Abstract
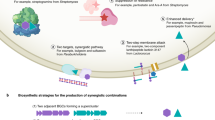
Antibiotic resistance has been an ongoing challenge that has emerged almost immediately after the initial discovery of antibiotics and requires the development of innovative new antibiotics and antibiotic combinations that can effectively mitigate the development of resistance. More than 35,000 people die each year from antibiotic resistant infections in just the United States. This signifies the importance of identifying other alternatives to antibiotics for which resistance has developed. Virtually, all currently used antibiotics can trace their genesis to soil derived bacteria and fungi. The bacteria and fungi involved in symbiosis is an area that still remains widely unexplored for the discovery and development of new antibiotics. This brief review focuses on the challenges and opportunities in the application of symbiotic microbes and also provides an interesting platform that links natural product chemistry with evolutionary biology and ecology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Gross M. The race against antibiotics resistance. Curr Biol. 2019;23:R859–61.
Centers for Disease Control and Prevention. Antibiotic/antimicrobial resistance (AR/AMR). 2020. https://www.cdc.gov/drugresistance/about.html.
Engl T, Kroiss J, Kai M, Nechitaylo TY, Svatoš A, Kaltenpoth M. Evolutionary stability of antibiotic protection in a defensive symbiosis. Proc Natl Acad Sci USA. 2018;115:E2020–9.
Centers for Disease Control and Prevention. Antibiotic resistance threats in the United States, 2019. 2019. https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf.
Staley JT, Castenholz RW, Colwell RR, Holt JG, Kane MD, Pace NR, et al. The microbial world: foundation of the biosphere. Washington, D.C.: American Academy of Microbiology; 1997.
Oulhen N, Schulz BJ, Carrier TJ. English translation of Heinrich Anton de Bary’s 1878 speech,‘Die Erscheinung der Symbiose’(‘De la symbiose’). Symbiosis. 2016;69:131–9.
Egerton FN. History of ecological sciences, Part 52: symbiosis studies. Bull Ecol Soc Am. 2015;96:80–139.
Blockley A, Elliott DR, Roberts AP, Sweet M. Symbiotic microbes from marine invertebrates: driving a new era of natural product drug discovery. Diversity. 2017;9:49.
Adnani N, Rajski SR, Bugni TS. Symbiosis-inspired approaches to antibiotic discovery. Nat Prod Rep. 2017;34:784–814.
Zhang X, Wei W, Tan R. Symbionts, a promising source of bioactive natural products. Sci China: Chem. 2015;58:1097–109.
Henkel T, Brunne RM, Müller H, Reichel F. Statistical investigation into the structural complementarity of natural products and synthetic compounds. Angew Chem, Int Ed. 1999;38:643–7.
Montaser R, Luesch H. Marine natural products: a new wave of drugs? Future Med Chem. 2011;3:1475–89.
Wang G. Diversity and biotechnological potential of the sponge-associated microbial consortia. J Ind Microbiol Biotechnol. 2006;33:545.
Reveillaud J, Maignien L, Eren AM, Huber JA, Apprill A, Sogin ML, et al. Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J. 2014;8:1198–209.
Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, et al. Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. ISME J. 2012;6:564–76.
Lee HS, Kwon KK, Kang SG, Cha S-S, Kim S-J, Lee J-H. Approaches for novel enzyme discovery from marine environments. Curr Opin Biotechnol. 2010;21:353–7.
Romano G, Costantini M, Sansone C, Lauritano C, Ruocco N, Ianora A. Marine microorganisms as a promising and sustainable source of bioactive molecules. Mar Environ Res. 2017;128:58–69.
Newman DJ. Predominately uncultured microbes as sources of bioactive agents. Front Microbiol. 2016;7:1832.
Macintyre L, Zhang T, Viegelmann C, Martinez IJ, Cheng C, Dowdells C, et al. Metabolomic tools for secondary metabolite discovery from marine microbial symbionts. Mar Drugs. 2014;12:3416–48.
Abdelmohsen UR, Bayer K, Hentschel U. Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat Prod Rep. 2014;31:381–99.
Bull AT, Stach JEM. Marine actinobacteria: new opportunities for natural product search and discovery. Trends Microbiol. 2007;15:491–9.
Wright GD. Antibiotics: a new hope. Chem Biol. 2012;19:3–10.
Fischbach MA, Walsh CT. Antibiotics for emerging pathogens. Science. 2009;325:1089–93.
Davies J, Davies D. Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev. 2010;74:417–33.
Fenical W, Jensen PR. Developing a new resource for drug discovery: marine actinomycete bacteria. Nat Chem Biol. 2006;2:666–73.
Jensen P, Fenical W. Strategies for the discovery of secondary metabolites from marine bacteria: ecological perspectives. Annu Rev Microbiol. 1994;48:559–84.
Freeman MF, Vagstad AL, Piel J. Polytheonamide biosynthesis showcasing the metabolic potential of sponge-associated uncultivated ‘Entotheonella’ bacteria. Curr Opin Chem Biol. 2016;31:8–14.
Senthilkumar K, Kim S-K. Marine invertebrate natural products for anti-inflammatory and chronic diseases. Evid-Based Compl Alt Med. 2013;2013.
Tsukimoto M, Nagaoka M, Shishido Y, Fujimoto J, Nishisaka F, Matsumoto S, et al. Bacterial production of the tunicate-derived antitumor cyclic depsipeptide didemnin B. J Nat Prod. 2011;74:2329–31.
Mehbub MF, Lei J, Franco C, Zhang W. Marine sponge derived natural products between 2001 and 2010: trends and opportunities for discovery of bioactives. Mar Drugs. 2014;12:4539–77.
Munro MHG, Blunt JW, Dumdei EJ, Hickford SJH, Lill RE, Li S, et al. The discovery and development of marine compounds with pharmaceutical potential. Prog Ind Microbiol. 1999;35:15–25.
Waters AL, Peraud O, Kasanah N, Sims JW, Kothalawala N, Anderson MA, et al. An analysis of the sponge Acanthostrongylophora igens’ microbiome yields an actinomycete that produces the natural product manzamine A. Front Mar Sci. 2014;1:54.
Mohamed NM, Rao V, Hamann MT, Kelly M, Hill RT. Monitoring bacterial diversity of the marine sponge Ircinia strobilina upon transfer into aquaculture. Appl Environ Microbiol. 2008;74:4133–43.
Jeewon R, Luckhun AB, Bhoyroo V, Sadeer NB, Mahomoodally MF, Rampadarath S, et al. Pharmaceutical potential of marine fungal endophytes. In: Jha S, editor. Endophytes and secondary metabolites. Reference series in phytochemistry. Springer; 2019. p. 1–23.
Sun W, Wu W, Liu X, Zaleta-Pinet DA, Clark BR. Bioactive compounds isolated from marine-derived microbes in China: 2009-18. Mar Drugs. 2019;17:339.
Wang R, Seyedsayamdost MR. Roseochelin B, an algaecidal natural product synthesized by the Roseobacter Phaeobacter inhibens in response to algal sinapic acid. Org Lett. 2017;19:5138–41.
Li Z-X, Wang X-F, Ren G-W, Yuan X-L, Deng N, Ji G-X, et al. Prenylated diphenyl ethers from the marine algal-derived endophytic fungus Aspergillus tennesseensis. Molecules. 2018;23:2368.
Zou J-X, Song Y-P, Ji N-Y. Deoxytrichodermaerin, a harziane lactone from the marine algicolous fungus Trichoderma longibrachiatum A-WH-20-2. Nat Prod Res. 2019:1–6.
Song Y-P, Shi Z-Z, Miao F-P, Fang S-T, Yin X-L, Ji N-Y. Tricholumin A, a highly transformed ergosterol derivative from the alga-endophytic fungus Trichoderma asperellum. Org Lett. 2018;20:6306–9.
El-Gendy MMAA, Yahya SMM, Hamed AR, Soltan MM, El-Bondkly AMA. Phylogenetic analysis and biological evaluation of marine endophytic fungi derived from Red Sea sponge Hyrtios erectus. Appl Biochem Biotechnol. 2018;185:755–77.
Suryanarayanan TS, Thirunavukkarasu N, Govindarajulu MB, Sasse F, Jansen R, Murali TS. Fungal endophytes and bioprospecting. Fungal Biol Rev. 2009;23:9–19.
Xu L, Meng W, Cao C, Wang J, Shan W, Wang Q. Antibacterial and antifungal compounds from marine fungi. Mar Drugs. 2015;13:3479–513.
Haygood MG, Schmidt EW, Davidson SK, Faulkner DJ. Microbial symbionts of marine invertebrates: opportunities for microbial biotechnology. J Mol Microbiol Biotechnol. 1999;1:33–43.
Rinkevich B. Cell cultures from marine invertebrates: obstacles, new approaches and recent improvements. J Biotechnol. 1999;70:133–53.
Bishara A, Rudi A, Goldberg I, Benayahu Y, Kashman Y. Novaxenicins A–D and xeniolides I–K, seven new diterpenes from the soft coral Xenia novaebrittanniae. Tetrahedron. 2006;62:12092–7.
Hamel C, Prusov EV, Gertsch J, Schweizer WB, Altmann K-H. Total synthesis of the marine diterpenoid blumiolide C. Angew Chem, Int Ed. 2008;47:10081–5.
Sata NU, Sugano M, Matsunaga S, Fusetani N. Sinulamide: an H, K-ATPase inhibitor from a soft coral Sinularia sp. Tetrahedron Lett. 1999;40:719–22.
Sung P-J, Chen Y-P, Hwang T-L, Hu W-P, Fang L-S, Wu Y-C, et al. Briaexcavatins C–F, four new briarane-related diterpenoids from the Formosan octocoral Briareum excavatum (Briareidae). Tetrahedron. 2006;62:5686–91.
Rocha J, Peixe L, Gomes NCM, Calado R. Cnidarians as a source of new marine bioactive compounds—an overview of the last decade and future steps for bioprospecting. Mar Drugs. 2011;9:1860–86.
Zhukova NV. Fatty acids of marine mollusks: Impact of diet, bacterial symbiosis and biosynthetic potential. Biomolecules. 2019;9:857.
Zhukova NV, Kharlamenko VI, Svetashev VI, Rodionov IA. Fatty acids as markers of bacterial symbionts of marine bivalve molluscs. J Exp Mar Biol Ecol. 1992;162:253–63.
Lopera J, Miller IJ, McPhail KL, Kwan JC. Increased biosynthetic gene dosage in a genome-reduced defensive bacterial symbiont. MSystems. 2017;2.
Liu Q-A, Shao C-L, Gu Y-C, Blum M, Gan L-S, Wang K-L, et al. Antifouling and fungicidal resorcylic acid lactones from the sea anemone-derived fungus Cochliobolus lunatus. J Agric Food Chem. 2014;62:3183–91.
Rao KV, Santarsiero BD, Mesecar AD, Schinazi RF, Tekwani BL, Hamann MT. New manzamine alkaloids with activity against infectious and tropical parasitic diseases from an Indonesian sponge. J Nat Prod. 2003;66:823–8.
Sakai R, Higa T, Jefford CW, Bernardinelli G. Manzamine A, a novel antitumor alkaloid from a sponge. J Am Chem Soc. 1986;108:6404–5.
Edrada RA, Proksch P, Wray V, Witte L, Müller WEG, Van Soest RWM. Four new bioactive manzamine-type alkaloids from the Philippine marine sponge Xestospongia ashmorica. J Nat Prod. 1996;59:1056–60.
Nakamura H, Deng S, Kobayashi J, Ohizumi Y, Tomotake Y, Matsuzaki T, et al. Keramamine-A and -B, novel antimicrobial alkaloids from the Okinawan marine sponge Pellina sp. Tetrahedron Lett. 1987;28:621–4.
Ang KKH, Holmes MJ, Higa T, Hamann MT, Kara UAK. In vivo antimalarial activity of the beta-carboline alkaloid manzamine A. Antimicrob Agents Chemother. 2000;44:1645–9.
El Sayed KA, Kelly M, Kara UAK, Ang KKH, Katsuyama I, Dunbar DC, et al. New manzamine alkaloids with potent activity against infectious diseases. J Am Chem Soc. 2001;123:1804–8.
Tsuda M, Kobayashi J. Structures and biogenesis of manzamines and related alkaloids. Heterocycles. 1998;46:765–94.
Magnier E, Langlois Y. Manzamine alkaloids, syntheses and synthetic approaches. Tetrahedron. 1998;54:6201–58.
Yousaf M, El Sayed KA, Rao KV, Lim CW, Hu J-F, Kelly M, et al. 12, 34-Oxamanzamines, novel biocatalytic and natural products from manzamine producing Indo-Pacific sponges. Tetrahedron. 2002;58:7397–402.
Bibi F, Faheem M, Azhar IE, Yasir M, Alvi SA, Kamal MA, et al. Bacteria from marine sponges: a source of new drugs. Curr Drug Metab. 2017;18:11–15.
Zheng L, Chen H, Han X, Lin W, Yan X. Antimicrobial screening and active compound isolation from marine bacterium NJ6-3-1 associated with the sponge Hymeniacidon perleve. World J Microbiol Biotechnol. 2005;21:201–6.
Burkholder PR. Studies on antimicrobial substances of sponges I. Isolation, purification, and properties of a new bromine-containing antibacterial substance. J Antibiot. 1967;20:200–3.
Pabel CT, Vater J, Wilde C, Franke P, Hofemeister J, Adler B, et al. Antimicrobial activities and matrix-assisted laser desorption/ionization mass spectrometry of Bacillus isolates from the marine sponge Aplysina aerophoba. Mar Biotechnol. 2003;5:424–34.
Carroll AR, Copp BR, Davis RA, Keyzers RA, Prinsep MR. Marine natural products. Nat Prod Rep. 2019;36:122–73.
Marty MJ, Vicente J, Oyler BL, Place A, Hill RT. Sponge symbioses between Xestospongia deweerdtae and Plakortis spp. are not motivated by shared chemical defense against predators. PloS ONE. 2017;12:e0174816.
Agarwal V, Blanton JM, Podell S, Taton A, Schorn MA, Busch J, et al. Metagenomic discovery of polybrominated diphenyl ether biosynthesis by marine sponges. Nat Chem Biol. 2017;13:537–43.
Genilloud O. Actinomycetes: still a source of novel antibiotics. Nat Prod Rep. 2017;34:1203–32.
Ramachandran G, Rajivgandhi G, Maruthupandy M, Manoharan N. Isolation and identification of antibacterial compound from marine endophytic actinomycetes against multi drug resistant bacteria. Ann Microbiol Immunol. 2018;1:1003.
Djinni I, Defant A, Kecha M, Mancini I. Antibacterial polyketides from the marine alga-derived endophitic Streptomyces sundarbansensis: a study on hydroxypyrone tautomerism. Mar Drugs. 2013;11:124–35.
El-Gendy MMA, Hawas UW, Jaspars M. Novel bioactive metabolites from a marine derived bacterium Nocardia sp. ALAA 2000. J Antibiot. 2008;61:379–86.
Kalinovskaya NI, Kalinovsky AI, Romanenko LA, Dmitrenok PS, Kuznetsova TA. New angucyclines and antimicrobial diketopiperazines from the marine mollusk-derived actinomycete Saccharothrix espanaensis An 113. Nat Prod Commun. 2010;5:597–602.
Rodríguez V, Martín J, Sarmiento-Vizcaíno A, De la Cruz M, García LA, Blanco G, et al. Anthracimycin B, a potent antibiotic against gram-positive bacteria isolated from cultures of the deep-sea actinomycete Streptomyces cyaneofuscatus M-169. Mar Drugs. 2018;16:406.
Chen M-H, Lian Y-Y, Fang D-S, Chen L, Jia J, Zhang W-L, et al. Identification and antimicrobial properties of a new alkaloid produced by marine-derived Verrucosispora sp. FIM06-0036. Nat Prod Res. 2019:1–7.
Braña AF, Sarmiento-Vizcaíno A, Pérez-Victoria I, Martín J, Otero L, Palacios-Gutiérrez JJ, et al. Desertomycin G, a new antibiotic with activity against Mycobacterium tuberculosis and human breast tumor cell lines produced by Streptomyces althioticus MSM3, isolated from the Cantabrian sea intertidal macroalgae Ulva sp. Mar Drugs. 2019;17:114.
Rajivgandhi G, Ramachandran G, Maruthupandy M, Saravanakumar S, Manoharan N, Viji R. Antibacterial effect of endophytic actinomycetes from marine algae against multi drug resistant gram negative bacteria. Exam Mar Biol Oceanogr. 2018;1:1–8.
Rajivgandhi G, Muneeswaran T, Maruthupandy M, Ramakritinan CM, Saravanan K, Ravikumar V, et al. Antibacterial and anticancer potential of marine endophytic actinomycetes Streptomyces coeruleorubidus GRG 4 (KY457708) compound against colistin resistant uropathogens and A549 lung cancer cells. Microb Pathog. 2018;125:325–35.
Abdalla MA, Sulieman S, McGaw LJ. Microbial communication: a significant approach for new leads. S Afr J Bot. 2017;113:461–70.
Kealey C, Creaven CA, Murphy CD, Brady CB. New approaches to antibiotic discovery. Biotechnol Lett. 2017;39:805–17.
Yang S-Q, Li X-M, Li X, Li H-L, Meng L-H, Wang B-G. New citrinin analogues produced by coculture of the marine algal-derived endophytic fungal strains Aspergillus sydowii EN-534 and Penicillium citrinum EN-535. Phytochem Lett. 2018;25:191–5.
Kaul S, Gupta S, Ahmed M, Dhar MK. Endophytic fungi from medicinal plants: a treasure hunt for bioactive metabolites. Phytochem Rev. 2012;11:487–505.
Wani ZA, Ashraf N, Mohiuddin T, Riyaz-Ul-Hassan S. Plant-endophyte symbiosis, an ecological perspective. Appl Microbiol Biotechnol. 2015;99:2955–65.
Parekh J, Chanda S. Antibacterial and phytochemical studies on twelve species of Indian medicinal plants. Afr J Biomed Res. 2007;10:175–81.
Cheesman MJ, Ilanko A, Blonk B, Cock IE. Developing new antimicrobial therapies: are synergistic combinations of plant extracts/compounds with conventional antibiotics the solution? Pharmacogn Rev. 2017;11:57–72.
Duraipandiyan V, Raja TW, Al-Dhabi NA, Savarimuthu I. Antimicrobial properties of traditional medicinal plants: status and potential. In: Goyal MR, Chauhan DN, editors. Plant- and marine-based phytochemicals for human health: attributes, potential, and use. CRC Press; 2018. p. 33–60.
Rao SR, Ravishankar GA. Plant cell cultures: chemical factories of secondary metabolites. Biotechnol Adv. 2002;20:101–53.
Cowan MM. Plant products as antimicrobial agents. Clin Microbiol Rev. 1999;12:564–82.
Zakaria ZA, Rofiee MS, Mohamed AM, Teh LK, Salleh MZ. In vitro antiproliferative and antioxidant activities and total phenolic contents of the extracts of Melastoma malabathricum leaves. J Acupunct Meridian Stud. 2011;4:248–56.
Ibrahim MA, Mansoor AA, Gross A, Ashfaq MK, Jacob M, Khan SI, et al. Methicillin-resistant Staphylococcus aureus (MRSA)-active metabolites from Platanus occidentalis (American sycamore). J Nat Prod. 2009;72:2141–4.
Nakayama M, Shimatani K, Ozawa T, Shigemune N, Tsugukuni T, Tomiyama D, et al. A study of the antibacterial mechanism of catechins: isolation and identification of Escherichia coli cell surface proteins that interact with epigallocatechin gallate. Food Control. 2013;33:433–9.
Miklasińska M, Kępa M, Wojtyczka RD, Idzik D, Dziedzic A, Wąsik TJ. Catechin hydrate augments the antibacterial action of selected antibiotics against Staphylococcus aureus clinical strains. Molecules. 2016;21:244.
Marchese A, Barbieri R, Coppo E, Orhan IE, Daglia M, Nabavi SF, et al. Antimicrobial activity of eugenol and essential oils containing eugenol: A mechanistic viewpoint. Crit Rev Microbiol. 2017;43:668–89.
Xu J-G, Liu T, Hu Q-P, Cao X-M. Chemical composition, antibacterial properties and mechanism of action of essential oil from clove buds against Staphylococcus aureus. Molecules. 2016;21:1194.
Qiu J, Feng H, Lu J, Xiang H, Wang D, Dong J, et al. Eugenol reduces the expression of virulence-related exoproteins in Staphylococcus aureus. Appl Environ Microbiol. 2010;76:5846–51.
Dhara L, Tripathi A. Antimicrobial activity of eugenol and cinnamaldehyde against extended spectrum beta lactamase producing enterobacteriaceae by in vitro and molecular docking analysis. Eur J Integr Med. 2013;5:527–36.
Marcos-Arias C, Eraso E, Madariaga L, Quindós G. In vitro activities of natural products against oral Candida isolates from denture wearers. BMC Complementary Altern Med. 2011;11:119.
Ahmad A, Khan A, Khan LA, Manzoor N. In vitro synergy of eugenol and methyleugenol with fluconazole against clinical Candida isolates. J Med Microbiol. 2010;59:1178–84.
de Oliveira Pereira F, Mendes JM, de Oliveira Lima E. Investigation on mechanism of antifungal activity of eugenol against Trichophyton rubrum. Med Mycol 2013;51:507–13.
Ikram M. A review on the chemical and pharmacological aspects of genus. Berberis Planta Med. 1975;28:353–8.
Neag MA, Mocan A, Echeverría J, Pop RM, Bocsan CI, Crişan G, et al. Berberine: Botanical occurrence, traditional uses, extraction methods, and relevance in cardiovascular, metabolic, hepatic, and renal disorders. Front Pharm. 2018;9:557.
Peng L, Kang S, Yin Z, Jia R, Song X, Li L, et al. Antibacterial activity and mechanism of berberine against Streptococcus agalactiae. Int J Clin Exp Pathol. 2015;8:5217–23.
Yu H-H, Kim K-J, Cha J-D, Kim H-K, Lee Y-E, Choi N-Y, et al. Antimicrobial activity of berberine alone and in combination with ampicillin or oxacillin against methicillin-resistant Staphylococcus aureus. J Med Food. 2005;8:454–61.
Kim W-S, Choi WJ, Lee S, Kim WJ, Lee DC, Sohn UD, et al. Anti-inflammatory, antioxidant and antimicrobial effects of artemisinin extracts from Artemisia annua L. Korean J Physiol Pharm. 2015;19:21–27.
Appalasamy S, Lo KY, Ch’ng SJ, Nornadia K, Othman AS, Chan L-K. Antimicrobial activity of artemisinin and precursor derived from in vitro plantlets of Artemisia annua L. BioMed Res Int. 2014;2014.
Macé S, Hansen LT, Rupasinghe HPV. Anti-bacterial activity of phenolic compounds against Streptococcus pyogenes. Medicines. 2017;4:25.
Kowalski KP, Bacon C, Bickford W, Braun H, Clay K, Leduc-Lapierre M, et al. Advancing the science of microbial symbiosis to support invasive species management: a case study on Phragmites in the Great Lakes. Front Microbiol. 2015;6:95.
Tanaka Y, Hosaka T, Ochi K. Rare earth elements activate the secondary metabolite–biosynthetic gene clusters in Streptomyces coelicolor A3(2). J Antibiot. 2010;63:477–81.
Traxler MF, Seyedsayamdost MR, Clardy J, Kolter R. Interspecies modulation of bacterial development through iron competition and siderophore piracy. Mol Microbiol. 2012;86:628–44.
Bhardwaj C, Cui Y, Hofstetter T, Liu SY, Bernstein HC, Carlson RP, et al. Differentiation of microbial species and strains in coculture biofilms by multivariate analysis of laser desorption postionization mass spectra. Analyst. 2013;138:6844–51.
Bertrand S, Schumpp O, Bohni N, Bujard A, Azzollini A, Monod M, et al. Detection of metabolite induction in fungal co-cultures on solid media by high-throughput differential ultra-high pressure liquid chromatography–time-of-flight mass spectrometry fingerprinting. J Chromatogr A. 2013;1292:219–28.
Du J, Zhou J, Xue J, Song H, Yuan Y. Metabolomic profiling elucidates community dynamics of the Ketogulonicigenium vulgare–Bacillus megaterium consortium. Metabolomics. 2012;8:960–73.
Derewacz DK, Covington BC, McLean JA, Bachmann BO. Mapping microbial response metabolomes for induced natural product discovery. ACS Chem Biol. 2015;10:1998–2006.
Goodwin CR, Covington BC, Derewacz DK, McNees CR, Wikswo JP, McLean JA, et al. Structuring microbial metabolic responses to multiplexed stimuli via self-organizing metabolomics maps. Chem Biol. 2015;22:661–70.
Goodwin CR, Sherrod SD, Marasco CC, Bachmann BO, Schramm-Sapyta N, Wikswo JP, et al. Phenotypic mapping of metabolic profiles using self-organizing maps of high-dimensional mass spectrometry data. Anal Chem. 2014;86:6563–71.
Watrous J, Roach P, Alexandrov T, Heath BS, Yang JY, Kersten RD, et al. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci USA. 2012;109:E1743–52.
Yang JY, Sanchez LM, Rath CM, Liu X, Boudreau PD, Bruns N, et al. Molecular networking as a dereplication strategy. J Nat Prod. 2013;76:1686–99.
Kersten RD, Yang Y-L, Xu Y, Cimermancic P, Nam S-J, Fenical W, et al. A mass spectrometry-guided genome mining approach for natural product peptidogenomics. Nat Chem Biol. 2011;7:794–802.
Liu W-T, Lamsa A, Wong WR, Boudreau PD, Kersten R, Peng Y, et al. MS/MS-based networking and peptidogenomics guided genome mining revealed the stenothricin gene cluster in Streptomyces roseosporus. J Antibiot. 2014;67:99–104.
Bibb MJ. Regulation of secondary metabolism in streptomycetes. Curr Opin Microbiol. 2005;8:208–15.
Hopwood DA. How do antibiotic‐producing bacteria ensure their self‐resistance before antibiotic biosynthesis incapacitates them? Mol Microbiol. 2007;63:937–40.
Davies J, Spiegelman GB, Yim G. The world of subinhibitory antibiotic concentrations. Curr Opin Microbiol. 2006;9:445–53.
Yim G, Wang HH, Davies J. Antibiotics as signalling molecules. Philos Trans R Soc, B. 2007;362:1195–1200.
Imai Y, Sato S, Tanaka Y, Ochi K, Hosaka T. Lincomycin at subinhibitory concentrations potentiates secondary metabolite production by Streptomyces spp. Appl Environ Microbiol. 2015;81:3869–79.
Netzker T, Fischer J, Weber J, Mattern DJ, König CC, Valiante V, et al. Microbial communication leading to the activation of silent fungal secondary metabolite gene clusters. Front Microbiol. 2015;6:299.
Macheleidt J, Mattern DJ, Fischer J, Netzker T, Weber J, Schroeckh V, et al. Regulation and role of fungal secondary metabolites. Annu Rev Genet. 2016;50:371–92.
Andersen RJ. Sponging off nature for new drug leads. Biochem Pharmacol. 2017;139:3–14.
Sagar S, Kaur M, Minneman KP. Antiviral lead compounds from marine sponges. Mar Drugs. 2010;8:2619–38.
Brinkmann CM, Marker A, Kurtböke DI. An overview on marine sponge-symbiotic bacteria as unexhausted sources for natural product discovery. Diversity. 2017;9:40.
Pandey S, Sree A, Dash SS, Sethi DP, Chowdhury L. Diversity of marine bacteria producing beta-glucosidase inhibitors. Microb Cell Fact. 2013;12:35.
Epstein SS. The phenomenon of microbial uncultivability. Curr Opin Microbiol. 2013;16:636–42.
Bhatnagar I, Kim S-K. Immense essence of excellence: Marine microbial bioactive compounds. Mar Drugs. 2010;8:2673–701.
Leal MC, Calado R, Sheridan C, Alimonti A, Osinga R. Coral aquaculture to support drug discovery. Trends Biotechnol. 2013;31:555–61.
Leal MC, Calado R. Marine natural products: biodiscovery, biodiversity, and bioproduction. In: Brahmachari G, editor. Bioactive natural products: chemistry and biology. Hoboken: Wiley Online Library; 2015. p. 473–90.
Leal MC, Sheridan C, Osinga R, Dionísio G, Rocha RJM, Silva B, et al. Marine microorganism-invertebrate assemblages: perspectives to solve the “supply problem” in the initial steps of drug discovery. Mar Drugs. 2014;12:3929–52.
Sweet MJ, Bulling MT. On the importance of the microbiome and pathobiome in coral health and disease. Front Mar Sci. 2017;4:9.
Sweet MJ, Smith D, Bythell JC, Craggs J. Changes in microbial diversity associated with two coral species recovering from a stressed state in a public aquarium system. J Zoo Aquar Res. 2013;1:52–60.
Trindade M, van Zyl LJ, Navarro-Fernández J, Abd Elrazak A. Targeted metagenomics as a tool to tap into marine natural product diversity for the discovery and production of drug candidates. Front Microbiol. 2015;6:890.
Acknowledgements
This study was provided by Abney Foundation, the Charles and Carol Cooper Endowment, and the South Carolina SmartState Programs and funding from NCCIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Dedicated to Professor William Fenical in recognition of his contributions to marine derived secondary metabolites.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gogineni, V., Chen, X., Hanna, G. et al. Role of symbiosis in the discovery of novel antibiotics. J Antibiot 73, 490–503 (2020). https://doi.org/10.1038/s41429-020-0321-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41429-020-0321-6
This article is cited by
-
Winners of the 2021 JA Ōmura Awards for excellence
The Journal of Antibiotics (2023)