Abstract
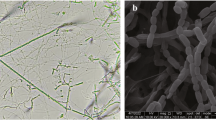
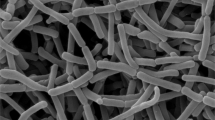
A novel actinomycete strain CYP1-5T was isolated from the mountain soil sample collected from Chaiyaphum province, Thailand and its taxonomic position was clarified by using a polyphasic taxonomic approach. The chemotaxonomic properties of strain CYP1-5T were consistent within the genus Actinomadura. Cell-wall peptidoglycan of this strain contained meso-diaminopimelic acid. Galactose, madurose, and ribose were presented as the diagnostic sugars in whole-cell hydrolysates. The major menaquinone was MK-9(H6). Major cellular fatty acids were iso-C16:0 and C16:0. Phosphatidylglycerol, diphosphatidylglycerol, phosphatidylinositol, and phosphatidylinositol mannoside were observed as predominant phospholipids. Based on the results of phylogenetic analyses of 16S rRNA gene sequence, strain CYP1-5T was constituent with the genus Actinomadura and was closely related to Actinomadura syzygii GKU157T (99.5%) and Actinomadura chibensis IFM 10266T (= JCM 14158T) (98.2%). The draft genome size of strain CYP1-5T was 9.30 Mb with 72.2 mol% of G + C content. Strain CYP1-5T showed ANIb values of 94.9% with A. syzygii GKU157T and 93.2% with A. chibensis JCM 14158T. Phenotypic characteristics, phylogenetic analysis and genome data support that strain CYP1-5T could be discriminated from its closest relatives, representing a novel species of the genus Actinomadura, for which the name Actinomadura decatromicini sp. nov. is proposed. The type strain is CYP1-5T (= JCM 16996T = KCTC 19916T = TISTR 2901T).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lechevalier MP, Lechevalier H. Chemical composition as a criterion in the classification of aerobic actinomycetes. Int J Syst Bacteriol. 1970;20:435–43.
Kroppenstedt RM, Stackebrandt E, Goodfellow M. Taxonomic revision of the actinomycete genera Actinomadura and Microtetraspora. Syst Appl Microbiol. 1990;13:148–60.
Kroppenstedt RM, Goodfellow M. The family Thermomonosporaceae: Actinocorallia, Actinomadura, Spirillispora and Thermomonospora. In: Dworkin M, Falkow S, Schleifer KH, Stackebrandt E, editors. The Prokaryotes, Archaea and Bacteria: Firmicutes, Actinomycetes.3rd ed, vol. 3. New York: Springer; 2006. pp. 682–724.
Parte AC. List of prokaryotic names with standing in nomenclature (bacterio.net), 20 years on. Int J Syst Evol Microbiol. 2018;68:1825–9.
Ara I, Matsumoto A, Bakir M, Kudo T, Omura S, Takahashi Y. Actinomadura maheshkhaliensis sp. nov., a novel actinomycete isolated from mangrove rhizosphere soil of Maheshkhali, Bangladesh. J Gen Appl Microbiol. 2008;54:335–42.
He J, Xu Y, Sahu M, Tian XP, Nie GX, Xie Q, et al. Actinomadura sediminis sp. Nov., a marine actinomycete isolated from mangrove sediment. Int J Syst Bacteriol. 2011;62:1110–16.
Qin S, Zhao GZ, Li J, Zhu WY, Xu LH, Li WJ. Actinomadura flavalba sp. nov., an endophytic actinomycete isolated from leaves of Maytenus austroyunnanensis. Int J Syst Evol Microbiol. 2009;95:2453–7.
Qin S, Chen HH, Zhao GZ, Li J, Zhu WY, Xu LH, et al. Abundant and diverse endophytic actinobacteria associated with medicinal plant Maytenus austroyunnanensis in Xishuangbanna tropical rainforest revealed by culture-dependent and culture-independent methods. Environ Microbiol Rep. 2012;4:522–31.
Rachniyom H, Matsumoto A, Indananda C, Duangmal K, Takahashi Y, Thamchaipenet A. Actinomadura syzygii sp. nov., an endophytic actinomycete isolated from the roots of a jambolan plum tree (Syzygium cumini L. Skeels). Int J Syst Evol Microbiol. 2015;65:1946–9.
Han XX, Cui CB, Gu QQ, Zhu W, Liu H, Gu JY, et al. ZHD-0501, a novel naturally occurring staurosporine analog from Actinomadura sp. 007. Tetrahedron Lett. 2005;46:6137–40.
Shaaban KA, Elshahawi SI, Wang X, Horn J, Kharel MK, Leggas M, et al. Cytotoxic indolocarbazoles from Actinomadura melliaura ATCC 39691. J Nat Prod. 2015;78:1723–9.
Takagi M, Motohashi K, Khan ST, Hashimoto J, Shin-ya K. JBIR-65, a new diterpene, isolated from a sponge-derived Actinomadura sp. SpB081030SC-15. J Antibiot. 2010;63:401–3.
Simmons L, Kaufmann K, Garcia R, Schwӓr G, Huch V, Muller R. Bendigoles D-F, bioactive sterols from the marine sponge-derived Actinomadura sp. SBMs009. Bioorg Med Chem. 2011;19:6570–75.
Mazzei E, Iorio M, Maffioli F, Sosio M, Donadio S. Characterization of madurastatin C1, a novel siderophore from Actinomadura sp. J Antibiot. 2012;65:267–9.
Igarashi Y, Iida T, Oku N, Watanabe H, Furihata K, Miyanouchi K. Nomimicin, a new spirotetronate-class polyketide from an actinomycete of the genus Actinomadura. J Antibiot. 2012;65:355–9.
Kornsakulkarn J, Saepua S, Boonruangprapa T, Suphothina S, Thongpanchang C. New β-carboline and indole alkaloids from actinomycete Actinomadura sp. BCC 24717. Phytochem Lett. 2013;6:491–4.
Intaraudom C, Dramae A, Supothina S, Konwijit S, Pittayakhajonwut P. 3-Oxyanthranilic acid derivatives from Actinomadura sp. BCC 27169. Tetrahedron. 2014;70:2711–6.
Kusserow K, Gulder TAM. Complete genome sequence of Actinomadura parvosata subsp. kistnae, a rich source of novel natural product (Bio-) chemistry. J Genom. 2017;5:75–6.
Küster E, Williams ST. Selection of media for the isolation of Streptomycetes. Nature. 1964;202:928–9.
Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. Int J Syst Bacteriol. 1966;16:313–40.
Kelly KL. Inter-Society Color Council—National Bureau of Standards Color Name Charts Illustrated with Centroid Colors. Washington, DC: US Government Printing Office; 1964.
Arai T. Culture media for Actinomycetes. Tokyo, Japan: The Society for Actinomycetes; 1975.
Williams ST, Cross T. Actinomycetes. In: Methods in microbiology. London, Academic Press; 1971. p. 295–334.
Staneck JL, Roberts GD. Simplified approach to identification of aerobic actinomycetes by thin-layer chromatography. Appl Microbiol. 1974;28:226–31.
Mikami H, Ishida Y. Post-column fluorometric detection of reducing sugar in high-performance liquid chromatography using arginine. Bunseki Kagaku. 32: E207-E.2101983.
Uchida K, Aida K. An improved method for the glycolate test for simple identification of the acyl type of bacterial cell walls. J Gen Appl Microbiol. 1984;37:463–4.
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, et al. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods. 1984;2:233–41.
Sasser M. Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. Newark, DE: MIDI, Inc; 1990.
Tomiyasu I. Mycolic acid composition and thermally adaptative changes in Nocardia asteroids. J Bacteriol. 1982;151:828–37.
Collins MD, Pirouz T, Goodfellow M, Minnikin DE. Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol. 1977;100:221–30.
Kudo T, Matsushima K, Itoh T, Sasaki J, Suzuki K. Description of four new species of the genus Kineosporia: Kineosporia succinea sp. nov., Kineosporia rhizophila sp. nov., Kineosporia mikuniensis sp. nov. and Kineosporia rhamnosa sp. nov., isolated from plant samples, and amended description of the genus Kineosporia. Int J Syst Bacteriol. 1998;48:1245–55.
Suriyachadkun C, Chunhametha S, Thawai C, Tamura T, Potacharoen W, Kirtikara K, et al. Planotetraspora thailandica sp. nov., isolated from soil in Thailand. Int J Syst Evol Microbiol. 2009;59:992–7.
Lane DJ. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. New York, Wiley; 1991. p. 115–75.
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98 NT. Nucleic acids Symp Ser. 1999;41:95–8.
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, et al. Introducing EzBiocloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol. 2017;67:1613–7.
Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16:111–20.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.
Fitch WM. Toward defining the course of evolution: minimum change for specific tree topology. Syst Zool. 1971;20:406–16.
Felsenstein J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol. 1981;17:368–76.
Kumar S, Stecher G, Tamura K. MEGA 7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–91.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012;19:455–77.
Aziz RK, Bartels D, Best AA, Dejongh M, Disz T, et al. The RAST Server: rapid annotations using subsystem technology. BMC Genom. 2008;9:75.
Meier-Kolthoff JP, Göker M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun. 2019;10:2182.
Ritcher M, Rosselló-Móra R. Shifting the genomics gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA. 2009;106:19126–31.
Ritcher M, Rosselló-Móra R, Glöckner FO, Peplies J. JspeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics. 2016;32:929–31.
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinforma. 2013;14:60.
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, et al. antiSMASH 5.0: update to the secondary metabolite genome mining pipeline. Nucleic Acid Res. 2019. https://doi.org/10.1093/nar/gkz310.
Goris J, Konstantinidis K, Klappenbach J, Coenye T, Vandamme P, et al. DNA-DNA hybridization values and their relationship to whole genome sequence similarities. Int J Syst Evol Microbiol. 2007;57:81–91.
Momose I, Iinuma H, Kinoshita N, Momose Y, Kunimoto S, Hamada M, et al. Decatromicins A and B, new antibiotics produced by Actinomadura sp. MK73-NF4. I. Taxonomy, isolation, physico-chemical properties and biological activities. J Antibiot. 1999;52:781–6.
Jørgensen H, Degnes KF, Sletta H, Fjaervik E, Dikiy A, Herfindal L, et al. Biosynthesis of macrolactam BE-14106 involves two distinct PKS systems and amino acid processing enzymes for generation of the aminoacyl starter unit. Chem Biol. 2009;16:1109–21.
Mascaretti OA, Chang CJ, Hook D, Otsuka H, Kreutzer EF, Floss HG. Biosynthesis of the macrolide antibiotic chlorothricin. Biochemistry. 1981;20:919–24.
Poralla K, Muth G, Härtner T. Hopanoids are formed during transition from substrate to aerial hyphae in Streptomyces coelicolor A3(2). FEMS Microbiol Lett. 2000;189:93–5.
Gottardi EM, Krawczyk JM, von Suchodoletz H, et al. Abyssomicin biosynthesis: formation of an unusual polyketide, antibiotic-feeding studies and genetic analysis. Chembiochem. 2011;12:1401–10.
Lefort V, Desper R, Gascuel O. FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol. 2015;32:2798–800.
Farris JS. Estimating phylogenetic trees from distance matrices. Am Nat. 1972;106:645–67.
Acknowledgements
This research is supported by Grant for International Research Integration: Research Pyramid, Ratchadaphiseksomphot Endowment Fund (GCURP_58_01_33_01). We also thank the Pharmaceutical Research Instrument Center, Faculty of Pharmaceutical Sciences, Chulalongkorn University, for providing research facilities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Songsumanus, A., Kuncharoen, N., Kudo, T. et al. Actinomadura decatromicini sp. nov., isolated from mountain soil in Thailand. J Antibiot 74, 51–58 (2021). https://doi.org/10.1038/s41429-020-0353-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-020-0353-y