Limberis J, Esmail A, Oelofse S, Gina P, Pietersen E, et al. Long-term bedaquiline-related treatment outcomes in patients with extensively drug-resistant tuberculosis from South Africa. Eur Respir J. 2018;51.
Marrakchi H, Laneelle MA, Daffe M. Mycolic acids: structures, biosynthesis, and beyond. Chem Biol. 2014;21:67–85.
Cruz JN, Costa JFS, Khayat AS, Kuca K, Barros CAL, Neto A. Molecular dynamics simulation and binding free energy studies of novel leads belonging to the benzofuran class inhibitors of Mycobacterium tuberculosis Polyketide Synthase 13. J Biomol Struct Dyn. 2019;37:1616–27.
Funding
This work was supported in part by a grant from Takeda Science Foundation to JT and a Grant-in-Aid for Scientific Research (C) (26460145) to SA and a Grant-in-Aid for Transformative Research Areas (A) (21H05887) to JT from Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Taira, J., Murakami, K., Monobe, K. et al. Identification of novel inhibitors for mycobacterial polyketide synthase 13 via in silico drug screening assisted by the parallel compound screening with genetic algorithm-based programs. J Antibiot 75, 552–558 (2022). https://doi.org/10.1038/s41429-022-00549-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-022-00549-z